Screening and Identification of Small Peptides Targeting Fibroblast Growth Factor Receptor2 using a Phage Display Peptide Library
Summary
Herein, we present a detailed protocol for screening small peptides that bind to FGFR2 using a phage display peptide library. We further analyze the affinity of the selected peptides toward FGFR2 in vitro and its ability to suppress cell proliferation.
Abstract
The human Fibroblast Growth Factor Receptor (FGFR) family comprises four members, namely, FGFR1, FGFR2, FGFR3, and FGFR4, that are involved in various biological activities including cell proliferation, survival, migration and differentiation. Several aberrations in the FGFR signaling pathway, owing to mutations or gene amplification events, have been identified in various types of cancers. Hence, recent research has focused on developing strategies involving therapeutic targeting of FGFRs. Current FGFR inhibitors at various stages of pre-clinical and clinical development include either small molecule inhibitors of tyrosine kinases or monoclonal antibodies, with only a few peptide- based inhibitors in the pipeline. Here, we provide a protocol using phage display technology to screen small peptides as antagonists of FGFR2. Briefly, a library of phage-displayed peptides was incubated in a plate coated with FGFR2. Subsequently, unbound phage was washed off by TBST (TBS + 0.1% [v/v] Tween-20), and bound phage was eluted with 0.2 M glycine-HCl buffer (pH 2.2). The eluted phage was further amplified and used as input for the next round of biopanning. Following three rounds of biopanning, the peptide sequences of individual phage clones were identified by DNA sequencing. Finally, the screened peptides were synthesized and analyzed for affinity and biological activity.
Introduction
Fibroblast growth factor receptors (FGFRs) play key roles in cell proliferation, wound healing, and angiogenesis in vivo1. Aberrant activation of FGFR signaling observed in a variety of tumors2,3,4,5 includes gene amplification, gene mutations, chromosomal aberrations, and excessive ligand secretion6. Many inhibitors targeting FGFRs have shown promising therapeutic effects in clinical trials and are mainly classified into three types: (1) small molecule kinase inhibitors, which bind to the intracellular domain of FGFR, (2) antagonists targeting the extracellular segment, and (3) FGF ligand traps6. Although several of the small molecule kinase inhibitors have good therapeutic effects both in vitro and in vivo7, most of them have poor target specificity and show adverse effects such as hypertension8. The majority of the antagonists are monoclonal antibodies9,10 and polypeptides11. Peptides have advantages over small molecules due to their specificity and lower side effects. They also retain cell permeability and do not accumulate in specific organs as compared to protein drugs12. Hence, targeted small peptides are both effective and prospective therapeutic agents.
Phage display technology is an easy but powerful tool for identifying small peptides which can bind to a given molecule13,14,15. We used a phage display peptide library that is based on a simple M13 phage with over 109 different peptide sequences displayed at the tail for binding to the target molecule (see Table of Materials)16. Due to the high affinity of phages towards the given molecule, unbound phages can be washed away, and only tightly bound phages with the desired short peptides are retained. The given molecular targets can be immobilized proteins17,18, carbohydrates, cultured cells, or even inorganic materials19,20. An exciting case was reported where organ-specific peptides were selected in vivo using phage display technology21. The advantages of phage display technology include high throughput, ease of operation, low cost and a wide range of applications22.
In this study, we provide a detailed protocol of screening small peptides binding to the immobilized protein (FGFR2) using a phage display library. The efficacy of the technology is also examined by measuring the affinity of the obtained peptide towards FGFR2 by Isothermal Titration Calorimetry (ITC), and the biological activity by a cell proliferation assay. The method may be extended for screening small peptides that bind to carbohydrates, cultured cells, or even inorganic materials.
Protocol
1. Reagent preparation
- LB (lysogeny broth) medium: Dissolve 1 g of tryptone, 0.5 g of yeast extract, and 0.5 g of NaCl in 100 mL of H2O. Autoclave and store at 4 °C.
- Tetracycline stock: Prepare 20 mg/mL in 1:1 ethanol:H2O. Store at -20 °C, and vortex before using.
- IPTG/X-gal solution: Mix 0.5 g of IPTG (isopropyl β-D-1-thiogalactopyranoside) and 0.4 g of X-gal (5-bromo-4-chloro-3-indolyl β-D-galactoside) in 10 mL of DMF (dimethyl formamide). The solution can be stored at -20 °C for one year in the dark.
- LB + Tetracycline (Tet) plates: Dissove 1 g of bacto-tryptone, 0.5 g of yeast extract, 0.5 g of NaCl, and 1.5 g of agar in 100 mL of H2O. Autoclave and cool to < 70 °C. Add 100 μL of the tetracycline stock and pour plates. Store plates at 4 °C in the dark.
- LB + IPTG/X-gal plates: Dissolve 1 g of bacto-tryptone, 0.5 g of yeast extract, 0.5 g of NaCl, and 1.5 g of agar in 100 mL of H2O. Autoclave and cool to < 70°C. Add 100 μL of the IPTG/X-gal solution and pour plates. Store plates at 4 °C in the dark.
- Glycine-HCI buffer (0.2 M): Dissolve 1.5014 g of glycine and 100 mg of BSA (bovine serum albumin) in 100 mL of H2O and adjust the pH to 2.2 with HCl. Filter the solution using a 0.22 μm filter and store at 4 °C.
- Tris-HCl buffer (1 M): Dissolve 12.11 g of Tris (tris(hydroxymethyl)aminomethane) in 100 mL of H2O and adjust the pH to 9.1 with HCl. Autoclave and store at 4 °C.
- Coating buffer (0.1 M): Dissolve 1.68 g of NaHCO3 in 200 mL of H2O and adjust the pH to 8.6 with NaOH. Filter the solution using a 0.22 μm filter and store at 4 °C.
- Blocking buffer: Prepare 0.1 M NaHCO3 (pH 8.6) with 5 mg/mL BSA. Filter the solution using a 0.22 μm filter and store at 4 °C.
- TBS: Prepare 50 mM Tris-HCl (pH 7.5) with 150 mM NaCl. Autoclave and store at room temperature.
- 0.05%, 0.1%, or 0.5% TBST: Prepare TBS with 0.05%, 0.1%, or 0.5% (v/v) Tween-20. Filter using a 0.22 μm filter and store at room temperature.
- PEG/NaCl: Prepare 20% (w/v) polyethylene glycol–8000 with 2.5 M NaCl. Autoclave and store at room temperature.
- TE solution: Mix 1 mL of 1 M Tris-HCl (pH8.0) and 2 mL of 50 mM EDTA (pH 8.0) and adjust the volume to 100 mL with H2O. Autoclave and store at room temperature.
- Iodide buffer: Dissolve 30 g of NaI in 50 mL of TE, pH 8.0. Store the solution at room temperature in the dark.
- Top Agar: Dissolve 1 g of tryptone, 0.5 g of yeast extract, 0.5 g of NaCl, and 0.7 g agarin in 100 mL of H2O. Autoclave and dispense into 15 mL aliquots. Store at room temperature for use.
2. First round of biopanning: Screening phage clones that bind to the extracellular domain of FGFR2
NOTE: Use aerosol-resistant pipette tips and wear gloves for all protocols in order to minimize contamination with environmental bacteriophages.
- Prepare a 10 µg/mL FGFR2 (Purity > 97%; see Table of Materials) solution in coating buffer. Add 1 mL of the prepared solution to a 35 cm2 dish and swirl repeatedly until the surface is completely wet. Incubate overnight at 4 °C with shaking.
NOTE: To avoid selection of decoy target binders, the target protein must be highly purified. - Inoculate 5 mL of LB + Tet medium with Escherichia coli (E. coli) ER2738 for 5–10 hours at 37 °C with shaking. The amplified ER2738 cells will be used for titration in step 3.3.
- Pour off the coating solution from the dish (the excess solution must be removed) and add the blocking solution. Incubate at 4 °C for at least 1 h.
- Discard the blocking solution and wash 6 times quickly with TBST (TBS + 0.05% (v/v] Tween-20). Avoid drying out the dish.
- Reconstitute the original phage library in 1 mL of TBST [final concentration of 1011 plaque forming units (PFU)], add to the coated dish, and rock gently for 2 h on a shaker at room temperature.
- Discard the supernatant and wash the dish 10x with TBST (TBS + 0.05% (v/v) Tween-20), as performed in step 2.4.
NOTE: The unbound phages must be removed thoroughly by vigorous washing. - Elute the bound phage by adding 1 mL of 0.2 M glycine-HCl buffer (pH 2.2) to the dish and rocking gently for 10 min at room temperature.
- Transfer the eluate into a sterilized microfuge tube and neutralize it with 100 µL of 1 M Tris-HCl, pH 9.1.
NOTE: In step 2.6, remove the unbound phage as much as possible by washing the dish 10x vigorously. This is a crucial step. The eluted phage can be stored at 4 °C for one week. However, it is best to go forward to the next step as soon as possible.
3. Titer determination of the eluted phages
- Pre-warm LB + IPTG/X-gal plates for at least 1 h at 37 °C before use.
- Prepare 100 µL of 10-fold serial dilutions (i.e., 101, 102, 103 and 104) of the eluate from step 2.8 in LB. Use tips with filter cartridges to avoid contamination.
- Let the culture from step 2.2 reach mid-log phase (OD600 ≈ 0.5). Dispense 200 µL of the culture into sterilized microfuge tubes, one for each eluate dilution.
- To initiate the infection, add 10 µL of each dilution from step 3.2 to individual microfuge tubes containing the E. coli ER2738 culture from step 3.3. Vortex quickly and incubate at room temperature for 5 min.
- Coat 90 μL of the mixture from step 3.4 into the LB +IPTG/X-gal plates with coating sticks.
- Optionally, melt top agar in a microwave, dispense 3 mL into sterile culture tubes, one for each eluate dilution, and maintain tubes at 45 °C. Transfer the mixture from step 3.4 to culture tubes containing 45 °C top agar. Mix quickly and immediately pour the culture onto a pre-warmed LB/IPTG/X-gal plate. Gently tilt and rotate the plate to spread the top agar evenly.
- After 5 min, invert and incubate plates overnight at 37 °C.
- Count plaques on plates with approximately 100 plaques. Multiply each number by the dilution factor for that plate to get the phage titer in PFU per 10 µL.
- Inoculate another 5 mL of LB + Tet medium with E. coli ER2738 at 37 °C overnight with shaking for phage amplification.
NOTE: Use the culture of ER2738 in mid-log phase (OD600 ≈ 0.5) for phage titer. If most of the plaques are white on X-gal/IPTG plates, it means that the pool of phage has been contaminated with wild bacteriophages. If contamination occurs, it is necessary to start again with the panning steps from the uncontaminated phage stock solution.
4. Phage amplification
- Dilute (1:100) the overnight culture from step 3.8 in 20 mL of LB in a 250 mL Erlenmeyer flask. Add the unamplified eluate and incubate with vigorous shaking for 4.5 h at 37 °C.
- Transfer the culture to a centrifuge tube and centrifuge for 10 min at 12,000 x g at 4 °C. Transfer the supernatant to a fresh tube, centrifuge again, and discard the pellet.
- Transfer the upper 80% of the supernatant to a fresh tube and add to it 1/6 volume of 20% PEG/2.5 M NaCl. Allow the phage to precipitate at 4 °C overnight.
NOTE: Mix the culture supernatant with PEG/2.5 M NaCl well. - Centrifuge the precipitated phage at 12,000 x g for 15 min at 4 °C. Discard the supernatant, centrifuge again, and remove the residual supernatant with a pipette.
- Resuspend the pellet in 1 mL of TBS and transfer the supernatant to a microfuge tube. Centrifuge at 12,000 x g for 5 min at 4 °C.
- Transfer the supernatant to a fresh microfuge tube and precipitate again by adding 1/6 volume of 20% PEG/2.5 M NaCl. Incubate on ice for 15–60 min. Centrifuge at 12,000 x g for 10 min at 4 °C. Discard the supernatant, re-spin, and remove the residual supernatant with a pipette.
- Resuspend the pellet in 200 μL of TBS and centrifuge for an additional minute to remove impurities. Transfer the supernatant to a fresh microfuge tube to obtain the amplified eluate.
- Titer the amplified phage as described in section 3.
NOTE: The dilution of the amplified eluate is higher compared to the unamplified eluate; serial dilutions of 108–1011 are typically suggested.
5. Second round of biopanning
- Repeat sections 2 to 4.
- Titer the second round of the amplified eluate on LB + IPTG/X-gal plates as described in section 3.
NOTE: Reduce the coating concentration of FGFR2 protein to 5 μg/mL. Use the amplified phage from step 4.7 as the input and keep the titer same as in the first round (1011 PFU). Shorten the incubation time of phage and FGFR2-coated dish to 1 h and increase the Tween concentration to 0.1% (v/v) in the wash steps.
6. Third round of biopanning
- Repeat steps 2.1-2.6.
- Elute the bound phage by adding 1 mL of 20 μM bFGF solution (a ligand for FGFR2) to the dish and rocking gently for 60 min at room temperature.
- Repeat steps 3.1-3.7.
NOTE: Reduce the coating concentration of the FGFR2 protein to 2.5 μg/mL. Use the second round of amplified phage from section 5 as the input and keep the titer same as in the first round (1011 PFU). Shorten the incubation time of phage and FGFR2-coated dish to 30 min and increase the Tween concentration to 0.5% (v/v) in the wash steps.
7. Acquisition of plaque DNA for sequencing
- Plaque amplification
- Dilute (1:100) an overnight culture of ER2738 in LB. Dispense 1 mL of diluted culture into a 2 mL tube, one for each clone to be characterized.
- Use a pipette tip to pick one well-separated blue colored plaque from a titer plate from section 6 and transfer to the tube containing the diluted culture. Pick a total of 15 plaques.
NOTE: Plates should be < 1–3 days old, stored at 4 °C, and have < 100 plaques. - Incubate the tubes with vigorous shaking for 4.5 h at 37 °C.
- Transfer the cultures to centrifuge tubes and centrifuge at 12,000 x g for 30 s. Transfer the supernatant to a fresh tube and centrifuge again.
- Using a pipette, transfer the upper 80% of the supernatant to a fresh tube. Label this as the amplified phage stock.
NOTE: Dilute the amplified phage with an equal volume of 100% sterile glycerol and store at -80 °C for later use.
- Extraction of plaque DNA
- Transfer 500 µL of the amplified phage to a fresh microfuge tube and add a 1/6 volume of 20% PEG/2.5 M NaCl. Invert to mix and let it stand for 10–20 min at room temperature.
- Centrifuge at 12,000 x g for 10 min at 4 °C and then pour off the supernatant.
NOTE: Phage pellet may not be visible. - Centrifuge again. Carefully remove any remaining supernatant.
- Resuspend the pellet thoroughly in 100 µL of the iodide buffer by vigorously tapping the tube.
- Add 250 µL of 100% ethanol and incubate for 15 min at room temperature.
- Centrifuge at 12,000 x g for 10 min at 4 °C and remove the supernatant. Wash the pellet with 0.5 mL of 70% ethanol, quickly centrifuge again, remove the supernatant, and briefly dry the pellet.
- Resuspend the pellet in 30 µL of TE buffer and use this for sequencing.
NOTE: Pre-cool the 70% ethanol at -20°C in advance of Step 7.2.6. The phage pellet must be thoroughly resuspended in the iodide buffer prior to the addition of ethanol.
8. Identification of the peptide sequence
- Use the -96 gIII sequencing primer (5´-CCC TCA TAG TTA GCG TAA CG –3´) for sequencing.
- Analyze the peptide sequences displayed on the phage based on the DNA sequencing results.
- Synthesize small peptides and analyze by high-performance liquid chromatography (HPLC) and mass spectrometry to confirm their purity (≥98%).
9. Detecting the affinity of the obtained small peptide and extracellular protein of FGFR2 by ITC
- Degas sterile water using an ultrasonic instrument. Dissolve the small peptide and the FGFR2 extracellular protein in sterile water. Centrifuge these samples to remove residual precipitate.
- Perform the ITC titration experiments at 25 °C. Titrate 40 μM of small peptide in the cell with 1.5 μM of FGFR2 protein from the syringe, with a stirring speed of 750 rpm. Add the FGFR2 protein in 2 μL aliquots (for a total of 19 injections) at 5-min intervals.
- Analyze the ITC data using a single-site binding model.
NOTE: Sterile water must be degassed, and all samples must be centrifuged to remove air bubbles and residual impurities. The experiment must be conducted at a constant temperature of 25 °C.
10. Verifying the biological activity of the obtained small peptide using a cell proliferation assay
- Seed BALB/c 3T3 cells (3.0 × 103 cells/well, 100 μL) in a 96-well plate with Dulbecco's Modified Eagle Medium (DMEM) with 10% fetal bovine serum (FBS) overnight at 37 °C and 5% CO2.
- Replace the supernatant with fresh DMEM with 0.5% FBS and starve the cells for 12 h.
- Make serial dilutions of the small peptide (0, 0.0064, 0.032, 0.16, 0.8, 4, 20, and 100 μM) with fresh DMEM supplemented with 0.5% FBS (6 duplicate wells /concentration). Remove the supernatant from the 96-well plate and add 100 μL of the small peptide solution per well. Incubate for 48 h at 37 °C and 5% CO2.
- Replace the supernatant with fresh DMEM containing 10% Cell Counting Kit (CCK)-8 solution and incubate for 2 h.
- Measure the absorbance of each well at 450 nm with a microplate reader and analyze cell viability.
Representative Results
Obtaining a high affinity small peptide targeting FGFR2.
To screen phages targeting FGFR2, a Ph.D.-7 library was used in this study. A schematic representation of the workflow is shown in Figure 1. In this process, the number of phage input (PFU) was kept unchanged, whereas the coating concentration of FGFR2 protein was reduced gradually. Results of the phage titer suggested that the number of recovered phages increased gradually, and after 3 rounds, there was a 65-fold increase as compared to that of the first round (Table 1).
Next, we picked phage clones randomly after the third round and extracted the phage DNA for sequencing. One representative peptide (WRARVPL) obtained by screening was named SP1. It was subsequently synthesized, and its molecular weight was measured by mass spectrometry.
The SP1 peptide showed high binding affinity towards FGFR2.
An ITC experiment was conducted to measure the affinity of the small peptide to FGFR2. The result indicated that the SP1 peptide has high affinity towards FGFR2 (Kd ≈ 1.4 μM; Figure 2). This date demonstrated the efficiency of the screening protocol.
The SP1 peptide inhibited the growth of fibroblasts.
To investigate the biological activity of the SP1 peptide, a fibroblast proliferation assay was performed using a CCK-8 kit. BALB/c 3T3 cells were incubated with the SP1 peptide at different concentrations (0, 0.0064, 0.032, 0.16, 0.8, 4, 20, and 100 μM). The result suggested that the SP1 peptide suppressed the growth of BALB/c 3T3 cells (Figure 3).
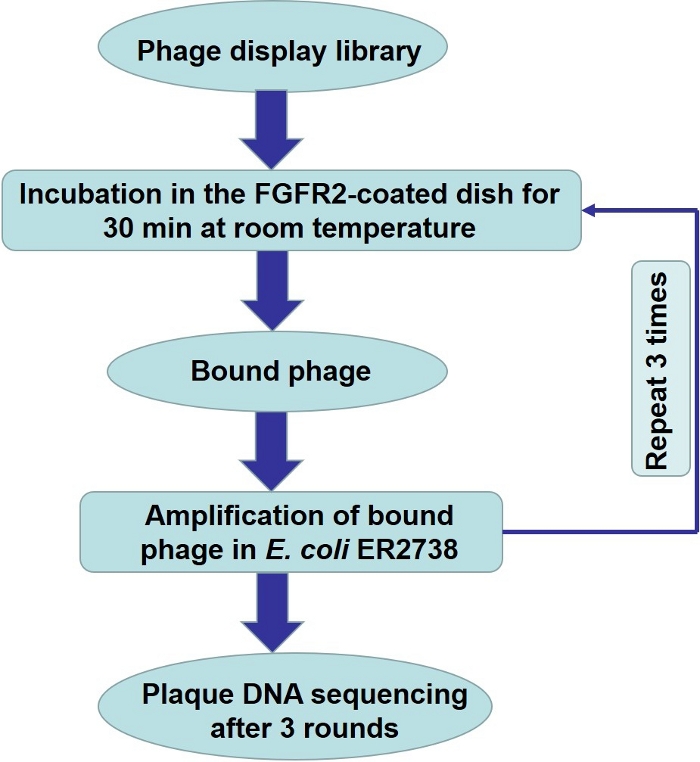
Figure 1. Biopanning for small peptide targeting FGFR2 using phage display library.
(A) Schematic representation of small peptide screening using phage display library. Incubate the library containing 1011 PFU in a dish coated with FGFR2. Discard unbound phages by washing, and elute bound phages. Amplify the eluted phages and use them as input for the next round of biopanning. Please click here to view a larger version of this figure.
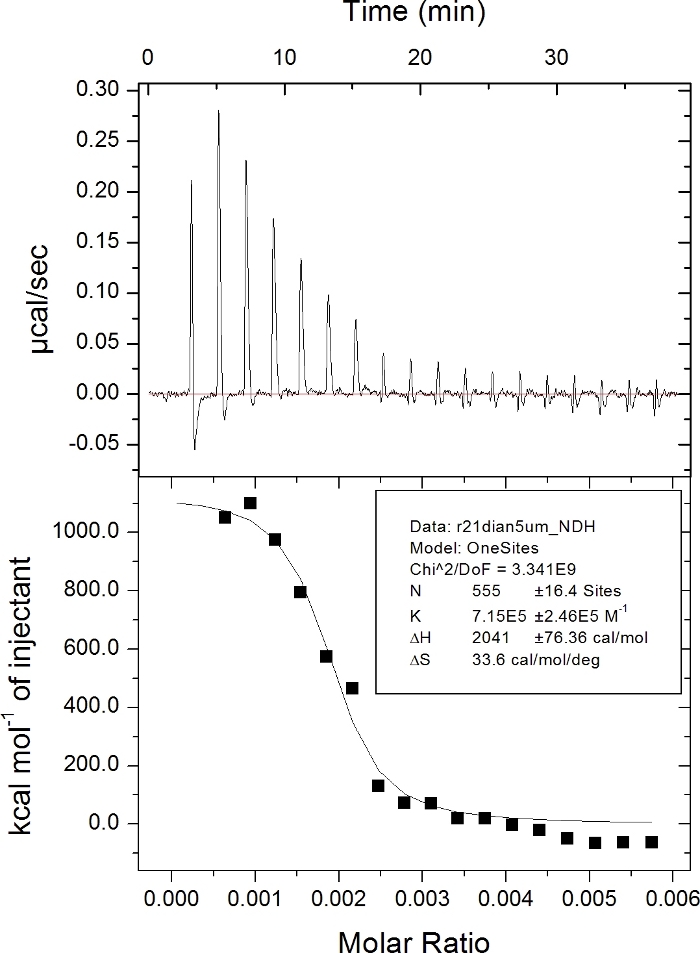
Figure 2. Measuring the affinity of interaction between the SP1 peptide and FGFR2 by ITC.
1.5 μM FGFR2 protein was titrated with 40 μM of the SP1 peptide. The ITC data was further analyzed using Origin 7.0 software. Please click here to view a larger version of this figure.
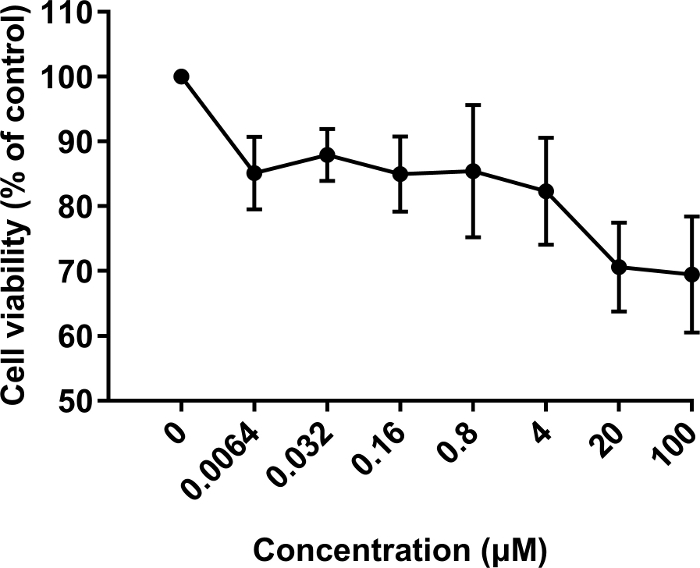
Figure 3. Effect of the SP1 peptide on the growth of fibroblasts.
BALB/c 3T3 cells were treated with the SP1 peptide at different concentrations (0, 0.0064, 0.032, 0.16, 0.8, 4, 20 and 100 μM). The results are expressed as the mean ± SD. Please click here to view a larger version of this figure.
| Round | FGFR2 (μg) | Input phage (PFU) | Output phage (PFU) | Recovery (%) | Enrichment |
| 1 | 10 | 2.0 x 1011 | 3.98 x 104 | 1.99 x 10-5 | 1 |
| 2 | 5 | 2.0 x 1011 | 8.1 x 105 | 4.05 x 10-4 | 20 |
| 3 | 2.5 | 2.0 x 1011 | 2.6 x 106 | 1.3 x 10-3 | 65 |
Table 1. Enrichment of phage targeting FGFR2 relative to round 1.
Discussion
A combinatorial phage library is a powerful and effective tool for high-throughput screening of novel peptides that can bind target molecules and regulate their function13. Currently, phage display peptide libraries have a wide range of applications. For example, they can be used for selecting bioactive peptides bound to receptor proteins23, non-protein targets24,25, disease-specific antigen mimics13, cell-specific peptides26,27, or tissue/organ-specific peptides21,28,29 and development of peptide-mediated drug delivery systems19,30. In brief, phage display peptide libraries are useful and efficient systems to identify specific peptides in basic research and development of translational medicine. In the study, a library was used for screening small peptides targeting FGFR2 for cell growth inhibition.
The critical steps of the protocol are mentioned below. In the panning step, contamination by wild-type phages must be avoided. The unbound phages must be removed thoroughly by vigorous washing. The LB + IPTG/X-gal plates for phage titer must be pre-warmed to 37 °C. For the phage titer, E. coli ER2738 must be grown till mid-log phase (OD600 ≈ 0.5). In step 4.3, the phage must be precipitated overnight at 4 °C. Well separated blue plaques must be picked from platea containing less than 100 plaques for DNA sequencing. In step 7.2.6, the 70% ethanol solution must be pre-cooled at 20 °C in advance. Lastly, for the ITC experiment, sterile water must be degassed, and all samples must be centrifuged to remove air bubbles and residual impurities. The experiment must be conducted at a constant temperature of 25 °C.
Some factors that can affect the quality of hits obtained22 are as follows: 1) In the first round of screening, the number of phage inputs needs to be 1011 PFU. However, in the next round of screening, the input can be lower. 2) A pure environment must be maintained to avoid wild-type phage contamination. 3) 3 or 4 rounds of screening are usually sufficient. Avoid over-panning the peptide library. 4) In each round, the amount of target protein is gradually reduced, whereas the content of Tween is gradually increased in the washing step. In addition, the incubation time of phage and FGFR2-coated dish is also gradually shortened. If most of the eluted phage plaques are white on X-gal/IPTG plates, this suggests contamination by wild-type phages. In order to avoid a low titer of amplified phage in the panning process, cultures must be well aerated and infected early in their growth phase (OD600 < 0.05).
In this study, the SP1 peptide showed high affinity towards FGFR2 (Kd ≈ 1.4 μM; Figure 2) and good biological activity (Figure 3). Thus, our results suggest that the protocol was effective in selecting peptides against the extracellular domain of FGFR2 protein, although note that due to the high sequence similarity of FGFR family members, the SP1 peptide may have some binding affinity with FGFR family members besides FGFR2, which could be responsible for the biological activity seen. Also, in the panning process, phage ELISA is an alternative to identify the positive phages qualitatively. However, in our lab, we always obtain the peptides by sequencing and evaluating their affinity by ITC assay quantitatively and have not had problems evaluating and selecting the candidate peptides.
The phage-display library has several advantages22. The libraries are of high capacity (up to 1011 PFU) and can be used in vitro, in vivo, and ex vivo. Libraries are highly efficient, easy to handle, inexpensive and commercially available. However, there are certain limitations of the technology. The libraries only contain proteinogenic amino acids and are only amenable to linear and simple cyclic peptides without complicated structures.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This work was supported by the Science and Technology Program of Guangzhou (No. 2016201604030039).
Materials
| 0.22 μm Filter | Merck Millipore | MPGP002A1 | |
| 35 cm2 Small dish | Thermo | 150460 | |
| 70% Ethanol | Guangzhou chemical reagent factory | 64-17-5 | |
| -96 gIII sequencing primer | Synthesis from Sangon Biotech (Shanghai) Co., Ltd. | ||
| 96-well plate | Nest | 701001-2 | |
| Agar | Beyotime | ST004D | |
| Bacto-Tryptone | Oxoid | L0037 | |
| BALB/c 3T3 cells | ATCC | CRL-6587 | |
| BSA | Biodragon | BD-M10110 | |
| CCK-8 kit | DOJINDO | CK04 | |
| DMEM | Hyclone | sh30243.01 | |
| DMF | Newprobe | PB10247 | |
| EDTA | Invitrogen | 15576028 | |
| FGF2 Protein | Sino Biological Inc. | 10014-HNAE | Purity >95% |
| Glycine | Sigma | G8898-1KG | |
| IPTG | Beyotime | ST097 | |
| ITC200 system | MicroCal Omega | ||
| NaCl | Sigma | S6191 | |
| NaHCO3 | Guangzhou chemical reagent factory | 144-55-8 | |
| NaI | Bidepharm | BD40879 | |
| NaOH | Guangzhou chemical reagent factory | 1310-73-2 | |
| PEG–8000 | Sigma | P2139-250 | |
| Ph.D.-7 phage display peptide library kit | New England BioLabs | E8100S | Containing the Ph.D.-7 phage library, E. coli ER2738 host strain and M13KE control phage |
| Recombinant FGFR2 extracellular domain proteins | Sino Biological Inc. | 10824-H08H | Purity > 97% |
| Small peptide | Synthesis from GL Biochem Ltd. (Shanghai, China) | ||
| Tetracycline | Sigma | S-SHS-5 | |
| Tris | Sigma | SLF-T1503 | |
| Tween-20 | Beyotime | ST825 | |
| X-gal | Beyotime | ST912 | |
| Yeast extract | Oxoid | LP0021 |
Riferimenti
- Eswarakumar, V. P., Lax, I., Schlessinger, J. Cellular signaling by fibroblast growth factor receptors. Cytokine & Growth Factor Reviews. 16 (2), 139-149 (2005).
- Turner, N., Grose, R. Fibroblast growth factor signaling: from development to cancer. Nature Reviews Cancer. 10 (2), 116-129 (2010).
- Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumors. Nature. 490 (7418), 61-70 (2012).
- Matsumoto, K., et al. FGFR2 gene amplification and clinicopathological features in gastric cancer. British Journal of Cancer. 106 (4), 727-732 (2012).
- Weiss, J., et al. Frequent and focal FGFR1 amplification associates with therapeutically tractable FGFR1 dependency in squamous cell lung cancer. Science Translational Medicine. 2 (62), 62ra93 (2010).
- Babina, I. S., Turner, N. C. Advances and challenges in targeting FGFR signaling in cancer. Nature Reviews Cancer. 17 (5), 318-322 (2017).
- Katoh, M. Fibroblast growth factor receptors as treatment targets in clinical oncology. Nature Reviews Clinical Oncology. 16 (2), 105-122 (2019).
- Soria, J. C., et al. Phase I/IIa study evaluating the safety, efficacy, pharmacokinetics, and pharmacodynamics of lucitanib in advanced solid tumors. Annals of Oncology. 25 (11), 2244-2251 (2014).
- French, D. M., et al. Targeting FGFR4 inhibits hepatocellular carcinoma in preclinical mouse models. PLoS ONE. 7 (5), e36713 (2012).
- Martinez-Torrecuadrada, J., et al. Targeting the extracellular domain of fibroblast growth factor receptor 3 with human single-chain Fv antibodies inhibits bladder carcinoma cell line proliferation. Clinical Cancer Research. 11 (17), 6280-6290 (2005).
- Palamakumbura, A. H., et al. Lysyl oxidase propeptide inhibits prostate cancer cell growth by mechanisms that target FGF-2-cell binding and signaling. Oncogene. 28 (38), 3390-3400 (2009).
- Ladner, R. C., Sato, A. K., Gorzelany, J., de Souza, M. Phage display-derived peptides as therapeutic alternatives to antibodies. Drug Discovery Today. 9 (12), 525-529 (2004).
- Wu, C. H., Liu, I. J., Lu, R. M., Wu, H. C. Advancement and applications of peptide phage display technology in biomedical science. Journal of Biomedical Science. 23, 8 (2016).
- Kay, B. K., Kasanov, J., Yamabhai, M. Screening phage-displayed combinatorial peptide libraries. Methods. 24 (3), 240-246 (2001).
- Rodi, D. J., Makowski, L. Phage-display technology – Finding a needle in a vast molecular haystack. Current Opinion in Biotechnology. 10, 87-93 (1999).
- Sidhu, S. S. Engineering M13 for phage display. Biomolecular Engineering. 18, 57-63 (2002).
- Hamzeh-Mivehroud, M., Mahmoudpour, A., Dastmalchi, S. Identification of new peptide ligands for epidermal growth factor receptor using phage display and computationally modeling their mode of binding. Chemical Biology & Drug Design. 79 (3), 246-259 (2012).
- Askoxylakis, V., et al. Peptide-based targeting of the platelet-derived growth factor receptor beta. Molecular Imaging and Biology. 15 (2), 212-221 (2013).
- Chen, Y., et al. Transdermal protein delivery by a coadministered peptide identified via phage display. Nature Biotechnology. 24 (4), 455-460 (2006).
- Azzazy, H. M., Highsmith, W. E. Phage display technology: clinical applications and recent innovations. Clinical Biochemistry. 35, 425-445 (2002).
- Pasqualini, R., Ruoslahti, E. Organ targeting In vivo using phage display peptide libraries. Nature. 380 (6572), 364-366 (1996).
- Liu, R., Li, X., Xiao, W., Lam, K. S. Tumor-targeting peptides from combinatorial libraries. Advanced Drug Delivery Reviews. 110-111, 13-37 (2017).
- Binetruy-Tournaire, R., et al. Identification of a peptide blocking vascular endothelial growth factor (VEGF)-mediated angiogenesis. The EMBO Journal. 19, 1525-1533 (2000).
- Peng, Y., Zhang, Y., Mitchell, W. J., Zhang, G. Development of a Lipopolysaccharide-Targeted Peptide Mimic Vaccine against Q Fever. Journal of Immunology. 189, 4909-4920 (2012).
- Lamichhane, T. N., Abeydeera, N. D., Duc, A. C., Cunningham, P. R., Chow, C. S. Selection of Peptides Targeting Helix 31 of Bacterial 16S Ribosomal RNA by Screening M13 Phage-Display Libraries. Molecules. 16, 1211-1239 (2011).
- Sahin, D., Taflan, S. O., Yartas, G., Ashktorab, H., Smoot, D. T. Screening and Identification of Peptides Specifically Targeted to Gastric Cancer Cells from a Phage Display Peptide Library. Asian Pacific. Journal of Cancer Prevention. 19 (4), 927-932 (2018).
- Kelly, K. A., et al. Targeted nanoparticles for imaging incipient pancreatic ductal adenocarcinoma. PLOS Medicine. 5 (4), e85 (2008).
- Sugihara, K., et al. Development of pro-apoptotic peptides as potential therapy for peritoneal endometriosis. Nature Communications. 5, 4478 (2014).
- Rafii, S., Avecilla, S. T., Jin, D. K. Tumor vasculature address book: Identification of stage-specific tumor vessel zip codes by phage display. Cancer Cell. 4, 331-333 (2003).
- Arap, W., Pasqualini, R., Ruoslahti, E. Cancer Treatment by Targeted Drug Delivery to Tumor Vasculature in a Mouse Model. Science. 279, 377-390 (1997).

