Reconstruction of Single-Cell Innate Fluorescence Signatures by Confocal Microscopy
Summary
Here, a protocol is presented for optically extracting and cataloging innate cellular fluorescence signatures (i.e., cellular autofluorescence) from every individual live cell distributed in a three-dimensional space. This method is suitable for studying the innate fluorescence signature of diverse biological systems at a single-cell resolution, including cells from bacteria, fungi, yeasts, plants, and animals.
Abstract
Described here is confocal reflection microscopy-assisted single-cell innate fluorescence analysis (CRIF), a minimally invasive method for reconstructing the innate cellular fluorescence signature from each individual live cell in a population distributed in a three-dimensional (3D) space. The innate fluorescence signature of a cell is a collection of fluorescence signals emitted by various biomolecules within the cell. Previous studies established that innate fluorescence signatures reflect various cellular properties and differences in physiological status and are a rich source of information for cell characterization and identification. Innate fluorescence signatures have been traditionally analyzed at the population level, necessitating a clonal culture, but not at the single-cell level. CRIF is particularly suitable for studies that require 3D resolution and/or selective extraction of fluorescence signals from individual cells. Because the fluorescence signature is an innate property of a cell, CRIF is also suitable for tag-free prediction of the type and/or physiological status of intact and single cells. This method may be a powerful tool for streamlined cell analysis, where the phenotype of each single cell in a heterogenous population can be directly assessed by its autofluorescence signature under a microscope without cell tagging.
Introduction
Diverse biomolecules within a cell1 emit autofluorescence signals, and the innate fluorescence signature of a cell consists of the assembly of these signals. This signature fluorescence reflects various cellular properties and also differences in physiological status. Analysis of innate fluorescence is minimally invasive and can complement traditional, more invasive microbiological probes that leave a range of traces from mild metabolic modification to complete cell destruction. While traditional techniques such as DNA or cell content extraction2,3, fluorescent in situ hybridization4, and the introduction of fluorescent reporter genes to the genome are effective in determining cell type or physiological status, they commonly require either manipulation of the cells or invasive tagging.
Studies of the innate fluorescence of various live and intact microbial colonies, including bulk microbial culture suspensions5,6, active sludges7, mammalian tissues8,9, and mammalian cells1,10, have shown that innate fluorescence analysis facilitates tag-free analysis of cell types and physiological status. Innate fluorescence signatures have been traditionally analyzed at the population level and not at the single-cell level, and thus necessitate a clonal culture. In contrast, the confocal reflection microscopy-assisted single-cell innate fluorescence analysis (CRIF) technique11 described here reconstructs and catalogues the innate cellular fluorescence signature of each individual live microbial cell. Moreover, CRIF can systematically collate the innate fluorescence signature of a single microbial cell within a population that is distributed in a three-dimensional (3D) space.
Protocol
1. Preparation of the sample
- Place a 1 mm thick silicone gasket with wells on a glass slide.
- Place a 1 mm thick 0.8% (w/v) agarose slab in the well of the silicone gasket.
- Dilute the cell density of an arbitrary microbial cell culture to an optical density at 600 nm (OD660) = 1.0.
- Place a 5 µL aliquot of cell suspension on the agarose slab.
- Cover gently with a glass coverslip.
2. Setup of a microscope
NOTE: The CRIF technique combines confocal reflection microscopy (CRM) and multichannel confocal microspectroscopy. CRM serves as the source of information for cellular morphology and spatial localization, which is independent from cellular innate fluorescence. Multichannel confocal microspectroscopy provides the spectral information of cellular innate fluorescence. In the following protocol, any image acquired with CRM or confocal fluorescence microspectroscopy is referred to as a CRM image or multichannel confocal microspectroscopy image, respectively.
- Connect a confocal microscope with descanned spectral channels to a photomultiplier tube (PMT) or GaAsP detector.
NOTE: These setups are available from several manufacturers. - Equip the microscope with a high numerical aperture (NA) objective with adequate magnification.
NOTE: A 63x objective with NA > 1.4 is recommended for analyzing bacterial cells. - Equip the microscope with a half-reflection mirror (e.g., NT 80/20) to accommodate CRM, which relies on the cellular scatter of incident light to visualize cell morphology.
- For multichannel confocal microspectroscopy, equip the microscope with dichroic mirrors. For example, use MBS InVis405, MBS458, MBS488, MBS458/514, MBS488/543, or MBS 488/543/633 beam splitters for 405, 458, 488, 514, 543, or 633 nm excitation, respectively.
- Adjust the illumination intensity for each excitation wavelength using a laser power meter. Keep the output under the microscope constant through excitation wavelengths (e.g., use 50 µW with the 63x objective).
3. Image acquisition
- Set the pinhole size to 1 AU using the microscope software.
- Set the pixel dwell time (i.e., scanning speed) for each excitation wavelength.
NOTE: An excessively long pixel dwell time can damage cells. Avoid excessively long pixel dwell time to minimize photodamage to the cells. For bacterial samples, a pixel dwell time <55.6 µs/µm2 (when the irradiance output under the microscope is ~17 µW/cm2) is usually suitable to avoid growth inhibition. This parameter may vary depending on the organisms and experimental setups. - Set the scanning resolution. For small cells such as bacteria, use a scanning area of 1,024 x 1,024.
- Set the Z-scanning range so that the region of interest is covered.
NOTE: For bacterial and yeast cell samples distributed on an agarose slab, a Z-scanning range of ~15 µm is usually sufficient. - Set the descanned detector to capture the visible wavelength range (e.g., 416−691 nm). Use a spectral window of 8−10 nm.
- Acquire multichannel confocal microspectroscopy images in a sequence from longest to shortest excitation wavelengths to create Z-stacks of fluorescence images.
- Acquire CRM images.
- Save the acquired images as 16-bit tiff files into a directory. Name the files using the naming convention XXXcYYzZZ.tif, where XXX is the excitation wavelength, YY is the detector channel number, ZZ is the Z-slice number, and “c” and “z” are prefixes for the detector number and the Z-slice number, respectively.
- For example, if a multichannel confocal microspectroscopy image is taken with an excitation wavelength of 405 nm, 1st detector channel of the detector array, and is the 5th slice of the Z-stack, name it “405c01z05.tif”. For CRM images, use the string “CRM” in place of XXX (e.g., “CRMc01z05.tif”).
NOTE: Decide whether 2D or 3D segmentation is most suitable for the image data. Use a 2D segmentation method in situations where small cells are constrained to a 2D plane (e.g., bacterial population adhering to a glass surface). Use the 3D segmentation method in situations where the cell population is distributed in a three-dimensional space (e.g., biofilms and tissue samples) or the cell sizes are larger than the thickness of the optical slice (e.g., yeast cells, mammalian cells). For 2D segmentation refer to section 4; for 3D segmentation, refer to section 5.
- For example, if a multichannel confocal microspectroscopy image is taken with an excitation wavelength of 405 nm, 1st detector channel of the detector array, and is the 5th slice of the Z-stack, name it “405c01z05.tif”. For CRM images, use the string “CRM” in place of XXX (e.g., “CRMc01z05.tif”).
4. 2D image analysis
- Equip a workstation with image analysis software (e.g., MATLAB).
- Perform cell segmentation and reconstruction of single-cell innate fluorescence signatures.
- Open the image analysis software.
- Double-click and open one of the provided scripts “Script2D.m”.
- Go to the Editor tab, then click Run. A folder selection window should appear.
- Select the directory created in step 3.8, then click Open to proceed. A dialogue box that prompts the input of the segmentation parameter will automatically appear.
NOTE: For test purposes select the provided dataset (“Sample_2D”). The sample dataset is provided as a compressed file and should be extracted in advance. - Input the segmentation parameters: Threshold of Image Binarization (0−1) for Image Binarization = 0.45, Upper Threshold For A Cell Region (in pixels) = 200, Lower Threshold for a Cell Region (in pixels) = 10, and The Number of Detectors = 32. Click OK to proceed.
NOTE: These parameters may require adjustment depending on the image quality. - A new image window presenting a CRM image should appear. Select an arbitrary background region (i.e., area where cells are absent) to use for background subtraction. Draw a rectangle within the CRM image by mouse dragging. Double-click within the selected region to confirm the selection.
- Find a new directory named Signature in the same directory selected in step 4.2.4.
NOTE: The provided code automatically creates this directory. The “Signature” directory stores the innate fluorescence signature of each microbial cell within a population as .png files that are serially numbered after a common prefix “Signature”.
5. 3D image analysis
- Equip a workstation with the image analysis software (Table of Materials).
- Perform cell segmentation and reconstruction of single-cell innate fluorescence signatures.
- Open the image analysis software.
- Double-click and open the provided script “Script3D.m”.
- Go to the Editor tab, then click Run. A folder selection window should appear.
- Select the directory created in step 3.8, then click Open to proceed. A dialogue box that prompts the input of the segmentation parameters will automatically appear.
NOTE: For test purposes select the provided dataset (“Sample_3D”). The sample dataset is provided as a compressed file and should be extracted in advance. - Input the segmentation parameters: Threshold of Image Binarization (0–1) for image binarization = 0.01, Upper Threshold for a Cell Volume (in pixels) = 1,000, Lower Threshold for a Cell Region (in pixels) = 20, X Pixel Size [μm/pixel] = 0.26, Y Pixel Size [μm/pixel] = 0.26, Z pixel Size [μm/pixel] = 0.42, and The Number of Detectors = 32. Click OK to proceed; a dialogue box that prompts the input for the number of excitation wavelengths will appear.
NOTE: These parameters may require adjustment depending on the image quality. - Input the number of wavelengths used for image acquisition (e.g., if 405, 488, 561, 630 are used, enter 4 in the dialogue box). Click OK.
- Enter the excitation wavelengths in a sequence from shortest (i.e., box name: Excitation No. 1) to longest wavelength into the dialogue boxes. Click OK to proceed; a new image window that presents a CRM image should pop up.
- Select the arbitrary background region (i.e., area where cells are absent) to be used for background subtraction. Draw a rectangle within the CRM image by mouse dragging. Double-click within the selected region to confirm the selection.
- Find the directory named Signature in the directory selected in step 5.2.4.
NOTE: The provided code automatically creates this directory. The “Signature” directory stores the innate fluorescence signature of each single microbial cell within a population as .png files that are serially numbered after a common prefix “Signature”.
6. Statistical analysis
NOTE: Perform dimensional reduction techniques (e.g., principal component analysis [PCA]) to visualize the distribution of hyperspectrums of the cell populations. The provided script (PCA.py) executes PCA for two cell populations (i.e., two classes).
- Equip a workstation with the programming language and accompanying libraries and modules (Table of Materials).
- Create an empty directory in the C drive (or equivalent) and name the directory “Parent_directory” (i.e., C:/ Parent_ directory).
- Store the fluorescence signatures (e.g., the .png files generated in step 4.2.7) of each of the two cell populations into two separate directories.
NOTE: The two directories should be both located in the “Parent_directory”.
C:/Parent_directory/
 putidaKT2440/
putidaKT2440/
 Signature01.png
Signature01.png
 Signature02.png
Signature02.png
 :
:
 putidaKT2442/
putidaKT2442/
 Signature01.png
Signature01.png
 Signature02.png
Signature02.png
 :
: - Download PCA.py into the “Parent_directory”.
- Open the command line interface of the workstation.
- Type “python C:/Parent_directory/PCA.py” in the command line interface.
- Select the “Parent_directory” after the message ‘Select target directory’ is displayed.
- In the “C:/Parent_directory”, find “PCA.png”, which contains a resulting PCA plot.
Representative Results
Figure 1A shows the typical single-cell fluorescence signature of a bacterial cell presented as a traditional spectrum plot (top) and as a heatmap (middle). Figure 1B shows the result of an accurate 2D cell segmentation superimposed over the original CRM image of a population of soil bacteria (Pseudomonas putida KT2440)12. The resulting innate fluorescence signatures for the population are presented as a heatmap in Figure 1D. Note that intrapopulation variability was relatively minor following successful cell segmentation. An example of inaccurate cell segmentation is shown in Figure 1C, which is superimposed onto the same population of P. putida as shown in Figure 1B. The impact of inaccurate cell segmentation on the innate fluorescence signatures of the population is readily apparent from the considerable number of outliers (Figure 1E, red triangles). Inaccurate cell segmentation resulted in a looser cluster after PCA compared to the tight cluster obtained following accurate cell segmentation (Figure 1F).
Typically, despite intraspecies variability, innate fluorescence signatures of different cell types form distinct clusters. Figure 2C presents the result of PCA analyses for a taxonomically close pair of strains (P. putida strain KT2440 and strain KT244213). Despite the minor variability observed within each separate population (Figure 2A,B), each population formed a distinct cluster on the PCA analysis plot (Figure 2C).
Figure 3A shows the result of accurate 3D cell segmentation superimposed over the original CRM image of a population of budding yeast Saccharomyces cerevisiae YM427114. Figure 3B shows the resulting innate fluorescence signatures for the population.
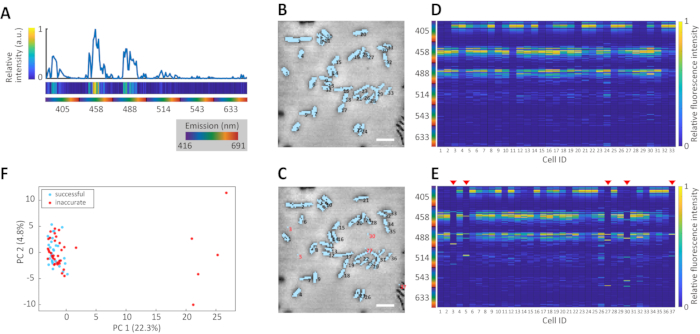
Figure 1: Accuracy of segmentation and apparent intraspecies variability. (A) Typical single-cell innate fluorescence signature of a bacterial cell (P. putida KT2440), presented as a spectrum plot (top) and a heat map (middle). The emission wavelength is indicated as a rainbow color map (bottom). The Y axis of the spectrum plot and the color of the heat map both indicate the relative fluorescence intensities. The numbers in the bottom indicate the excitation wavelength. Visual representation of accurate (B) and inaccurate (C) cell region recognition by the image analysis algorithm. Light blue shading indicates cell regions detected for a population of soil bacterium P. putida KT 2440. Blobs with red numbering in panel C are examples of false detection, where the algorithm classified non-cell regions (i.e., background noise) as cell regions due to inappropriate binarization threshold settings. Scale bars = 1 µm throughout. Panels D and E depict the innate fluorescence signatures for the same population, generated with accurate and inaccurate cell segmentation, respectively. Each column shows a reconstructed single-cell hyperspectrum presented as a 1 x 192 matrix, where the yellow-to-blue color map indicates relative fluorescence intensity. (F) The variance of innate fluorescence signatures in accurate (light blue) and inaccurate (red) segmentations visualized using PCA. Each axis label denotes the component number and its cumulative percentage contribution to the PCA analysis. Please click here to view a larger version of this figure.
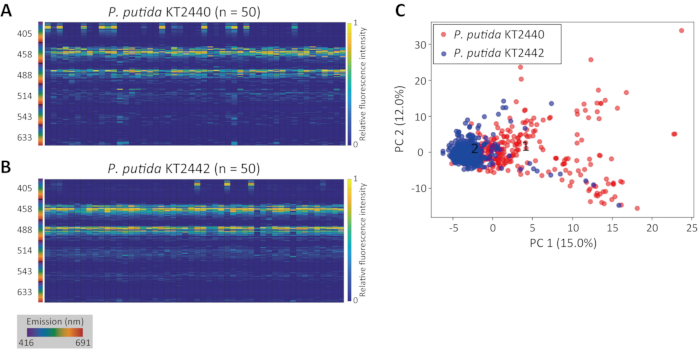
Figure 2: Intra- and interspecies variability of innate fluorescence signatures. Innate fluorescence signatures presented as heat maps for a population of P. putida KT2440 (A) and P. putida KT2442 (B). (C) Projection of the two innate fluorescence signature pairs visualized using PCA, corresponding to P. putida KT2440 (red, n = 288) and P. putida KT2442 (blue, n = 373). X- and Y-axes represent PC1 and PC2, respectively. The inset number indicates the center of each cluster (1: P. putida KT2440, 2: P. putida KT2442). Please click here to view a larger version of this figure.
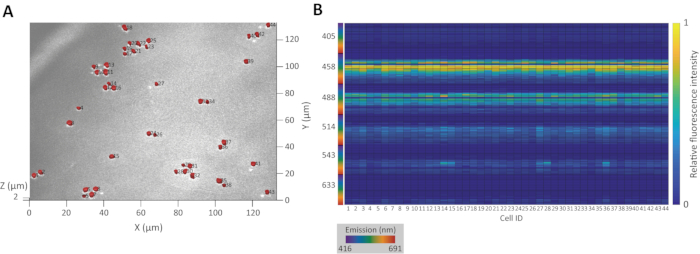
Figure 3: Example of 3D segmentation and innate fluorescence signature extraction. (A) 3D projection of regions recognized as cell populations of budding yeast S. cerevisiae YM4271. The inset numbers are the identification numbers. (B) Heat map of innate fluorescence signatures extracted from the 3D regions. The X-axis number corresponds to the identification number. Please click here to view a larger version of this figure.
Discussion
There are two critical points in this method that need to be closely followed to obtain reproducible results: 1) keep the laser power output under the microscope objective consistent through excitation wavelengths and experiments, and 2) perform accurate cell segmentation.
The first point is particularly important when comparing the innate fluorescence signature among different experiments. Avoid simply applying the same “percent output” settings to the excitation wavelengths (i.e., using 5% power output for all of 405, 488, 514, and 530 nm laser lines), because the maximum power output can differ up to an order of magnitude among laser lines. In addition, the objective and the internal optics usually have uneven optical absorption characteristics, which also need to be accounted for. For these reasons, actually measuring the output under the objective using a laser power meter (Table of Materials) is recommended. Taking this measurement every few experiments, or periodically, is also recommended, because the output from laser lines can significantly decay over a few years.
The second point, accurate cell segmentation, becomes particularly important in situations where intra- or interspecies variability is a concern. Inaccurate cell segmentation (Figure 1D) results in outlier data points (Figure 1E, indicated by red triangles) and greater apparent intraspecies variability. The segmentation parameters should be determined carefully, checking the cell segmentation accuracy by overlaying the segmentation results onto the original CRM image, before proceeding to any further analysis or training machine learning models. If working with a poor-quality image, additional image processing could be required to achieve accurate segmentation. Using either a ‘Gaussian filter’ or ‘median filter’ can remove grainy artifacts, and bilateral filter processing counters any striped background.
With poor quality optics (e.g., a non-confocal grade objective) or insufficient detector sensitivity, dealing with the detection of faint innate fluorescence signatures might be challenging. Note that using the setup described here and previously11, we did not encounter any type of cell sample whose innate fluorescence was too faint to reconstruct an innate fluorescence signature. Although higher emission output for excitation and longer pixel dwell time can be used to obtain stronger emission signals, this may cause damage to the cell samples. A potential solution for cases where faint innate fluorescence is an issue is to reduce the background signal intensity, for example by suspending cells in buffer or synthetic media instead of broth. Types of cell containers, such as glass bottom dishes and microfluidic devices, are compatible and appropriate for this technique, although an agarose slab was used to retain cells in position here and in a previous study15.
The confocal microscope method offers spatial resolution and in situ analysis for adherent cell populations, biofilms, and tissue samples. A potential trade-off of this methodology is the maximum throughput compared to a flow cytometer, for example. A complete set of confocal scans collecting a few hundred to a thousand innate fluorescence signatures with a single set of scans can take up to a few minutes. A flow cytometer is less sensitive due to its optical design but can potentially achieve a throughput several orders of magnitude greater.
To rigorously identify a compound that is responsible for a specific peak in an innate fluorescent signature is typically challenging, due to its complicated nature. However, it has been suggested that a number of biologically relevant molecules, such as vitamins (e.g., flavin adenine dinucleotide [FAD]), coenzymes (e.g., nicotinamide adenine dinucleotide [NADH]), and lipofuscin pigments could be major fluorophores in cells1,16. For example, FAD has an excitation maximum between 350–450 nm and an emission peak at approximately 525 nm16. Free NADH has an excitation maximum and emission peak at 340 nm and 460 nm, respectively, although the emission spectra of protein-bound NADH differs16,17,18. Lipopigments that are known to associate with stressed or aged cells have a broad excitation range (350–500 nm) and emission range (450–600 nm)16,18.
The use of CRM offers an independent source of information to identify cell contours and therefore provides another significant advantage of CRIF: the selective extraction of fluorescence signals from individual cells distributed in a 3D space. This represents a step forward from the traditional analysis of the fluorescence characteristics of a whole microbial colony or a bulk culture suspension, which is an averaged mixture of signals from a vast number of cells contaminated with non-cell signals from medium components, secreted metabolites, and extracellular matrixes. An analysis of single-cell innate fluorescence signatures allows predictive characterization of the physiological status of intact cells, providing a potential solution for resolving temporal development of a cell's distribution and physiological state.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This study was supported in part by a grant-in-aid for scientific research from the Ministry of Education, Culture, Sports, and Technology of Japan (18K04843) to Y. Yawata, the JST ERATO (JPMJER1502) to N. Nomura.
Materials
| Agarose | Wako Chemicals | 312-01193 | |
| Beam splitters | Carl Zeiss, Nikon | MBSInVis405, MBS458, MBS488, MBS458/514, MBS488/543, or MBS 488/543/633 beam splitters (Carl Zeiss) | |
| Confocal microscope | Carl Zeiss, Nikon | Model LSM 880 (Carl Zeiss), Model A1R (Nikon) | |
| Cover slips | Matsunami Glass | C024601 | |
| Glass slides | Matsunami Glass | S011120 | |
| Half-reflection mirror | Carl Zeiss, Nikon | NT80/20 | |
| Laser power meter | Thorlabs | PM400 (power meter console) and S175C (sensor) | |
| LB Broth | Nacalai tesque | 20066-95 | For bacteria culture |
| Image analysis software | The MathWorks | MATLAB version 2019a or later, Image Processing Toolbox is needed | |
| Microscope objective | Carl Zeiss, Nikon | 440762-9904 | e.g. 63x plan Apochomat NA = 1.4 (Carl Zeiss) |
| Microscope software | Carl Zeiss, Nikon | ZEN (Carl Zeiss),NIS-elements (Nikon) | |
| PBS(-) | Wako Chemicals | 166-23555 | |
| Programming language | Python and libraries, modules (numpy, scikit-learn, scikit-image, os, glob, matplotlib, tkinter) are rquired to run the supplied PCA script. | ||
| Silicone gasket | ThermoFisher Scientific | P24744 | |
| Workstation | A high-performance workstation with discrete GPUs is recommended. | ||
| Yeast extract-peptone-dextrose (YPD) agar medium | Sigma-Aldrich | Y1500-250G | For yeast culture |
| YPD medium | Sigma-Aldrich | Y1375-250G |
Riferimenti
- Monici, M. Cell and tissue autofluorescence research and diagnostic applications. Biotechnology Annual Review. 11, 227-256 (2005).
- Tang, J. Microbial metabolomics. Current Genomics. 12, 391-403 (2011).
- Woo, P. C., Lau, S. K., Teng, J. L., Tse, H., Yuen, K. Y. Then and now: use of 16S rDNA gene sequencing for bacterial identification and discovery of novel bacteria in clinical microbiology laboratories. Clinical Microbiology and Infection. 14, 908-934 (2008).
- Amman, R., Fuchs, B. M. Single-cell identification in microbial communities by improved fluorescence in situ hybridization techniques. Nature Reviews Microbiology. 6, 339-348 (2008).
- Giana, H. E., Silveira, L., Zângaro, R. A., Pacheco, M. T. T. Rapid identification of bacterial species by fluorescence spectroscopy and classification through principal components analysis. Journal of Fluorescence. 13 (6), 489-493 (2003).
- Leblanc, L., Dufour, E. Monitoring the identity of bacteria using their intrinsic fluorescence. FEMS Microbiology Letters. 211 (2), 147-153 (2002).
- Hou, X., Liu, S., Feng, Y. The autofluorescence characteristics of bacterial intracellular and extracellular substances during the operation of anammox reactor. Scientific Reports. 7, 39289 (2017).
- Ramanujam, N., et al. In vivo diagnosis of cervical intraepithelial neoplasia using 337-nm-excited laser-induced fluorescence. Proceeding National Academy of Sciences of the United States of America. 91 (21), 10193-10197 (1994).
- Zhang, J. C., et al. Innate cellular fluorescence reflects alterations in cellular proliferation. Lasers in Surgery and Medicine. 20 (3), 319-331 (1997).
- Gosnell, M. E., et al. Quantitative non-invasive cell characterisation and discrimination based on multispectral autofluorescence features. Scientific Reports. 6, 23453 (2016).
- Yawata, Y., et al. Intra- and interspecies variability of single-cell innate fluorescence signature of microbial cell. Applied and Environmental Microbiology. 85, 00608-00619 (2019).
- Nelson, K. E., et al. Complete genome sequence and comparative analysis of the metabolically versatile Pseudomonas putida KT2440. Environmental Microbiology. 4 (12), 799-808 (2002).
- Bagdasarian, M., et al. Specific-purpose plasmid cloning vectors. II. Broad host range, high copy number, RSF1010-derived vectors, and a host-vector system for gene cloning in Pseudomonas. Gene. 16 (1-3), 237-247 (1981).
- Luo, Y., Vijaychander, S., Stile, J., Zhu, L. Cloning and analysis of DNA-binding proteins by yeast one-hybrid and one-two-hybrid systems. Biotechniques. 20 (4), 564-568 (1996).
- Yawata, Y., et al. Monitoring biofilm development in a microfluidic device using modified confocal reflection microscopy. Journal of Bioscience and Bioengineering. 110 (3), 377-380 (2010).
- Lakowicz, J. R. . Principles of Fluorescence Spectroscopy. Third edition. , (2006).
- Blacker, T. S., Duchen, M. R. Investigating mitochondrial redox state using NADH and NADPH autofluorescence. Free Radical Biology & Medicine. 100, 53-65 (2016).
- Croce, A. C., Bottiroli, G. Autofluorescence spectroscopy and imaging: A tool for biomedical research and diagnosis. European Journal of Histochemistry. 58 (4), 2461 (2014).

