High-throughput Screening of Chemical Compounds to Elucidate Their Effects on Bacterial Persistence
Summary
In this method paper, we present a high-throughput screening strategy to identify chemical compounds, such as osmolytes, that have a significant impact on bacterial persistence.
Abstract
Bacterial persisters are defined as a small subpopulation of phenotypic variants with the capability of tolerating high concentrations of antibiotics. They are an important health concern as they have been associated with recurrent chronic infections. Although stochastic and deterministic dynamics of stress-related mechanisms are known to play a significant role in persistence, mechanisms underlying the phenotypic switch to/from the persistence state are not completely understood. While persistence factors triggered by environmental signals (e.g., depletion of carbon, nitrogen and oxygen sources) have been extensively studied, the impacts of osmolytes on persistence are yet to be determined. Using microarrays (i.e., 96 well plates containing various chemicals), we have designed an approach to elucidate the effects of various osmolytes on Escherichia coli persistence in a high throughput manner. This approach is transformative as it can be readily adapted for other screening arrays, such as drug panels and gene knockout libraries.
Introduction
Bacterial cultures contain a small subpopulation of persister cells that are temporarily tolerant to unusually high levels of antibiotics. Persister cells are genetically identical to their antibiotic-sensitive kins, and their survival has been attributed to transient growth inhibition1. Persister cells were first discovered by Gladys Hobby2 but the term was first used by Joseph Bigger when he identified them in penicillin-treated Staphylococcus pyogenes cultures3. A seminal study published by Balaban et al.4 discovered two persister types: type I variants that are primarily formed by passage through the stationary phase, and type II variants that are continuously generated during the exponential growth. Persisters are detected by clonogenic survival assays, in which culture samples are taken at various intervals during antibiotic treatments, washed, and plated on a typical growth medium to count the surviving cells that can colonize in the absence of antibiotics. The existence of persisters in a cell culture is assessed by a biphasic kill curve4,5 where the initial exponential decay indicates the death of antibiotic-sensitive cells. However, the killing trend decreases over time, eventually leading to a plateau region which represents the surviving persister cells.
Persister cells have been associated with various diseases such as tuberculosis6, cystic fibrosis7, candidiasis8 and urinary tract infections9. Almost all microorganisms tested so far were found to generate persister phenotypes, including highly pathogenic Mycobacterium tuberculosis6, Staphylococcus aureus10, Pseudomonas aeruginosa7 and Candida albicans8. Recent studies also provide evidence of the rise of multidrug-resistant mutants from persister subpopulations11,12. Substantial efforts in this field have revealed that persistence mechanisms are highly complex and diverse; both stochastic and deterministic factors associated with the SOS response13,14, reactive oxygen species (ROS)15, toxin/antitoxin (TA) systems16, autophagy or self-digestion17 and ppGpp-related stringent response18 are known to facilitate persister formation.
Despite significant progress in understanding the persistence phenotype, the effects of osmolytes on bacterial persistence have not been fully understood. Since the maintenance of optimal osmotic pressure is a necessity for cells’ growth, proper functioning and survival, an in-depth study of osmolytes could lead to potential targets for anti-persister strategies. Although laborious, high-throughput screening is a very effective approach for identifying metabolites and other chemicals that play a crucial role in the persistence phenotype19,20. In this work, we will discuss our published method19, where we have used microarrays, i.e., 96 well plates containing various osmolytes (e.g., sodium chloride, urea, sodium nitrite, sodium nitrate, potassium chloride), to identify osmolytes that significantly influence E. coli persistence.
Protocol
1. Preparation of growth medium, ofloxacin solution and E. coli cell stocks
- Regular Luria-Bertani (LB) medium: Add 10 g/L of tryptone, 10 g/L of sodium chloride (NaCl) and 5 g/L of yeast extract in deionized (DI) water. Sterilize the medium by autoclaving.
- LB agar plates: Add 10 g/L of tryptone, 10 g/L of NaCl, 5 g/L of yeast extract and 15 g/L agar in DI water and sterilize the medium by autoclaving. At the desired temperature (~55 °C), pour ~30 mL of agar medium into square plates (10 x 10 cm). Dry the plates and store at 4 °C.
- Modified LB medium: Add 10 g/L of tryptone and 5 g/L of yeast extract in DI water. Sterilize the medium by autoclaving.
- Modified LB medium including an osmolyte: Mix 2x modified LB medium with a 2x osmolyte solution in equal volumes. To prepare 2x modified LB medium, add 20 g/L of tryptone and 10 g/L of yeast extract in DI water and sterilize by autoclaving. To prepare the 2x osmolyte solution, dissolve the osmolyte of interest (2x amount) in DI water, and then filter sterilize the solution.
NOTE: The osmolyte of interest and its concentrations can be determined from the Phenotype Microarray screening assays (see Protocol 4). However, the tested osmolyte and the associated 2x concentration can be adjusted depending on the nature of the research. For example, to study the impact of 1 mM sodium chloride, a 2x osmolyte solution would be a 2 mM sodium chloride solution. - Ofloxacin stock solution (5 mg/mL): Add 5 mg of ofloxacin (OFX) salt in 1 mL of DI water. Add 10 µL of 10 M sodium hydroxide to increase the solubility of OFX in water, and then filter sterilize the solution. Prepare aliquots and store at -20 °C.
NOTE: Ofloxacin is a quinolone antibiotic that has been widely used for both growing and non-growing bacterial cells14,21. The minimum inhibitory concentration (MIC) of OFX for E. coli MG1655 cells is within the range of 0.039–0.078 μg/mL19,20. Also note that other antibiotics such as ampicillin and kanamycin are commonly used in persister research. The choice of antibiotics is dependent on the nature of the study. - E. coli MG1655 cell stocks: Inoculate 2 mL of regular LB medium with a single colony in a 14 mL test tube (snap-capped) and culture the cells in an orbital shaker at 250 rpm and 37 °C. When the cells reach the stationary phase, mix 500 μL of the cell culture with 500 μL of 50% glycerol (sterile) in a cryogenic vial and store at -80 °C.
2. Propagation of cells to eliminate pre-existing persisters
- To prepare an overnight culture, scrape a small amount of cells from a frozen cell stock with a sterile pipette tip (do not thaw the glycerol cell stock), and inoculate the cells in 2 mL of modified LB medium in a 14 mL snap-capped test tube. Culture the cells in an orbital shaker at 250 rpm and 37 °C for 12 h.
- First propagation
- After 12 h, transfer 250 µL of overnight culture to 25 mL of fresh modified LB medium in a 250 mL baffled flask covered with a sterile aluminum foil.
- Grow the cell culture in an orbital shaker at 250 rpm and 37 °C until cells reach the mid-exponential phase (OD600 = 0.5).
- Measure the optical density at 600 nm (OD600) using a microplate reader every 30 min.
- Second propagation
- At OD600 = 0.5, dilute 250 µL of cell culture from the first flask into 25 mL of fresh modified LB medium in a 250 mL baffled flask.
- Grow the second cell culture in an orbital shaker at 250 rpm and 37 °C until OD600=0.5.
- Measure the optical density every 30 min.
NOTE: An overnight culture may have a significant amount of persister cells4,5,22. The dilution/growth cycle method5 described above can be used to eliminate these pre-existing persisters before transferring the cells to microarrays. Their elimination can be validated by quantifying persister levels of overnight cultures with the assay described in Protocol 3. This method has been already validated in a previous study19.
3. Validating the elimination of pre-existing persister cells
- After the second propagation (see step 2.3), dilute 250 µL of the cell culture (OD600 = 0.5) in 25 mL of fresh modified LB medium in a 250 mL baffled flask.
NOTE: For controls, dilute 250 µL of overnight culture (see step 2.1) in 25 mL of fresh modified LB medium in a 250 mL baffled flask. Before transferring the cells to the flask, the cell density of the overnight culture should be adjusted in fresh modified LB medium to obtain OD600 = 0.5. - Add 25 µL of OFX stock solution (5 mg/mL) into the cell suspension and shake the flask gently to make the assay culture homogeneous. The final concentration of OFX is 5 µg/mL in the assay culture.
- Incubate the assay culture in an orbital shaker at 250 rpm and 37 °C.
- At every hour during the treatment (including 0 h, the time point before adding OFX into the assay culture), transfer 1 mL of the assay culture from the flask to a 1.5 mL microcentrifuge tube.
- Centrifuge the assay culture in the microcentrifuge tube at 17,000 x g for 3 min.
- Carefully remove 950 µL of the supernatant without disturbing the cell pellet.
- Add 950 µL of phosphate-buffered saline (PBS) solution into the microcentrifuge tube.
- Repeat the washing steps 3.5-3.7 for 3x in total until the antibiotic concentration is below the minimum inhibitory concentration (MIC).
- After the final wash, resuspend the cell pellet in 100 µL of PBS solution, resulting in a 10x concentrated sample.
- Take 10 µL of the cell suspension and serially dilute six times in 90 µL of PBS solution using a 96 well round bottom plate.
- Spot 10 µL of diluted cell suspensions on antibiotic-free fresh agar plates. To increase the limit of detection, plate the remaining 90 µL of cell suspension on a fresh agar plate.
- Incubate the agar plates at 37 °C for 16 h, and then count the colony-forming units (CFUs). Account for the dilution rates while calculating the total number of CFUs in 1 mL of assay culture. Kill curves are generated by plotting the logarithmic CFU values with respect to the duration of antibiotic treatment.
NOTE: A 6 h OFX treatment is sufficient to obtain a biphasic kill curve for E. coli cells19,20. The persister level of propagated cells (as measured by CFU counts at 6 h) should be significantly less than that of the overnight culture. The procedures described in Protocol 2 and 3 can be performed with regular LB depending on the research design.
4. Microarray plate screenings
- Preparing microarray-cell cultures
- Transfer 250 µL of exponential-phase cells to 25 mL of fresh modified LB medium in a 50 mL centrifuge tube. Gently mix the cell suspension to make it homogeneous.
NOTE: The exponential-phase cells (OD600 = 0.5) are obtained from the second propagation step (see step 2.3). - Transfer the diluted cell suspension into a sterile 50 mL reservoir.
- Using a multichannel pipette, transfer 150 µL of the cell suspension to each well of a microarray, i.e., a 96 well plate containing various osmolytes.
NOTE: Wells that do not have osmolytes serve as controls. - Cover the microarray with a gas-permeable sealing membrane.
- Incubate the plate in an orbital shaker at 37 °C and 250 rpm for 24 h.
NOTE: In these experiments, commercially available plates were used, such as Phenotype Microarrays (PM-9 and PM-10) that include a wide range of osmolytes, pH buffers and other chemicals in a dried state at various concentrations. These microarrays are in half-area 96 well plate formats. Culture volumes should be adjusted depending on the type of plates being used. Microarrays can also be generated manually (see below).
- Transfer 250 µL of exponential-phase cells to 25 mL of fresh modified LB medium in a 50 mL centrifuge tube. Gently mix the cell suspension to make it homogeneous.
- Manual preparation of microarray plates
- Transfer 75 μL of 2x osmolyte solutions into the wells of a half-area 96 well plate.
- Transfer 500 µL of exponential-phase cells to 25 mL of 2x modified LB medium in a 50 mL centrifuge tube. Gently mix the cell suspension to make it homogeneous.
NOTE: The inoculation rate was adjusted to be consistent with the microarray screening protocol described in 4.1. - Add 75 μL of the cell suspension into each well of the half-area 96 well plate containing 2x osmolyte solutions.
- Incubate the plate in an orbital shaker at 37 °C and 250 rpm for 24 h.
- Preparing persister assay plates
- Prepare 25 mL of modified LB medium containing 5 µg/mL of OFX in a 50 mL centrifuge tube and transfer this medium to a sterile reservoir.
- Transfer 190 µL of modified LB medium with OFX from the reservoir into each well of a generic flat-bottom 96 well plate (persister-assay plate) using a multichannel pipette.
- Remove the microarray from the shaker (after culturing for 24 h) and transfer 10 µL of cell cultures from the microarray to the wells of the persister-assay plate, containing modified LB medium with OFX.
- Take 10 µL of cell suspensions from the persister-assay plate and serially dilute three times in 290 µL of PBS solution using a round-bottom 96 well plate and a multichannel pipette.
- Following the serial dilution, spot 10 µL of all serially diluted cell suspensions on antibiotic-free fresh agar plates using a multichannel pipette.
- Incubate the persister-assay plate (prepared in step 4.3.3) in an orbital shaker at 37 °C and 250 rpm for 6 h after covering the plate with a gas-permeable sealing membrane.
- After 6 h incubation in a shaker, take the persister-assay plate out and repeat the steps 4.3.4-4.3.5.
- Incubate the agar plates for 16 h at 37 °C, and then count CFUs. The CFU levels before and 6 h after the antibiotic treatment enable to calculate the persister fraction in each well. The CFU counts before the OFX treatment also help assess the effects of osmolytes and on E. coli viability.
5. Validating the identified conditions
- Transfer 250 µL of the exponential phase cells from Step 2.3 to 25 mL of fresh modified LB medium containing the osmolyte identified from the microarray screening (see Protocol 4).
- Incubate the flask in an orbital shaker at 250 rpm and 37 °C for 24 h.
- After 24 h, remove the flask from the shaker and transfer 250 µL of the cell culture to 25 mL of fresh modified LB medium in a 250 mL baffled flask.
- Add 25 µL of OFX stock solution (5 mg/mL) in the cell suspension and shake the flask gently to make the assay culture homogenous. Incubate the flask in a shaker at 37 °C and 250 rpm.
- At every hour during the treatment, transfer 1 mL of the assay culture from the flask to a 1.5 mL microcentrifuge tube.
- Centrifuge the assay culture in the microcentrifuge tube at 17,000 x g for 3 min.
- Remove 950 µL of supernatant and add 950 µL of PBS.
- Repeat the washing steps 5.6 and 5.7 for 3x.
- After the final wash, resuspend the cell pellet in 100 µL of PBS solution.
- Take 10 µL of the cell suspension and serially dilute 6x in 90 µL of PBS solution using a 96 well round bottom plate.
- Spot 10 µL of the diluted cell suspensions on an antibiotic-free fresh agar plate. To increase the limit of detection, plate the remaining 90 µL of cell suspension on a fresh agar plate.
- Incubate the agar plate at 37 °C for 16 h, and then count CFUs.
Representative Results
Figure 1 describes our experimental protocol. The dilution/growth cycle experiments (see Protocol 2) were adapted from a study conducted by Keren et al.5 to eliminate the persisters originating from the overnight cultures. Figure 2A is a representative image of agar plates used to determine CFU levels of cell cultures before and after OFX treatment. In these experiments, cells were cultured in modified LB medium with osmolytes in half-area 96 well plates as described in step 4.2. After incubating the plate in an orbital shaker for 24 h, the persister assay was performed using a generic flat bottom 96 well plate (see step 4.3). The osmolytes and the concentration being tested here were chosen based on our previous study19, where we performed steps 4.1 and 4.3 using the PM-9 plate that includes various osmolytes at different concentrations (sodium chloride, potassium chloride, sodium sulfate, ethylene glycol, sodium formate, urea, sodium lactate, sodium phosphate, sodium benzoate, ammonium sulfate, sodium nitrite and sodium nitrate). The first column in Figure 2A shows the CFU counts of the control group. The second column represents a condition where cells were cultured in 100 mM sodium nitrate; this condition was previously found to slightly increase the persister levels19. The third column represents a condition where cells were cultured in 60 mM sodium nitrite, and this condition was previously found to significantly decrease the persister levels compared to controls19. Figure 2B is a graphical representation of the CFU data obtained from the agar plates. Figure 2C shows the persister fractions of the cell cultures tested in 96 well plates. To calculate the fractions, persister counts were normalized to the cell counts obtained before the antibiotic treatments. Figure 3 shows the biphasic kill curves and the persister fractions, respectively, for the assay cultures performed in baffled flasks. In these experiments, cells were first cultured in 25 mL of modified LB medium with the indicated osmolytes in 250 mL baffled flasks for 24 h, and then the cells were transferred to persister-assay flasks for persister enumeration as described in Protocol 5.
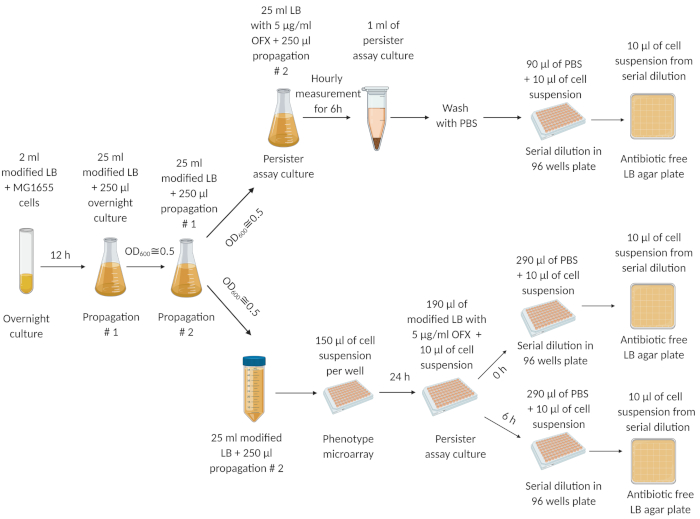
Figure 1: The experimental procedure. Cells from a frozen cell stock were grown overnight (12 h) in fresh LB medium. At 12 h, the overnight culture was diluted (1:100) in 25 mL of modified LB medium and grown until OD600=0.5. This propagation step was repeated twice. After the final propagation step, the exponential phase cells (at OD600=0.5) were diluted (1:100) in fresh modified LB medium in a 250 mL baffled flask and a 50 mL centrifuge tube, respectively. The cell suspension in the 250 mL baffled flask was treated with 5 μg/mL of OFX to quantify persisters. The cell suspension in the 50 mL centrifuge tube was transferred to microarrays and incubated for 24 h in an orbital shaker at 250 rpm and 37 °C. The cells from the microarrays were then transferred to persister-assay plates to quantify persisters. This figure was created using biorender.com. This figure has been modified from our previous publication19. Please click here to view a larger version of this figure.
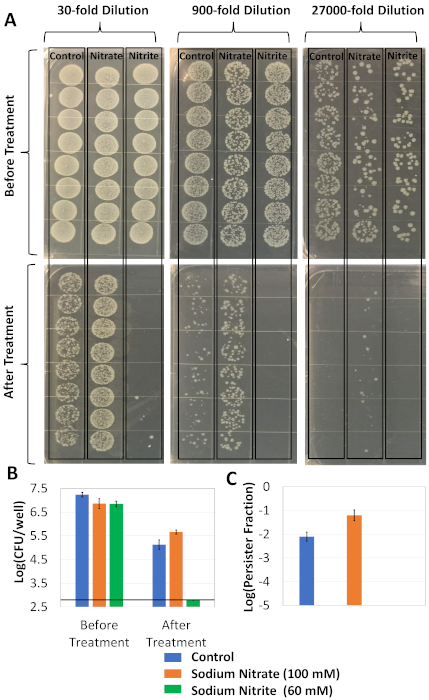
Figure 2: Microarray experiments. (A) Cells were first grown in modified LB medium with or without indicated osmolytes in a half-area 96 well plate for 24 h. The cells were then transferred to a generic flat-bottom 96 well plate and treated with 5 μg/mL of OFX for 6 h. Before and after 6 h treatment, cells were serially diluted in PBS, plated on agar plates and incubated at 37 °C for 16 h. Each condition has 8 technical replicates. (B) CFU measurements were performed to assess the effects of osmolytes on cell viability and persistence, respectively. The straight line indicates the limit of detection (600 CFUs). (C) The graph represents the persister fractions of the cell cultures, calculated by taking the ratio of the CFU counts after and before OFX treatment. The persister fraction of the cell culture that has 60 mM sodium nitrite was not calculated as its persister level is below the limit of detection. Each data point was denoted by mean value ± standard deviation, calculated from 8 technical replicates. Please click here to view a larger version of this figure.
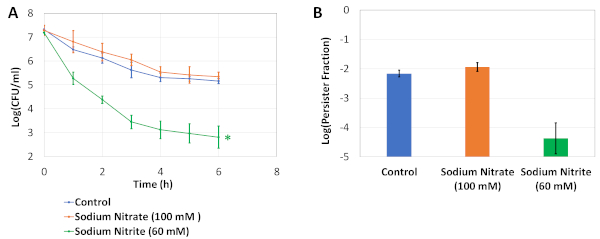
Figure 3: Validation experiments. (A) Exponential-phase cells were transferred to 25 mL of fresh modified LB medium with indicated osmolytes in 250 mL baffled flasks and cultured for 24 h. Then, the cells were diluted (1:100) in 250 mL baffled flasks containing 25 mL of modified LB medium and treated with 5 μg/mL OFX for 6 h. The CFU counts in the assay cultures were monitored hourly to generate biphasic kill curves. * indicates the condition that significantly affects the OFX persister levels compared to no-osmolyte controls (two-tailed unequal variance t-test, p<0.05). (B) The graph represents the persister fractions of the cultures. Each data point was denoted by mean value ± standard deviation, calculated from 3 biological replicates. This figure has been modified from our previous publication19. Please click here to view a larger version of this figure.
Discussion
The high throughput persister assay described here was developed to elucidate the effects of various chemicals on E. coli persistence. In addition to commercial PM plates, microarrays can be constructed manually as described in step 4.2. Moreover, the protocol presented here is flexible and can be used to screen other microarrays, such as drug panels and cell libraries, that are in 96 well plate formats. The experimental conditions including the growth phase, inoculation rate and medium can be adjusted to test these libraries. For instance, if one wants to screen a cell library, such as Keio E. coli knockout collection23, they can first transfer the strains from the library to 96 well plates that include fresh media, using a multichannel pipette. Once the cells reach the desired growth phase (e.g., exponential or stationary phase), the cells are then transferred to persister-assay plates as described in step 4.3 to enumerate the persister levels of the knockout strains. Similarly, this strategy can be used to screen the E. coli Promoter collection24 (a library of fluorescent reporter strains in 96 well plate formats) to identify promoters that are activated by antibiotics. These promoters can be readily detected by measuring the fluorescence signals of strains in persister-assay plates during the antibiotic treatment.
In our experiments, we used a modified LB (lacking NaCl) to avoid any additional effect that could arise from NaCl, considering that the microarrays already have various osmolytes. Although NaCl in regular LB medium is known to be good for preserving the membrane integrity of cells, it has been reported that NaCl at 0-1% range has a minimal effect on cell growth25. Moreover, results from propidium iodide (PI) staining and persisters assays in our previous study have shown that the absence of NaCl does not significantly affect the membrane integrity and the persistence of E. coli cells19. This modification was made specifically to address the concerns in our study and can be changed based on the nature of the research.
We have also adapted a method developed by Keren et al.5 to eliminate pre-existing persisters in our cell cultures prior to inoculation into the microarrays. Type I persisters are known to be generated during the stationary phase; therefore, the direct inoculation of the cells from overnight cultures into microarray plates would transfer a significant number of persisters, which may hinder the effects of osmolytes. With the dilution/growth cycle experiments (see Protocol 2), we were able to significantly reduce these preexisting persisters arising from the overnight cultures19. We note that we have cultured the cells in microarray plates for 24 h to be able to count all persisters, including type I variants that are formed during the stationary phase in the presence of osmolytes. However, these adapted techniques can always be modified depending on the nature of the study.
We treated the cells in microarrays with 5 µg/mL OFX to enumerate the persisters. Since the washing procedure to remove the antibiotic from 96 samples would be very labor-intensive, cells after the treatment were serially diluted in PBS without washing (see step 4.2). This procedure diluted the OFX concentration more than 30-fold. The diluted cell suspensions were then plated on antibiotic-free agar plates where the OFX was further dispersed. Our preliminary studies where we repeated these experiments with and without washing have verified that this serial-dilution method did not affect the persister levels19.
During the multi-channel pipetting in 96 well plates, one should be carefully mixing the cell suspensions to maintain a homogenous cell distribution. To do this, depending on the working volume, it would be beneficial to note the minimum number of pipetting that is needed to make the solution homogeneous. For this purpose, we conducted a simple control experiment where we monitored the pipetting (up and down) by dispersing a dye molecule in PBS and medium under the conditions studied here. Additionally, if working with more complex environments such as pH buffers, we would suggest measuring the pH of the cell culture before and during the incubation in a shaker as the cell culture tends to be very alkaline (pH ≈ 8) over time. This can give an idea of the quality as well as the pH range of the buffer used. Additionally, in order to obtain countable CFUs on agar plates from the high-throughput assays, certain conditions, such as the cell inoculation rate, age of cultures, and serial-dilution parameters (PBS volume, dilution rates and the number of dilution steps, etc.) should be optimized before testing the microarrays. Finally, the results obtained from the microarrays should be further validated in flasks as the volume of the culture, surface area and aeration in a 96 well plate could have additional effects on the observed results.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
We would like to thank the members of Orman Lab for their valuable inputs during this study. This study was funded by the NIH/NIAID K22AI125468 career transition award and a University of Houston startup grant.
Materials
| 14-ml test tube | Fisher Scientific | 14-959-1B | |
| E. coli strain MG1655 | Princeton University | Obtained from Brynildsen lab | |
| Flat-bottom 96-well plate | USA Scientific | 5665-5161 | |
| Gas permeable sealing membrane | VWR | 102097-058 | Sterilized by gamma irradiation and free of cytotoxins |
| Half-area flat-bottom 96-well plate | VWR | 82050-062 | |
| LB agar | Fisher Scientific | BP1425-2 | Molecular genetics grade |
| Ofloxacin salt | VWR | 103466-232 | HPLC ≥97.5 |
| Phenotype microarray (PM-9 and PM-10) | Biolog | N/A | PM-9 and PM-10 plates contained various osmolytes and buffers respectively |
| Round-bottom 96-well plate | USA Scientific | 5665-0161 | |
| Sodium chloride | Fisher Scientific | S271-500 | Certified ACS grade |
| Sodium nitrate | Fisher Scientific | AC424345000 | ACS reagent grade |
| Sodium nitrite | Fisher Scientific | AAA186680B | 98% purity |
| Square petri dish | Fisher Scientific | FB0875711A | |
| Tryptone | Fisher Scientific | BP1421-500 | Molecular genetics grade |
| Varioskan lux multi mode microplate reader | Thermo Fisher Scientific | VLBL00D0 | Used for optical density measurement at 600 nm |
| Yeast extract | Fisher Scientific | BP1422-100 | Molecular genetics grade |
Riferimenti
- Lewis, K. Persister cells, dormancy and infectious disease. Nature Reviews Microbiology. 5 (1), 48-56 (2007).
- Hobby, G. L., Meyer, K., Chaffee, E. Observations on the Mechanism of Action of Penicillin. Experimental Biology and Medicine. 50 (2), 281-285 (1942).
- Bigger, J. Treatment of staphylococcal infections with penicillin by intermittent sterilisation. The Lancet. 244 (6320), 497-500 (1944).
- Balaban, N. Q., Merrin, J., Chait, R., Kowalik, L., Leibler, S. Bacterial persistence as a phenotypic switch. Science. 305 (5690), 1622-1625 (2004).
- Keren, I., Kaldalu, N., Spoering, A., Wang, Y., Lewis, K. Persister cells and tolerance to antimicrobials. FEMS Microbiology Letters. 230 (1), 13-18 (2004).
- Keren, I., Minami, S., Rubin, E., Lewis, K. Characterization and transcriptome analysis of mycobacterium tuberculosis persisters. mBio. 2 (3), (2011).
- Mulcahy, L. R., Burns, J. L., Lory, S., Lewis, K. Emergence of Pseudomonas aeruginosa Strains Producing High Levels of Persister Cells in Patients with Cystic Fibrosis. Journal of Bacteriology. 192 (23), 6191-6199 (2010).
- LaFleur, M. D., Kumamoto, C. A., Lewis, K. Candida albicans biofilms produce antifungal-tolerant persister cells. Antimicrobial Agents and Chemotherapy. 50 (11), 3839-3846 (2006).
- Allison, K. R., Brynildsen, M. P., Collins, J. J. Metabolite-enabled eradication of bacterial persisters by aminoglycosides. Nature. 473 (7346), 216-220 (2011).
- Lechner, S., Lewis, K., Bertram, R. Staphylococcus aureus persisters tolerant to bactericidal antibiotics. Journal of Molecular Microbiology and Biotechnology. 22 (4), 235-244 (2012).
- Barrett, T. C., Mok, W. W. K., Murawski, A. M., Brynildsen, M. P. Enhanced antibiotic resistance development from fluoroquinolone persisters after a single exposure to antibiotic. Nature Communications. 10 (1), 1177 (2019).
- Windels, E. M., et al. Bacterial persistence promotes the evolution of antibiotic resistance by increasing survival and mutation rates. ISME Journal. 13 (5), 1239-1251 (2019).
- Dörr, T., Lewis, K., Vulić, M. SOS response induces persistence to fluoroquinolones in Escherichia coli. PLoS Genetics. 5 (12), 1000760 (2009).
- Völzing, K. G., Brynildsen, M. P. Stationary-phase persisters to ofloxacin sustain DNA damage and require repair systems only during recovery. mBio. 6 (5), (2015).
- Grant, S. S., Kaufmann, B. B., Chand, N. S., Haseley, N., Hung, D. T. Eradication of bacterial persisters with antibiotic-generated hydroxyl radicals. Proceedings of the National Academy of Sciences. 109 (30), 12147-12152 (2012).
- Gerdes, K., Maisonneuve, E. Bacterial Persistence and Toxin-Antitoxin Loci. Annual Review of Microbiology. 66 (1), 103-123 (2012).
- Orman, M. A., Brynildsen, M. P. Inhibition of stationary phase respiration impairs persister formation in E. coli. Nature Communications. 6 (1), 7983 (2015).
- Korch, S. B., Henderson, T. A., Hill, T. M. Characterization of the hipA7 allele of Escherichia coli and evidence that high persistence is governed by (p)ppGpp synthesis. Molecular Microbiology. 50 (4), 1199-1213 (2003).
- Karki, P., Mohiuddin, S. G., Kavousi, P., Orman, M. A. Investigating the effects of osmolytes and environmental ph on bacterial persisters. Antimicrobial Agents and Chemotherapy. 64 (5), 02393 (2020).
- Mohiuddin, S. G., Hoang, T., Saba, A., Karki, P., Orman, M. A. Identifying Metabolic Inhibitors to Reduce Bacterial Persistence. Frontiers in Microbiology. 11, 472 (2020).
- Brooun, A., Liu, S., Lewis, K. A dose-response study of antibiotic resistance in Pseudomonas aeruginosa biofilms. Antimicrobial Agents and Chemotherapy. 44 (3), 640-646 (2000).
- Luidalepp, H., Jõers, A., Kaldalu, N., Tenson, T. Age of inoculum strongly influences persister frequency and can mask effects of mutations implicated in altered persistence. Journal of Bacteriology. 193 (14), 3598-3605 (2011).
- Baba, T., et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: The Keio collection. Molecular Systems Biology. 2, 0008 (2006).
- Zaslaver, A., et al. A comprehensive library of fluorescent transcriptional reporters for Escherichia coli. Nature Methods. 3 (8), 623-628 (2006).
- Hajmeer, M., Ceylan, E., Marsden, J. L., Fung, D. Y. C. Impact of sodium chloride on Escherichia coli O157:H7 and Staphylococcus aureus analysed using transmission electron microscopy. Food Microbiology. 23 (5), 446-452 (2006).

