Genome Engineering of Primary Human B Cells Using CRISPR/Cas9
Summary
Here we provide a detailed, step-by-step protocol for CRISPR/Cas9-based genome engineering of primary human B cells for gene knockout (KO) and knock-in (KI) to study biological functions of genes in B cells and the development of B-cell therapeutics.
Abstract
B cells are lymphocytes derived from hematopoietic stem cells and are a key component of the humoral arm of the adaptive immune system. They make attractive candidates for cell-based therapies because of their ease of isolation from peripheral blood, their ability to expand in vitro, and their longevity in vivo. Additionally, their normal biological function—to produce large amounts of antibodies—can be utilized to express very large amounts of a therapeutic protein, such as a recombinant antibody to fight infection, or an enzyme for the treatment of enzymopathies. Here, we provide detailed methods for isolating primary human B cells from peripheral blood mononuclear cells (PBMCs) and activating/expanding isolated B cells in vitro. We then demonstrate the steps involved in using the CRISPR/Cas9 system for site-specific KO of endogenous genes in B cells. This method allows for efficient KO of various genes, which can be used to study the biological functions of genes of interest. We then demonstrate the steps for using the CRISPR/Cas9 system together with a recombinant, adeno-associated, viral (rAAV) vector for efficient site-specific integration of a transgene expression cassette in B cells. Together, this protocol provides a step-by-step engineering platform that can be used in primary human B cells to study biological functions of genes as well as for the development of B-cell therapeutics.
Introduction
B cells are a subgroup of the lymphocyte lineage derived from hematopoietic stem cells. They perform a critical role in the adaptive humoral immune system by producing large amounts of antibodies in response to immune challenges1. B cells are also precursors of memory B cells and the terminally differentiated, long-lived plasma cells, thereby providing lasting humoral immunity2. Plasma cells, in particular, are unique among immune cells in their ability to produce large amounts of a specific antibody while surviving for years or decades3. Additionally, the ease of isolation from peripheral blood makes the B-cell lineage an excellent candidate for novel cell-based therapies4.
Previously, random integration methods, such as those using lentiviral vectors or a Sleeping Beauty transposon, have been used to engineer B cells for transgene delivery and expression5,6,7,8. However, the non-specific nature of these approaches makes it difficult to study the biological functions of a specific gene in the B cells and carries an inherent risk of insertional mutagenesis and variable transgene expression and/or silencing in the therapeutic setting.
The CRISPR/Cas9 system is a powerful genome engineering tool that allows researchers to precisely edit the genome of various cells in numerous species. Recently, two groups, including our own, have successfully developed methods for ex vivo expansion and targeted genome engineering of primary human B cells9,10. We will describe the process of purifying primary human B cells from a leukaphoresis sample. After that, we will describe our updated protocol for B-cell expansion and activation of isolated B cells. We will then describe a process for knocking out cluster of differentiation 19 (CD19), a specific B-cell receptor and a hallmark of B cells, by electroporation to introduce CRISPR/Cas9 mRNA together with CD19 sgRNA into activated B cells.
Cas9 mRNA gets translated and binds to the CD19 sgRNA to form a CRISPR/Cas9-sgRNA ribonucleoprotein complex (RNP). Subsequently, sgRNA in the complex leads Cas9 to create double-strand break (DSB) at the target sequence on exon 2 of the gene. The cells will repair the DSB by “non-homologous end joining” by introducing or deleting nucleotides, leading to frameshift mutation and causing the gene to be knocked out. We will then measure the loss of CD19 by flow cytometry and analyze indel formation by tracking of indels by decomposition (TIDE) analysis.
We will then describe the process of using CRISPR/Cas9 together with a recombinant AAV6 vector (rAAV6, a donor template for homology-directed repair (HDR)) to mediate site-specific insertion of enhanced green fluorescent protein (EGFP) at the adeno-associated virus integration site 1 (AAVS1) gene. The AAVS1 gene is an active locus without known biological functions and an AAV viral integration site on the human genome; therefore, it is considered a “safe harbor” for genome engineering. Here, we report that expansion and activation of B cells allowed up to 44-fold expansion in 7 days of culture (Figure 1). Electroporation of B cells showed a slight reduction of overall cell health (Figure 2A) at 24 h post-transfection. Scatter plot analysis of the CD19 marker (Figure 2B) showed up to 83% reduction in the edited cells (Figure 2C).
TIDE analysis of the chromatographs (Figure 3A) revealed that the % indels was similar to the % protein loss by flow cytometry (Figure 3B). Flow cytometry analysis of the KI experiment showed that the cells that received AAV vector (Figure 4), together with RNP, expressed up to 64% EGFP-positive cells (Figure 5A) and later displayed successful integration by junction polymerase chain reaction (PCR) (Figure 5B). Cell counts showed that all samples quickly recovered within 3 days post-engineering (Figure 5C).
Protocol
Leukapheresis samples from healthy donors were obtained from a local blood bank. All experiments described here were determined to be exempt for research by the Institutional Review Board (IRB) and were approved by the Institutional Biosafety Committee (IBC) at the University of Minnesota.
NOTE: All experiments were performed in compliance with the universal precaution for bloodborne pathogens, with sterile/aseptic techniques and proper biosafety level-2 equipment.
1. Prepare supplements for B-cell expansion medium
- Reconstitute CpG oligonucleotide to a concentration of 1 mg/mL.
- Reconstitute CD40 ligand (CD40L) to a concentration of 100 μg/mL.
- Reconstitute recombinant human IL-10 (rhIL-10) to a concentration of 50 μg/mL.
- Reconstitute recombinant human IL-15 (rhIL-15) to a concentration of 10 μg/mL.
NOTE: Keep each supplement in small aliquots at -20 °C to -80 °C for up to 6 months.
2. Prepare basal medium
- Combine B-cell basal medium with 5% (v/v) media supplement for in vitro immune cell expansion (e.g., CTS Immune Cell SR) and 1% (v/v) penicillin and streptomycin.
- Filter-sterilize the basal medium using a 0.22 µm filter adaptor into a sterilized bottle.
- Keep the basal medium at 4 °C for up to 1 month.
3. Prepare B-cell expansion medium
- Transfer the required amount of the basal medium into a sterile container to culture the B cells at 5 × 105 cells/mL.
- Supplement the basal medium with 1 μg/mL CpG, 100 ng/mL CD40L, 50 ng/mL rhIL-10, and 10 ng/mL rhIL-15.
- Filter the B-cell expansion medium using a 0.22 µm filter.
- Equilibrate the B-cell expansion medium in the tissue culture incubator at 37 °C, 5% CO2 with humidity for at least 30 min before use.
NOTE: Prepare fresh B-cell expansion medium to use for one day. Do not prepare the B-cell expansion medium to use for multiple days. This media recipe encourages proliferation of memory B cells and activated primary human B cells.
4. Human B-cell purification and expansion
NOTE: Add 99–100% isopropyl alcohol to a temperature-controlled freezing container, following the manufacturer’s instruction, and chill the freezing container at 4 °C before starting step 4.1.
- Isolate PBMCs from a leukaphoresis sample
- Transfer a leukaphoresis sample (approximate 8–10 mL) to a sterile 50 mL conical tube.
- Bring up the volume to 35 mL with sterile 1x phosphate-buffered saline (PBS).
- Carefully layer a 35 mL leukaphoresis sample on 15 mL of density-gradient medium.
- Centrifuge at 500 × g for 25 min without brake, remove the plasma layer without disturbing the buffy coat (PBMC layer), collect the PBMCs from the interface, and transfer to a new sterile 50 mL conical tube.
- Bring up the PBMCs to 50 mL with 1x PBS.
- Centrifuge at 500 × g for 5 min without brake. Remove the supernatant without disturbing the PBMC pellet, which may appear red.
- Add 7 mL of ammonium-chloride-potassium lysis buffer, pipette 3 times to mix well, and incubate at room temperature (RT) for 3 min.
- Bring up the volume to 50 mL with 1x PBS.
- Centrifuge at 400 × g for 5 min with low-resistance brake. Remove the supernatant without disturbing the pellet. The pellet should look pinkish or white.
NOTE: To continue to culture the freshly isolated B cells, prepare the B-cell expansion medium before starting B-cell isolation (step 4.2).
- B-cell isolation from PBMCs using human primary B-cell negative isolation kit
- Resuspend PBMCs in the isolation buffer to a concentration of 5 × 107 cells/mL.
NOTE: If the total number of PBMCs is less than 5 × 107 cells, scale down the volume of isolation buffer to maintain 5 × 107 cells/mL. Minimum and maximum volumes of cell suspension are 0.25 mL and 8 mL, respectively. - Transfer up to 8 mL (5 × 107 cells/mL) to a sterile, polypropylene, round-bottom tube with cap.
- Add 50 μL/mL of Cocktail Enhancer to the PBMCs.
- Add 50 μL/mL of Isolation Cocktail to the PBMCs, cap the tube, and invert 2–3 times to mix.
- Incubate at RT for 5 min; at the 4th minute of incubation, vortex magnetic microbeads for at least 30 s.
- Transfer 50 μL of magnetic microbeads per 1 mL of PBMCs, cap the tube, and invert 2–3 times to mix.
- Top up to 10 mL with isolation buffer and gently pipette up and down 2–3 times.
- Place the tube in a magnetic station and incubate at RT for 3 min.
- Hold the magnet and tube together, and in one motion, invert the magnet and tube together to pour the cell suspension into the new tube. Discard the old tube.
- Repeat step 4.2.8 (reduce the incubation time to 2 min) and pour the B-cell suspension into a clean conical tube.
- The enriched B cells are ready to use. If cells will be used immediately, continue to section 4.3 (Human B-cell expansion). Check the purity of the isolated B cells by flow cytometry (optional). If cells are to be frozen before use, continue to step 4.2.12.
- To freeze the B cells, centrifuge at 400 × g for 5 min, and discard the supernatant without disturbing the pellet.
- Resuspend the cells in freezing medium at 107 cells/mL, and aliquot 1 mL/cryovial.
- Place the cryovial in the chilled freezing container, and store at -80 °C overnight; then, transfer the frozen cryovial to a liquid nitrogen tank; keep frozen cells up to 1 year.
NOTE: Expected yield of isolated B cells is 2%–8% of the total PBMCs, with 95%–99% viability.
- Resuspend PBMCs in the isolation buffer to a concentration of 5 × 107 cells/mL.
- Human B-cell expansion
NOTE: If using freshly isolated B cells, skip steps 4.3.1–4.3.5. Count the cells and transfer the required number of cells into a sterile conical tube and continue with step 4.3.6.- Pre-warm fetal bovine serum (FBS) in a water bath prior to thawing the B cells. Prepare 20 mL of B-cell expansion medium, transfer to a T25 flask, and pre-equilibrate the medium in a tissue-culture incubator (at 37 °C, 5% CO2, with humidity) at least 15 min before use.
- Thaw B cells in a 37 °C water bath. While waiting, transfer 2 mL of pre-warmed FBS into a sterile 15 mL conical tube.
- After the B cells are completely thawed, immediately add 1 mL of pre-warmed FBS, drop-wise, into the sample. Incubate at RT for 1 min.
- Gently pipette to resuspend the sample and transfer the whole volume, dropwise, into a conical tube containing 2 mL of pre-warmed FBS.
- Bring up the volume to 15 mL with sterile 1x PBS, cap, and invert the tube gently 2–3 times.
- Centrifuge at 400 × g for 5 min.
- Discard the supernatant without disturbing the cell pellet, resuspend the cell pellet with 1 mL of pre-equilibrated B-cell expansion medium, and count the cells. The total cell number should be approximately 107 cells.
- Transfer the cells into a flask containing 20 mL of the pre-equilibrated B-cell expansion medium. The final concentration of the cells should be approximately 5 x 105 cells/mL.
- Incubate the flask vertically in a tissue-culture incubator.
- Refresh the expansion medium completely every 2 days by transferring the whole volume of B cells in to a sterile conical tube and repeat steps 4.3.6–4.3.9.
NOTE: T25 flask can hold 10–20 mL of medium; T75 flask can hold up to 20–60 mL of medium.
5. Primary human B-cell engineering
- Engineer B cells at 48 ± 2 h after expansion/activation for optimal results. Prepare the B-cell expansion medium, aliquot 1 mL of the medium into a 48-well tissue-culture plate, and pre-equilibrate in a tissue-culture incubator at least 15 min prior to use.
NOTE: When designing a CRISPR/Cas9 sgRNA for a gene of interest, follow the steps outlined below.
– Design sgRNAs using an online tool11.
– Design sgRNAs on an exon that is common to all isoforms of the protein.
– Order chemically modified sgRNA from reputable companies.
– sgRNA usually comes in lyophilized form; reconstitute sgRNA in sterile DNase/RNase-free Tris-EDTA (TE) buffer to a concentration of 1 μg/μL. - Prepare CRISPR/Cas9 transfecting substrate by mixing 1 μL (1 μg/μL) of chemically modified sgRNA with 1.5 μL (1 μg/μL) of chemically modified Streptococcus pyogenes Cas 9 (S.p. Cas9) nuclease per transfection reaction. For control, use 1 μL of TE buffer instead of sgRNA.
NOTE: When CD19 sgRNA is used for a gene KO experiment, see Figure 2 for results. When AAVS1 sgRNA is used for a gene KI experiment, see Figure 5 for results.- Keep all the components on ice.
- Mix gently and transfer 2.5 μL of CRISPR/Cas9 transfecting substrate per reaction into a tube of a 0.2 mL 8-tube strip; set aside at RT.
- Turn on a nucleofector (electroporator), and prepare transfection reagents as shown in Table 1.
NOTE: This is a good step for a pause, if necessary, by putting all reagents on ice. Remove all reagents from ice when ready to resume the experiment. When using S.p. Cas9 protein, the investigator MUST pre-complex CRISPR/Cas9-sgRNA ribonucleoprotein by mixing 1 μg sgRNA with 5 μg of S.p. Cas9 protein, avoiding any bubbles, and incubating the mixture at RT for at least 20 min before use for optimal results. - Count and transfer 106 B cells per transfection reaction into a sterile conical tube.
- Bring up the volume to 15 mL with sterile 1x PBS, and centrifuge at 400 × g for 5 min. While waiting, prepare the primary cell transfection reagent (Table 2) and set aside at RT.
- Discard the supernatant without disturbing the cell pellet.
- Resuspend the cell pellet with 10 mL of sterile 1x PBS and centrifuge at 400 × g for 5 min.
- Discard the supernatant completely without disturbing the cell pellet.
- Transfer 0.5 μg of chemically modified GFP mRNA (as a transfection reporter) per 106 B cells to the cell pellet (optional).
- Resuspend the cell pellet with 20 μL primary cell transfection reagent per 106 B cells; mix gently by pipetting 5–6 times. Transfer 20.5 μL per transfection reaction into the 0.2 mL tube of the 8-tube strip containing 2.5 μL of the CRISPR/Cas9 transfection substrate.
- Pipette up and down once to mix and transfer the entire volume (23 μL) into a transfection cuvette. Cap and tap the cuvette on the bench gently to ensure that the liquid covers the bottom of the cuvette.
- Use human primary B-cell protocol on the nucleofector for transfection.
NOTE: The nucleofector (electroporator) can be placed and be used outside the tissue-culture hood. Cap the cuvette to ensure sterility. - Rest the electroporated cells in the cuvette at RT for 15 min.
- Transfer 80 μL of the pre-equilibrated B-cell expansion medium from the tissue culture plate into the transfection reaction in the cuvette. Place the cuvette in the tissue culture incubator for 30 min.
- Gently pipette a couple times to mix and transfer the whole volume of the sample from the cuvette to an appropriate well of a 48-well tissue-culture plate containing 1 mL of the B-cell expansion medium. The final concentration of the cells should be 106 cells/mL.
- If performing a gene KI experiment, transfer rAAV6 vector at 500,000 multiplicity of infection into the appropriate well containing electroporated cells (approximately 45 min post-electroporation). See example rAAV6 vector construct in Figure 4A.
NOTE: For example: A control sample will be electroporated without CRISPR/Cas9 or the rAAV6 vector. A vector-only sample will be electroporated without CRISPR/Cas9 and then be transduced with the rAAV6 vector. A KI sample will be electroporated with CRISPR/Cas9 and then be transduced with the rAAV6 vector. rAAV6 must contain homology arms up- and downstream of the targeted DSB site for HDR. - Place the plate in a tissue culture incubator at 37 °C and 5% CO2 with humidity.
- Count the cells and record viability at day 1 post-engineering.
- Refresh the B-cell expansion medium every 2 days by counting the cells and then transferring the whole volume of the cells into a clean 1.5 mL microcentrifuge tube. Centrifuge at 400 × g for 5 min, and discard the supernatant without disturbing the pellet. Resuspend the cells with 100 µL of fresh B-cell expansion medium, and transfer to a well of a 24-well tissue culture plate. Bring up the medium volume to achieve a final cell concentration at 5 × 105 cells/mL.
NOTE: A 48-well plate can hold up to 1 mL medium/well; a 24-well plate can hold up to 2 mL medium/well; a 12-well plate can hold up to 4 mL medium/well, and a 6-well plate can hold up to 8 mL medium/well. - Allow the engineered cells to expand for at least 5 days before performing downstream analyses such as for flow cytometry analysis, TIDE analysis, and junction PCR.
Representative Results
The updated expansion and activation protocol enabled the rapid expansion of B cells up to 44-fold in 7 days (Figure 1; n =3 donors). In the KO experiment, the B-cell count using Trypan blue staining showed more than 80% viable cells with a slight reduction in cell recovery in both the control and the CD19 KO samples at 24 h post-electroporation (Figure 2A; p ≥ 0.05, n = 3 donors). This result indicates that electroporation slightly affected overall B-cell health. B cells were collected on day 5 post-transfection for flow cytometry and TIDE analyses. Representative scatter plots of the control and KO sample showed 14% and 95% CD19-negative cells, respectively (Figure 2B). Quantitation of the flow plots showed significant reduction in CD19 expression in the KO samples when compared to the control (Figure 2B; p ≤ 0.0001, n = 3 donors). Chromatograms of genomic sequencing (see primer sequences in Table 3) showed double peaks in the CD19 KO B cells, indicating insertions/deletions of nucleotides post-CRISPR/Cas9-mediated DSB, whereas single peaks were observed in the control, indicating no DSB occurred in this sample (Figure 3A). Indel analysis of the chromatographs (using a free online TIDE analysis tool) of the KO samples showed high % of indel formation (>90%) at the CD19 locus, which is consistent with % CD19 protein loss detected by flow cytometry (Figure 3B; p ≥ 0.05, n = 3 donors). These results indicate that CRISPR/Cas9 efficiently generated a CD19 KO in B cells. B cells from the KI experiment were collected on day 12 post-engineering for flow cytometry and junction PCR analyses (Table 4). Scatter plots showed 64% of EGFP-positive cells in the sample that received the rAAV6 vector (Figure 4) together with RNP, whereas no EGFP-positive cell was observed in the control; minimal EGFP-positivity was observed in the sample that received AAV vector only (Figure 5A). A junction PCR amplification (see primer sequences in Table 3) showed 1.5 Kbps amplicons in the KI sample (Figure 5B), whereas no PCR product was observed in either the control or vector-only sample. Cell counts showed that the engineering process affects cell recovery in the KI sample more than the control or the vector-only samples (Figure 5B). However, all samples quickly rebounded within 3 days after engineering (Figure 5B). Together, these results indicate that successful integration of EGFP at the AAVS1 locus leads to stable expression of EGFP at least 12 days post-engineering.
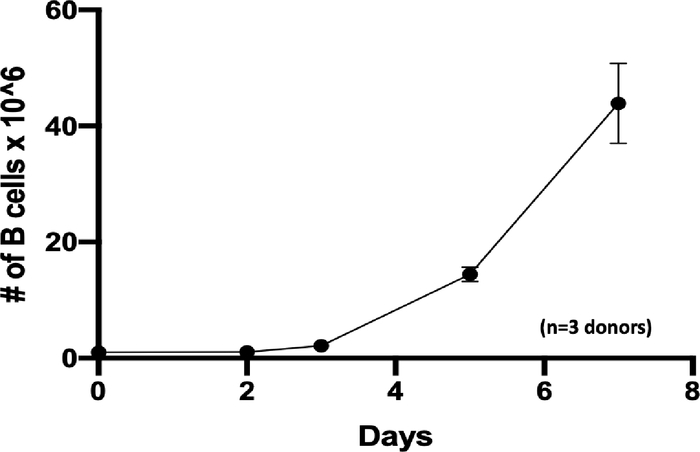
Figure 1: B cell expansion in vitro. B cells were seeded at 1 × 106 cells on day 0 (zero) at a density of 5 × 105 per mL and expanded 44-fold in 7 days (n=3 independent donors). Please click here to view a larger version of this figure.
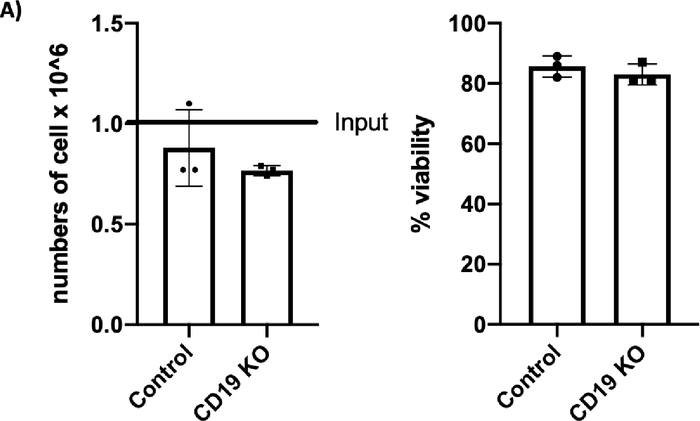
Figure 2A: Please click here to view a larger version of this figure.
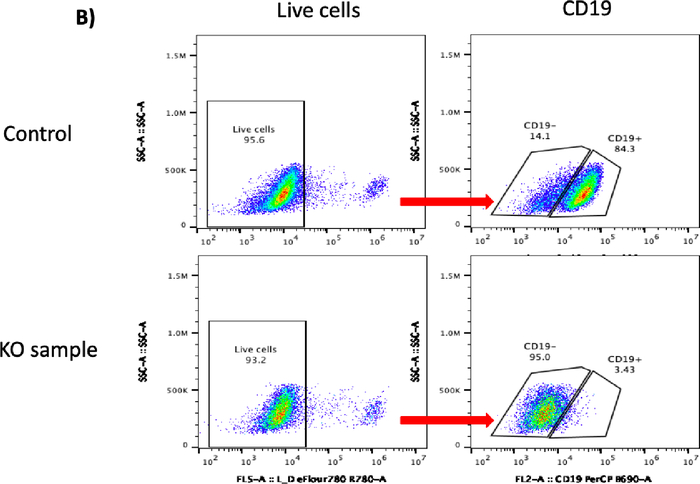
Figure 2B: Please click here to view a larger version of this figure.
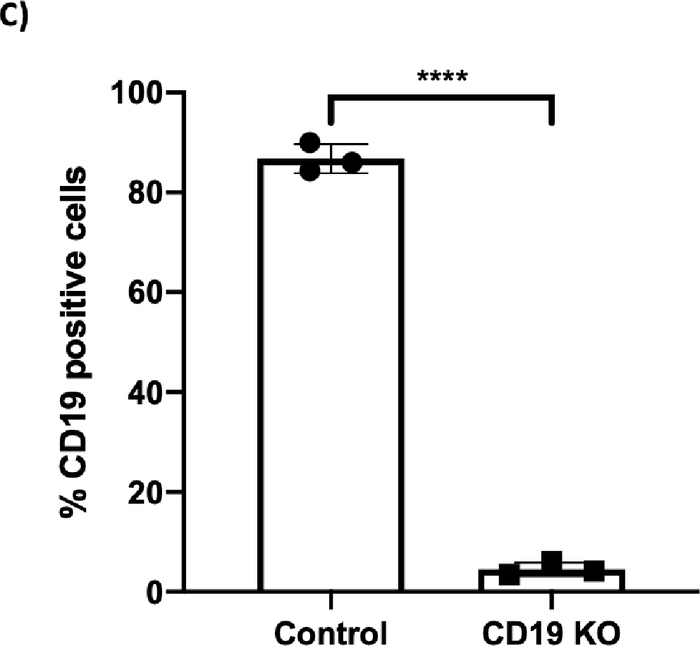
Figure 2C: CRISPR/Cas9-mediated CD19 knockout (KO) in B cells. (A) Bar graph shows >70% cell recovery (left panel) and >80% viability (right panel) of cells post transfection were observed in both the control and the CD19 KO samples at 24 hours post electroporation. (B) Representative flow plots of CD19 gating of live cells shows 84.3% and 3.43% CD19-positive cells in the control sample and the CD19 KO sample, respectively. (C) Bar graph shows significant reduction of CD19 in the CRISPR/Cas9-mediated CD19 KO group (p ≤ 0.0001). Please click here to view a larger version of this figure.
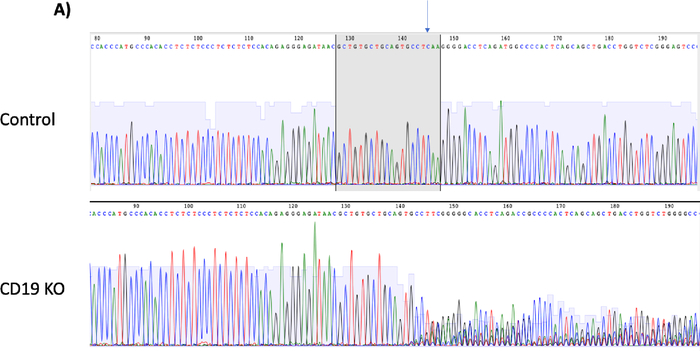
Figure 3A: Please click here to view a larger version of this figure.
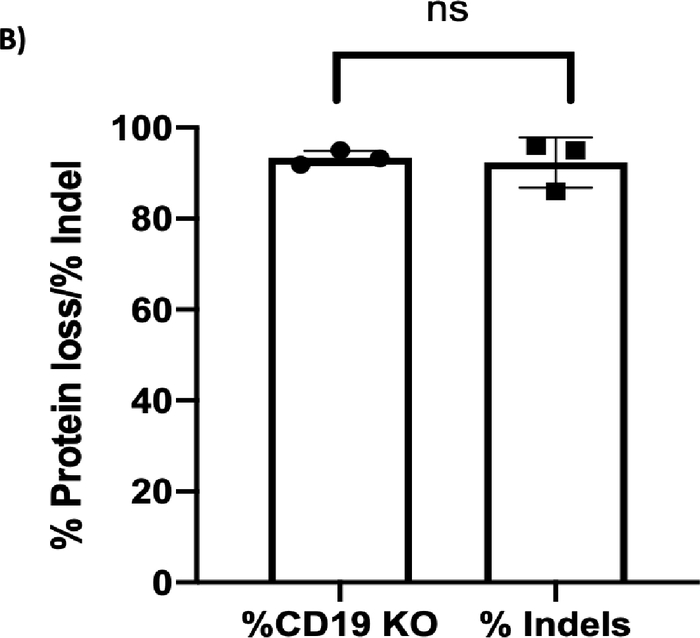
Figure 3B: CD19 protein loss vs indel formation. (A) Chromatograms depict sequencing peaks of the control and CD19 knockout (KO) sample. The gray box on the control peaks highlights the target sequence of CD19 gRNA with the predicted cut site indicated by an arrow. CD19 KO showed “double peaks” sequencing around the predicted cut site, indicating insertion/deletions of nucleotides after the double-stranded break. (B) Bar graph showing consistent results between % CD19 protein loss and % indel formation at the CD19 locus (p ≥ 0.05). Please click here to view a larger version of this figure.

Figure 4: rAAV6 AAVS1 MND-GFP vector construct. Expression cassette contains a strong synthetic promoter (MND) sequence, immediately followed by an enhanced green fluorescence protein (EGFP) coding sequence and poly adenylation (Poly A) sequence. AAVS1 homology arms flank upstream of the MND promoter and downstream of the poly A sequences. EGFP will be expressed under the regulation of the MND promoter. Sequence lengths are indicated above each component of the construct. Please click here to view a larger version of this figure.
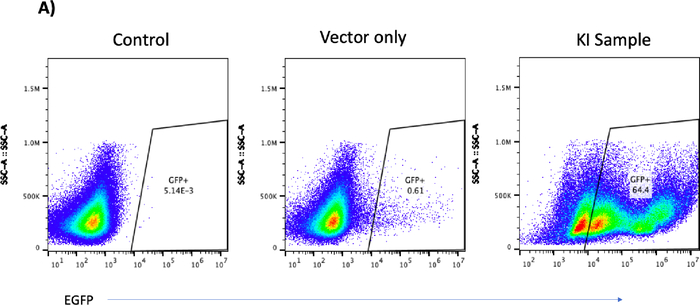
Figure 5A: Please click here to view a larger version of this figure.
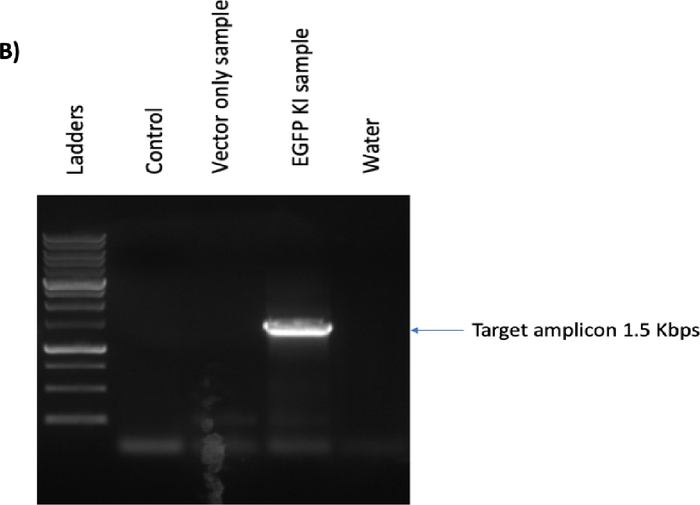
Figure 5B: Please click here to view a larger version of this figure.
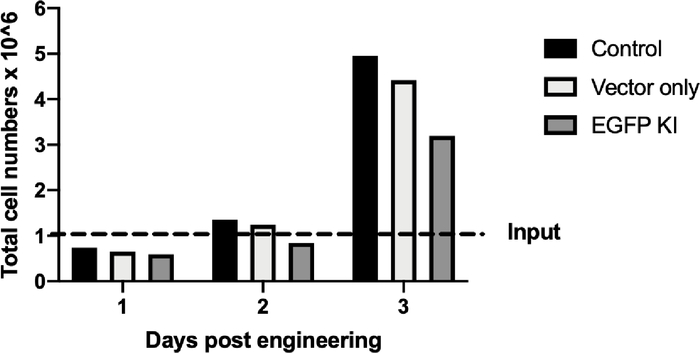
Figure 5C: CRISPR/Cas9- and rAAV6-mediated site-specific integration of the EGFP reporter cassette in B cells at day 12 post-engineering. (A) Representative flow plot shows no EGFP-positive B cells in either the control or the vector-only sample versus 64.4% of the EGFP-positive B cells from the knockin (KI) sample. (B) Junction polymerase chain reaction of KI sample shows the predicted 1.5 Kbps band; no band was found in the control or vector-only sample. Water was used to ensure no contamination during the PCR process. (C) Bar graph depicts cell growth of the control, the vector-only, and the EGFP-KI samples over a period of 3 days after engineering. Broken line indicates the 1 × 106 cell input. Please click here to view a larger version of this figure.
| Name | gRNA sequence |
| CD19 | 5’-GCTGTGCTGCAGTGCCTCAA-3’ |
| AAVS1 | 5’-GTCACCAATCCTGTCCCTAG-3’ |
Table 1: gRNA sequences.
| Reagents | Volume per 1 reaction |
| P3 Primary Cell solution | 16.4 mL |
| Supplement 1 | 3.6 mL |
| Total | 20 mL |
Table 2: Preparation of Nucleofection reagent mix.
| Description | Sequence | Purpose |
| CD19 forward primers | 5’-AAATTCAGGAAAGGGTTGGAAG-3’ | Amplify the CD19 locus for Indel analysis |
| CD19 Reverse primer | 5’-GCGGACCTCTTCTGTCCATG-3’ | Amplify the CD19 locus for Indel analysis |
| Junction PCR forward primer | 5’-GGACGAGCTGTACAAGTAACG-3’ | Junction PCR |
| Junction PCR Reverse primer | 5’-GAGACAGTGACCAACCATCC-3 | Junction PCR |
Table 3: Primers used.
| Description | Fluorophore | Clone |
| Anti-human CD19 | PerCP | HIB19 |
| Viability Dye | eFlour 780 | – |
Table 4: Flow cytometry antibody and viability dye.
Discussion
Precise genome engineering in primary human B cells has been challenging until recently9,10. We had previously published protocols using CRISPR/Cas9 to engineer primary human B cells9. Here, we outline improved protocols for B-cell isolation, expansion, and engineering to allow for efficient KO of CD19 or for knocking-in EGFP.
Here we demonstrate that our expansion protocol allows the rapid expansion of B cells in culture for up to 44-fold expansion in 7 days (Figure 1). This protocol showed a faster expansion rate than those reported by Johnson et al.9 and Hung et al.10 The critical steps to ensure healthy and rapid expansion of primary B cells are replenishing the medium with fresh activation factors every two days and ensuring that the number of total B cells in culture does not exceed 2 × 106 cell/mL.
We also found that our improved protocol for transfecting the CRISPR/Cas9 system for CD19 KO resulted in high CD19 KO efficiency (Figure 2 and Figure 3). Similar to a previous study9, we observed a slight reduction in the cell recovery and viability post-electroporation at 24 h. This indicates that electroporation affects the overall cell health of primary B cells; however, the cells eventually rebound within 48 h (data not shown).
The benefit of using this electroporator protocol is that we can electroporate samples at a much faster rate (16 reactions in less than 1 min), whereas previous protocols9,10 take approximately 10–12 min for 16 reactions. In addition, this transfection system eliminates the potential electric arc of the electroporator observed in previous studies9,10. Furthermore, this engineering method can be scaled up for larger numbers of B cells using larger, commercially available cuvettes (data not shown).
A few critical steps to ensure optimal overall cell health and KO efficiencies: First, ensure that the B-cell expansion medium is prepared and pre-equilibrated for at least 15 min before use. Second, the total volume of transfection substrate should not exceed 20% (v/v) of the nucleofection reagent. Third, the cells must be expanded/activated for 48 ± 2 h for optimal results. Fourth, tap the electroporation cuvette gently before placing in the electroporator to ensure that the transfection reaction solution covers the bottom of the cuvette. Fifth, be sure to rest the electroporated cells in the cuvette for only 15 min at RT; leaving cells in the nucleofection reagent after electroporation for too long can harm the overall cell survival. Sixth, be sure to incubate the cells with media in the cuvette for no longer than 30 min. Seventh, placing 106 electroporated cells in 1 mL for the first 48 h tends to help cells recover more quickly than culturing them at lower density (data not shown). Lastly, using either Cas9 mRNA or Cas9 protein will result in similar editing efficiencies (data not shown); however, we used Cas9 mRNA for convenience and cost.
Two major concerns when using CRISPR/Cas9 are off-target effects and chromosomal translocations. Off-target effects caused by mismatched base pairs of sgRNA to the target sequence, lead to numerous possible binding sites on the genome and create multiple, unwanted gene KOs. Therefore, the predicted off-target score should be taken into consideration along with the on-target score to minimize this issue. Chromosomal translocation can occur due to off-target effects or when knocking out multiple genes. This can cause catastrophic events experimentally and clinically. Base editors12 alter a single nucleotide (two classes: cytosine base editor and adenine base editor) to disrupt the splicing element, to create a premature stop codon, or to create a point mutation, leading to target gene KO without DSB (reviewed extensively elsewhere12). Thus, the base-editing approach can be used for single or multiple-gene KOs to circumvent chromosomal translocations.
We also demonstrate that CRISPR/Cas9, together with rAAV6, can be used to efficiently mediate site-specific KI and expression of EGFP at the AAVS1 locus. We observed EGFP expression for at least 12 days post-engineering. We also observed two EGFP-positive populations: a high- and an intermediate-EGFP expression in the KI sample on day 12 (Figure 5A), whereas the vector-only sample showed a minimal percentage of cells with intermediate EGFP expression. We speculate that this “high and intermediate populations” phenomenon is due to biallelic integration of the vector. Further investigation using spanning PCR13 of the intermediate- and high-EGFP cells can be done to confirm this hypothesis. Two caveats on using AAV as a vector: First, AAV vectors have a small cargo capacity, up to 4.7 kilobases, causing issues when knocking in a large gene. Reducing the size of homology arms will allow accommodation of a larger gene, which in turn will reduce KI efficiency (data not shown). Alternatively, simultaneous or sequential integration of multiple gene-loading vectors can be used14,15. Studies have reported an immune response and a clearance of AAV-transduced cells in immunocompetent animal models16,17. Alternatively, a non-viral-based donor HDR template can be explored to circumvent this issue.
In summary, we have demonstrated comprehensive, step-by-step processes of isolation, expansion, and engineering of B cells that resulted in high gene modification efficiencies. This engineering method can be used for gene KO and to study the functions of genes in B cells. In addition, this method can be used to engineer B cells to express a recombinant antibody to fight against infection. Lastly, this method can be applied to engineer B cells to express and secrete therapeutic enzymes that can be used as an autologous cell-based therapy to treat enzymopathies.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This work was funded by the Children’s Cancer Research Fund (CCRF) and NIH R01 AI146009 to B.S.M.
Materials
| Alt-R S.p. Cas9 Nuclease V3 protein, 500 ug | IDT | 1081059 | smaller size is also available |
| Amaxa P3 primary cell 4D- Nucleofector X Kit S (32 RCT) | Lonza | V4XP-3032 | |
| Ammonium-chloride-potassium (ACK) lysing buffer | Quality Biological | 118-156-101 | |
| CleanCap chemically modified Cas9 mRNA | Trilink Biotechnology | L-7206-1000 | |
| CpG ODN 2006 (ODN 7909) 5 mg | Invivogen | TLRL-2006-5 | different sizes available |
| Cryostor CS10, 100 mL | STEMCELL Technology | 7930 | |
| CTS Immune Cell SR | Thermo Fisher Scientific | A2596101 | |
| EasySep human B cells isolation kit | STEMCELL Technology | 17954 | |
| eBioscience fixable viability dye eFlour 780 | eBiosciences | 65-0865-14 | |
| Excellerate B cell media, Xeno-free | R&D Systems | CCM031 | B-cell basal medium |
| Falcon 14 mL Polypropylene Round-bottom Tube | Corning | 352059 | |
| Fetal Bovine Serum (FBS) | R&D Systems | S11550 | For thawing B cells only |
| Ficoll-Paque Plus | GE Healthcare | 17-1440-03 | |
| GeneMate SnapStrip® 8-Strip 0.2 mL PCR Tubes with Individual Attached Dome Caps | BioExpress | T-3035-1 / 490003-692 | |
| Hyclone 0.0067M PBS (No Ca, No Mg) or 1x PBS | GE lifesciences | SH30256.01 | |
| Lonza 4D Nucleofector core unit | Lonza | AAF-1002B | |
| Lonza 4D Nucleofector X unit | Lonza | AAF-1002X | |
| Mega CD40 Ligand | Enzo Life Sciences | ALX-522-110-C010 | |
| Mr. Frosty | Sigma-Aldrich | C1562-1EA | For freezing cells |
| Pen/Strep 100X | Sigma-Aldrich | TMS-AB2-C | |
| PerCP anti-human CD19 clone HIB19 | biolegend | 302228 | smaller size is also available |
| rAAV6 SA-GFP pakaging ( with our SA-GFP cassette see Figure 4.) | Vigene Biosciences | N/A | large scale packaging, 1e13 GC/mL, 500 mL |
| Recombinant human IL-10 protein 250 ug | R&D Systems | 217-IL-250 | different sizes available |
| Recombinant human IL-15 protein 25 ug | R&D Systems | 247-ILB-025 | different sizes available |
| The Big Easy EasySep Magnet | STEMCELL Technology | 18001 | different sizes available |
| Tris-EDTA (TE) buffer | Fisher Scientific | BP2476100 |
Riferimenti
- LeBien, T. W., Thomas, T. F. B lymphocytes: how they develop and function. Blood. 112 (5), 1570-1580 (2008).
- Nutt, S. L., Hodgkin, P. D., Tarlinton, D. M., Corcoran, L. M. The generation of antibody-secreting plasma cells. Nature Reviews Immunology. 15, 160-171 (2015).
- Slifka, M. K., Antia, R., Whitmire, J. K., Ahmed, R. Humoral immunity due to long-lived plasma cells. Immunity. 8 (3), 363-372 (1998).
- Spriggs, M. K., et al. Recombinant human CD40 ligand stimulates B cell proliferation and immunoglobulin E secretion. Journal of Experimental Medicines. 176 (6), 1543-1550 (1992).
- Fusil, F., et al. A lentiviral vector allowing physiologically regulated membrane-anchored and secreted antibody expression depending on B-cell maturation status. Molecular Therapy. 23 (11), 1734-1747 (2015).
- Luo, X. M., et al. Engineering human hematopoietic stem/progenitor cells to produce a broadly neutralizing anti-HIV antibody after in vitro maturation to human B lymphocytes. Blood. 113 (7), 1422-1431 (2008).
- Mock, U., Thiele, R., Uhde, A., Fehse, B., Horn, S. Efficient lentiviral transduction and transgene expression in primary human B cells. Human Gene Therapy Methods. 23 (6), 408-415 (2012).
- Heltemes-Harris, L. M., et al. Sleeping Beauty transposon screen identifies signaling modules that cooperate with STAT5 activation to induce B-cell acute lymphoblastic leukemia. Oncogene. 35 (26), 3454-3464 (2016).
- Johnson, M. J., Laoharawee, K., Lahr, W. S., Webber, B. R., Moriarity, B. S. Engineering of primary human B cells with CRISPR/Cas9 targeted nuclease. Scientific Reports. 8, 12144 (2018).
- Hung, K. L., et al. Engineering protein-secreting plasma cells by homology-directed repair in primary human B cells. Molecular Therapy. 26 (2), 456-467 (2018).
- Cui, Y., Xu, J., Cheng, M., Liao, X., Peng, S. Review of CRISPR/Cas9 sgRNA design tools. Interdisciplinary Sciences. 10 (2), 455-465 (2018).
- Rees, H. A., Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nature Reviews Genetics. 19 (12), 770-788 (2018).
- Spiegel, A., Bachmann, M., Jurado Jiménez, G., Sarov, M. CRISPR/Cas9-based knockout pipeline for reverse genetics in mammalian cell culture. Methods. 164-165, 49-58 (2019).
- Suzuki, T., Kazuki, Y., Oshimura, M., Hara, T. A novel system for simultaneous or sequential integration of multiple gene-loading vectors into a defined site of a human artificial chromosome. PLoS One. 9, 110404 (2014).
- Bak, R. O., Porteus, M. H. CRISPR-mediated integration of large gene cassettes using AAV donor vectors. Cell Reports. 20 (3), 750-756 (2017).
- Shirley, J. L., de Jong, Y. P., Terhorst, C., Herzog, R. W. Immune responses to viral gene therapy vectors. Molecular Therapy. 28 (3), 709-722 (2020).
- Martino, A. T., Markusic, D. M. Immune response mechanisms against AAV vectors in animal models. Molecular Therapy – Methods and Clinical Development. 17, 198-208 (2020).

