Application of Flow Vermimetry for Quantification and Analysis of the Caenorhabditis elegans Gut Microbiome
Summary
Caenorhabditis elegans is a powerful model to examine the molecular determinants driving host-microbiome interactions. We present a high throughput pipeline profiling the single animal levels of gut microbiome colonization together with key aspects of the C. elegans physiology.
Abstract
The composition of the gut microbiome can have a dramatic impact on host physiology throughout the development and the life of the animal. Measuring compositional changes in the microbiome is crucial in identifying the functional relationships between these physiological changes. Caenorhabditis elegans has emerged as a powerful host system to examine the molecular drivers of host-microbiome interactions. With its transparent body plan and fluorescent-tagged natural microbes, the relative levels of microbes within the gut microbiome of an individual C. elegans animal can be easily quantified using a large particle sorter. Here we describe the procedures for the experimental setup of a microbiome, collection, and analysis of C. elegans populations in the desired life stage, operation, and maintenance of the sorter, and statistical analyses of the resulting datasets. We also discuss considerations for optimizing sorter settings based on the microbes of interest, the development of effective gating strategies for C. elegans life stages, and how to utilize sorter capabilities to enrich animal populations based on gut microbiome composition. Examples of potential applications will be presented as part of the protocol, including the potential for scalability to high-throughput applications.
Introduction
Animal evolution is under constant microbial influence1. From diverse microbes in the environment, animal hosts acquire specific partners2 that extend the capabilities of the host and drive its physiology and susceptibility to disease3. For example, metagenomic analyses of the gut microbiome uncovered enriched metabolic classes of microbial genes that may confer greater energy harvest and storage in obese mice4, many of which are also found in the human gut microbiome5. There is still a great need to establish causal relationships and pinpoint the molecular determinants of the microbiome impact, though progress has been hampered by the microbiome complexities and tractability of host systems to large-scale screening.
The model organism C. elegans provides a platform to advance molecular understanding of links between microbiome and host physiology. C. elegans possesses 20 intestinal cells with a mucosal layer and villi structures. These cells are equipped with abundant chemoreceptor genes that sense microbial products and produce antimicrobial molecules that potentially regulate their gut colonizers6,7. This conserved biology of C. elegans has led to a tremendous number of discoveries in host signaling that regulate gut microbes, including insulin signaling, TGF-beta, and MAP Kinase8,9,10.
C. elegans utilize microbes as both their diet for growth during development and microbiome as adults. With old age, some microbes may over-accumulate in the gut lumen and the host-microbe relationship shifts from symbiosis to pathogenesis11. In their natural habitats, C. elegans encounters a wide array of bacterial species12,13. Sequencing 16S rDNA from representative samples collected in natural habitats (rotten fruits, plant stem, and animal vectors) revealed that the natural microbiome of C. elegans is dominated by four bacterial phyla: Proteobacteria, Bacteroidetes, Firmicutes, and Actinobacteria. Within these divisions lies great variation in the diversity and richness of bacteria based on the habitat12,13,14,15. Several defined communities have been established, including the 63-member (BIGbiome)16 and 12-member (CeMbio) collections representing the top microbiome genera created for the C. elegans research community17. Both microbiomes and component strains can have a diverse impact on the physiology of C. elegans such as body size, growth rates, and stress responses9,16,17. These studies provide resources and examples to establish C. elegans as a model for microbiome research.
Here a large particle sorter (LPS) based workflow (Figure 1) is presented that utilizes the C. elegans system to simultaneously measure microbiome composition and basic measures of host physiology at the population scale. From the microbial side, the workflow is adaptable to assemble a defined microbiome or single microbes to test the robustness and plasticity of the community with increasing microbial interactions. From the host side, the workflow enables high throughput assays to measure colonization levels of fluorescent microbes in the microbiome and host physiological readout in terms of development, body size, and reproduction. Taken together, the C. elegans microbiome model enables high throughput screens to pinpoint the metabolic and genetic determinants modulating host physiology.
Protocol
1. Preparation of microbiome mixture
- Stamp or streak out bacteria from a glycerol freezer stock onto a lysogeny broth (LB) plate, or appropriate growth medium, and grow overnight at an optimal temperature based on bacterial strains of interest (typically 25 °C for C. elegans natural microbes).
- From the LB plate, use a single colony from each bacterial isolate (e.g., 12 bacteria of the CeMbio collection) to inoculate 800 µL of LB medium in separate wells of a 1 mL deep-well plate. Incubate overnight at the optimal temperature with shaking at 250 rpm.
NOTE: This protocol is optimized for use with defined natural microbiomes (e.g., CeMbio), but can be easily adjusted for individual microbes by growth under appropriate conditions or other culture vessels (tubes). - Assess the growth of each microbe by spectrophotometry. Transfer a 20 µL aliquot to 80 µL of LB and add the solution to a clear flat bottom 96-well plate. Measure the OD600 on a plate reader.
- After overnight growth, centrifuge the deep-well culture plate at 4,000 x g for 10 min to pellet bacteria and remove the supernatant by aspiration using a 96-well aspiration manifold or multichannel pipette.
- Using OD600 values, normalize the concentration of each microbe to a final OD600 of 1.0 using sterile M9 medium.
- If combining microbes, determine the amount of bacterial culture needed for the experiment and create a microbiome master mix by combining equal volumes of each bacterial strain into a tube of sufficient size (e.g., 5 mL or 15 mL tubes).
- Spot 30-50 µL of the microbes with a final OD600 of 1.0 onto the center of each nematode growth medium (NGM) agar-containing plate or well18. Allow the seeded microbiome to grow overnight at an optimal temperature (25 °C here).
NOTE: Prepare NGM agar plates in advance according to the instructions in the Wormbook or using pre-mixed NGM powder18. For example, 3 mL of NGM is added for 12-well plates, and 1.5 mL is added for 24-well plates.
2. Preparation of synchronized C. elegans for growth on the microbiome
- Grow approximately 100 worms on NGM plates (6 cm diameter) seeded with Escherichia coli OP50, until worms reach gravid adulthood. For the purposes of this protocol, the C. elegans N2 strain was used, though the protocol can easily be adapted to other nematode strains.
- Add 6 mL of M9 buffer (plus 0.01% triton X-100; M9-TX) to the plate, pipetting up and down to wash off approximately 100 gravid adult worms into a 15 mL conical tube.
- Prepare freshly made bleach solution by mixing two parts of bleach with one part of 5 M NaOH.
- Centrifuge the worms at 3,000 x g for 30 s to pellet the worms. Aspirate to bring the volume down to 4 mL, and then add 2 mL of bleach solution.
- Close the lid tightly and shake the tube. Monitor the adult worms in the solution under a stereo microscope at 10x magnification. When adult bodies start to break apart, typically after 3.5 min, stop shaking and centrifuge at 3,000 x g for 30 s.
- Aspirate to a minimal volume with a pipette. Resuspend, centrifuge, and aspirate four times in M9-TX to wash out the bleach.
- Resuspend in 4-6 mL of M9-TX and place the tube on a rotator overnight, allowing eggs to hatch and synchronize at the L1 stage.
- The next day, take 10 µL of L1 worms in M9-TX from the tube and drop them on a clean Petri dish. Count the number of living L1 worms under a stereo microscope at 20x total magnification.
- Based on the L1 density, pipette the appropriate volume to drop ~50 synchronized L1 larvae on the edge of the microbiome seeded NGM plates and incubate at 20 °C. Allow worms to grow until the desired age (Day 2 or Day 3 of adulthood).
3. Collecting worm population for gut microbiome analyses
- Wash the worms off the bacterial lawn with 1-2 mL of M9-TX to a sterilized 2 mL 96-well deep plate.
NOTE: For microbial strains that create significant biofilms, shake the plate with liquid for 5 min to break up microbial debris. - Centrifuge the deep-well plate at 300 x g for 1 min to pellet the worms, and then remove the liquid using an aspirating manifold.
NOTE: Users can use a multichannel pipette to manually remove the liquid if an aspirating manifold is not available. Make sure the pipette tips do not touch the bottom of the wells to avoid the loss of worm samples. - Add 1.8 mL of M9-TX to each well of the deep-well plate using a 1.2 mL multichannel pipette. Mix by pipetting up and down a couple of times, and centrifuge at 300 x g for 1 min. Repeat this step four times to remove bacteria in the liquid.
- After the final wash, bring the volume in each well to 100 µL using the aspirating manifold.
NOTE: To separate adults from the larvae using gravity, leave the plate on the bench for 30-60 s, allowing adult worms to sink to the bottom of the well. Then, aspirate the plate to remove larvae from the suspension. - Add 100 µL of 10 mM levamisole in M9-TX to each well and allow the worms to paralyze for 5 min. After the levamisole treatment, add 4% of the bleach solution described in step 2.3 to each well for 2 min to reduce bacterial clumps, if needed. Wash with M9-TX twice following the bleach treatment and aspirate to a final volume of 100 µL after the second wash.
- Transfer worms to a flat bottom 96-well plate containing 150 µL of 10 mM levamisole in M9-TX. Keep the worm density at 50-100 per well and split the worms into multiple wells if the population is too crowded.
4. Setting up the large particle sorter and autosampler
- Turn on the air compressor, computer, and LPS instrument. Check and empty the waste tank. Then, add 500 mL of bleach to the empty waste tank. Check and refill sheath and water tanks if the volume is low. Ensure that the 250 µm fluidic and optical core assembly (FOCA) is in place.
- Run the control particles for quality control. Open the LPS instrument software. On the software window, check Enable Lasers to turn on the 488 nm and 561 nm lasers.
NOTE: It is possible to skip steps 4.2-4.5 if the control particles have been run within the previous 2 weeks and FOCA has not been changed in that time. - Select Set Up > Control Particles. Select No in the prompt to go to the default settings. Vortex the control particles (10 mL dH2O and 10 mL control particles in a 50 mL conical tube) before each use and load them onto the sample port.
- When ready, click on Acquire to record 500 events. Ensure the event rate reads between 15–30 events/s.
NOTE: If the event rate is not 15–30 events/s, an engineer should be contacted to adjust machine settings. - Examine coefficient of variation (CV) for all parameters. If CV < 15%, the quality control is passed and the LPS software can be closed. If CV > 15%, run again and adjust micrometers with freshly made control particles to align the laser with the flow cell.
- Connect and turn on the autosampler. Open the autosampler instrument software. This opens the autosampler and the LPS software. On the LPS software window, go to File > New Experiment then File > New Sample. Check Enable Lasers to turn on 488 nm and 561 nm lasers if it has not been done already.
- Create templates for acquisition dot plots using time of flight (TOF), extinction (EXT), and fluorescent channels in the LPS software window. A good starting maximum number to capture all the worms is 8,000 for TOF and EXT. Fluorescence scales will vary based on each microbe. Include control worm samples grown on non-fluorescent microbe (e.g., OP50) and each fluorescent microbe alone (if mixing microbes) as controls with every experiment.
- Right-click within the body of a graph to modify the scaling. Use control animals such as Day 1 adults or synchronized L1s (or population of interest) to assist with TOF and EXT gating selection.
- On the autosampler window, select Prime. Go to File > Open Script to select the correct built-in script, and then click on OK.
- For analysis only, select the script with Acquire. For sorting, select the script with Dispense. For acquisition from 96-well plates, select the script with 96 well; for acquisition from 384-well plates, select the script with 384 well. For sample volume (40 or 100 µL), select the script with the corresponding volume.
- Go to Plate Template on the autosampler software menu to select the desired wells for analysis. Load the control samples. After the plate is loaded onto the autosampler stage and secured, press Run Plate on the autosampler window. Save the file when prompted.
NOTE: Control samples and settings can be done with a 50 mL conical tube on the LPS sample port before attaching the LP sampler. - When the acquisition is finished, click on the Store Data button on the top ribbon in the LPS software window and save the data again.
5. Analysis of C. elegans features and gut microbiome levels per animal
- Raw data is stored in three file types: .lmd, .bxr3, .txt. Load the text file labeled .txt file into the R program for analysis. Complete the steps below in R19.
- Use the package tidyverse20 as a framework for the analysis.
- Using the ggplot function (included in tidyverse) plot the LPS data using the TOF values as x-axis and EXT values as y-axis. A normal plot will show two clouds of dots. Ensure one is close to the origin and x-axis, indicating smaller particles such as larvae and small debris and the second group of particles is toward the upper right of the plot and represents adult nematodes.
- Using this grouping as a guide, choose adequate TOF and EXT cut-off values. Gating may vary by C. elegans strains on different microbes based on changes in physiology (e.g., dark vs. clear animals or changes in egg carriage may alter EXT coefficients).
- Use the function subset to separate adults and larvae by TOF and EXT gating. In the example here, use TOF > 1,200 and EXT > 1,000 settings to gate for adult animals.
- Analyze the resulting datasets for statistical differences; use the t-test function in R for this experiment.
- To determine the impact of microbiome on the size and optical density of C. elegans, compare the TOF and EXT values, respectively, between conditions. TOF is a proxy for worm size and can indicate changes in the growth rates of the animals at specific times. EXT changes are greatest between the end of development and adulthood due to the light scattered by eggs and can be used to assess changes in the timing of that transition and fecundity. Assess the proportion and distributions of the progeny if they were retained during the analyses.
- To determine changes in gut microbiome colonization levels, first normalize fluorescence values (e.g., red or green columns) by dividing with individual TOF values for each animal. As worms grow in size and become adults, the volume of their gut gets larger as well. Normalization helps to reduce artifacts from differences in sample animal size distributions. Use normalized animals to assess changes in gut microbiome colonization across conditions.
NOTE: A raw data example and corresponding R scripts can be found in the Supplementary File 1 JOVE_worm_microbiome.txt and Supplementary File 2 JOVE_worm_microbiome.r. See Figure 2 and Figure 3 for a representative example of these analyses of the host and the microbiome using two fluorescent bacteria.
6. Sorting of C. elegans animals by gut microbiome features
- Collect a C. elegans population grown on fluorescent-tagged microbes (e.g., RFP) as described in Step 3. Worms should be placed into 50 mL conical tubes with a minimum of 5 mL of sample. Aim for approximately 2,000 worms/mL including all adults and offspring.
NOTE: This can also be done in a flat bottom 96-well plate using the LP sampler. Keep the worm density at 50–100 per well.- If LP sampler has been used then turn off and disconnect the LP sampler. Reconnect the sample and purge lines to the LPS before sorting.
- Open the LPS software, go to File > New Experiment then File > New Sample. Create a dot plot with TOF on the x-axis and Extinction on the y-axis. Create another dot plot with TOF on the x-axis and Red on the y-axis. Use the same axis scale and laser settings as used previously. Check Enable Lasers to turn on 488 nm and 561 nm lasers
NOTE: If these plots have been created previously go to File > Open Experiment and File > Open Sample to load graphs.- Place the control sample tube onto the sample port. In the Acquire/Dispense dialog box click Acquire. Save the file when prompted. After enough worms have been measured to distinguish populations click Abort.
NOTE: Flow rates should be approximately 15–30 events/s. If not, adjust worm concentrations to ensure that individual animals are analyzed or sorted per droplet. In LPS, worms pass through the path of the laser in different positions (straight versus curled or bent). They can take a different amount of time to pass the laser and detector depending on their positions. - In the TOF vs Extinction dot plot draw a gate around the adult population. In the TOF vs Red dot plot draw gates around the areas with high and low red values as regions of interest for adult worms with different levels of microbiome colonization. Go to View > Gating Hierarchy and ensure that the fluorescent gates are listed under the adult population gate. After settings are optimized go to File > Save Experiment and File > Save Sample.
NOTE: In the example here, select animals based on TOF/EXT gates for adults and sort into pools that exhibit either high or low colonization with red fluorescent protein-expressing bacteria. However, any combination of parameters measured by the LPS can be used to identify the population(s) of interest.
- Place the control sample tube onto the sample port. In the Acquire/Dispense dialog box click Acquire. Save the file when prompted. After enough worms have been measured to distinguish populations click Abort.
- Prior to sorting, ensure that the proper collection apparatus has been loaded. To load the 96-well plate onto the collection apparatus, select Load Plate A > Move under the Move Stage section in the Acquire/Dispense dialog box to allow the collection stage to move into a position primed for loading a 96-well plate.
- For bulk sorting, select the appropriate tube size and click Move to load either a 15 mL or 50 mL conical tube. In the Sorting section of the Acquire/Dispense dialog box choose the sorting gate from the drop-down list of the regions created. Input the number of objects from the defined region to dispense to a collection tube directly below the flow cell nozzle. Click on the Bulk Sort button under the Acquire/Dispense dialog box.
- For dispensing to a 96-well plate, go to Setup > Plates > Calibrated Plates > 96-well Plate. Then go to View > Plate Template, select the wells to which the worms will be sorted, input the number of objects to be sorted into each well, and select the gated regions of interest. If more than one gated region will be sorted into the same plate check the box Gate Each Well. Click OK to save changes.
- After sorting numbers and locations have been assigned, click on the Fill Plate buttons from the Acquire/Dispense dialog box to initiate dispensing to a 96-well plate.
- Before filling the entire 96-well plate perform test sorting of a subset of animals or controls into a 96-well plate and analyze using a stereomicroscope to ensure appropriate numbers and desired animal populations have been sorted. Once accurate sorting has been confirmed repeat steps 6.3-6.6 for experimental samples. Click on the Store Data button on the top ribbon in the LPS software window to save data again.
NOTE: All the measurements (EXT, TOF, RFP, GFP, etc.) can also be collected for sorted animals and compared to those that were not sorted. This can allow for analysis and sorting modes to be run at the same time if desired.
- Before filling the entire 96-well plate perform test sorting of a subset of animals or controls into a 96-well plate and analyze using a stereomicroscope to ensure appropriate numbers and desired animal populations have been sorted. Once accurate sorting has been confirmed repeat steps 6.3-6.6 for experimental samples. Click on the Store Data button on the top ribbon in the LPS software window to save data again.
Representative Results
Defining adult and larvae population gates
Here, synchronized C. elegans L1s were grown on an NGM plate seeded with E. coli OP50 (Eco), a standard laboratory diet. C. elegans populations were collected for LPS analysis after 96 h or 120 h of growth at 20 °C (Figure 2A). A dot plot of extinction (EXT, a proxy of body density) versus time-of-flight (TOF, a proxy of body length) creates two visually separated clouds of animals. Each dot represents a single animal where higher EXT and TOF values are observed from adults compared to larvae (Figure 2B). These two parameters are valuable inferences for population growth and physiology. For example, a density plot of larvae TOF can visualize the distribution of larval stages. Progenies from 2-day-old adults were dominated by L1 and L2 stages with a TOF below 200, while most progenies from 3-day-old adults reached L3 and L4 stages (Figure 2C). Additionally, 2D density and box-whisker plots are useful to visualize changes in adult body size and density since values of TOF and EXT increase on Day 3 compared to Day 2 when grown on E. coli (Figure 2D–F). This relationship is typically a linear one during adulthood, but some changes in physiology may impact one feature more than another (e.g., adults without eggs may have lower EXT values without affecting TOF).
Profiling gut microbiome composition using fluorescently tagged microbes
In order to illustrate different levels of colonization, we compare a dominant colonizer of the natural C. elegans microbiome dTomato-tagged Ochrobactrum BH3 (Och) and green fluorescent protein (GFP)-tagged E. coli OP50. These two were seeded individually and in an equal mixture based on OD (1:1 mix) on the NGM plate. Synchronized C. elegans L1s were grown on the three conditions and collected at 120 h to examine colonization dynamics in 3-day-old adults (Figure 3A). 2D density and box-whisker plots showed that there are differences in adult TOF and EXT values when C. elegans is grown on Ochrobactrum BH3 and mix cultures (Figure 3B–E). Gut colonization of bacteria can be inferred by the fluorescence level detected in the individual nematodes. Box-whisker plots of red fluorescent readings show increased Ochrobactrum BH3 colonization in the mix condition than in Ochrobactrum BH3 alone. In contrast, green-fluorescent values indicate lower OP50 colonization in the mix condition than in OP50 alone. Similar trends are observed in TOF-normalized fluorescent signals, which removes the effect of body size and reduces variation within the population (Figure 3F–I). For a defined microbiome with multiple fluorescent-tagged microbes, a dot plot can illustrate colonization patterns for these microbes. For example, in the two-member mix microbiome, a dot plot of red fluorescent protein (RFP) versus GFP channels shows that worms are heavily skewed toward the y-axis (RFP), suggesting OP50 colonization is low in most worms while levels of Ochrobactrum BH3 colonization are evenly distributed in the population (Figure 3J). Similarly, a dot plot of RFP versus EXT can reveal the relationship between Ochrobactrum BH3 colonization levels and host development such as body density (Figure 3K). In addition, the differences in reproduction patterns can be observed by plotting the density plot of the respective larvae population on the three conditions (Figure 3L).
Enrichment for targeted populations based on microbiome colonization
The 3-day-old adults grown on the two-member mix microbiome exhibit a wide range of RFP intensity, indicating individual variations in Ochrobactrum BH3 colonization within the group (Figure 4A). To further separate these sub-groups, sorting gates for high and low RFP were manually drawn to sort 15 individuals from each gate into a 96-well plate, as shown by the RFP image of the whole well (Figure 4B). Under higher magnification, overlay images of bright fields and RFP channels confirm that the sorting method selects high and low Ochrobactrum BH3-colonized worms, which allows for further characterization of phenotypic consequences and molecular drivers (Figure 4C).
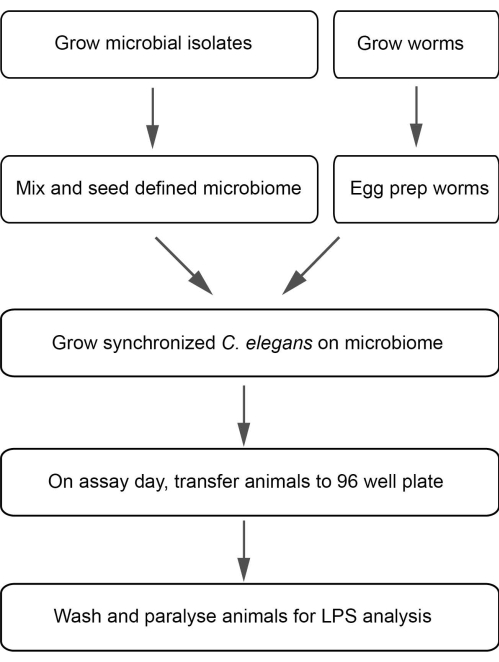
Figure 1: A flow chart for flow vermimetry-based methods to assess gut microbiome and host physiology. Please click here to view a larger version of this figure.
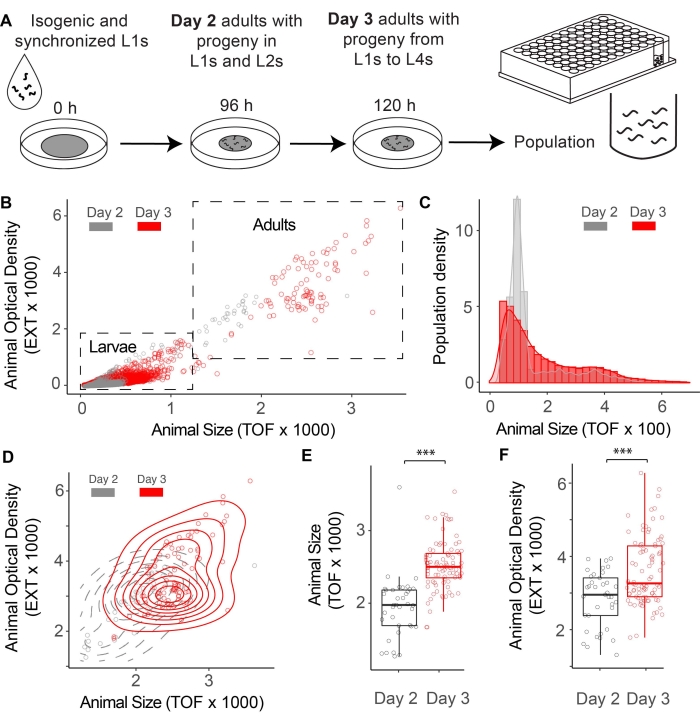
Figure 2: Defining adult and larvae populations. (A) A workflow to collect C. elegans populations on Day 2 and Day 3 of adulthood. (B) Dot plot of time-of-flight (TOF) versus extinction (EXT) for C. elegans populations on Day 2 (Gray) and Day 3 (Red) of adulthood. (C) Density plot of larvae TOF on Day 2 and Day 3 of adulthood. (D–E) Dot and box-and-whisker plots of TOF versus EXT for adults on Day 2 and Day 3 of adulthood. (F) Box-and-whisker plot of adult extinction on Day 2 and Day 3 of adulthood. P-values were calculated with the student's t-test (*** p < 0.001; Day 2 n = 38; Day 3 n = 88). Please click here to view a larger version of this figure.
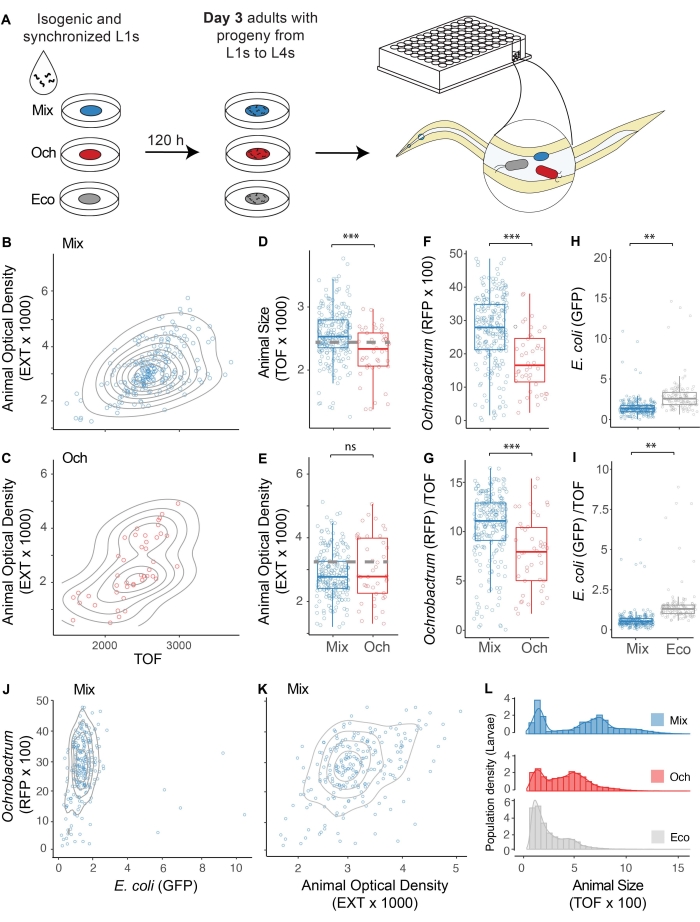
Figure 3: Profiling gut microbiome colonization using fluorescent-tagged microbes. (A) Collection of 3-day-old adult populations grown on Ochrobactrum BH3 (dTomato-expressing; Och), E. coli OP50 (GFP-expressing; Eco) or a 1:1 mix of the two bacteria (mix). (B) Dot plot of TOF versus EXT for Day 3 adults grown on the Mix. (C–E) Dot and box-and-whisker plots of TOF versus EXT for 3-day-old adults grown on Ochrobactrum BH3. Gray dash lines indicate the mean value of the population grown on OP50. (F–G) Box-and-whisker plot of raw and TOF-normalized dTomato (RFP) values for 3-day-old adults grown on Mix and Ochrobactrum BH3 alone. (H–I) Box-and-whisker plots of raw and TOF-normalized GFP for 3-day-old adults grown on mix and OP50. (J) Dot plot of Ochrobactrum BH3 (RFP) versus E. coli OP50 (GFP) for 3-day-old adults grown on mix. (K) Dot plot of RFP versus EXT for 3-day-old adults grown on mix. (L) Density plot of larvae from 3-day-old adults grown on mix, Ochrobactrum BH3, and OP50. All P-values were generated from student's t-test (*** p < 0.001; ** p < 0.01; n.s., not significant; mix n = 230; Och n = 45). Please click here to view a larger version of this figure.
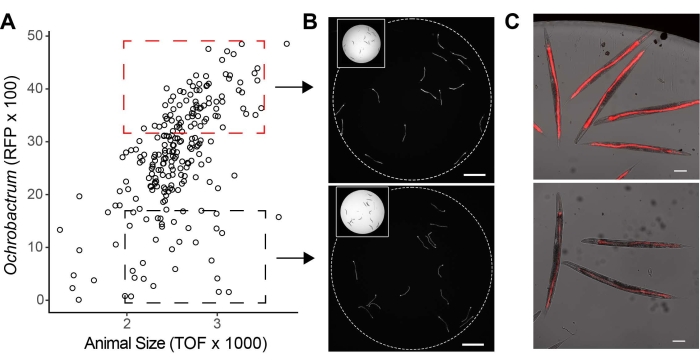
Figure 4: Enrich targeted populations based on microbiome colonization. (A) Dot plot of TOF versus RFP (Ochrobactrum BH3) for 3-day-old adults. High (red box) and low (black box) RFP gates are drawn to enrich C. elegans with high and low Ochrobactrum BH3 colonization. (B) Representative RFP images (4x) of the wells containing 15 sorted worms from high and low RFP gates (Bar = 1 mm). (C) Representative images (10x) of individual worms from high and low RFP gates (Bar = 100 µm). Please click here to view a larger version of this figure.
Supplementary File 1: Representative dataset generated by large particle sorter for N2 populations in Day 2 and Day 3 adulthoods grown on E. coli OP50 and Ochrobactrum BH3. Please click here to download this File.
Supplementary File 2: Scripts used in the analysis and figure generation of representative dataset in the R environment. Please click here to download this File.
Discussion
Flow vermimetry has been used to characterize C. elegans genes and pathways against pathogen colonization and toxicity in several studies21,22. Here, a high throughput amenable approach is presented that uses C. elegans to investigate how intestinal microbiomes modulate their host physiology. Compared to existing methods using colony forming units (CFU) or 16S rRNA amplicon sequencing9,16,17,23,24, this approach does not require labor-intensive counting or introduce potential PCR bias. In addition to measuring microbiomes in a whole population, this approach allows users to visualize individual variations within the population. This approach is limited only by the availability of fluorescent protein expression or staining of microbes and the number of detection channels in the LPS. Additional considerations for experimental design are discussed below so that users can create a customized workflow based on their needs.
Considerations for creating and seeding a defined microbiome
First, each bacterial culture is normalized to an OD600 value of 1 before mixing different strains. It is worth noting that, depending on the complexity of the community and the interspecies relationships, the starting density can have various impacts on the established microbiome composition23. While OD can be easily measured by spectrophotometers, one caveat is that different bacteria will have different concentrations (e.g., the number of cells per mL) for the same OD value and should be adjusted accordingly. Second, variation in microbiome composition in lawns should be accounted for. Once the microbiome is seeded on the NGM plate (or other media), the microbiome is allowed to grow overnight at room temperature before dropping synchronized L1 C. elegans populations onto the plates. Due to variations in growth rates and interspecies interactions among bacterial strains, microbiomes grown on NGM plates or any plate with substrates for growth will differ in composition from the original seed mixture. Using a peptone-free NGM plate can preserve the original microbial community13. However, due to limited microbiome growth on the peptone-free plate, a higher seeding OD or a reduced number of synchronized L1s per plate may be needed. This prevents the population from starving before reaching the desired age for assay. Another approach is to sequence or otherwise determine the proportion of microbes in the lawn.
Measuring microbiome colonization and C. elegans physiology
First, the extinction (EXT) and time-of-flight (TOF) gating thresholds for adults may vary based on age, C. elegans strain background, and the microbes on which they are grown. Having a 1-day-old adult population in the experiment for each worm/bacterial condition can be useful to determine the TOF and extinction cutoffs as well as to control the variation due to growth media and environment. Besides gating for adults, TOF and EXT can be applied to gate for larvae, and a density plot of the progeny can estimate the timing and output of reproduction at the population level (Figure 3L). Second, PMT gains and voltage on the LPS need to be adjusted to maximize the signal-to-noise ratio and set the signal range to avoid saturation. This is dependent upon the fluorescent intensity of the bacterial strains and their colonization levels. Default PMT at 350 works well for high colonizers such as Ochrobactrum BH3, but users may need to increase PMT or voltage values for low colonizers such as E. coli OP50. Third, due to differences in the properties of the fluorophores and their levels of expression in transgenic bacterial strains, fluorescent values do not reflect the absolute number of bacteria present in the microbiome. Therefore, GFP and RFP values cannot be used to compare colonization between two different fluorescent microbes directly. Disruption and plating of animals to identify live bacterial counts can help to better resolve these relationships24.
Potential applications for enrichment-based strategies
This method provides an amenable high throughput platform to investigate the impact of microbiome changes on host physiology at a population level. On the microbial side, the increasing availability of tools and resources to edit bacterial genomes25,26,27 will increase the number of fluorescent and functional microbial mutants related to the C. elegans natural microbiome. From the host side, numerous fluorescent reporters are available to explore the link between cell signaling and microbiome composition. The ability to enrich a host population or mutant from a mutagenized population (e.g., EMS forward mutagenesis)28 or pooled wild strains29, with specific microbiome colonization features can greatly advance the ability to connect host genes that regulate the microbiome impact. Furthermore, the selection of populations based on host and/or microbiome characteristics can facilitate a cadre of omics-based profiling modalities as well. This approach has great flexibility and promises to advance the understanding of the molecular mediators of host-microbiome interactions.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This work was supported by NIH grants DP2DK116645 (to B.S.S.), Dunn Foundation pilot award and NASA grant 80NSSC22K0250 (to B.S.S.). This project was also supported by the Cytometry and Cell Sorting Core at Baylor College of Medicine with funding from the CPRIT Core Facility Support Award (CPRIT-RP180672), the NIH (S10 OD025251, CA125123, and RR024574), and the assistance of Joel M. Sederstrom, plus an instrumentation grant for the LPS NIH grant (S10 OD025251). Some strains were provided by the CGC, which is funded by NIH Office of Research Infrastructure Programs (P40 OD010440).
Materials
| 15 mL conical bottom centrifuge tubes | VWR | 10026-076 | |
| 96 deep-well plates (1 mL) | Axygen | P-DW-11-C | |
| 96 deep-well plates (2 mL) | Axygen | P-DW-20-C | |
| 96-well Costar plate | Corning | 3694 | |
| Agar | Millipore Sigma | Standard bacteriology agar is also sufficient. | |
| Aspirating manifold | V&P scientific | VP1171A | |
| Bleach | Clorox | ||
| Bleach solution | Mix Bleach with 5M Sodium hypochlorite 2:1 (v/v) | ||
| Cell Imaging Multimode Reader | Biotek | Cytation 5 | Bacterial OD measurement |
| Centrifuge | Thermo scientific | Sorvall Legend XTR | For 96 well plate and conical tubes |
| Fluorescent Microscope | Nikon | TiE | |
| ggplot: Various R Programming Tools for Plotting Data. | R package | Version 3.3.2 | |
| Large Particle Autosampler | Union Biometrica | LP Sampler | |
| Large Particle Sorter | Union Biometrica | COPAS Biosorter | |
| Levamisole | Fisher | AC187870100 | |
| Lysogeny Broth (LB) | RPI | L24066 | Standard LB home-made recipes using Bacto-tryptone, yeast extract, and NaCl are also sufficient. |
| M9 solution | 22 mM KH2PO4 monobasic, 42.3 mM Na2HPO4, 85.6 mM NaCl, 1 mM MgSO4 | ||
| Nematode Growth Medium | RPI | N81800-1000.0 | 1 mM CaCl2, 25 mM KPO4 pH 6.0, 1 mM MgSO4 added after autoclaving. |
| RStudio | GNU | Version 1.3.1093 | |
| Sodium hypochlorite | Sigma-Aldrich | 5M NaOH | |
| Stereo Microscope | Nikon | SMZ745 | |
| Sterile 10 cm diameter petri dishes | Corning | 351029 | |
| Sterile 12-well plates | VWR | 10062-894 | |
| Sterile 24-well plates | VWR | 10062-896 | |
| Sterile 6 cm diameter petri dishes | Corning | 351007 | |
| Triton X-100 | Sigma-Aldrich | T8787 |
Riferimenti
- McFall-Ngai, M., et al. Animals in a bacterial world, a new imperative for the life sciences. Proceedings of the National Academy of Sciences of the United States of America. 110 (9), 3229-3236 (2013).
- Seedorf, H., et al. Bacteria from diverse habitats colonize and compete in the mouse gut. Cell. 159 (2), 253-266 (2014).
- Bäckhed, F., et al. The gut microbiota as an environmental factor that regulates fat storage. Proceedings of the National Academy of Sciences of the United States of America. 101 (44), 15718-15723 (2004).
- Turnbaugh, P. J., et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature. 444 (7122), 1027-1031 (2006).
- Gill, S. R., et al. Metagenomic analysis of the human distal gut microbiome. Science. 312 (5778), 1355-1359 (2006).
- Bargmann, C. I. Chemosensation in C. elegans. WormBook: The Online Review of C. elegans Biology. , 1-29 (2006).
- Couillault, C., et al. TLR-independent control of innate immunity in Caenorhabditis elegans. by the TIR domain adaptor protein TIR-1, an ortholog of human SARM. Nature Immunology. 5 (5), 488-494 (2004).
- Kim, D. H., et al. A conserved p38 MAP kinase pathway in Caenorhabditis elegans innate immunity. Science. 297 (5581), 623-626 (2002).
- Berg, M., et al. TGFβ/BMP immune signaling affects abundance and function of C. elegans gut commensals. Nature Communications. 10 (1), 1-12 (2019).
- Garsin, D. A., et al. Long-lived C. elegansdaf-2 mutants are resistant to bacterial pathogens. Science. 300 (5627), 1921 (2003).
- Cabreiro, F., Gems, D. Worms need microbes too: microbiota, health and aging in Caenorhabditis elegans. EMBO Molecular Medicine. 5 (9), 1300-1310 (2013).
- Samuel, B. S., Rowedder, H., Braendle, C., Félix, M. -. A., Ruvkun, G. Caenorhabditis elegans responses to bacteria from its natural habitats. Proceedings of the National Academy of Sciences of the United States of America. 113 (27), 3941-3949 (2016).
- Dirksen, P., et al. The native microbiome of the nematode Caenorhabditis elegans: gateway to a new host-microbiome model. BMC Biology. 14 (1), 38 (2016).
- Berg, M., et al. Assembly of the Caenorhabditis elegans gut microbiota from diverse soil microbial environments. The ISME Journal. 10 (8), 1998-2009 (2016).
- Zhang, F., et al. Caenorhabditis elegans as a model for microbiome research. Frontiers in Microbiology. 8, 485 (2017).
- Zhang, F., et al. Natural genetic variation drives microbiome selection in the Caenorhabditis elegans gut. Current biology: CB. 31 (12), 2603-2618 (2021).
- Dirksen, P., et al. CeMbio – The Caenorhabditis elegans microbiome resource. G3. 10 (9), 3025-3039 (2020).
- Stiernagle, T. Maintenance of C. elegans. WormBook: The Online Review of C. elegans Biology. , 1-11 (2006).
- R Core. Team R: A language and environment for statistical computing. R Core. , (2018).
- Wickham, H., et al. . tidyverse. , (2019).
- Anderson, Q. L., Revtovich, A. V., Kirienko, N. V. A high-throughput, high-content, liquid-based C. elegans pathosystem. Journal of Visualized Experiments. (137), e58068 (2018).
- Twumasi-Boateng, K., Berg, M., Shapira, M. Automated separation of C. elegans variably colonized by a bacterial pathogen. Journal of Visualized Experiments. (85), e51090 (2014).
- Portal-Celhay, C., Bradley, E. R., Blaser, M. J. Control of intestinal bacterial proliferation in regulation of lifespan in Caenorhabditis elegans. BMC Microbiology. 12 (1), 49 (2012).
- Zhang, F., et al. High-Throughput assessment of changes in the Caenorhabditis elegans gut microbiome. Aging: Methods and Protocols. 144, 131-144 (2020).
- Wiles, T. J., et al. Modernized tools for streamlined genetic manipulation and comparative study of wild and diverse proteobacterial lineages. mBio. 9 (5), 01877 (2018).
- Ronda, C., Chen, S. P., Cabral, V., Yaung, S. J., Wang, H. H. Metagenomic engineering of the mammalian gut microbiome in situ. Nature Methods. 16 (2), 167-170 (2019).
- Leonard, S. P., et al. Genetic engineering of bee gut microbiome bacteria with a toolkit for modular assembly of broad-host-range plasmids. ACS Synthetic Biology. 7 (5), 1279-1290 (2018).
- Kutscher, L. M., Shaham, S. Forward and reverse mutagenesis in C. elegans. WormBook: The Online Review of C. elegans Biology. , 1-26 (2014).
- Mok, C. A., et al. MIP-MAP: High-Throughput mapping of Caenorhabditis elegans temperature-sensitive mutants via molecular inversion probes. Genetica. 207 (2), 447-463 (2017).

