Visualizing Low-Abundance Proteins and Post-Translational Modifications in Living Drosophila Embryos via Fluorescent Antibody Injection
Summary
This protocol describes the customized antibody-based fluorescence labeling and injection into early Drosophila embryos to enable live imaging of low-abundance proteins or post-translational modifications that are challenging to detect using traditional GFP/mCherry-tag approaches.
Abstract
Visualization of proteins in living cells using GFP (Green Fluorescent Protein) and other fluorescent tags has greatly improved understanding of protein localization, dynamics, and function. Compared to immunofluorescence, live imaging more accurately reflects protein localization without potential artifacts arising from tissue fixation. Importantly, live imaging enables quantitative and temporal characterization of protein levels and localization, crucial for understanding dynamic biological processes such as cell movement or division. However, a major limitation of fluorescent tagging approaches is the need for sufficiently high protein expression levels to achieve successful visualization. Consequently, many endogenously tagged fluorescent proteins with relatively low expression levels cannot be detected. On the other hand, ectopic expression using viral promoters can sometimes lead to protein mislocalization or functional alterations in physiological contexts. To address these limitations, an approach is presented that utilizes highly sensitive antibody-mediated protein detection in living embryos, essentially performing immunofluorescence without the need for tissue fixation. As proof of principle, endogenously GFP-tagged Notch receptor that is barely detectable in living embryos can be successfully visualized after antibody injection. Furthermore, this approach was adapted to visualize post-translational modifications (PTMs) in living embryos, allowing the detection of temporal changes in tyrosine phosphorylation patterns during early embryogenesis and revealing a novel subpopulation of phosphotyrosine (p-Tyr) underneath apical membranes. This approach can be modified to accommodate other protein-specific, tag-specific, or PTM-specific antibodies and should be compatible with other injection-amenable model organisms or cell lines. This protocol opens new possibilities for live imaging of low-abundance proteins or PTMs that were previously challenging to detect using traditional fluorescent tagging methods.
Introduction
Immunofluorescence is a cornerstone technique of modern cell biology originally developed by Albert Coons, which enables the detection of molecules at their native cellular compartments and characterization of the molecular compositions of subcellular organelles or machineries1. Coupled with genetic manipulations, immunofluorescence helps establish the now well-accepted concept that protein localization is essential for its function2. Aside from specific primary antibodies and bright fluorescent dyes, the success of this technique relies on a preliminary process named fixation and permeabilization, which preserves cellular morphologies, immobilizes antigens, and increases the accessibility of antibodies into intracellular compartments. Inevitably, the fixation and permeabilization process would kill cells and terminate all biological processes3. Therefore, immunofluorescence only provides snapshots of the life journey of proteins. However, many biological processes such as cell migration and divisions are dynamic in nature, requiring investigation of protein behaviors in a spatial-temporally resolved manner4,5.
To examine protein dynamics in living organisms, live imaging methods based on genetically encoded fluorescent proteins such as green fluorescent protein (GFP)6 and high-speed confocal microscopes have been developed. Briefly, the protein of interest can be genetically manipulated to be fused with GFP7, and then ectopically expressed from viral or yeast promoters such as cytomegalovirus (CMV)8 or upstream activation sequence (UAS)9. Because GFP is autofluorescent in nature, no fluorophore-coupled antibodies are required to reveal the localization of target proteins, which bypasses the necessity of preliminary processes of fixation or permeabilization. Over the last two decades, fluorescent tags spanning the whole spectrum of wavelength have been developed10, enabling multi-color live imaging of several target proteins at the same time. However, compared to chemically engineered fluorescent dyes such as AlexaFluor or ATTO, the autofluorescence of these genetically encoded fluorescent proteins is relatively weak and unstable when expressed from endogenous promoters, especially during live imaging over longer time scales10. While this shortfall can be mitigated by over-expressing fluorescently tagged target proteins, many with enzymatic activities such as kinases and phosphatases severely disrupt normal biological processes if not expressed at physiological levels.
This protocol presents a method that enables photostable antibody-based target illumination in a live image setup, essentially allowing immunofluorescence without the process of fixation or permeabilization (Figure 1). Through a simple NHS-based primary amine reaction11, one can conjugate fluorescent dyes such as AlexaFluor 488 or 594 with essentially any primary antibody or GFP/HA/Myc nanobody12. Taking advantage of a developmental feature that all Drosophila embryonic cells share a common cytoplasm during the syncytium stage13, one can achieve antigen binding and illumination across entire embryos after the injection of dye-conjugated antibodies. With expanding libraries of endogenously tagged proteins available in Drosophila and other model systems14, this method can potentially broaden applications of these libraries by revealing dynamics of low-abundance fluorescently tagged proteins and other non-fluorescently tagged (HA/Myc-tagged) proteins in living tissues.
Protocol
The experiments were conducted in accordance with the guidelines and approval of the School of Life Sciences, SUSTech University. The organism used is Drosophila melanogaster, and the genotypes are Notch-Knockin-GFP (Chromosome X) and Sqh-sqh-GFP (Chromosome II), generously provided by the labs of Dr. Francois Schweisguth (Institute Pasteur) and Dr. Jennifer Zallen (Sloan Kettering Institute), respectively. While this protocol mainly focuses on aspects of antibody labeling and live imaging, please refer to published reports for more detailed descriptions of Drosophila embryo collection and injection15,16.
1. Fluorescent labeling of antibodies
- Preferably, use monoclonal antibodies or nanobodies for the protein of interest. Prepare the stock concentration of antibodies at 1 mg/mL or higher.
NOTE: In this study, GFP nanobody (see Table of Materials) is used as an example. The commercially available GFP nanobody is packaged at a concentration of 1.0 mg/mL and a volume of 250 µL. Additionally, a commercially available antibody labeling kit (see Table of Materials) is utilized, which conjugates Alexa Fluor 594 to primary amines of proteins through a succinimidyl ester reaction11. Importantly, this labeling process does not significantly alter the antibody concentration. - Prepare a 1 M sodium bicarbonate solution by resuspending Component B (provided in the antibody labeling kit) in 1 mL of deionized water.
- Adjust the antibody concentration to 1.0 mg/mL, then add 1/10th volume (10 µL for 100 µL GFP nanobody) of the 1 M sodium bicarbonate solution.
- Add 110 µL of the antibody mix (from step 1.3) directly to the tube containing Alexa Fluor 594 dye. Invert to mix (do not vortex) and incubate on a rotator for 1 h at room temperature.
- Assemble the purification column from the conjugation kit (see Table of Materials). Prepare a 1.5 mL resin bed (provided in the antibody labeling kit) and centrifuge the column at 1100 x g for 3 min at room temperature (RT) to remove excess liquid from the resin.
- Add the reaction mix (from step 1.4) dropwise to the top of the resin column and centrifuge at 1100 x g for 5 min at 4 °C to collect the labeled antibody. The collected antibody should appear pink in color, and the volume should be slightly less than 100 µL. Store in foil-wrapped or dark-colored tubes at 4 °C.
2. Preparation of Drosophila embryos
- Place 200 newly hatched adult Drosophila in a plastic cylinder-shaped cage (diameter = 5 cm, height = 8 cm) with a male-to-female ratio of 1: 10. Seal one side with porous metal mesh for ventilation and the other side with a fruit-juice/yeast-paste agar plate.
- Change the plate every 12 h for three days before embryo collection. This allows synchronization of Drosophila egg laying and improves collection efficiency.
- On the day of injection, change the cage to a fruit-juice plate with minimal yeast paste and collect embryos at 25 °C for 1 h. If the first round of collection does not yield sufficient embryos (<50 embryos), discard the first plate and repeat this step for a second round of collection until more than 100 embryos are laid within 1 h.
- Remove the plate from the cage to stop the egg laying and incubate at 25 °C for another 2 h to obtain embryos that are 2-3 h old. The correct stage of embryos is critical for the success of antibody injection.
- To remove the eggshell of embryos, add 2 mL of 50% bleach to the fruit-juice plate and gently dislodge embryos from the plate using a paintbrush (Figure 2A-4). Often, embryos will be laid directly on top of the yeast paste. In this case, it is acceptable to brush the yeast paste into the bleach solution to collect all the embryos.
- Incubate the floating embryos in 50% bleach for 2 min and swirl the fruit-juice plate every 30 s to improve the efficiency of eggshell removal.
- Transfer the embryo/bleach mix by pouring it into a 70 µm nylon cell strainer (Figure 2A-7). Wash the embryos thoroughly using a water bottle to remove any residual bleach and yeast paste. Dry the cell strainer completely with paper towels, ensuring that any excess water around the embryos is removed.
3. Embryo alignment and desiccation
- Prepare a 5 cm x 5 cm x 0.5 cm (width, length, height) 4% agarose gel (Figure 2A-9) and allow the gel to cool down to room temperature. Cut a 1 cm x 3 cm x 0.5 cm (width, length, height) agarose block using a clean razor blade and place the block on a glass slide (Figure 2A-2). Examine the gel block under the dissecting scope to ensure that the edges are cut straight, and the surface is flat and dry.
- Transfer embryos from the strainer onto the gel block using a paintbrush. Spread the embryos evenly along the midline of the block by brushing gently. Ideally, clusters of embryos are separated into single ones or doublets without touching each other.
- Prepare a tweezer with two legs taped tightly together to create a single fine tip (Figure 2A-5). Under a dissecting scope, pick up individual embryos using the tweezer and place them along the long edge of the gel block. Align 20-30 embryos, ensuring their anterior-posterior axis is parallel to the edge (Figure 2C).
- Pipette 10 µL heptane glue (see Table of Materials) at the center of a 24 mm x 50 mm coverslip (Figure 2A-1) and spread the glue into a 0.5 cm x 3 cm rectangular area using a pipette tip. Wait for the glue to completely dry before use.
- Place the glass slide with the gel block at the edge of the desk, with embryos facing outward. Lift the coverslip with glue using a flat-tip tweezer (Figure 2A-6) and hold it still on top of the embryos, with the glue side facing toward the embryos at a tilted angle of around 45 degrees.
NOTE: Gently press the coverslip against the gel block so that the embryos will be in contact with the glue, and then quickly release the pressure and lift the coverslip. The amount of tension applied is critical so that embryos can be stably attached to the glue without being pressed too hard and bursting. - Pipette 10 µL water at the center of a new glass slide and place the coverslip with the embryos on top of it, with the side of the embryo facing upwards (Figure 2B). Ensure to use water instead of nail polish or other adhesive glue for attaching the coverslip to the glass slide, as this side of the coverslip will be placed directly on top of the confocal lens for live imaging.
- Dry the embryos in a desiccation chamber (Figure 2A-3) (see Table of Materials) for 10-15 min until the vitelline membrane wrinkles (Figure 2E). The required time may vary with the humidity of the room and should be experimentally tested for each lab condition.
- Pipette 40 µL of halocarbon oil (a mix of type 27 and type 700 at a 1: 1 ratio in volume, see Table of Materials) at one end of the embryo strip. Tilt the slide until the halocarbon oil covers the entire surface of the embryos.
4. Antibody Injection and imaging
- Prepare a few injection glass needles using a micropipette puller (see Table of Materials for parameter settings) and load each needle with 5 µL of AlexaFluor-labeled antibody solution (from step 1.6).
- Place a 25 mm x 25 mm coverslip on top of a glass slide. Pipette 40 µL of halocarbon oil and spread it along the edge of the coverslip.
- Under the injection scope, align the tip of the needle against the edge of the coverslip under oil. Adjust the injection air pressure of the picopump (see Table of Materials) so that one pump generates one bubble filled with antibodies. The bubble diameter should be limited to 20-50 µm (Figure 2F).
- Replace the empty glass slide with one containing embryos under oil. Align the injection needle tip perpendicularly against the anterior-posterior axis of the embryos (Figure 2D,G).
- Check the stage of the embryos to ensure that most have initiated cellularization but have not yet started gastrulation (stage 4 and stage 5), remaining as a syncytium (Figure 2F,G).
NOTE: The hallmark of this stage is the absence of any grooves or folds and the presence of an oval-shaped, dark-colored yolk sac visible at the center of the embryo. Morphological characterization of the embryo stage under oil is based on the book chapter "Stages of Drosophila Embryogenesis" by Volker Hartenstein17. - Use the X-Y stage manipulator to move embryos toward the needle until the tip arrives at the center of the yolk. Pump two to three times to inject antibodies into the yolk. Successful injection is indicated by the quick disappearance of the vitelline membrane wrinkle as embryos gain volume from the antibody solution (Figure 2G).
- Use the X-Y stage manipulator to move embryos away from the needle after injection and move downward to the next embryo. Repeat step 6 until all embryos on the slide are injected.
- Transfer the glass slide with injected embryos to a humidity chamber (Figure 2A-8). Incubate at 25 °C until the desired stage of embryo development is achieved.
- Lift the 24 mm x 50 mm coverslip off the glass slide and place it directly into the slide holder of the microscope. The side without embryos will be facing the objectives. Locate the embryos using a low-magnification lens under bright field illumination and switch to a 40x or 63x lens for high-resolution fluorescent live imaging.
Representative Results
To demonstrate the advantages of the antibody injection method over fluorescent-tag-based live imaging or immunofluorescence, two case studies are provided that characterize the dynamic localization of a low-abundance transmembrane receptor, Notch, and a type of post-translational modification called tyrosine phosphorylation in living embryos.
Notch signaling activity plays a major role in cell fate determination during embryogenesis and adult organ homeostasis18,19. Upon activation by its ligands Delta/Jagged20, the intracellular domain of the transmembrane receptor Notch is cleaved and released into the nucleus21, initiating downstream transcriptional programs to drive cell fate changes22. The static localization of the Notch receptor has been well-characterized by immunofluorescence in formaldehyde-fixed tissues. However, the dynamic localization of Notch during ligand binding or the intracellular cleavage process remains largely unknown23, due to the lack of a method for live imaging this relatively low-abundance protein in a high-speed manner. Here, we injected AlexaFluor-conjugated GFP nanobody into embryos expressing GFP-tagged Notch from the endogenous locus20. Without injection, Notch-GFP is barely detectable under standard live imaging conditions, and the fluorescent signal quickly bleaches during time-lapse imaging. After injection, the signal-to-noise ratio of the Notch receptor significantly improves, comparable to the signal quality of immunofluorescence (Figure 3A). Moreover, antibody injection allows for the temporal characterization of Notch localization at 45 s intervals, without an apparent loss of signal intensity over a 5 min imaging window (Figure 3B).
Tyrosine phosphorylation is a major type of post-translational protein modification that mediates signal transduction in many biological pathways24. Highly specific monoclonal antibodies (such as PY20 and 4G10) against phosphotyrosine (p-Tyr) have been developed to characterize the localization and levels of overall tyrosine phosphorylation using immunofluorescence and western blots25. While no fluorescent tags can track phosphorylation changes, tissues or cells need to be fixed and stained or lysed and blotted at different time points to provide snapshots of the phosphorylation status over time, to study the kinetics of tyrosine phosphorylation upon signal activation26 (e.g., growth factor treatment). The time interval of this approach is at least a few minutes long and inherently inaccurate due to the variable time required for procedures such as fixation or cell lysis.
Here, evidence is presented that the antibody injection method enables the direct visualization of the phosphorylation status in living embryos, tracking the localization and intensity changes of tyrosine phosphorylation at regular, seconds-level time intervals. AlexaFluor-conjugated PY20 antibody was injected into embryos expressing GFP-tagged myosin light chain and performed dual-color live imaging at 45 s intervals. As it was previously shown, tyrosine phosphorylation is highly enriched at tricellular junctions27, a pattern that is also recapitulated by immunofluorescence (Figure 4A). Interestingly, live imaging also revealed a novel, second population of p-Tyr signal underneath the center of the apical membrane, which is not observed using immunofluorescence (Figure 4B). Through dual-color imaging, it was found that this population of p-Tyr signal is in close vicinity to medial myosin (Figure 4B, close-ups), a subpopulation of myosin28 that is similarly only pronounced in live imaging conditions but barely detectable using immunofluorescence. In addition, the medial population of p-Tyr exhibits similar pulsatile coalescence and dissipation patterns (Figure 4C), as previously shown for medial myosin28. The identity and function of a medial subpopulation of p-Tyr are still unknown. Together, these results demonstrate that the antibody injection method could greatly complement traditional approaches to characterize behaviors of low-abundance proteins and reveal novel localization patterns that might have been disrupted during the process of immunofluorescence.
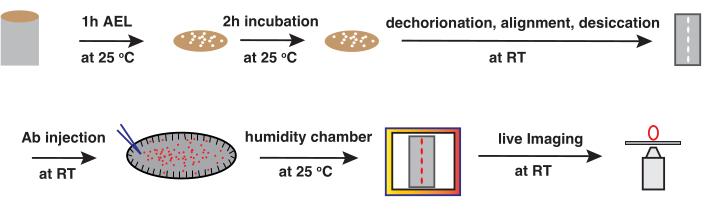
Figure 1: Antibody injection workflow. Schematic workflow illustrating the steps involved in the antibody injection method. The entire process, from embryo collection to antibody injection, typically takes around 4-5 h to complete. After antibody injection, embryos can be incubated in a humidity chamber to the desired stage of development before live imaging. AEL, after egg laying; RT, room temperature; Ab, antibody. Please click here to view a larger version of this figure.
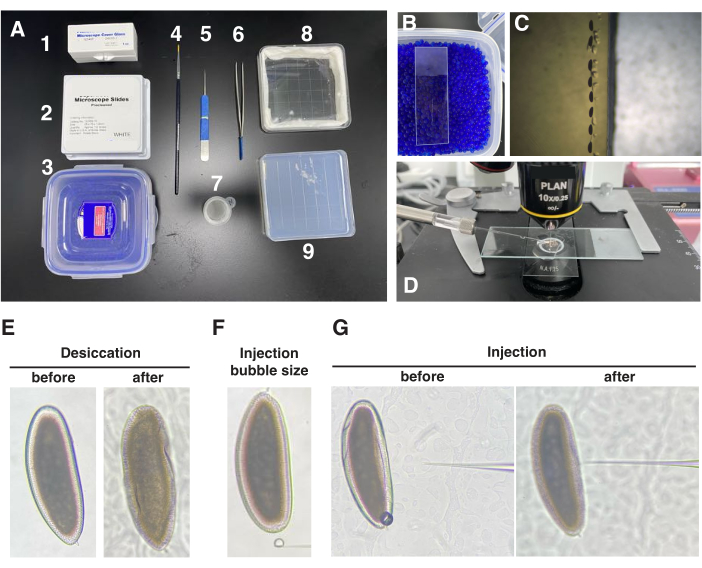
Figure 2: Embryo alignment and injection. (A) Overview of the items required before injection, including a coverslip, glass slide, desiccation box, paintbrush, tweezers, cell strainer, humidity chamber, and agarose gel. (B) Aligned embryos attached to heptane glue in the center of the coverslip and placed on top of desiccation beads. (C) Alignment of the embryos' anterior-posterior axis in parallel with the edge of the agarose gel. (D) Overview of the injection setup. (E) Comparison of embryo morphology before and after the desiccation process, with a focus on the wrinkle of the vitelline membrane after desiccation. (F) Size of the injection bubble after a single press of the picopump. (G) Comparison of embryo morphology before and after antibody injection, highlighting the disappearance of membrane wrinkles post-injection. (E–G) were captured under a 10x magnification objective lens using bright-field microscopy. Please click here to view a larger version of this figure.
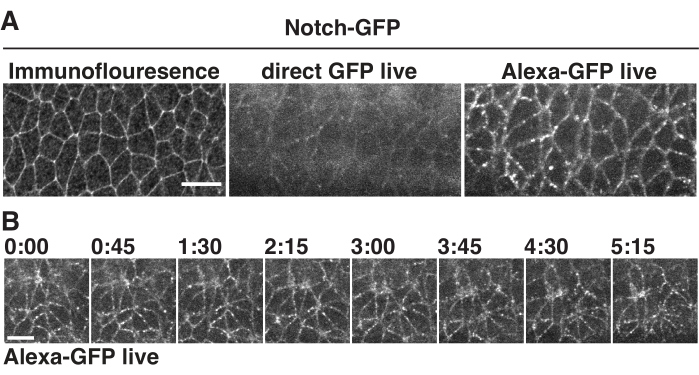
Figure 3: Live imaging of Notch receptor in early embryos. (A) Localization of endogenously GFP-tagged Notch receptor through immunofluorescence (left), direct live imaging based on GFP autofluorescence (middle), and AlexaFluor 594-conjugated GFP nanobody injection (right). (B) Dynamic localization of Notch-GFP imaged at 45 s intervals after antibody injection. All images were acquired with the anterior of the embryos to the left and the ventral side facing down. Scale bars = 10 µm. Please click here to view a larger version of this figure.
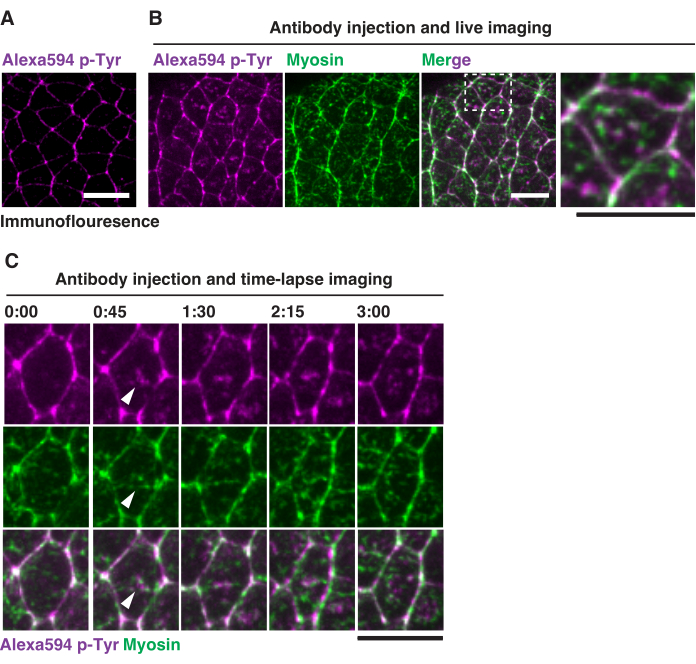
Figure 4: Live imaging of embryonic phosphotyrosine patterns. (A) Localization of phosphorylated tyrosine (p-Tyr) in fixed embryos through immunofluorescence. (B) Localization of p-Tyr (magenta) in live embryos expressing GFP-tagged myosin light chain (green). The white dashed box indicates close-up views of the medial population of phosphotyrosine and myosin under the apical membrane. Scale bars = 10 µm. (C) Localization of p-Tyr and GFP-myosin imaged every 45 s in live embryos. White arrows indicate the medial population of p-Tyr and myosin. All images were captured with the anterior of the embryos to the left and the ventral side facing down. Scale bars = 10 µm. Please click here to view a larger version of this figure.
Discussion
This presented procedure outlines the specialized method of fluorescence labeling with custom antibodies and subsequent injection into early-stage Drosophila embryos. This technique facilitates real-time visualization of proteins or post-translational modifications that exist in low quantities and are typically difficult to observe through conventional GFP/mCherry tagging methods.
Caution should be exercised when extending this method to make quantitative comparisons between wild-type and mutant embryos. While the concentration of primary and secondary antibodies can be kept the same between control and experimental groups in immunofluorescence, the amount of antibody injected and the labeling efficiency could vary between embryos. Therefore, quantitative analysis should be limited to tracking fluorescent intensity changes over time or conducting correlation analysis with signals of other channels in the same embryo when performing multi-color live imaging. For example, colocalization and correlation analysis can be performed using p-Tyr and myosin intensities27, whereas p-Tyr intensities cannot be directly compared between wild-type and gene-X knockdown embryos using the antibody injection method.
Similar to other fluorescent-tag-based approaches to examine protein dynamics, antibody binding could potentially alter the activity, trafficking, or localization of target proteins. Therefore, monoclonal antibodies or nanobodies are preferred over polyclonal antibodies for injection. Because the epitopes of monoclonal antibodies or nanobodies are well defined, whether blocking these epitopes with antibodies alters protein activity could be modeled based on alpha-fold structures12,25. On the other hand, epitopes of polyclonal antibodies are not precisely delineated, and their binding could potentially alter protein localization or activity if catalytic pockets or signal peptides of target proteins are blocked. Second, antibodies can recognize non-specific epitopes other than target proteins, especially considering that there are no "washout" steps here, as in immunofluorescence, to remove lower-affinity non-specific bindings. Therefore, it is important to have control groups, such as injecting antibodies into cells lacking the epitope, to verify whether the signal observed truly reflects the target proteins or other non-specific epitopes.
The injection of antibodies is no different from injecting siRNA or CRISPR gRNAs in terms of experimental setup. Therefore, most cellular systems that are amenable to injections should be compatible with the antibody injection method. In Drosophila embryos, Alexa Fluor-conjugated antibodies are stable during development, and fluorescent signals remain detectable after hours of embryogenesis. For other systems such as Xenopus or Zebrafish embryos, the concentration and injection volume of antibodies should be empirically tested through serial dilutions to achieve ideal labeling efficiency. In addition, alternative fluorescent dyes such as ATTO10 could potentially offer a better signal-to-noise ratio and water solubility.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
We would like to thank Dr. Jennifer A. Zallen for providing the Sqh-GFP Drosophila line and support for the initial development of this technique, and Dr. Francois Schweisguth for providing the Notch-GFP Drosophila line. This work was supported by funding from the National Natural Science Foundation of China (32270809) to H.H.Yu, generous financial and staff support from the School of Life Sciences, SUSTech, and funding to Y. Yan from Shenzhen Science and Technology Innovation Commission/JCYJ20200109140201722.
Materials
| Agarose | Sangon Biotech | A620014 | |
| Alexa Fluor 594 Antibody Labeling Kit | Invitrogen | A20185 | Purification column from step 1.6 is included in this kit |
| Biological Microscope | SOPTOP | EX20 | Eyepiece lens: PL 10X/20. Objective lens: 10x/0.25 |
| Bleach | Clorox® | ||
| Borosilicate Glass Capillaries | World Precision Instruments | TW100F-4 | |
| Centrifuge | Eppendorf | 5245 | |
| Cell Strainer | FALCON | 352350 | |
| Desiccation chamber | LOCK&LOCK | HSM8200 | 320ml |
| Dissecting Microscope | Mshot | MZ62 | Eyepiece lens: WF10X/22mm. |
| Double-sided Tape | Scotch | 665 | |
| Fine Super Tweezer | VETUS | ST-14 | |
| Fisherbrand™ Cover Glasses: Rectangles | Fisherbrand | 12-545F | |
| Fisherbrand™ Superfrost™ Plus Microscope Slides | Fisherbrand | 12-550-15 | |
| Forcep | VETUS | 33A-SA | |
| Halocarbon oil 27 | Sigma-Aldrich | H8773-100ML | |
| Halocarbon oil 700 | Sigma-Aldrich | H8898-100ML | |
| Heptane | Sigma-Aldrich | H2198-1L | Heptane glue is made of double-sided tape immersed in heptane |
| Dehydration reagent | TOKAI | 1-7315-01 | Fill to 90% volume of the dessication chamber |
| Manual Micromanipulator | World Precision Instruments | M3301R | |
| Micropipette puller | World Precision Instruments | PUL-1000 | Procedure: step 1, Heat: 290, Force:300, Distance:1.00, Delay:50. Step 2, Heat: 290, Force:300, Distance:2.21, Delay:50 |
| Pneumatic picopump | World Precision Instruments | PV 830 | Eject: 20 psi; Range: 100ms; Duration: timed |
| PY20 | Santa Cruz | SC-508 | |
| Square petri dishes | Biosharp | BS-100-SD | |
| GFP nanobody | Chromotek | gt |
Riferimenti
- Coons, A. H. The beginnings of immunofluorescence. Journal of Immunology (Baltimore, Md). 87, 499-503 (1961).
- Arthur, G., et al. Harnessing the power of the antibody. The Lancet Respiratory Medicine. 4 (3), 181-182 (2016).
- Im, K., Mareninov, S., Diaz, M. F. P., Yong, W. H. Biobanking, methods and protocols. Methods in Molecular Biology. 1897, 299-311 (2018).
- Ragkousi, K., Gibson, M. C. Epithelial integrity and cell division: Concerted cell cycle control. Cell Cycle. 17 (4), 399-400 (2018).
- Herszterg, S., Leibfried, A., Bosveld, F., Martin, C., Bellaiche, Y. Interplay between the dividing cell and its neighbors regulates adherens junction formation during cytokinesis in epithelial tissue. Developmental Cell. 24 (3), 256-270 (2013).
- Shimomura, O., Johnson, F. H., Saiga, Y. Extraction, purification and properties of aequorin, a bioluminescent protein from the luminous hydromedusan, aequorea. Journal of Cellular and Comparative Physiology. 59 (3), 223-239 (1962).
- Chalfie, M., Tu, Y., Euskirchen, G., Ward, W. W., Prasher, D. C. Green fluorescent protein as a marker for gene expression. Science. 263 (5148), 802-805 (1994).
- Boshart, M., et al. A very strong enhancer is located upstream of an immediate early gene of human cytomegalovirus. Cell. 41 (2), 521-530 (1985).
- Brand, A. H., Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development. 118 (2), 401-415 (1993).
- Rodriguez, E. A., et al. The growing and glowing toolbox of fluorescent and photoactive proteins. Trends in Biochemical Sciences. 42 (2), 111-129 (2016).
- Berg, E. A., Fishman, J. B. Labeling antibodies using N -Hydroxysuccinimide (NHS)-fluorescein. Cold Spring Harbor Protocols. 2019 (3), (2019).
- Saerens, D., et al. Identification of a universal VHH framework to graft non-canonical antigen-binding loops of camel single-domain antibodies. Journal of Molecular Biology. 352 (3), 597-607 (2005).
- Loncar, D., Singer, S. J. Cell membrane formation during the cellularization of the syncytial blastoderm of Drosophila. Proceedings of the National Academy of Sciences. 92 (6), 2199-2203 (1995).
- Lye, C. M., Naylor, H. W., Sanson, B. Subcellular localisations of the CPTI collection of YFP-tagged proteins in Drosophila embryos. Development. 141 (20), 4006-4017 (2014).
- Iordanou, E., Chandran, R. R., Blackstone, N., Jiang, L. RNAi interference by dsRNA injection into Drosophila embryos. Journal of Visualized Experiments. 50, e2477 (2011).
- Figard, L., Sokac, A. M. Imaging cell shape change in living Drosophila embryos. Journal of Visualized Experiments. 49, e2503 (2011).
- Campos-Ortega, J. A., Hartenstein, V. The embryonic development of Drosophila melanogaster. , 9-102 (1997).
- Ho, D. M., Artavanis-Tsakonas, S. The Notch-mediated proliferation circuitry. Current Topics in Developmental Biology. 116, 17-33 (2016).
- Kopan, R., Ilagan, X. G., Ma, The canonical Notch signaling pathway: unfolding the activation mechanism. Cell. 137 (2), 216-233 (2009).
- Fehon, R. G., et al. Molecular interactions between the protein products of the neurogenic loci Notch and Delta, two EGF-homologous genes in Drosophila. Cell. 61 (3), 523-534 (1990).
- Struhl, G., Greenwald, I. Presenilin is required for activity and nuclear access of Notch in Drosophila. Nature. 398 (6727), 522-525 (1999).
- Henrique, D., Schweisguth, F. Mechanisms of Notch signaling: a simple logic deployed in time and space. Development. 146 (3), dev172148 (2019).
- Trylinski, M., Mazouni, K., Schweisguth, F. Intra-lineage fate decisions involve of notch receptors basal to the midbody in drosophila sensory organ precursor cells. Current Biology. 27 (15), 2239.e3-2247.e3 (2017).
- Hunter, T. Protein kinases and phosphatases: The Yin and Yang of protein phosphorylation and signaling. Cell. 80 (2), 225-236 (1995).
- Glenney, J. R., Zokas, L., Kamps, M. P. Monoclonal antibodies to phosphotyrosine. Journal of Immunological Methods. 109 (2), 277-285 (1988).
- Hunter, T., Cooper, J. A. Epidermal growth factor induces rapid tyrosine phosphorylation of proteins in A431 human tumor cells. Cell. 24 (3), 741-752 (1981).
- Yu, H. H., Zallen, J. A. Abl and Canoe/Afadin mediate mechanotransduction at tricellular junctions. Science. 370 (6520), eaba5528 (2020).
- Martin, A. C., Kaschube, M., Wieschaus, E. F. Pulsed contractions of an actin–myosin network drive apical constriction. Nature. 457 (7228), 495-499 (2009).

