Dissection, Histological Processing, and Gene Expression Analysis of Murine Supraclavicular Brown Adipose Tissue
Summary
Here, we provide a practical procedure for dissecting and performing histological and gene expression analyses of murine supraclavicular brown adipose tissue.
Abstract
Brown adipose tissue (BAT)-mediated thermogenesis plays an important role in the regulation of metabolism, and its morphology and function can be greatly impacted by environmental stimuli in mice and humans. Currently, murine interscapular BAT (iBAT), which is located between two scapulae in the upper dorsal flank of mice, is the main BAT depot used by research laboratories to study BAT function. Recently, a few previously unknown BAT depots were identified in mice, including one analogous to human supraclavicular brown adipose tissue. Unlike iBAT, murine supraclavicular brown adipose tissue (scBAT) is situated in the intermediate layer of the neck and thus cannot be accessed as readily.
To facilitate the study of newly identified mouse scBAT, presented herein is a protocol detailing the steps to dissect intact scBAT from postnatal and adult mice. Due to scBAT's small size relative to other adipose depots, procedures have been modified and optimized specifically for processing scBAT. Among these modifications is the use of a dissecting microscope during tissue collection to increase the precision and homogenization of frozen scBAT samples to raise the efficiency of subsequent qPCR analysis. With these optimizations, the identification of, morphological appearance of, and molecular characterization of the scBAT can be determined in mice.
Introduction
The increasing prevalence of obesity in the U.S. and worldwide has ignited great interest in understanding its etiology and identifying potential treatments1,2. Adipose tissue plays a vital role in metabolism, and dysregulation of the adipose tissue can lead to the development of obesity. Generally, there are two types of adipose tissues, white and brown adipose tissue. While white adipose tissue (WAT) can store chemical energy and secrete endocrine factors, brown adipose tissue (BAT) can use chemical energy to generate heat and maintain body temperature in the cold3,4. Because of this unique ability, activation of BAT can also increase energy expenditure and improve insulin sensitivity5.
BAT exerts its function through non-shivering thermogenesis, a process mediated by uncoupling protein 1 (UCP1)6. Mammals, including mice and humans, possess varying amounts of BAT. The classical view of BAT is that these adipose tissues are more abundant in mice and infants than in adult humans. iBAT, located in the upper dorsal flank between the scapulae, is the most studied BAT depot in mice. By applying radioisotope imaging and biopsy tests, recent studies identified several BAT depots in adult humans. Some of them, including the depots found in the deep neck and supraclavicular region, had not previously been identified in mice or other model animals7,8,9,10,11. Among these BAT depots, the scBAT is the most frequently seen depot in adult humans. To better understand the origin and molecular contribution of these newly found BAT depots in humans, it is essential to identify equivalent depots in mice that allow genetic and molecular manipulations to trace and test the functional role of these depots. Thus, we and others identified a few previously unknown BAT depots in different anatomical locations in mice, including scBAT12,13, thoracic perivascular BAT14,15, perirenal BAT16, and periaortic BAT17. Mouse scBAT anatomically resembles human scBAT and morphologically resembles classical iBAT, expressing high levels of UCP112.
Unlike mouse iBAT, which can be readily dissected, scBAT is situated in the intermediate layer of the mouse neck, beneath the salivary glands and along the external jugular vein. Isolation of this depot for histological and molecular analyses can be challenging. Here, we describe in detail the procedure for dissecting scBAT from postnatal and adult mice and processing this depot for histology and gene expression analysis.
Protocol
The animal procedures were approved by the Institutional Animal Care and Use Committee at Baylor College of Medicine. All procedures were performed on C57BL/6J male mice aged 3 weeks and 3 months old. Prior to the dissection, all mice were euthanized using the approved rodent carbon dioxide euthanasia procedure. See the Table of Materials for details related to all materials, reagents, and instruments used in this protocol.
1. Dissection of scBAT
- Place the mouse carcass on the workbench and clean the surgical scissors and superfine point forceps with 70% ethanol.
- Spray the mouse's ventral neck with 70% ethanol to wet the fur and minimize fur contamination.
- Make a bilateral incision, approximately 1.5 cm, along the clavicles.
- From the ends of the previous incision, cut longitudinally towards the ears, approximately 1 cm long. This will form a squared U-shaped incision around the neck (Figure 1A,B).
NOTE: scBAT lies in the intermediate layer of the neck. The superficial layer, composed of the skin, subcutaneous adipose tissue, and salivary glands, is exposed during this step. The intermediate layer, along with scBAT, contains the jugular veins and neck muscles. The deep layer of the neck, containing deep neck BAT and artery and jugular veins, is exposed by removing the skeletal muscle. - Place the mouse under a dissecting microscope and bring the exposed neck into focus (Figure 1C).
NOTE: This added step greatly increases the precision of tissue collection, necessary given the small size of scBAT. - Expose the salivary glands fully by lifting the incised skin sections away with forceps.
- Using surgical scissors and forceps, disconnect the tissue connecting the salivary glands to the tissue around the clavicles and to each other.
- Expose the scBAT depots by lifting the salivary glands. Look for the BAT along the external jugular vein and possibly connected to the underside of the salivary glands. Identify it by its orangish color, which stands out against the surrounding tissue.
- Hold the target adipose tissue with forceps and gently peel it away from the salivary glands.
- Once detached from the salivary glands, continue to disconnect the scBAT from the external jugular vein and the musculature of the neck until the depots on either side of the neck can be removed whole.
- Inspect each dissected scBAT sample under the dissecting microscope (Figure 1D,E), using forceps to remove any remaining connective tissue.
- Place each sample into a glass scintillation vial containing 10 mL of phosphate-buffered saline solution (PBS) and remove the mouse from under the microscope.
2. Processing and hematoxylin and eosin (H&E) staining of scBAT (illustrated in Figure 2A)
- Replace the PBS (step 1.12) with 10 mL of cold 4% paraformaldehyde (PFA) and place the vial on a nutator, rocking at 4 °C overnight to fix the scBAT.
NOTE: Perfusion fixation may be applied in postnatal, especially adult mice, when further processing for immunohistochemistry analysis is needed. - The next day, remove the PFA solution from the vial with a sterile transfer pipette and replace it with 10-15 mL of PBS. Place the vial on a nutator and rock for 30 min at room temperature. Replace PBS with 0.85% saline solution and continue rocking for another 30 min at room temperature.
NOTE: The saline solution is 0.85% saline (NaCl) w/v in DEPC-treated water (RNase-free). PBS can be substituted for the saline solution in this and the following step. - Remove the 0.85% saline with a transfer pipette and add 10 mL of 70% Ethanol/0.85% saline solution to the vial. Store the vial in a 4 °C fridge overnight or until ready for paraffin embedding.
NOTE: PBS can be used if 0.85% saline is not available. scBAT should be processed as soon as possible for easier sectioning and better-preserved tissue morphology. - Prior to paraffin embedding, dehydrate scBAT through a series of ethanol exchanges to remove water from the tissue.
- To begin, remove 70% ethanol/0.85% saline from the vial with a transfer pipette and add 10-15 mL of 95% Dehydrant Alcohol, rocking the vial on a nutator at room temperature for 1 h. Repeat this step one time.
NOTE: Ethanol can be used in place of dehydrant alcohol solutions. - Next, replace 95% Dehydrant Alcohol with 10-15 mL of 100% Dehydrant Alcohol and continue rocking on a nutator for 1 h. Refill with new 100% Dehydrant Alcohol and continue to rock for several hours or overnight, depending on the quantity of scBAT in the vial.
- To begin, remove 70% ethanol/0.85% saline from the vial with a transfer pipette and add 10-15 mL of 95% Dehydrant Alcohol, rocking the vial on a nutator at room temperature for 1 h. Repeat this step one time.
- To clear scBAT for embedding, add 10 mL of toluene to the vial and continue rocking on a nutator until scBAT is transparent, approximately 6-8 h.
NOTE: The duration of time necessary to achieve satisfactory clearing is dependent on the size of the scBAT. It is recommended to begin clearing immediately in the morning so that infiltration and embedding can take place in the afternoon. - To embed scBAT in paraffin wax, remove toluene and add 10 mL of melted infiltration paraffin wax to the vial. Place the vial in an oven at 65 °C for 1 h. Repeat this step with new wax.
NOTE: Begin melting wax pellets in an oven overnight before embedding begins to ensure the wax is fully liquid. - Replace infiltration paraffin wax with embedding paraffin wax and place the vial at the same temperature for 1 h. Repeat this step once.
NOTE: The infiltration stage can be interrupted and continued later. Remove the vial from the oven and store it at room temperature until ready to continue. - Embed the tissue by first placing it in a tissue embedding mold, add melted embedding wax to the mold, and reorient the tissue under a stereomicroscope. Then, place an embedding cassette on top of the wax and continue to pour embedding wax over the top of it until it is full and the embedding cap is fastened to the wax block. Transfer the block to a 4 °C fridge overnight to let the wax stiffen and shrink before removing the metal mold.
- Section the scBAT with a microtome to a thickness of 5-6 µm18, three to four sections per microscope slide, and place in a warm paraffin section floatation bath at ~42 °C to allow the tissue sections to smooth out.
- Transfer the sections to slides and place the slides upright against the floatation bath to allow water to drip off the slides.
- Transfer the slides to a stainless staining rack and place the rack onto a slide drying bench to dry overnight.
NOTE: The paraffin slides can be stored long-term at room temperature before further processing is achieved. - Perform the H&E staining19. To begin, place the slides in a staining rack, then place the rack in a staining jar with xylene for 40 s. Repeat this step once.
NOTE: If paraffin is still visible after the two stages, wait until it has completely dissolved before continuing. - To rehydrate the tissue, place the slide rack in a staining jar filled with 100% Dehydrant Alcohol for two 20 s stages, followed by a 15 s stage in 95% Dehydrant Alcohol, and a final 15 s rinse in ddH2O.
- Place the slides in a staining dish filled with hematoxylin solution for 75 s to achieve nuclear staining, then rinse the slides in a staining dish with ddH2O for 45 s.
NOTE: Time can be adjusted depending on the desired stain color. - Differentiate the staining by dipping slides in a staining dish filled with HCl-ethanol solution for 15 s, then transfer slides to another ddH2O rinse for 15 s.
NOTE: The HCl-ethanol solution is prepared according to a published method19. - For cytoplasmic staining, place the slide in an Eosin Y counterstain solution for 15 s.
- Dehydrate the tissue by dipping the slides for 15 s each in three sequential 95% Dehydrant Alcohol solution stages, then two sequential 100% Dehydrant Alcohol baths-the first for 15 s and the second for 45 s-to finish dehydration.
- Finally, transfer the slides to a xylene bath for 45 s to reacclimate the tissue to organic solvents. After this step, the staining is completed; remove the tissue from the solution.
- Apply mounting medium to each slide and coverslip it inside a chemical fume hood. Dry overnight before imaging. Imaging results are shown in Figure 2B.
3. Gene expression analysis of scBAT
- Tissue Collection (grinding)
- After dissecting scBAT, immediately put the tissue into a microcentrifuge tube, poke a hole in the top of the tube with a sterile needle, and snap freeze in liquid nitrogen.
NOTE: The major steps of the protocol outlined here are illustrated in Figure 3A. Poking a hole in the top of the tube before dropping it into liquid nitrogen prevents the tube from bursting due to the sudden pressure change. - Fill a plastic beaker with liquid nitrogen and leave a pestle and thin spatula to soak.
NOTE: It is important to keep the temperature of all tools and tissue samples as cold as possible, close to the temperature of liquid nitrogen, during the entire procedure to prevent the tissue from thawing. Thawed adipose tissue is very adherent and makes the powdering more challenging. Only remove tools and samples from liquid nitrogen baths immediately before use. - Take one snap-frozen tissue sample at a time and pour it from its microcentrifuge tube into the mortar.
- Add a small amount of liquid nitrogen to the mortar, then use the pestle to grind the frozen tissue until it is completely pulverized.
NOTE: If the tissue starts becoming soft or sticky at any point, it is thawing; pour more liquid nitrogen into the mortar. - Use the thin spatula to carefully scrape and transfer the pulverized tissue back into the microcentrifuge tube. Pour liquid nitrogen into the mortar while scraping to gather leftover bits of tissue before transferring with a spatula. Repeat the transfer steps until all the tissue is removed from the mortar.
- Clean the mortar with a light-duty tissue wiper and 70% ethanol before dumping the next sample to grind. Store the powdered tissue in a -80 °C freezer long-term.
- After dissecting scBAT, immediately put the tissue into a microcentrifuge tube, poke a hole in the top of the tube with a sterile needle, and snap freeze in liquid nitrogen.
- Total RNA isolation
- Preparation
- Clean the workbench, pipettes, tip holders, and tube racks with both 70% ethanol and surface decontaminant to remove RNase.
- Run the centrifuge machine to reach 4 °C.
- Warm the elution solution to 70 °C.
- Dilute the DNase I by mixing 5 μL of reconstituted DNase I with 75 μL of DNase dilution solution per sample. Place the DNase I solution on ice until use.
- Procedure
- For each sample, label two microcentrifuge tubes, two column holding tubes (two capless graduated microcentrifuge tubes), and one binding mini column.
- Add 500 μL of RNA isolation solution to each sample and sonicate using a pellet pestle motor for 30 s.
- Get a syringe for each sample and pipet up and down 10x to further lyse the tissue. Ensure that no solids are visible after syringing.
- Add 250 μL of chloroform and vortex occasionally while waiting 5 min for the samples to incubate at room temperature.
- Centrifuge at 21,000 × g for 20 min at 4 °C.
- Transfer the top aqueous portion to a new microcentrifuge tube and add an equivalent amount of 70% ethanol (~500 μL).
NOTE: The top aqueous portion should be clear, the bottom should be the red RNA isolation solution, and between the two layers should be some unwanted solids. - Transfer 500 μL of the new lysate into the binding column. Centrifuge at 18,000 × g for 60 s and then discard the filtrate.
- Repeat the previous step with the remaining lysate to get all the RNA into the binding column.
- Add 700 μL of low-stringency wash to the binding column, centrifuge for 30 s, and discard the filtrate.
- Add 80 μL of diluted DNase I to each binding column. Incubate for 15 min at room temperature.
- Add 700 μL of high stringency wash, centrifuge for 30 s, and discard the filtrate.
- Add 700 μL of low stringency wash, centrifuge for 60 s, and discard the filtrate.
- Move the binding columns into the 2nd new capless column holding tube and centrifuge for 2 min to ensure all liquids are removed from the binding column.
- Move the binding columns to a new microcentrifuge tube and add 30 μL of warmed elution solution. Incubate at room temperature for 1 min.
- Centrifuge for 2 min to collect the eluted RNA at the bottom of the microcentrifuge tube.
- Measure the total RNA concentration using a spectrophotometer.
NOTE: If total RNA purity is low, indicated by a 260/280 value below 2.0 or 260/230 value below 1.8, then after step 3.2.2.12., the samples can be washed once more with low stringency wash followed by being washed with 80% ethanol before continuing with step 3.2.2.13. - Store the RNA in a -80 °C freezer.
- Reverse transcription (RT)
- In a PCR tube strip, add 0.5-1 μg of total RNA diluted in DEPC-treated water to a total of 8 μL for each RNA sample to a different well.
NOTE: The amount of RNA used for reverse transcription can be scaled up or down according to the manufacturer's instructions. - Add 1 μL Oligo dT and 1 μL of dNTPs to each sample in the PCR tube. Ensure that the total volume is 10 μL per sample.
- Place the PCR tube strip in a thermocycler and set it to 65 °C for 5 min followed by 4 °C indefinitely.
- After 5 min at 65 °C, take out the PCR tube strip and place the tube on ice for at least 2 min. After incubation on ice, add five reagents: 4 μL of 5x First-Strand buffer, 2 μL of 25 mM MgCl2, 2 μL of 0.1 M DTT, 1 μL of RNase inhibitor, and 1 μL of reverse transcriptase to each sample. Ensure that the total volume is now 20 μL per sample.
- Place the PCR tube strip back into the thermocycler and set the program to 50 °C for 50 min, then 85 °C for 5 min, then 4 °C indefinitely.
- Once the program reaches 4 °C, take out the strip and add 1 μL of Ribonuclease H (RNase H).
NOTE: RNase H is used to remove any remaining RNA template in the RT reaction; by this step, the desired RNA should already be converted to cDNA. - Place the strip back into the thermocycler and run 37 °C for 20 min and then 4 °C indefinitely.
- Reverse transcription is complete; measure cDNA concentration using a spectrophotometer.
- Dilute the cDNA to 250 ng/μL; store the cDNA in either a -80 °C or -20 °C freezer.
- In a PCR tube strip, add 0.5-1 μg of total RNA diluted in DEPC-treated water to a total of 8 μL for each RNA sample to a different well.
- Quantitative PCR (qPCR)
- Make a spreadsheet to organize the position of every primer and sample on the 96-well qPCR plate.
NOTE: One of the primers needs to be a reference gene to normalize the expression of all other genes of interest, such as 36B4. - Make a primer master mix for each "primer + sample" combination. To each qPCR reaction well, add 3 μL of Molecular Biology Grade Water, 5 μL of qPCR enzyme master mixture, 0.5 μL of forward primer, and 0.5 μL of reverse primer, which adds up to 9 μL per reaction. Add 1 μL of cDNA per reaction to make a total of 10 μL per reaction well.
NOTE: The standard is to have three technical replicates for each "primer + sample" combination, so each master mix should be 4x of the above quantities. - Add 1 μL of cDNA for each replicate to each primer master mix. If each master mix is 4x, then pipet 4 μL of each sample cDNA into their own labeled tube.
- Finally, pipet 10 μL of each reaction mix into the 96-well qPCR plate in the positions determined by the spreadsheet.
- Seal the plate with a thick adhesive seal, and then centrifuge the 96-well plate at 450 × g at 20 °C for 2 min.
NOTE: It is critical that the plate is fully sealed to avoid samples evaporating during thermocycling. Use light-duty tissue wipers to press the seal down onto the plate, especially the edges. - Run the plate in a real-time PCR instrument. Program the PCR reaction according to the manufacturer's instructions: 2 min at 50 °C, followed by another 2 min at 95 °C, and finally 40 cycles of 15 s at 95 °C and 30 s at 60 °C. The results of qPCR gene expression analysis are shown in Figure 3B.
- Make a spreadsheet to organize the position of every primer and sample on the 96-well qPCR plate.
- Preparation
Representative Results
Unlike iBAT, which is situated in the subcutaneous layer of the back between two scapulae, scBAT is situated in the intermediate layer of the neck, extending deep between layers of skeletal muscle and the salivary gland as it grows along the external jugular vein (Figure 1A). Dissecting scBAT is not as straightforward as iBAT. Here, we provide a detailed procedure including crucial steps for dissecting intact scBAT from postnatal and adult mice (Figure 1B,C). After opening the neck using the provided protocol, a thin layer of scBAT can be identified under a dissecting microscope and peeled off from the connected salivary gland and external jugular vein with a pair of forceps (Figure 1D,E). To assess the depot's morphology, freshly isolated scBAT was processed using the provided processing procedure combined with a previously published H&E staining procedure19 (Figure 2A). As shown in Figure 2B, scBAT possesses tissue structures typical of BAT depots and is composed of many small, multilocular adipocytes in healthy postnatal and adult mice. To assess gene expression levels in scBAT, RNA was extracted from isolated scBAT depots using the provided procedures (Figure 3A). Expression levels of genes of interest can then be assessed by standard RT-qPCR methods (Figure 3A). As shown in Figure 3B, the differential expression levels of genes involved in mediating scBAT function, including Pparg, the master regulator of the BAT development; Fabp4 and Glut4, two nutrient transporters; and Ucp1 and Ppargc1a, two genes involved in thermogenesis, can be readily determined. For comparison, RNA was extracted from isolated iBAT, and the expression of the above-listed genes was assessed using the provided procedures as well (Figure 3A). The expression levels of these genes are relatively similar between these two depots. (Figure 3B).
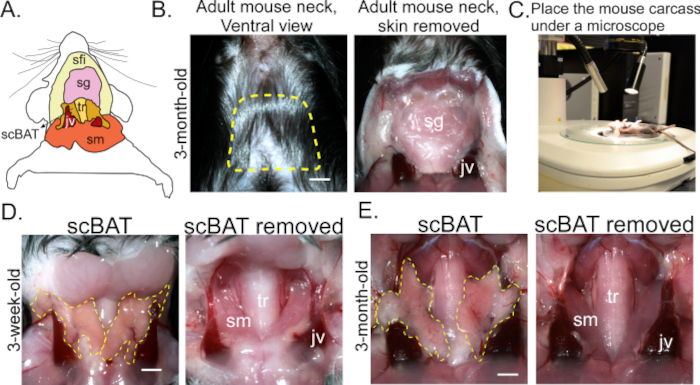
Figure 1: Anatomical location of scBAT and process of its dissection. (A) scBAT's location in the intermediate layer of the neck. (B) The superficial layer of the neck before and after the skin is removed. Scale bar = 250 μm. Dotted yellow lines outline the area that should be exposed after making the U-shaped incision. (C) Image of a mouse carcass placed beneath a dissecting microscope to increase visual clarity during dissection. (D,E) Representative images of the intermediate layer of the neck before and after scBAT is removed from mice aged 3 weeks and 3 months. Scale bar = 250 μm. Dotted yellow lines outline exposed bilateral scBAT depots; n = 2. Abbreviations: scBAT = supraclavicular brown adipose tissue; sfi = superficial layer; sg = salivary gland; tr = trachea; jv = external jugular vein; sm = skeletal muscle. Please click here to view a larger version of this figure.
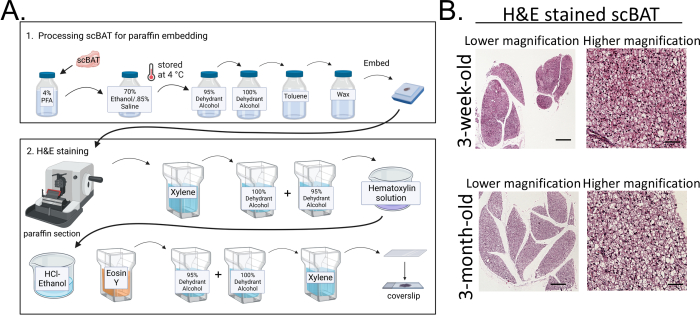
Figure 2: Processing scBAT for H&E staining. (A) Flowchart illustrating the major steps of scBAT processing for H&E staining. After dissecting the tissue, it is sequentially fixed, dehydrated, embedded, sectioned, and stained. (B) Representative images of H&E-stained scBAT from mice aged 3 weeks and 3 months; n = 3 for each developmental stage; scale bar = 250 µm for lower magnification images; scale bar = 50 µm for higher magnification images. Abbreviations: scBAT = supraclavicular brown adipose tissue; PFA = paraformaldehyde; H&E = hematoxylin and eosin. Please click here to view a larger version of this figure.
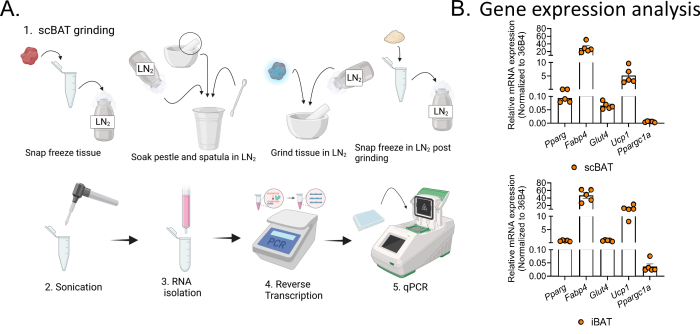
Figure 3: Preparation of RNA from scBAT for gene expression analysis. (A) Flowchart illustrating the stages of RNA isolation and RT-qPCR analysis of scBAT gene expression. Due to its small size and soft texture, snap freezing the scBAT depot and grinding it up in liquid nitrogen is crucial to successful RNA isolation. (B) Relative expression of marker genes, including Pparg, Fabp4, Glut4, Ucp1, and Ppargc1a, in scBAT and iBAT isolated from male mice aged 3 months, n = 5. Data are presented as mean ± SEM. 36B4 was used as a housekeeping gene for normalization. Abbreviations: scBAT = supraclavicular brown adipose tissue; RT-qPCR = reverse transcription-quantitative PCR; LN2 = liquid nitrogen. Please click here to view a larger version of this figure.
Discussion
In this protocol, we present in detail the procedures for dissecting and processing scBAT for H&E and gene expression analyses. Because scBAT resides in the intermediate layer of the neck and lies along the large veins, the isolation of this depot requires precise technique. Specifically, to gain a clear view of the depot, we recommend placing the mouse under a dissecting microscope after the neck has been opened. Using a pair of superfine point forceps to peel scBAT off the salivary gland and surrounding veins, care should be taken to avoid puncturing the veins. Excessive bleeding can make it more difficult to locate the scBAT. To process scBAT for H&E staining, we adapted the use of a published protocol19 with slight modifications to include the use of 4% PFA, dehydrant alcohols, and an organic solvent, toluene, for tissue processing as shown in the protocol section. The entire procedure takes ~6 days to complete, and H&E-stained slides can be imaged the next day after completion of staining.
RT-qPCR using RNA as the starting material is the most frequently used method for gene expression analysis in the field of adipose tissue biology. Because scBAT is a relatively small BAT depot compared to iBAT, obtaining sufficient RNA for gene expression can be difficult. To increase RNA yield, we recommend applying sequential lysate preparation steps as outlined in the protocol section, starting by powdering the tissue using a pestle and mortar while frozen. Using this lysate preparation method, we have successfully obtained a high-yield and high-quality RNA sample for gene expression analysis of scBAT from one adult mouse. This lysate preparation method can also be applied to obtain high-quality protein lysates from scBAT for western blotting with the use of protein lysis buffer.
Using mouse iBAT, researchers have gained substantial knowledge regarding BAT function in thermogenesis and metabolism. The recent identification of a few previously unrecognized BAT depots in mice and humans, including scBAT, revealed the need for more studies before we can fully understand the physiological contribution of BAT in mice and adult humans. Particularly, studies illuminating the origins, functions, and involvement of these newly found BAT depots in thermogenesis and regional or whole-body metabolism are warranted.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work is supported by NIDDK of the NIH under Award Number R01DK116899, USDA/ARS under Award Number 3092-51000-064-000D, and a pilot award from the Baylor College of Medicine Cardiovascular Research Institute. The flowcharts were produced using BioRender.
Materials
| 95% Dehydrant Alcohol (Flex 95) | Epredia | 8201 | |
| 100% Dehydrant Alcohol (Flex 100) | Epredia | 8101 | |
| 96-well PCR plate | Bio-Rad | MLL9601 | |
| Aurum Binding Mini Column | Bio-Rad | 7326826 | |
| Aurum High Stringency Wash | Bio-Rad | 7326803 | |
| Aurum Low Stringency Wash | Bio-Rad | 7326804 | |
| Base Molds (for embedding) | Tissue-Tek | 4122 | |
| BD PrecisionGlide Needle 21g x 1 1/2" | Becton Dickinson | 305167 | |
| C1000 Touch Thermal Cycler | Bio-Rad | 1840148 | |
| Capless Microcentrifuge Tubes 2 mL | Fisherbrand | 02-681-453 | |
| Centrifuge | Eppendorf | 5430R | |
| CFX Opus 96 Real-Time PCR Instrument | Bio-Rad | 12011319 | |
| Chloroform | Thermo Scientific Chemicals | 383760010 | |
| Cytoseal 60 Low-viscosity mounting medium | Epredia | 83104 | |
| DEPC-Treated Water | Ambion | AM 9906 | |
| Dissecting Microscope | Nikon | SMZ1500 | |
| DNase Dilution Solution | Bio-Rad | 7326805 | |
| DNase I | Bio-Rad | 7326828 | |
| dNTPs | Invitrogen | 18427013 | |
| Elution solution | Bio-Rad | 7326801 | |
| EM 400 embedding medium paraffin | Leica Biosystems | 3801320 | |
| Eosin Y (0.5% w/v) | RICCA | 2858-16 | |
| Formula R Infiltration medium paraffin | Leica Biosystems | 3801470 | |
| Genemark Nutator Gyromixer 349 | Bio Express | S-3200-2 | |
| Gill #3 Hematoxylin | Sigma-Aldrich | GHS332-1L | |
| HCl (for HCL-Ethanol) | Fisher Chemical | A142212 | |
| IP VI Embedding Cassettes | Leica Biosystems | 39LC-550-5-L | |
| Koptec's Pure Ethanol – 200 Proof (for 70% Ethanol) | Decon Labs | V1001 | |
| MgCl2 (25 mM) | Thermo Fisher Scientific | R0971 | |
| Microcentrifuge Tubes 1.7 mL | Avantor | 87003-294 | |
| Microseal 'B' Seals (adhseive seals) | Bio-Rad | MSB1001 | |
| Microtome | Leica Biosystems | RM2245 | |
| Molecular Biology Grade Water | Corning | 46-000-CM | |
| Mortar Coors Tek | Thomas Scientific | 60310 | |
| NaCl (for 0.85% saline) | Fisher Bioreagents | BP358-212 | |
| NanoDrop Spectrophotometer | NanoDrop Technologies | ND-1000 UV/Vis | |
| Oligo dT | Invitrogen | 18418020 | |
| Paraffin Section Flotation Bath | Boekel Scientific | 14792V | |
| Paraformaldehyde (PFA) | Sigma-Aldrich | P6148-500G | |
| PCR Tube Strip | Avantor | 76318-802 | |
| Pestle by Coors Tek | Thomas Scientific | 60311 | |
| Pestle Pellet Motor | Kimble | 749540-0000 | |
| Phosphate Buffer Saline (PBS) | Sigma-Aldrich | D8537-500ML | |
| Precision Model 19 Vacuum Oven | Thermo Fisher Scientific | CAT# 51221162 | |
| Primer: 36B4 (forward) 10 μM 5' TGA AGT GCT CGA CAT CAC AGA GCA 3’ |
Chen lab Oligo database | ||
| Primer: 36B4 (reverse) 10 μM 5' GCT TGT ACC CAT TGA TGA TGG AGT GT 3’ |
Chen lab Oligo database | ||
| Primer: Fabp4 (forward) 10 μM 5’ ACA CCG AGA TTT CCT TCA AAC TG 3’ |
Chen lab Oligo database | ||
| Primer: Fabp4 (reverse) 10 μM 5’ CCA TCT AGG GTT ATG ATG CTC TTC A 3’ |
Chen lab Oligo database | ||
| Primer: Glut 4 (forward primer) 10 μM 5’ CTG ATT CTG CTG CCC TTC TGT CCT 3’ |
Chen lab Oligo database | ||
| Primer: Glut 4 (reverse) 10 μM 5’ GAC ATT GGA CGC TCT CTC TCC AAC TT 3’ |
Chen lab Oligo database | ||
| Primer: PPARg (forward) 10 μM 5’ AGG GCG ATC TTG ACA GGA AAG ACA 3’ |
Chen lab Oligo database | ||
| Primer: PPARg (reserve) 10 μM 5’ AAA TTC GGA TGG CCA CCT CTT TGC 3’ |
Chen lab Oligo database | ||
| Primer: Ppargc1a (reverse) 10 μM 5' ATG TTG CGA CTG CGG TTG TGT ATG 3’ |
Chen lab Oligo database | ||
| Primer: Ppargc1a(forward) 10 μM 5' ACG TCC CTG CTC AGA GCT TCT CA 3’ |
Chen lab Oligo database | ||
| Primer: Ucp1 (forward) 10 μM 5’ AGC CAC CAC AGA AAG CTT GTC AAC 3’ |
Chen lab Oligo database | ||
| Primer: Ucp1 (reverse) 10 μM 5’ ACA GCT TGG TAC GCT TGG GTA CTG 3’ |
Chen lab Oligo database | ||
| RNA isolation solution (PureZol) | Bio-Rad | 7326880 | |
| RNase Away (surface decontaminant) | Thermo Scientific | 1437535 | |
| RNase H | NEB | M0297S | |
| Rnase inhibitor (RNase Out) | Invitrogen | 10777019 | |
| Scintillation Vial (glass) | Electron Microscopy Sciences | 72632 | |
| Slide drying bench | Electrothermal (Cole-Parmer) | MH6616 | |
| Stainless staining rack | Electron Microscopy Sciences | 70312-54 | |
| Stereo microscope (for embedding) | Olympus | SZ51 | |
| Sugical scissors | McKesson | 43-1-104 | |
| Superfine point Straight Dissecting Forceps | Avantor | 82027-402 | |
| Superfrost Plus Microscope Slides | Fisher Scientific | 12-550-15 | |
| Superscript III Reverse Transcriptase (Includes 5x First-Strand Buffer and 0.1M DTT) | Invitrogen | 18080044 | |
| SUR-VET syringe with needle 25 G x 5/8", 1 mL | Terumo | 100281 | |
| SYBR Green (qPCR enzyme master mixture) | Applied Biosystems | A25778 | |
| Tissue-Tek Manual Slide Staining Set (jars) | Electron Microscopy Sciences | SKU: 62540-01 | |
| Toluene | Fisher Chemical | T324-1 | |
| Transfer pipette | Avantor | 414004-005 | |
| Xylene | Fisher Chemical | X3P-1GAL |
References
- Boutari, C., Mantzoros, C. S. A 2022 update on the epidemiology of obesity and a call to action: as its twin COVID-19 pandemic appears to be receding, the obesity and dysmetabolism pandemic continues to rage on. Metabolism. 133, 155217 (2022).
- Hales, C. M., Carroll, M. D., Fryar, C. D., Ogden, C. L. Prevalence of obesity and severe obesity among adults: United States, 2017-2018. NCHS Data Brief. (360), 1-8 (2020).
- Berry, D. C., Stenesen, D., Zeve, D., Graff, J. M. The developmental origins of adipose tissue. Development. 140 (19), 3939-3949 (2013).
- Wang, W., Seale, P. Control of brown and beige fat development. Nat Rev Mol Cell Biol. 17 (11), 691-702 (2016).
- Maliszewska, K., Kretowski, A. Brown adipose tissue and its role in insulin and glucose homeostasis. Int J Mol Sci. 22 (4), 1530 (2021).
- Cannon, B., Nedergaard, J. Brown adipose tissue: function and physiological significance. Physiol Rev. 84 (1), 277-359 (2004).
- Cypess, A. M., et al. Identification and importance of brown adipose tissue in adult humans. N Engl J Med. 360 (15), 1509-1517 (2009).
- van Marken Lichtenbelt, W. D., et al. Cold-activated brown adipose tissue in healthy men. N Engl J Med. 360 (15), 1500-1508 (2009).
- Virtanen, K. A., et al. Functional brown adipose tissue in healthy adults. N Engl J Med. 360 (15), 1518-1525 (2009).
- Cypess, A. M., et al. Anatomical localization, gene expression profiling and functional characterization of adult human neck brown fat. Nat Med. 19 (5), 635-639 (2013).
- Leitner, B. P., et al. Mapping of human brown adipose tissue in lean and obese young men. Proc Natl Acad Sci U S A. 114 (32), 8649-8654 (2017).
- Mo, Q., et al. Identification and characterization of a supraclavicular brown adipose tissue in mice. JCI Insight. 2 (11), e93166 (2017).
- Shi, Y., et al. Gene Expression Analysis of Environmental Temperature and High-Fat Diet-Induced Changes in Mouse Supraclavicular Brown Adipose Tissue. Cells. 10 (6), 1370 (2021).
- Chang, L., et al. Loss of perivascular adipose tissue on peroxisome proliferator-activated receptor-gamma deletion in smooth muscle cells impairs intravascular thermoregulation and enhances atherosclerosis. Circulation. 126 (9), 1067-1078 (2012).
- Ye, M., et al. Developmental and functional characteristics of the thoracic aorta perivascular adipocyte. Cell Mol Life Sci. 76 (4), 777-789 (2019).
- de Jong, J. M., Larsson, O., Cannon, B., Nedergaard, J. A stringent validation of mouse adipose tissue identity markers. Am J Physiol Endocrinol Metab. 308 (12), E1085-E1105 (2015).
- Fu, M., et al. Neural crest cells differentiate into brown adipocytes and contribute to periaortic arch adipose tissue formation. Arterioscler Thromb Vasc Biol. 39 (8), 1629-1644 (2019).
- Tucker, D. K., Foley, J. F., Bouknight, S. A., Fenton, S. E. Sectioning mammary gland whole mounts for lesion identification. J Vis Exp. (125), e55796 (2017).
- Berry, R., et al. Imaging of adipose tissue. Methods Enzymol. 537, 47-73 (2014).

