High-throughput Detection Method for Influenza Virus
Summary
This method describes the use of Infrared dye based imaging system for detection of H1N1 in bronchioalveolar lavage (BAL) fluid of infected mice at a high sensitivity. This methodology can be performed in a 96- or 384-well plate, requires <10 μl volume of test material and has the potential for concurrent screening of multiple pathogens.
Abstract
Influenza virus is a respiratory pathogen that causes a high degree of morbidity and mortality every year in multiple parts of the world. Therefore, precise diagnosis of the infecting strain and rapid high-throughput screening of vast numbers of clinical samples is paramount to control the spread of pandemic infections. Current clinical diagnoses of influenza infections are based on serologic testing, polymerase chain reaction, direct specimen immunofluorescence and cell culture 1,2.
Here, we report the development of a novel diagnostic technique used to detect live influenza viruses. We used the mouse-adapted human A/PR/8/34 (PR8, H1N1) virus 3 to test the efficacy of this technique using MDCK cells 4. MDCK cells (104 or 5 x 103 per well) were cultured in 96- or 384-well plates, infected with PR8 and viral proteins were detected using anti-M2 followed by an IR dye-conjugated secondary antibody. M2 5 and hemagglutinin 1 are two major marker proteins used in many different diagnostic assays. Employing IR-dye-conjugated secondary antibodies minimized the autofluorescence associated with other fluorescent dyes. The use of anti-M2 antibody allowed us to use the antigen-specific fluorescence intensity as a direct metric of viral quantity. To enumerate the fluorescence intensity, we used the LI-COR Odyssey-based IR scanner. This system uses two channel laser-based IR detections to identify fluorophores and differentiate them from background noise. The first channel excites at 680 nm and emits at 700 nm to help quantify the background. The second channel detects fluorophores that excite at 780 nm and emit at 800 nm. Scanning of PR8-infected MDCK cells in the IR scanner indicated a viral titer-dependent bright fluorescence. A positive correlation of fluorescence intensity to virus titer starting from 102-105 PFU could be consistently observed. Minimal but detectable positivity consistently seen with 102-103 PFU PR8 viral titers demonstrated the high sensitivity of the near-IR dyes. The signal-to-noise ratio was determined by comparing the mock-infected or isotype antibody-treated MDCK cells.
Using the fluorescence intensities from 96- or 384-well plate formats, we constructed standard titration curves. In these calculations, the first variable is the viral titer while the second variable is the fluorescence intensity. Therefore, we used the exponential distribution to generate a curve-fit to determine the polynomial relationship between the viral titers and fluorescence intensities. Collectively, we conclude that IR dye-based protein detection system can help diagnose infecting viral strains and precisely enumerate the titer of the infecting pathogens.
Protocol
1. MDCK cells culture
- Culture 5 million MDCK cells overnight in a T-75 cm2 flask in 20 ml of RPMI1640 complete medium with 10% FBS, 100 U/ml penicillin, 100 μg/ml streptomycin, 1 mM sodium pyruvate, 5% of 7.5% sodium bicarbonate solution and 0.001% β-mercaptoethanol in a 37 °C incubator infused with 5.2% CO2.
2. PR8 infection and BAL fluid collection
- Challenge C57BL/6 mice intranasally with 5,000 PFU of PR8 virus in sterile PBS in a total volume of 30 μl through one nostril. Carry out mock infections using only sterile PBS without the virus.
- Euthanize mice 0, 2, 4 and 7 days after infection and cut the thoracic cavity open and make a 1 cm incision parallel to the trachea through the fur of the mouse to expose it.
- Make a midline incision on the proximal aspect of the trachea.
- Infuse the trachea with 0.5 ml of PBS-1% BSA and aspirate the BAL fluid. BAL fluids can be stored in -20 °C freezer until used.
3. Virus infection and detection of viral matrix protein (M2) with the LI-COR Odyssey
- To quantify the viral titers using infra red dye-conjugated antibodies, harvest the MDCK cells from T-75 cm2 flasks and plate them in optical flat bottom black 96-well (10,000 cells per well) or 384-well (5,000 cells per well) plates. Culture the MDCK cells for overnight at 37 °C in RPMI1640 complete medium and wash twice with serum-free DMEM containing BSA (0.2%, weight/volume), penicillin (100 U/ml), streptomycin (100 μg/ml), sodium pyruvate (1 mM), sodium bicarbonate solution (5% of 7.5%) and β-mercaptoethanol (0.001%).
- Incubate the MDCK cells with BAL fluid (50 μl for 96-well plate or 10 μl for 384-well plate) or PR8 virus with known titers in each well with equal volume (50 μl for 96-well plate or 10 μl for 384-well plate) of DMEM medium, containing L-1-tosylamido-2-phenylethyl chloromethyl ketone (TPCK)-treated trypsin (0.2 μg/ml).
- Following 1 h of infection, add 100 μl (96-well plate) or 20 μl (384-well plate) of 10% FBS-containing RPMI1640 complete medium to each well.
- Continue culturing MDCK cells for another 16 h of infection; wash the MDCK cells twice with 100 μl (96-well plate) or 20 μl (384-well plate) of PBS-1% BSA solution and fix with 100 μl (96-well plate) or 20 μl (384-well plate) of 1% paraformaldehyde for 5 min.
- Following fixing, incubate the MDCK cells further for 30 min with 100 μl (96-well plate) or 20 μl (384-well plate) of PBS-1% BSA for blocking.
- After blocking incubate the MDCK cells for 1 h with primary antibody against M2 protein (1:1000, diluted in PBS-1% BSA). Use 50 μl per well and 20 μl of diluted antibody per well in 96-well or 384-well plates, respectively.
- Wash MDCK cells thrice with 100 μl (96-well plate) or 20 μl (384-well plate) of PBS-containing 1% BSA and incubate for 1 h with 50 μl (96-well plate) or 20 μl (384-well plate) goat anti-mouse IRDye@800 secondary antibody (1:200 dilution).
- Wash MDCK cells thrice with 100 μl of PBS-containing 1% BSA. Place the stained plates on the reading glass platform of the LI-COR Odyssey IR scanner and set the reader for a 96- or 384-well plate reading.
- Blank negative control wells can be set using the LI-COR Odyssey software. Quantify whole plate or selected wells within plate using LI-COR Odyssey software.
- Using Auto Shape Tool, draw the boundaries of a target region (ROI) in the middle of a test well of 96- or 384-well plates. Creation of ROI allows the software to compare the defined background wells to the virus-titrated samples.
- To set the baseline value for the whole assay, use a background ROI. Introduce the two opposing cross hairs provided by the Odyssey software within the ROI to measure the intensity of fluorescence across the well.
- Scan plates using 780 nm channel for detection and 680 nm as reference wavelength in a LI-COR Odyssey IR scanner.
- Once the ‘Read’ command is activated, the fluorescent intensities at uniform intervals of the ROI will be measured and the collected data points integrated. A standard deviation multiplier will determine the level of signal over the baseline that is included in the ROI determination.
- Background fluorescence will be quantified from the mock-infected or secondary antibody alone controls and will be used to estimate the integrated intensity in test wells.
- Chose ‘integrated intensity’ for calculations because it represents net pixel volume for a defined individual spot and is independent of feature size. Use the standard curve from the known viral titers to calculate the viral titers of unknown samples. Use a long interpolation curve to precisely calculate the viral titers. Calculate the viral titers in the test samples by comparing the intensities of the test samples with the standard curve.
4. Representative Results
Standardization of IR dye-based high throughput viral detection system
PR8 virus can infect MDCK cells; therefore, we used these cells to develop this assay. We measured the viral titers using a primary antibody against an influenza viral matrix protein (M2). In contrast to other methodologies, we used a secondary antibody that is conjugated to a near-IR dye. This method provides a simple, automated PR8 virus titer determination using fluorescence intensity that is generated while detecting M2 protein. After influenza virus infection, M2 protein is translated, transported and accumulates in the host cell including the cell surface. Thus, the quantity of the M2 protein represents a measurable parameter to determine the quantity of the virus. Comparing the intensities from the test samples with the standard curve provides the precise measurement of viral titers.
To determine the efficacy of the fluorescence-based method, we cultured 104 MDCK cells/well in chamber slides overnight. PR8 viral stocks of known titers were added to duplicate wells with different plaque forming units (PFU) and allowed to adsorb onto the cells for one hour. Cells were incubated for 16 h to allow the virus to propagate. After this, the MDCK cells in the chamber slides were fixed with 1% paraformaldehyde, washed and stained with anti-M2 antibody for 1 h, followed by AlexaFluor 488-conjugated secondary antibody for an additional 1 h. Confocal microscopic analyses of infected MDCK cells demonstrated the adherent monolayer was largely intact and the PR8-derived M2 protein was abundantly present inside the infected MDCK cells (Figure 1A). Fluorescing cells could be detected in infections with viral titers as low as 102. These analyses also revealed that the number of fluorescing MDCK cells per well were directly proportional to the increasing PR8 viral titers. Although, the optical efficiency of argon laser-based fluorochrome excitation provides vivid cellular images, its narrow signal to noise ratio and the autofluorescence of the cells render viral quantifications extremely difficult, particularly at lower titrations.
Therefore, to establish an accurate viral estimation method we employed an IR dye-based detection and quantification system. MDCK cells (104/well) were cultured in 96-well plates, infected with PR8 and viral proteins were detected using anti-M2 followed by an IR dye-conjugated secondary antibody. The use of anti-M2 antibody allowed us to use the antigen-specific fluorescence intensity as a direct metric of viral quantity. To enumerate the fluorescence intensity, we used the LI-COR Odyssey-based IR scanner. This system uses two channel laser-based IR detections to identify fluorophores and differentiate them from background noise. The first channel excites at 680 nm and emits at 700 nm to help quantify the background. The second channel detects fluorophores that excite at 780 nm and emit at 800 nm. Scanning of PR8-infected MDCK cells in LI-COR Odyssey indicated a viral titer-dependent bright fluorescence (Figure 1B). M2 positive fluorescent MDCK cells were clearly visible inside the wells (Figure 1B). A positive correlation of fluorescence intensity to virus titer starting from 102-105 PFU could be consistently observed. PR8 viral titrations higher than 106 PFU lead to cell death, which sets an upper limit for the sensitivity of the assay. Minimal but detectable positivity consistently seen with 102-103 PFU PR8 viral titers demonstrated the high sensitivity of the near-IR dyes. The signal-to-noise ratio was determined by comparing the mock-infected or isotype antibody-treated MDCK cells. Both of these controls resulted in negligible or undetectable levels of fluorescence (Figure 1B).
To further improve the sensitivity and to reduce the test sample volume requirements, we infected 5×103 MDCK cells/well in 384-well plates. Different PFUs of PR8 were tested as serial one log dilutions (Figure 1C). Detection sensitivity from 384-well plates was comparable to that of the 96-well plate assays. Both 101 and 102 PFU worked consistently better in the 384-well plates compared to 96-well plates. Using the fluorescence intensities from 96- or 384-well plate formats, we constructed standard titration curves (Figure 1D). In these calculations, the first variable is the viral titer while the second variable is the fluorescence intensity. Therefore, we used the exponential distribution to generate a curve-fit to determine the polynomial relationship between the viral titers and fluorescence intensities. We have also validated our observations using another antibody that is directed against the nucleoprotein (NP) of PR8 virus (data not shown). Distinct sources of PR8 strains were also used to infect MDCK cells that were tested with anti-NP or anti-M2 antibodies (data not shown). Collectively, we conclude that IR dye-based protein detection system can help diagnose infecting viral strains and precisely enumerate the titer of the infecting pathogens.
Mathematical considerations for calculating viral titers
The mathematical calculations of fluorescence intensity and titer determinations are done as described below. Following the staining procedure, whole plate or selected wells within plate are quantified using LI-COR Odyssey software. Using Auto Shape Tool, boundaries of a target region (ROI) was drawn in the middle of the test wells of 96- or 384-well plates (Figure 2A and B). Creation of ROI allows the software to compare the defined background wells to the virus-titrated samples. A background ROI was used to set the baseline value for the whole assay. Two opposing cross hairs (white) were introduced within the ROI to measure the intensity of fluorescence across the well (Figure 2C). Curves to the right side and below the representative wells in Figure 2C represent the intensity of the pixels along these cross hairs. The fluorescent intensities at uniform intervals of the ROI were measured and the collected data points were integrated. A standard deviation multiplier determined the level of signal over the baseline that was included in the ROI determination. Background fluorescence was quantified from the mock-infected or secondary antibody alone controls and used to estimate the integrated intensity in test wells. We chose integrated intensity for calculations because it represents net pixel volume for a defined individual spot and is independent of feature size. In addition, integrated intensity is substantially independent of resolution. Total intensity/pixel (I) corresponds to the signal intensity arising from the selected well in the pixel area (Is) plus the signal arising from the background of the pixel area (b). Therefore, for pixel ‘i’:
Ii = Isi + bi
Pixel volume represents both the magnitude of the signal and the area in which it is distributed. Signal area is related to the distribution of sample that is generating the signal. The pixel volume is equal to total signal measured in pixel ‘i’ in the area (a) of the pixel times its height (I). So for pixel ‘i’:
vi = aIi
Total pixel volume is the summation of total signal from the entire area thus:
| n | n | ||
| V= | Σvi | = | aΣIi |
| i=1 | i=1 |
Integrated intensity is the sum of the intensity values of all pixels enclosed by feature, multiplied by the area of the circle/rectangle (count mm2). Therefore, the integrated intensity =
a(ΣIi– b )
Here, b stands for the average background pixel intensity. This formula calculates the integrated signal intensities of the control or experimental wells and thereby establishes a standard curve. Viral titers in the test samples were calculated using this standard curve. Concentration (of intensity) is defined as the quantity of fluorescence present in a defined ROI. Concentrations in test samples are calculated relative to the defined concentrations of the standards in the same image. To calculate the precise viral titers, the intensity of each concentration standard is plotted and fitted with a long interpolation curve. The concentrations of the test samples are calculated by comparing the intensity of the area within the standard curve.
Determination of viral titers from BAL fluid of influenza infected mice
Detection of live influenza viral particles in clinical and laboratory specimens is of critical significance. Therefore, we next examined whether we can utilize this method to determine the viral titers in laboratory samples. BAL fluids were collected from non-immunized mice and spiked with a known amount of PR8 virus. Spiked BAL fluids were linearly titrated for enumerating the viral titers. Aliquots of spiked BAL fluids were used to infect MDCK cells in 96-well plates, fixed and stained with anti-M2 and IR dye-conjugated secondary antibodies. Results shown in Figure 3A demonstrate that the viral titers in the spiked BAL fluid were detectable and correlated with the PR8 titers in the standard curve. Using the standard curve, precise viral numbers in the spiked and titrated BAL fluid was quantified. Exponential curve fit calculations provided a measure to calculate the viral titers in the spiked BAL fluid (Figure 3B). Through this method we obtained excellent correlations between the calculated and spiked viral titers, validating this approach (Figure 3C).
Next, we analyzed the BAL fluid from PR8-infected mice. PR8 has been extensively used in murine models to understand human pathology and anti-viral immunity3. Groups of mice were intranasally infected with 5,000 PFU of PR8. Mice were monitored for weight loss, appearance of hunched back, ruffled fur and other clinical symptoms. On days 0, 2, 4, 7 and 10 of post infection, mice were sacrificed and BAL fluids were collected. To utilize these laboratory samples, MDCK cells were incubated with serial dilutions of BAL fluid in a final volume of 50 μl for 17 h in 96-well plates. Control wells of MDCK cells infected with known titers of stock PR8 virus served to generate a standard curve. After infection, cells were washed, fixed and stained with anti-M2 primary antibody followed by near-IR dye-conjugated secondary antibody. MDCK cells that were mock-infected or treated with isotype controls after infection provided the background fluorescence intensities for titer calculations.
Analyses of the BAL fluid demonstrated that PR8 virus in the laboratory specimen is detectable through the IR dye-based assay system. A significant increase in the viral load in the BAL fluid was observed on days 2 and 4 post infection (Figure 4A). Calculations of integrated intensity indicated that BAL fluids from days 2 and 4 of post infection contained significant levels of PR8 virus (Figure 4B). Our results show a gradual but significant weight loss during the early phase of PR8 infection, demonstrating the severity of the disease. Nevertheless, most mice started recovering from disease symptoms and gaining weight 7 days post infection (Figure 4C). Mice at day 10 post infection lacked any detectable PR8 indicating the clearance of the virus, which also correlated with the increase in body weight and considerable reductions in the disease symptoms.
To further extend the application of this assay, we tested a 2009 A(H1N1) human influenza isolate with IR dye-based antibody detection system and compared it with real-time PCR assay. This clinical sample was isolated from a combined nasopharyngeal/throat swab obtained from suspected patient. The isolate was propagated in MDCK cell line at the Milwaukee Health Department Laboratory. The test results were compared to check the sensitivity of the IR dye-based method to real-time PCR assays (Figure 5). The results indicate that the IR dye-based antibody detection system has the potential to detect viral load as low as 103 TCID50/ml, which is comparable to PCR-based assay (10 TCID50/ml) and fall within the clinical relevance range for most patient samples. These results provide strong evidence that the IR dye-based assay system has sufficient sensitivity and potential to be applied for viral titer enumeration in research laboratory and clinical settings.
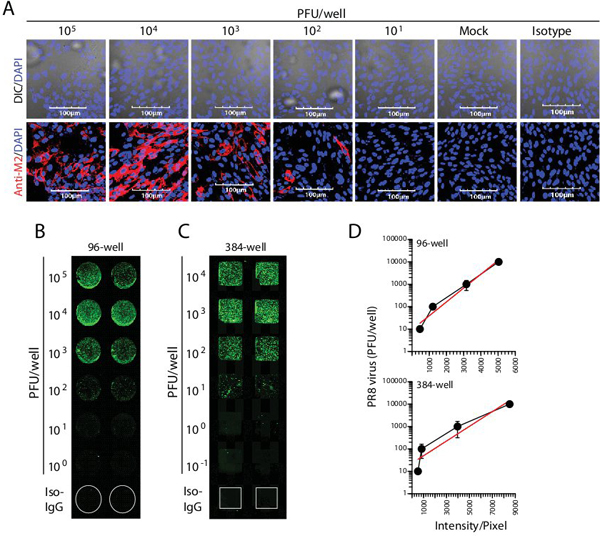
Figure 1. IR dye-based immune detection of influenza virus. (A) Confocal microscopic analyses of influenza-infected MDCK cells in 8-well chamber slides. Immunofluorescence microscopic analyses demonstrate the adherent MDCK monolayer is largely intact and loaded with virus-derived M2 protein. (B-C) Quantification of viral titers based on IR dye and LI-COR Odyssey system. (D) Generation of standard curves and exponential curve-fit profiles. Data presented in A-D are representative of five independent experiments.
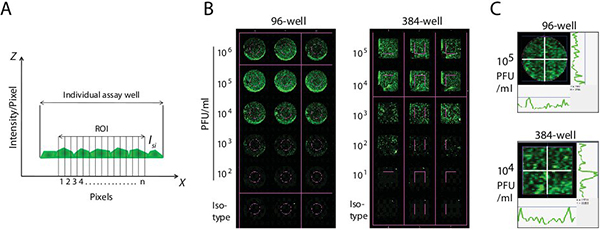
Figure 2. Methodology of determining viral titers using IR dyes. (A) Integrated fluorescence intensity measurements. A defined area inside the test wells (ROI) was marked to measure the fluorescence intensities. Using Auto Shape Tool, boundaries of the ROI were drawn in the middle of the test wells of 96- or 384-well plates. Fluorescence was measured as individual pixels (pixel=n) within the ROI. Integrated intensity is the sum of the intensity values of all pixels enclosed by feature, multiplied by the area of the circle/rectangle (count mm2). (B) Determining the integrated fluorescence intensities in assay wells. Creation of ROI allowed the LI-COR scanner to compare the defined background wells to the virus-titrated samples. Background fluorescence was quantified from the mock-infected or secondary antibody alone controls and used to estimate the integrated intensity in test wells. (C) Collection of data points within the ROI. Two opposing cross hairs (white) were introduced within the ROI to measure the intensity of fluorescence across the well. Curves to the right side and below the representative wells represent the intensity of the pixels along these cross hairs.
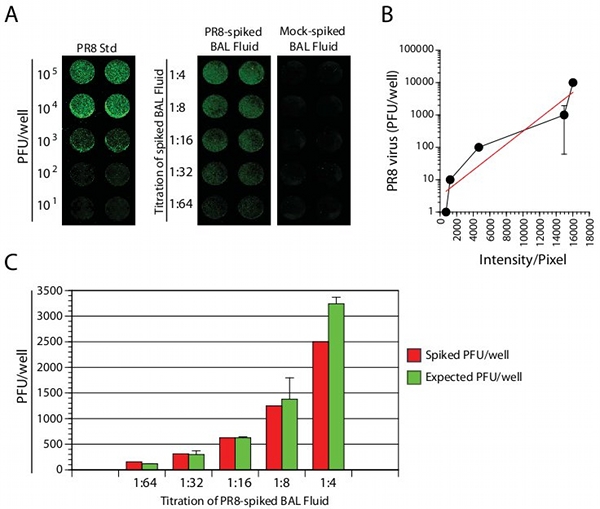
Figure 3. Detection and quantification of virus titers in PR8 spiked BAL fluids. (A) MDCK cells were cultured in 96-well plates overnight and added with PR8-spiked BAL fluid. Plates were read in the LI-COR Odyssey system. (B) Exogenously spiked BAL fluid was compared with standard curves. (C) Comparison of calculated to expected viral titers. Exponential curve fitting provided y=9.4784e0.0014x . Data presented in A-C are representative of three independent experiments.
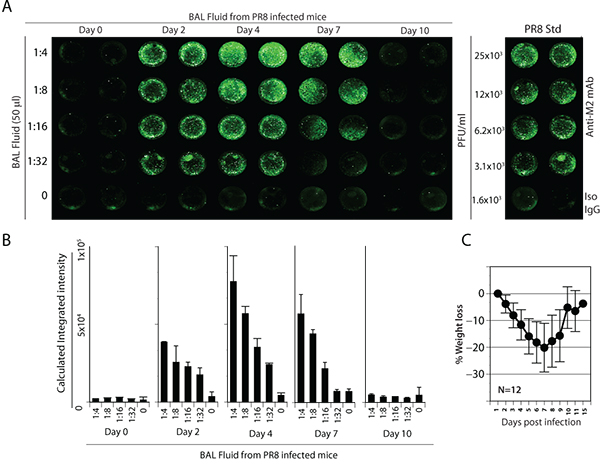
Figure 4. Detection and quantification of influenza virus in BAL fluids from PR8 infected mice. (A) Groups of mice were intranasally infected with 5,000 PFU of PR8. MDCK cells were incubated with serial dilutions of BAL fluid in a final volume of 50 μl for 17 h in 96-well plates. After infection, cells were washed, fixed and stained with anti-M2 primary antibody followed by IR dye-conjugated secondary antibody. The rightmost panel represents the standard curve. (B) Quantification of viral titers in the BAL fluid of infected mice. (C) Weight loss of mice during PR8 infection correlated with the viral load. Data presented in A-C are representative of three independent experiments.
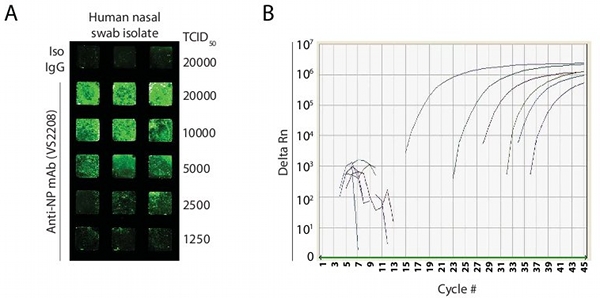
Figure 5. Detection and quantification of 2009 A(H1N1) pandemic (pdm) influenza isolate from a human patient. (A) The IR dye-based antibody fluorescence system detection for half-fold titration of infected MDCK cell line. The fluorescence results indicate the assay has potential to detect viral load as low as 103 TCID50/ml. (B) Nucleic acid extracted from culture isolate along with serial ten-fold dilutions of a known titered H1N1 isolate were analyzed with the CDC real-time PCR assay12. Limit of detection for real-time PCR was 10 TCID50/ml.
Discussion
The recent epidemic outbreak of avian influenza in Southeast Asia and a global fear of the swine flu pandemic impose enormous concerns to public health and safety. Critical focus has been dedicated in generating vaccines for existing and newer strains of influenza virus. Availability of rapid high-throughput technologies for precise detection and accurate viral titer calculations will tremendously improve treatment. The most commonly used techniques employed to calculate virus titer include plaque forming and influenza hemagglutination assays. The plaque forming assay was first developed to estimate bacteriophage titers and was adopted by Renato Dulbecco to calculate animal viral titers 4. To perform this assay, 10-fold dilutions of influenza stock or animal specimens such as BAL fluid are prepared and 0.1 ml aliquots are inoculated onto the monolayer of MDCK cells in 6-well plates. The virus is allowed to adsorb onto the MDCK cells followed by the addition of a nutrient medium and agarose. During the incubation, the original infected cells release viral progeny that spread only to the neighboring cells. The cytopathogenic effect of the influenza viral infection produces a circular zone of infected cells called a plaque. Each plaque is presumably formed after infection of a single cell with a single virus particle followed by replication of virus. Contrast between the living cells and the plaques can be enhanced by dyes such as crystal violet. A major drawback of this assay is lack of reproducibility and huge variations within experiments. Another assay used to determine the influenza viral titer is the hemagglutination assay. The hemagglutinin protein present on the surface of the influenza virus efficiently binds to N-acetylneuraminic acid-containing proteins on mammalian and avian red blood cells 1. This interaction between the influenza virus and the erythrocytes leads to agglutination in the form of a lattice. The level of agglutination is used to estimate the titer of the influenza virus in test specimens. Erythrocyte agglutination assay is used only to determine the titration of a known virus and cannot be used for strain identification purposes. Since this assay does not differentiate between the live and the dead viruses, it is difficult to obtain a precise calculation of viral titers.
Current clinical diagnoses of influenza infections are based on serologic testing, polymerase chain reaction, direct specimen immunofluorescence and cell culture 1,6. The new method described here also has clinical and laboratory diagnostic potential. Influenza detection kits Directigen FLU-A (BD Biosciences, San Jose, CA) and BinaxNow (Binax, Inc., Scarborough, ME) use immunochromatography assay to detect viral antigen such as nucleoprotein 7. Using these kits, detection of virus can be achieved within 15 min; however, all negative samples have to be further screened by the cell culture method. Various studies have also reported low sensitivity of these kits, particularly, when infection was mild or low 8. In our method, we consistently detect as low as 100 PFU of virus. This suggests our technique is useful in detecting H1N1 virus in clinical samples, even when the infection is mild. A fluorescence-based method to detect viral antigens has been reported, where either nasopharyngeal aspirate specimens were directly analyzed or were inoculated onto a monolayer of susceptible cells for virus multiplication and subsequent analyses 9. However, these methods are limited for diagnostic purposes and are not suitable for virus titer calculations. Q-PCR helps to identify the presence of viral RNA and does not estimate the infectivity of live viruses. Compared to these assays, IR dye-based assay system will help in the detection and viral titer enumeration in clinical and laboratory samples. In case of influenza outbreak, it is critical to diagnose thousands of samples in a short time period. Our method based on a 384-well plate and IR scanner can scan 6-plates at a time (>2,000 samples), therefore offering a greater advantage over other direct fluorescent methods. Feasibility of rapid processing of thousands of clinical samples could reduce test variability, length of assay time and costs. The use of an M2-specific antibody offers the advantage of being independent of strain differences in H and N proteins. The M2-protein is relatively conserved in most influenza strains infecting humans. If antibodies specific to circulating stains are combined with our system one can measure both the titer of virus and identify the strain. However, since M2 is the target for the Amantadine-like drugs, mutants can arise that might change the binding site for the mAb used here. This represents a potential limitation for this system in individuals who are being treated by these drugs.
Multiple assays are used to enumerate influenza titers in test samples. To calculate virus titer, 50% tissue culture infectious dose (TCID50) and egg allantoic fluid-based assays are widely used 1,10. However, requirements of multiple days make these methods less reliable. Further, in these methods calculations based on the 50% change in the number of plaques provide theoretical but not accurate viral titrations. Plaque forming assays can be performed within 96 hours; however, lack of reproducibility is a major concern in viral titer calculations. The present technique described here has several distinct advantages. First, it is a highly reproducible technique with consistent outcomes. Second, viral titer calculations are based on simple quantification of fluorescence intensities that are compared against defined standard curves. Third, two or more primary antibodies can be used to diagnose multiple serotypes in a given sample. Also, IR dye-conjugated antibodies have minimal autofluorescence and have been used for cell imaging 11. UV or visual radiation (300-600 nM) results in high cellular autofluorescence due to the presence of high quantum yielding endogenous fluorophores. These include aromatic amino acids, lipo-pigments, pyridinic nucleotides (NADPH), elastin, collagen and flavin coenzymes. This intrinsic property of cells makes it difficult to utilize these sources of radiation as probes to quantify the expression of specific proteins. In contrast, near-IR dyes excite and emit between 670-1000 nm. Many host cellular molecules and polyaromatic hydrocarbons do not excite at this wavelength and due to reduced light scattering, emit extremely low background fluorescence. Further, the window between the background and specific detection is vastly improved due to a high extinction coefficient of the near-IR fluorophores. Photostability, superior signal-to-noise ratio, is extremely relevant in the quantification of viral titers. Use of microfluidic chambers will automate the screening process through robotic controls, and could allow us to test highly contagious clinical samples with ease. Thus, the utilization of near-IR dyes in these methods is ideal and highly reliable to enumerate viral titers in tissue samples.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This work is supported in part NIH grants R01 A1064826-01, U19 AI062627-01, NO1-HHSN26600500032C (to S.M.). We thank our laboratory members for discussion and technical help, Tina Heil for the critical review of the manuscript. We would like to thank David Bina, Milwaukee Health Department Laboratory for virus culture and PCR.
Materials
| Name of the reagent | Company | Catalogue number | Comments |
| Bovine serum albumin | Atlanta Biologicals | S11750 | |
| PR8 virus | From Dr Thomas M Moran | ||
| Goat anti-mouse IRDye@800 | LI-COR biosciences | 926-32210 | 1:200 dilution |
| LI-COR Odyssey IR scanner | LI-COR-biosciences | 9201-01 | |
| 384-wells flat bottom plate | Nalge Nunc international | 164730 | |
| 96-wells flat bottom plate | Nalge Nunc international | 165305 | |
| DMEM medium | Invitrogen | 11965126 | |
| RPMI medium | Invitrogen | 11875135 | |
| Sodium bicarbonate | Invitrogen | 25080 | |
| L-glutamine 100x | Invitrogen | 25030 | |
| Trypson-EDTA | Invitrogen | R001100 | |
| Sodium Pyruvate | Invitrogen | 11360070 | |
| Anti-M-2 antibody | From Dr Thomas M Moran | ||
| Anti-NP mAb | CDC | V2S2208 | Obtained with an MTA |
Riferimenti
- Hirst, G. K. The agglutination of red cells by allantoic fluid of chick embryos infected with influenza virus. Science. 94, 22-23 (1941).
- Jonsson, N., Gullberg, M., Lindberg, A. M. Real-time polymerase chain reaction as a rapid and efficient alternative to estimation of picornavirus titers by tissue culture infectious dose 50% or plaque forming units. Microbiol. Immunol. 53, 149-154 (2009).
- Kalter, S. S. Hemagglutinating Behavior of Mouse and Egg-adapted Type A (PR8) Influenza Virus. Science. 110, 184-184 (1949).
- Dulbecco, R. Production of Plaques in Monolayer Tissue Cultures by Single Particles of an Animal Virus. Proc. Natl. Acad. Sci. U. S. A. 38, 747-752 (1952).
- Deyde, V. M., Nguyen, T., Bright, R. A., Balish, A., Shu, B., Lindstrom, S., Klimov, A. I., Gubareva, L. V. Detection of molecular markers of antiviral resistance in influenza A (H5N1) viruses using a pyrosequencing method. Antimicrob. Agents Chemother. 53, 1039-1047 (2009).
- Jonsson, N., Gullberg, M., Lindberg, A. M. Real-time polymerase chain reaction as a rapid and efficient alternative to estimation of picornavirus titers by tissue culture infectious dose 50% or plaque forming units. Microbiol. Immunol. 53, 149-154 (2009).
- Rahman, M., Kieke, B. A., Vandermause, M. F., Mitchell, P. D., Greenlee, R. T., Belongia, E. A. Performance of Directigen flu A+B enzyme immunoassay and direct fluorescent assay for detection of influenza infection during the 2004-2005 season. Diagnostic Microbiology and Infectious Disease. 58, 413-418 (2007).
- Landry, M. L., Ferguson, D. Suboptimal detection of influenza virus in adults by the Directigen Flu A+B enzyme immunoassay and correlation of results with the number of antigen-positive cells detected by cytospin immunofluorescence. J. Clin. Microbiol. 41, 3407-3409 (2003).
- Herrmann, B., Larsson, C., Zweygberg, B. W. Simultaneous detection and typing of influenza viruses A and B by a nested reverse transcription-PCR: comparison to virus isolation and antigen detection by immunofluorescence and optical immunoassay (FLU OIA). J. Clin. Microbiol. 39, 134-138 (2001).
- Reed, L. J., Muench, H. A simple method of estimating fifty percent endpoints. Am. J. Epidemiol. 27, 493-497 (1938).
- Doerr, A. Fluorescent proteins: into the infrared. Nat. Meth. 6, 482-483 (2009).
- Shu, B., Wu, K. H., Emery, S., Villanueva, J., Johnson, R., Guthrie, E., Berman, L., Warnes, C., Barnes, N., Klimov, A., Lindstrom, S. Design and performance of the CDC real-time reverse transcriptase PCR swine flu panel for detection of 2009 A (H1N1) pandemic influenza virus. J. Clin. Microbiol. 49, 2614-2619 (2011).

