Procedure for Decellularization of Porcine Heart by Retrograde Coronary Perfusion
Summary
A method to rapidly and completely remove cellular components from an intact porcine heart through retrograde perfusion is described. This method yields a site specific cardiac extracellular matrix scaffold which has the potential for use in multiple clinical applications.
Abstract
Perfusion-based whole organ decellularization has recently gained interest in the field of tissue engineering as a means to create site-specific extracellular matrix scaffolds, while largely preserving the native architecture of the scaffold. To date, this approach has been utilized in a variety of organ systems, including the heart, lung, and liver 1-5. Previous decellularization methods for tissues without an easily accessible vascular network have relied upon prolonged exposure of tissue to solutions of detergents, acids, or enzymatic treatments as a means to remove the cellular and nuclear components from the surrounding extracellular environment6-8. However, the effectiveness of these methods hinged upon the ability of the solutions to permeate the tissue via diffusion. In contrast, perfusion of organs through the natural vascular system effectively reduced the diffusion distance and facilitated transport of decellularization agents into the tissue and cellular components out of the tissue. Herein, we describe a method to fully decellularize an intact porcine heart through coronary retrograde perfusion. The protocol yielded a fully decellularized cardiac extracellular matrix (c-ECM) scaffold with the three-dimensional structure of the heart intact. Our method used a series of enzymes, detergents, and acids coupled with hypertonic and hypotonic rinses to aid in the lysis and removal of cells. The protocol used a Trypsin solution to detach cells from the matrix followed by Triton X-100 and sodium deoxycholate solutions to aid in removal of cellular material. The described protocol also uses perfusion speeds of greater than 2 L/min for extended periods of time. The high flow rate, coupled with solution changes allowed transport of agents to the tissue without contamination of cellular debris and ensured effective rinsing of the tissue. The described method removed all nuclear material from native porcine cardiac tissue, creating a site-specific cardiac ECM scaffold that can be used for a variety of applications.
Protocol
1. Tissue Preparation and Experiment Setup
- Harvest porcine organ immediately after euthanasia from an abattoir or research facility and rinse off excess blood. Trim the heart of excess fat and tissue, keeping the atria and aorta intact. Trim away fat to separate the pulmonary artery from the aorta. If there are any cuts in the tissue, discard appropriately.
- Wrap each heart individually in freezer paper and store all tissue in a -80 °C freezer for at least 24 hr to ensure complete freezing.
- When ready for use (usually less than 3 months), thaw one intact frozen porcine heart in Type 1 water overnight submerged in a 4 L beaker at 4 °C.
- After the heart is completely thawed, pat the heart dry, weigh the heart, and record the weight. The heart of a market weight pig should weigh approximately 375-450 g.
- Connect size 18 Masterflex tubing to the ¼” end of a barbed reducer. Insert the barbed reducer and tubing inside the aorta. Place 2 hose clamps or secure zip ties around the aorta, just below the brachiocephalic trunk. The reducer and tubing must remain above the aortic valve, so the coronary arteries can be perfused (Figure 1).
- Use a 30 or 60 ml syringe to fill the tubing with Type I water. Insert the tubing within the cartridge of a Masterflex roller pump at its approximate midpoint. Submerge the inflow end of the tubing in the bottom of a 4 L beaker filled with 2.5 L of water and secure the tubing.
- Place the heart in the beaker filled with water, and prime the pump to remove air bubbles. If bubbles are observed coming from the aorta where the tubing is inserted, the aorta may need to be repositioned or secured with additional ties. An airtight seal is important to maintain adequate pressure during the decellularization process (Figure 2).
- Place the 4 L beaker containing 3 L of a 0.2% Trypsin/0.05% EDTA/0.05% NaN3 solution on stir plate and warm it to 37 °C in preparation of the decellularization process.
2. Tissue Rinses
- Set the pump to a flow rate of 400 ml/min, ensuring that the correct tubing size is selected. Flush the heart with Type I water for 15-25 min. As the pump is started, the heart should swell and effuse blood from the ventricles. Fresh solution should be substituted every 5-10 min, or as needed based on the amount of blood removed from the heart. If blood is not effused from the heart, adjust the tubing and clamps as necessary.
- Stop the pump and transfer the heart to a separate beaker filled with 2X Phosphate Buffered Saline (PBS). After the tubing is submerged in solution, start the pump and increase the flow rate to 700 ml/min. The heart should remain in solution for 15 min, changing the solution every 5 min. Each solution change requires the pump to be stopped temporarily while the tissue and tubing is moved to the new beaker.
- Transfer the heart to Type I water for 10 min and increase the flow rate to 750 ml/min.
3. Decellularization and Solution Perfusion
- Transfer the heart to the beaker containing 0.2% Trypsin/0.05% EDTA/0.05% NaN3 at 37 °C. Increase the pump speed to 1,200 ml/min and start the pump. Use a stir bar placed at the bottom of the beaker to circulate solution in the beaker. The heart should remain in the 0.2% Trypsin/0.05% EDTA/0.05% NaN3 solution at 37 °C for a total of three hours. After 1 hr, increase the pump speed to 1,500 ml/min. After an additional hour, increase the pump speed to 1,800 ml/min. The tissue is slowly subjected to increased perfusion speeds to condition the tissue and prevent rupture of the vessels. The heart will swell and nearly double in size during this step of the protocol. The tissue will lose its natural color, progressing from the atria to the apex throughout the protocol (Figure 3).
- After each solution perfusion, a two step rinse is performed to remove cellular debris, chemical residue, and aid cell lysis. Each rinse consists of a 10 min rinse in Type I water followed by a 10 min rinse with 2X PBS solution at room temperature. Each wash consists of removal of solution from the original beaker, adding rinse solutions, and circulating the perfusate within the beaker containing the submerged heart. After the 0.2% Trypsin/0.05% EDTA/0.05% NaN3 solution, perfuse water at 1,900 ml/min and then perfuse 2X PBS at 1950 ml/min.
- Transfer the heart to a solution of 3% Triton X-100/0.05% EDTA/0.05% NaN3 at room temperature. Increase the pump speed to 2,000 ml/min and perfuse solution for one hour. Remove the solution from the beaker and replace with fresh solution, increase the pump speed to 2100 ml/min, and perfuse the fresh solution for an additional hour and a half, bringing the total time in 3% Triton X-100/0.05% EDTA/0.05% NaN3 to 2.5 hr.
- Rinse the tissue in Type I water at 2150 ml/min and 2X PBS at 2180 ml/min for 10 min each.
- Transfer the heart to a 4% Sodium Deoxycholate solution at room temperature. Increase the pump speed to 2,200 ml/min and perfuse solution for 3 hr.
- Rinse the tissue in Type I water at and 2X PBS at 2,200 ml/min for 15 min each, changing the solutions after 5-10 min for each solution. The described perfusion steps may be split over multiple days by performing the rinse step twice and storing the heart with attached tubing overnight at 4 °C and submerged in Type I water.
- The following day, perform a 5 min rinse with Type I water at 750 ml/min, followed by a 5 min rinse in 1X PBS at 1,500 ml/min. The protocol may then be continued at the described flow rate in the proper solution.
4. Disinfection and Final Processing
- Transfer the heart to a 0.1% peracetic acid/4% ethanol solution and perfuse solution for 1.5 hr at 2,200 ml/min.
- The final rinses for the tissue are all performed at 2,200 ml/min. Perfuse the tissue with 1X PBS for 15 min, followed by two 5 min washes in Type I water. This series of rinses is repeated once more in order to complete the solution perfusion procedure.
- Turn the pump off and remove the heart from solution to drain the heart. Cut the ties from the aorta, remove all tubing, and place the heart in an empty beaker to drain for 1 hr. Excess liquid will need to be drained periodically. Lay the heart on an absorbent pad to fully drain the heart (Figure 4).
- After most of the water is removed, record the weight of the cardiac extracellular matrix (C-ECM). The heart can be expected to lose approximately 20-25% of its initial weight during the decellularization process.
- Dissect the right and left ventricles, as well as the ventricular septum for DNA quantification and histological processing in order to confirm complete decellularization of the tissue (Figure 5).
- Freeze the C-ECM at -80 °C for at least 2 hr before lyophilization.
Representative Results
The effect of decellularization on whole porcine hearts naturally varies due to differences in size, pressures, and vessel arrangement. Therefore, the exact composition of the derived extracellular matrix scaffolds will not be the same from heart to heart. The completion of the described protocol will yield a heart that appears white or translucent, indicating the loss of cellular material. However, it is widely accepted that a tissue can be considered “decellularized” based on the combination of a few more quantitative parameters 8. A successful decellularization protocol will produce a matrix with less than 50 ng of double stranded DNA per mg of tissue (Figure 6). In order to avoid a host immune response upon implantation of the matrix, the remaining DNA should also contain less than 200 base pairs (Figure 7). To confirm these findings, Hematoxylin and Eosin staining should reveal the absence of nuclear staining in representative sections of the ventricles and ventricular septum (Figure 8). Masson’s Trichrome further confirms the loss of cardiac muscle bundles and retention of collagen networks (Figure 9).
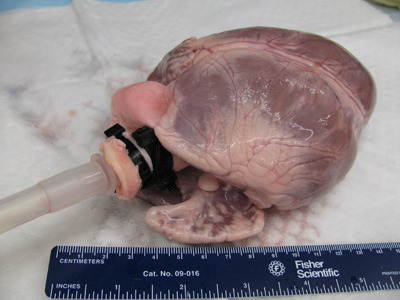
Figure 1. The barbed end of the tubing is inserted into the aorta of the native heart. The tubing must be secured with hose clamps or zip ties above the aortic valve to ensure perfusion through the coronary arteries.
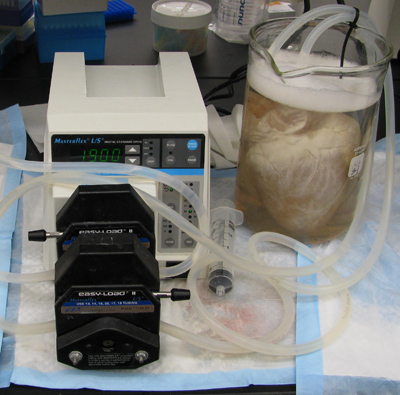
Figure 2. The heart is submerged in water in a 4L beaker and air bubbles must be removed from the tubing. If bubbles are observed emerging from the aorta near the tubing, additional ties must be used to secure the tubing to the aorta in order to maintain adequate pressure in the tissue.
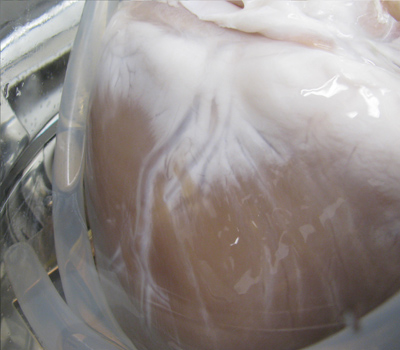
Figure 3. As solutions are perfused through the coronary arteries, the heart will lose its native color, progressing from the atria to the apex of the heart and localized around the coronaries.
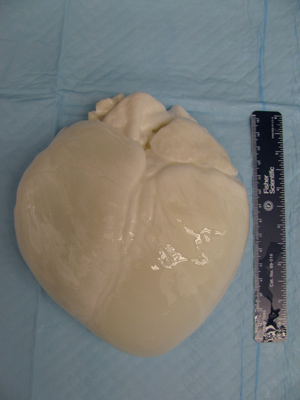
Figure 4. After completion of the disinfection and rinse steps of the protocol, the tubing is removed and the heart is placed on an absorbent pad to allow the excess water to drain out of the heart. This ensures an accurate measurement when weighing the tissue and also allows the tissue to relax before sectioning.
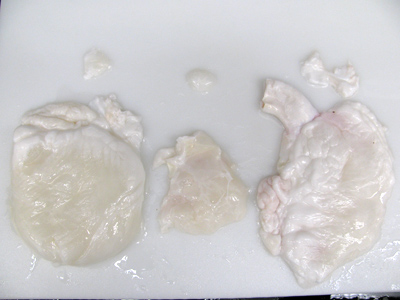
Figure 5. The left ventricle (LV), right ventricle (RV), and ventricular septum are all removed from the decellularized heart for histologic processing, freezing and lyophilization, and DNA quantification.
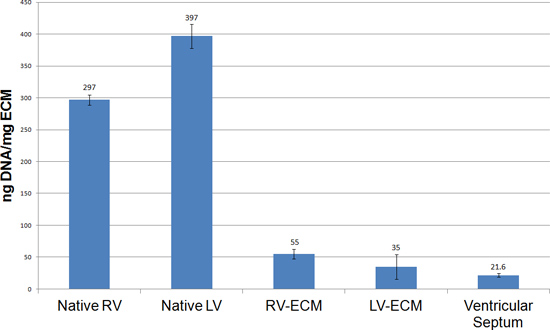
Figure 6. Quantitative analysis of DNA content using a Pico Green assay. The ventricles from cECM hearts show a significant decrease in DNA content when compared to native ventricles. The DNA values observed from this protocol are observed at or below the 50 ng/mg standard for decellularized tissues.
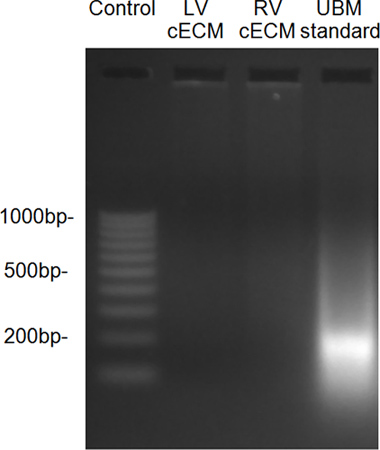
Figure 7. DNA fragment size, as determined by ethidium bromide gel, showed little residual DNA in the decellularized ventricles when compared to a urinary bladder matrix (UBM) standard.
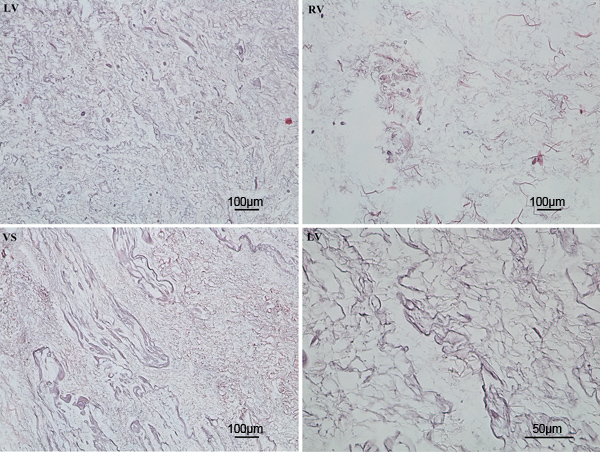
Figure 8. Hematoxylin and Eosin staining showed complete removal of nuclear material from the ventricles following completion of the decellularization protocol.
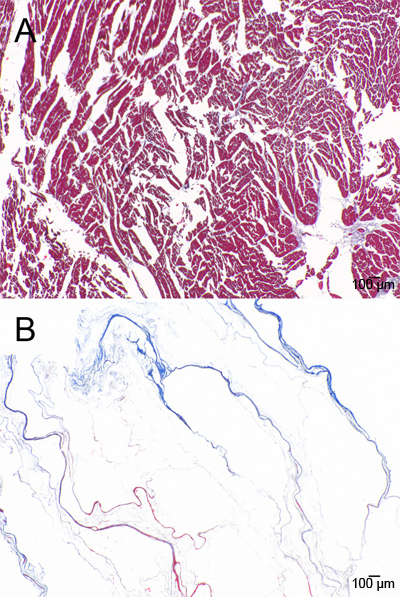
Figure 9. Masson’s Trichrome staining of A) native and B) decellularized ventricle.
Discussion
The current study described methodology for consistent and efficient decellularization of a porcine heart. The protocol was a modification to a previously published report 1, and included longer exposure to flow and increased pressure, which provided more repeatable results. The resulting decellularized tissue met all of the published criteria for successful decellularization of tissue 2. Frequent solution changes were performed to limit the reintroduction of cellular material to the tissue, and the duration of exposure to each decellularization agent was minimized to reduce adverse effects on the ECM. During the beginning stages of the protocol, the perfusion rate was gradually increased to condition the tissue and allow for higher flow rates during the later stages of the protocol. Without conditioning the tissue in the early stages, the vasculature of the heart can rupture, making perfusion of the heart impossible. The protocol was used due to its efficiency, and no claims are made to its superiority over other protocols. The precise composition of decellularization agents and rates of perfusion may conceivably be varied to yield a protocol with better mechanical or biologic characteristics, but the general principles for delivery of the agents to the heart are applicable.
The preservation of the native three-dimensional structure of the heart was attributed to several procedures performed throughout the decellularization protocol. First, the tissue was trimmed and frozen upon arrival. Freezing promoted cell lysis and was important for pre-conditioning the tissue for the perfusion cycles. The tissue was thoroughly inspected for cuts and 2 cm intact intact aorta superior to the aortic valve. If any pericardium or epicardium was cut, the organ was discarded because the perfusate did not reach downstream regions of the heart, and the heart was not fully decellularized. Next, the tissue was fully thawed in type I water before use. The water allowed the tissue to relax as it thawed and also aided the removal of residual blood clots within the heart. Finally, as the tubing was inserted, care was taken to ensure that the aortic valve remained intact so that it formed a water-tight seal around the tubing, so that a proper pressure was maintained and that the solution entered the coronary arteries.
After each decellularization protocol was completed, a series of quality control measures were completed to ensure complete removal of cellular material. The current study verified that the protocol eliminated histologic staining for cell nuclei, showed that less than 50 ng of DNA was present per mg of dry weight of the tissue, and that any DNA was less than 200 bp in size 6. Previously published methods for cardiac decellularization showed similar levels of decellularization in DNA staining and quantification 2,5,9,10. Complete decellularization was accomplished in these studies using similar treatments of enzymes and detergents. However, in the present study, the length of exposure to each chemical was increased, there were more solution changes, and the flow rates were increased. The present protocol also increased the length of chemical rinses, potentially leading to more efficient removal of chemical residues from the extracellular matrix.
Recellularization of decellularized rat hearts with cardiac specific cells has yielded promising results in vitro 2,5. Whole organ perfusion decellularization allowed for maintenance of the native vasculature, which is critical in recellularization of the tissue. The inherent growth factors, matrix proteins, and three-dimensional fiber architecture also promoted proper cell attachment, migration, and signaling to reconstitute contractile myocardial tissue. Porcine cardiac extracellular matrix will be more difficult to recellularize due to the size of the scaffold and the number of cells necessary for proper cell communication and nutrient transport. However, patches cut from the cardiac matrix may be useful for in vivo applications. Multiple studies have shown the potential advantage of using a site-specific matrix for reconstruction of damaged tissue in animal models 11-13. Thus, an extracellular matrix scaffold derived from cardiac tissue is desirable for myocardial reconstruction applications. The inherent architecture of the cardiac tissue may present advantages over an ECM scaffold derived from another organ or an artificial biomaterial. A site-specific scaffold may support host cell infiltration and promote a constructive remodeling response, as opposed to scar tissue formation. To date, cardiac ECM patches have been investigated in vivo to reconstruct a defect created in the myocardial wall 14. Future studies will be performed in vitro to examine the ability of the scaffold to support cardiac cells seeded and cultured on the matrix. The methods described herein may also be applicable to decellularization of human hearts.
In conclusion, porcine heart decellularization is possible and the methods are straight-forward. Continued investigation of this material will provide insight into its potential for clinical use.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
The authors would like to acknowledge Brogan Guest, Michelle Weaver, and Kristen Lippert. Funding for this study was provided by NIH Grant R03EB009237, as well as NIH Training Grants T32EB001026-06 from the National Institute of Biomedical Imaging And Bioengineering and T32HL076124-05.
Materials
| Name of Reagent/Material | Company | Catalogue Number | Comments |
| Trypsin | Gibco | 15090 | |
| EDTA | Fisher | BP120-500 | |
| NaN3 | Sigma | S2002-500G | |
| Triton X-100 | Sigma | X100-1L | |
| 10X PBS | Fisher | BP399-20 | |
| Sodium Deoxycholate | Sigma | D6750-500G | |
| Peracetic Acid | Pfaltz and Bauer | P05020 | 35% CAS# 79-21-0 |
| Ethanol | Pharmco | 111000200 | |
| Masterflex Pump Drive | Cole Parmer | SI-07524-50 | |
| Masterflex Tubing | Cole Parmer | 96400-18 | Size 18 |
| Barbed Reducer | Cole Parmer | EW-30612-20 | |
| 4L Beaker | Fisher Scientific | 02-540T |
Riferimenti
- Ott, H. C., et al. Regeneration and orthotopic transplantation of a bioartificial lung. Nat. Med. 16, 927-933 (2010).
- Ott, H. C., et al. Perfusion-decellularized matrix: using nature’s platform to engineer a bioartificial heart. Nat. Med. , (2008).
- Petersen, T. H., et al. Tissue-engineered lungs for in vivo implantation. Science. 329, 538-541 (2010).
- Uygun, B. E., et al. Organ reengineering through development of a transplantable recellularized liver graft using decellularized liver matrix. Nat. Med. , (2010).
- Wainwright, J. M., et al. Preparation of cardiac extracellular matrix from an intact porcine heart. Tissue Eng. Part C Methods. 16, 525-532 (2010).
- Crapo, P. M., Gilbert, T. W., Badylak, S. F. An overview of tissue and whole organ decellularization processes. Biomaterials. 32, 3233-3243 (2011).
- Gilbert, T. W. Strategies for tissue and organ decellularization. Journal of cellular biochemistry. , (2012).
- Gilbert, T. W., Sellaro, T. L., Badylak, S. F. Decellularization of tissues and organs. Biomaterials. 27, 3675-3683 (2006).
- Akhyari, P., et al. The quest for an optimized protocol for whole-heart decellularization: a comparison of three popular and a novel decellularization technique and their diverse effects on crucial extracellular matrix qualities. Tissue Eng. Part C Methods. 17, 915-926 (2011).
- Weymann, A., et al. Development and evaluation of a perfusion decellularization porcine heart model–generation of 3-dimensional myocardial neoscaffolds. Circulation journal : official journal of the Japanese Circulation Society. 75, 852-860 (2011).
- Cortiella, J., et al. Influence of acellular natural lung matrix on murine embryonic stem cell differentiation and tissue formation. Tissue Eng. Part A. 16, 2565-2580 (1089).
- Remlinger, N. T. Hydrated xenogeneic decellularized tracheal matrix as a scaffold for tracheal reconstruction. Biomaterials. 31, 3520-3526 (2010).
- Sellaro, T. L., Ravindra, A. K., Stolz, D. B., Badylak, S. F. Maintenance of hepatic sinusoidal endothelial cell phenotype in vitro using organ-specific extracellular matrix scaffolds. Tissue Eng. 13, 2301-2310 (2007).
- Wainwright, J. M. Right ventricular outflow tract repair with a cardiac biologic scaffold. Cells, tissues, organs. 195, 159-170 (2012).

