A Versatile Automated Platform for Micro-scale Cell Stimulation Experiments
Summary
We have developed an automated cell culture and interrogation platform for micro-scale cell stimulation experiments. The platform offers simple, versatile, and precise control in cultivating and stimulating small populations of cells, and recovering lysates for molecular analyses. The platform is well suited to studies that use precious cells and/or reagents.
Abstract
Study of cells in culture (in vitro analysis) has provided important insight into complex biological systems. Conventional methods and equipment for in vitro analysis are well suited to study of large numbers of cells (≥105) in milliliter-scale volumes (≥0.1 ml). However, there are many instances in which it is necessary or desirable to scale down culture size to reduce consumption of the cells of interest and/or reagents required for their culture, stimulation, or processing. Unfortunately, conventional approaches do not support precise and reproducible manipulation of micro-scale cultures, and the microfluidics-based automated systems currently available are too complex and specialized for routine use by most laboratories. To address this problem, we have developed a simple and versatile technology platform for automated culture, stimulation, and recovery of small populations of cells (100 – 2,000 cells) in micro-scale volumes (1 – 20 μl). The platform consists of a set of fibronectin-coated microcapillaries (“cell perfusion chambers”), within which micro-scale cultures are established, maintained, and stimulated; a digital microfluidics (DMF) device outfitted with “transfer” microcapillaries (“central hub”), which routes cells and reagents to and from the perfusion chambers; a high-precision syringe pump, which powers transport of materials between the perfusion chambers and the central hub; and an electronic interface that provides control over transport of materials, which is coordinated and automated via pre-determined scripts. As an example, we used the platform to facilitate study of transcriptional responses elicited in immune cells upon challenge with bacteria. Use of the platform enabled us to reduce consumption of cells and reagents, minimize experiment-to-experiment variability, and re-direct hands-on labor. Given the advantages that it confers, as well as its accessibility and versatility, our platform should find use in a wide variety of laboratories and applications, and prove especially useful in facilitating analysis of cells and stimuli that are available in only limited quantities.
Introduction
The study of cells maintained in culture (in vitro analysis) has provided invaluable insight into the fundamental principles and molecular mechanisms governing complex biological systems and human health. The conventional methods for culture, stimulation, and collection of cells for analysis, which utilize Petri dishes and microtiter plates, were designed for study of large populations of cells (≥105) in milliliter-scale culture volumes (≥0.1 ml). However, there are many instances in which only limited quantities of cells are available (e.g. primary cells), or small populations of cells are desirable (e.g. to reduce cell-to-cell variability across the population), or required reagents are difficult to obtain or prohibitively expensive (e.g. purified cell-secreted factors). Such issues can be successfully addressed by scaling down culture size, which has the added benefit of reducing consumption of all reagents required for in vitro analysis1,2. Unfortunately, conventional equipment and methods do not support precise and reproducible manipulation of micro-scale cultures and the microfluidics-based automated systems currently available3-11 are too complex and specialized for routine use by most laboratories.
In this report, we describe assembly and use of a simple and versatile technology platform for automated culture, stimulation, and recovery of small populations of cells (100 – 2,000 cells) in micro-scale volumes (1 – 20 μl). The platform architecture (Figure 1) is modular in design: a set of fibronectin-coated microcapillaries (“cell perfusion chambers” module) serves as the site for establishment, maintenance, and stimulation of micro-scale cultures; and a digital microfluidics (DMF)12,13 device outfitted with “transfer” microcapillaries (“central hub” module)14,15 routes cells and reagents to and from the perfusion chambers. DMF enables the user to individually address multiple droplets simultaneously and to change or re-order the manipulations (i.e. reconfigure sample processing trains) without altering the device hardware. Its tremendous flexibility is evident in its recent emergence as a key technology in a wide range of applications, including cell culture16,17, enzyme assays18,19, immunoassays20,21, DNA analysis22,23, protein processing,24,25 and clinical specimen processing.26,27 Our central hub takes advantage of the flexibility inherent to DMF devices, and further enhances it through the addition of microcapillary interfaces, which provides opportunity to carry out a subset of manipulations (e.g. cell culture) in specialized peripheral modules, rather than on the DMF device itself. Compartmentalization of processing trains in this way also simplifies design of the platform architecture (no need to build a DMF device that can carry out all processing steps) and facilitates its evolution as new functions are required (simply integrate new peripheral modules as necessary). Transport of cells and reagents within the central hub is driven by electrowetting forces generated by sequential activation of electrodes within the DMF device13,28; transport to, from, and within the perfusion chambers is powered by pressure changes generated by a high-precision syringe pump. All of these fluid movements are controlled via a simple electronic interface and automated through use of pre-determined scripts.
As a representative example, we demonstrate the use of the platform for the study of transcriptional responses elicited in immune cells upon challenge with bacteria (Figure 2). Carrying out these experiments on the platform enabled us to work with small numbers of cells (~1,000 per experimental condition), minimize experiment-to-experiment variability, conserve reagents, and re-direct hands-on labor. Given the advantages that it confers, as well as its accessibility and versatility, this platform should find use in a wide variety of laboratories and applications and prove especially useful in facilitating analysis of cells and stimuli that are available in only limited quantities.
Protocol
This general protocol is designed to support application of the platform to a wide variety of studies; aspects specific to the representative study described in this report are separated out in brackets. Figure 2 illustrates a representative study carried out using the protocol. Note that in the protocol, all “instruct” commands are automated through use of pre-determined scripts. Note too that Step 2 can be carried out in parallel with Step 1 (e.g. during Steps 1.7 and 1.8), and that Step 2.1 is often bypassed (because patterned/coated DMF device plates can be washed and re-used; see Step 5.2).
1. Assemble and Populate the Cell Perfusion Chambers
- Coat the interior surfaces of sterilized (autoclaved) microcapillaries (fused-silica; 679 μm o.d., 534 μm i.d; 15 cm long) with fibronectin, by instructing the syringe pump to draw a sterile fibronectin solution (30 μl of 50 μg/ml) into each microcapillary, incubating (37 °C for 2 hr), instructing the syringe pump to withdraw the solution, and letting the microcapillaries dry (room temperature (RT) for 6 hr).
- Connect each fibronectin-coated microcapillary (perfusion chamber) to polycarbonate tubing (500 μm o.d., 100 μm i.d.) and the tubing to the multi-port syringe pump, using CapTite fittings (6 perfusion chambers, 8-port syringe pump).
- Using tape, secure the body of each perfusion chamber to a heat block set to desired temperature (37 °C).
- Immerse the open ends of the perfusion chambers in a reservoir (microcentifuge tube or microtiter plate well) containing cells (P388D.1 murine macrophages) suspended in fresh growth medium (RPMI-1640, 10% FBS, 500 units/ml penicillin, 500 μg/ml streptomycin) at desired concentration (106 cells/ml).
- Instruct the syringe pump to draw cells (10 μl, 104 cells) into each perfusion chamber followed by enough air to move the liquid plug (~4.5 cm) to the section secured to the heat block.
- Allow the cells to adhere to the fibronectin-coated interior surfaces of the perfusion chambers (37 °C for 1 – 2 hr). In the meantime, transfer the open ends of the perfusion chambers from the cells reservoir to a new reservoir containing fresh growth medium.
- After the adherence period, instruct the syringe pump to send the liquid plugs (conditioned media and unattached cells) to waste, withdraw fresh medium (10 μl), and follow with enough air to position the new liquid plugs (fresh medium) over the adhered cells.
- Continue to carry out medium exchanges (i.e. repeat Step 1.7) at regular intervals (every 2 hr) until the cell populations are sufficiently equilibrated and expanded (16 – 24 hr of micro-scale culture generated populations of ~1,000 cells/chamber).
2. Fabricate the DMF Device and Assemble the Hub Architecture
- As described previously14,29, pattern the bottom plate of the DMF device with forty-six indium tin oxide (ITO) electrodes (drivers of droplet actuation) by photolithography and etching, and coat with 5 μm of Parylene-C and 50 nm of Teflon-AF. Form the top plate of the DMF device by coating an unpatterned ITO glass substrate with Teflon-AF, as above.
- Using metal compression frames, fix the DMF plates into a polymer cast14,30 with recesses that maintain a 400 μm spacing between the plates; this spacing, combined with the actuation electrode size (2.5 mm2), defines the droplet volume (2.5 μl per electrode). Note that DMF assembly can be accomplished by simpler means24.
- Insert Teflon-coated transfer microcapillaries (360 μm o.d., 100 μm i.d.; 3.5 – 4.0 cm long) into the space between the DMF plates, positioning each to extend to the edge of its cognate actuation electrode.
- To the opposite end of each transfer microcapillary affix a CapTite fitting.
- Engage the electrical connectors to supply voltage to the ITO electrodes. Electrode activation sequences are automated through use of a computer-controlled electronic interface running pre-determined scripts.
- Optional: Move the assembled DMF hub onto the translational stage of a microscope (SZ-6145TR (Olympus, Japan)) equipped with a charge-coupled device (CCD) camera (DCR-HC96 (Sony, Japan)) to facilitate tracking of droplets.31
3. Connect the Perfusion Chambers to the DMF Hub and Stimulate the Cell Populations
- Remove the open ends of the perfusion chambers from the medium reservoir (see Step 1.6) and insert them into the CapTite fittings affixed to the ends of the transfer microcapillaries of the DMF hub (see Step 2.4).
- Instruct the syringe pump to send the liquid plugs (conditioned media) in the perfusion chambers to waste.
- Instruct the DMF hub to draw stimulus (E. coli BioParticles at 100 μg/ml in fresh medium with 0.1% Pluronic F127 w/v) from an on-board reservoir and deliver a droplet of appropriate size (10 μl) to each transfer microcapillary. The on-board reservoirs are comprised of cylindrical-shaped compartments (1.5 x 1.5 cm each) and connected to the DMF Hub via tubing.
- Instruct the syringe pump to draw the stimulus plugs into the transfer microcapillaries, and follow with enough air to position the plugs over the cell populations in the perfusion chambers.
- Incubate the cells with stimulus (37 °C for 1 – 4 hr). Optional: Exchange for fresh stimulus (i.e. repeat Steps 3.2-3.4) at regular intervals.
4. Terminate Stimulation and Recover Cell Lysates for Analysis
- Optional: If a post-stimulation period is required, exchange the stimulus-containing liquid plugs for fresh medium (i.e. repeat Steps 3.2 – 3.4, substituting fresh medium for stimulus), and continue to incubate and exchange as needed.
- Exchange the liquid plugs in the perfusion chambers for wash buffer (Prelude Direct Lysis Module Buffer B) (i.e. repeat Steps 3.2 – 3.4, substituting wash buffer for stimulus), and incubate as needed (5 min).
- Exchange the liquid plugs (wash buffer) for lysis solution (Prelude Direct Lysis Module Buffer A with 0.1% Pluronic F127 w/v) (i.e. repeat Steps 3.2 – 3.4, substituting lysis solution for stimulus), and incubate as needed (5 min).
- Instruct the syringe pump to send the liquid plugs (cell lysates) to the DMF hub.
- Disassemble the DMF hub, remove the top plate of the DMF device (using care not to tilt the plate sideways, as this can result in droplet mixing), and collect the lysates using a Pipetman or syringe for off-platform analysis (gene expression profiling via qPCR).
5. Clean Platform Hardware for Re-use
- Immerse the transfer microcapillaries and CapTite fittings in 10% bleach (v/v in water) for 10 min at RT, rinse well with deionized water, and dry using nitrogen gas.
- Immerse the DMF device plates in 10% bleach for 10 min at RT, rinse well with isopropanol followed by deionized water, and dry using nitrogen gas followed by baking at 160 °C for 10 min.
- The syringe pump’s polycarbonate tubing, and the DMF device’s polymer cast and compression frame, can be re-used without cleaning. The perfusion chamber microcapillaries are discarded after each experiment.
Representative Results
As a demonstration of the automated platform, we used it to carry out a study in which small populations of immune cells were grown in micro-scale cultures, challenged with bacteria, and lysed for off-platform analysis of pro-inflammatory responses (Figure 2).
Each of six cell perfusion chambers was seeded with 104 immune cells (P388D.1 murine macrophages) resuspended in 10 μl of growth medium. After an adherence period (37 °C for 2 hr) and a medium exchange, ~500 cells remained attached to the interior surfaces of each chamber (Figure 3A), as measured through analysis of replicate microscale cultures (cells released via chelator treatment and counted via hemacytometer and light microscopy). Further incubation at 37 °C for 16 hr, with medium exchanges at 2 hr intervals, expanded the cell populations to ~1,000 cells per chamber (Figure 3B) (as measured through analysis of replicate microscale cultures, as described above). In four of the six perfusion chambers, the cell populations were challenged with E. coli bacteria (pHrodo BioParticles), resuspended in growth medium, and delivered at a multiplicity of infection (MOI) of 20 (i.e. each cell in the population was challenged with 20 BioParticles, on average). In the remaining two chambers, the cell populations received a mock challenge (i.e. growth medium only). After incubation at 37 °C for 1 hr (two bacteria-challenged cultures) or 4 hr (two bacteria-challenged, and two mock-challenged, cultures), the cells were washed and lysed, and the lysates recovered from the DMF hub. Conventional bench-scale methods were used to prepare cDNA from the lysates, and to measure transcript levels for inflammation mediators: Chemokines MIP-1β (CCL4) and RANTES (CCL5), inducible cyclooxygenase COX-2 (PTGS2), and cytokine TNFα (TNF).
Figure 4 summarizes the results from three independently replicated experiments. We found that immune cells grown and challenged in micro-scale cultures (red bars) mounted pro-inflammatory transcriptional responses that were essentially identical to those elicited in cells grown and challenged in conventional cultures (blue bars). Moreover, the on-platform (micro-scale) experiments showed less variability (as indicated by smaller error bars) and consumed smaller quantities of cells and reagents (Table 1).
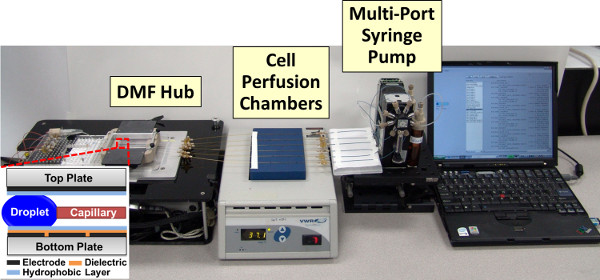

Figure 1. Photograph of Automated Platform for Micro-Scale Cell Stimulation Experiments.
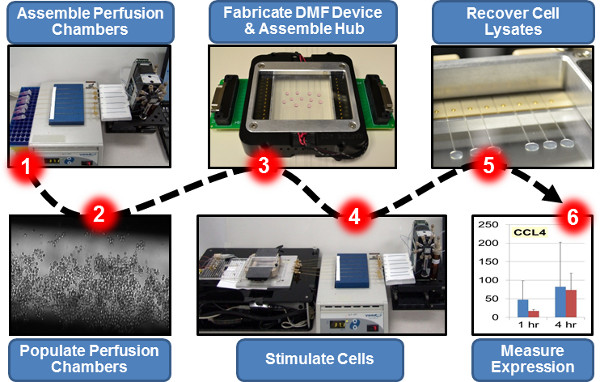

Figure 2. Overview of a Representative Platform-Enabled Experiment: Transcriptional Profiling of Immune Cells Challenged with Bacteria in Micro-Scale Cultures.
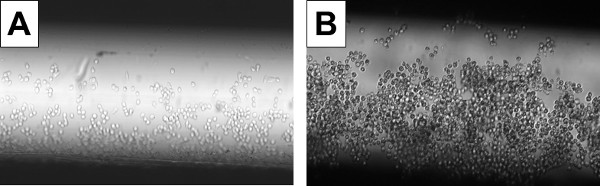
Figure 3. Photographs of Immune Cells Cultured in a Micro-scale Perfusion Chamber. Murine macrophages (P388D.1 cell line) were resuspended in growth medium at a concentration of 106 cells/ml, and 10 μl (104 cells) used to seed a fibronectin-coated microcapillary (perfusion chamber). Bright-field photographs were taken using a MVX10 microscope (Olympus) equipped with a CCD camera (Qimaging). To measure population size, cells from replicate micro-scale cultures were released from the perfusion chamber surfaces using Cell Stripper, stained using trypan blue, and counted using a hemacytometer and light microscope. A. After a 2 hr adherence period (incubation at 37 °C without disturbance) and a medium exchange, ~500 cells remained attached to the interior surfaces of the perfusion chamber. B. After 16 hr of culture (incubation at 37 °C with medium exchanges every 2 hr), the population had expanded to ~1,000 cells.
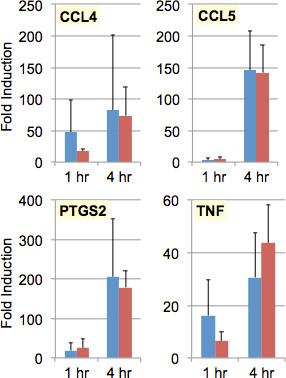
Figure 4. Transcriptional Profiles of Immune Cells Cultured and Stimulated in Conventional Vessels versus Micro-scale Perfusion Chambers. Murine macrophages (P388D.1 cell line) were resuspended in growth medium at a concentration of 106 cells/ml. For conventional cultures, 50 μl (5 x 104 cells) were used to seed 150 μl of growth medium in a well of a 96-well microtiter plate; for micro-scale cultures, 10 μl (104 cells) were used to seed a fibronectin-coated microcapillary (perfusion chamber). After incubation at 37 °C for 16 hr, the cells (~5 x 104 per conventional culture, ~103 per micro-scale culture) were challenged with E. coli bacteria (pHrodo BioParticles) at a multiplicity of infection (MOI) of 20 (that is, 20 BioParticles for every cell in the culture); negative control (mock-challenged) cultures received only fresh media. After further incubation at 37 °C for 1 or 4 hr, the cells were washed and lysed using reagents from the Prelude Direct Lysis Module. Lysate RNA was amplified and converted to cDNA using Ovation PicoSL WTA, the larger (≥200 nt) cDNA products purified using Agencourt RNAClean XP, and these products used as template in TaqMan qPCR assays to measure transcript levels for MIP-1β (CCL4), RANTES (CCL5), COX-2 (PTGS2), and TNFα (TNF), as well as GAPDH (for normalization of cDNA levels between lysates). Each qPCR assay was carried out in triplicate, and the results averaged. Each experiment was independently replicated three times. Bars indicate average fold increase in transcript levels in conventional cultures (blue) or micro-scale cultures (red) challenged with bacteria, as compared to negative control (mock-challenged) cultures; error bars indicate standard deviation across the three experiments.
| Requirement | Conventional Experiment | Micro-Scale Experiment |
| Cells | 3,000,000 | 60,000 |
| Growth Medium | 2.4 ml | 0.7 ml |
| BioParticles | 200 μg | 4 μg |
| Wash & Lysis Reagents | 300 μl each | 60 μl each |
Table 1. Requirements for Conventional versus Micro-Scale Cell Stimulation Experiments. Numbers are drawn from the representative experiment described in this report (Figure 2), in which murine macrophages (P388D.1 cell line) were grown in six conventional (96-well microtiter plate) versus micro-scale (perfusion chamber) cultures, challenged with E. coli bacteria (pHrodo BioParticles) at an MOI of 20, and lysed for qPCR analysis.
Discussion
We have developed a simple and versatile automated platform for micro-scale cell culture and stimulation experiments. The platform enables us to work with small culture volumes and cell populations (1 – 20 μl and 100 – 2,000 cells, per chamber); culture sizes could be further reduced through use of microcapillaries of smaller diameter. Working at these scales reduces the cost of routine studies and makes feasible studies that require use of precious reagents and/or cells. Platform-executed experiments also show less variability, presumably due to the precision and reproducibility with which cells and fluids are handled by the automated platform.
The current configuration of the platform draws cells and media from off-board reservoirs (microcentrifuge tubes or microtiter plate wells) for seeding, equilibration, and expansion of cell populations in the perfusion chambers. This is a simple way to address the mismatch between growth medium requirements (e.g. 700 μl consumed in the representative experiment described above) and the capacity of the DMF device’s on-board reservoirs (24 μl each). However, this configuration necessitates mid-experiment manual intervention, to switch between two reservoirs (Step 1.6) and connect the perfusion chambers to the DMF hub (Step 3.1). Future configurations could include large-capacity (e.g. 1 ml) on-board reservoirs, which would enable routing of all materials through the DMF hub, thereby eliminating need for mid-experiment manual intervention.
Our experiments required only short-term (<24 hr) culture of cells on the platform. We have verified that cell viability on the platform is comparable to that in conventional cultures for at least 24 hr (data not shown). However, we have not yet attempted to maintain cultures on the platform for longer periods of time. Evaporation is not a major concern because the platform’s perfusion chambers and growth medium reservoirs are partially closed systems and the liquid plugs covering the cells are easily replenished via medium exchange. On the other hand, to maintain pH in long-term cultures it may be necessary to house the growth medium reservoirs in a 5 – 10% CO2 atmosphere or switch to a growth medium that does not include a CO2-bicarbonate based pH buffer. Additionally, long-term culture of actively dividing cells would expand populations to the point of confluency (i.e. coverage of all available surfaces within the perfusion chamber), which would necessitate passaging of the cells (release, wash, dilute, and re-seed). Others have accomplished this in microfluidic regimes3,5,6,9,17 and in its current configuration the platform should be able to carry out all of the required manipulations. In summary, we anticipate that with modest modification (primarily to protocols, rather than hardware) our platform should be capable of supporting long-term mammalian cell culture. This would seem a particularly promising future direction because conventional methods are time-consuming and labor-intensive; adding a long-term culture capability to the repertoire of our automated platform would further enable its users to re-direct their labor more productively.
Although we have tested only a few different cell types thus far (e.g. murine macrophage cell line RAW264.7; murine bone marrow derived macrophages (BMDM)), the platform should be able to accommodate virtually any adherent cell type. In some cases, it may be necessary to change the perfusion chamber coating from fibronectin to another extracellular matrix component (or mixture of components), or to change the growth medium, but these modifications are easily made. Work with non-adherent cell types could be more challenging, but cell tethering strategies have been successful in similar contexts32-34. It should also be possible to create mixed cultures comprised of multiple cell types; indeed, the precision with which the platform handles cells and fluids should ensure that construction of mixed cultures is highly reproducible.
With regard to stimuli, virtually any substance that can be transported within an aqueous liquid plug should be compatible with our platform5,13,16,35. In the case of large or dense particles, which can settle out over time, it may be necessary to periodically resuspend them within the reservoirs; in the past, we have addressed this issue through use of high-density carrier fluid36 or a simple agitation system (data not shown), solutions that should be easy to implement on our platform.
Thus far, our characterization of cells cultured and stimulated on the platform has primarily focused on qRT-PCR analysis of hallmark pro-inflammatory responses. As a next step, comprehensive analysis of the cell population’s transcriptome (via DNA microarrays or Generation Sequencing (SGS) (i.e. RNA-Seq37,38)), is well within reach; with modest modification of the cDNA preparation step, our platform and protocols should be fully capable of supporting global transcriptional profiling studies. Similarly, analysis of cell population responses at the protein level (via ELISA, immunoprecipitation, mass spectrometry, or protein microarrays) should require only modification of the lysate preparation steps. The platform also supports imaging-based analysis of cell responses. Cells can be imaged within the perfusion chamber (e.g. Figure 3); use of flat-surfaced (rather than rounded) microcapillaries for the perfusion chamber module would facilitate this type of analysis. Additionally, cells can be imaged on the DMF hub14,15, prior to seeding of the perfusion chambers and/or after release from them. Released cells also may be imaged off of the platform. Thus far, we have used light and fluorescence microscopy to image cells at all of these stages in order to analyze their morphology, viability, proliferation rate, and physical interactions with stimuli (e.g. BioParticles) and each other (data not shown). Microscopy also should enable analysis of metabolite content (assessed through use of fluorescent dyes), transcriptional activity (assessed through use of reporter constructs), and protein levels/localization (assessed through use of reporter constructs, fluorescent protein fusion constructs, and/or immunocytochemistry). Released cells could be analyzed using flow cytometry as well. Many of these analyses could be carried out entirely on the platform in its current configuration; others would require off-platform analysis or integration of specialized microfluidics-based modules (e.g. see References 36,39-41 ) into the platform via connection to the DMF hub. Taken together, it seems likely that our platform will prove valuable in enabling multi-dimensional analysis of small populations of cells, which otherwise can be achieved only through use of very complex and expensive robotic systems.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
The authors thank Ronald F. Renzi and Michael S. Bartsch for their contributions to the design and development of DMF devices and the DMF hub. This research was fully supported by the Laboratory Directed Research and Development program at Sandia National Laboratories. Sandia is a multi-program laboratory managed and operated by Sandia Corporation, a wholly owned subsidiary of Lockheed Martin Corporation, for the U.S. Department of Energy’s National Nuclear Security Administration under contract DE-AC04-94AL85000.
Materials
| Name of Reagent/Material | Company | Catalog Number | Comments |
| Prelude Direct Lysis Module | NuGEN | 1400-24 | |
| Trypan Blue (0.4% w/v) | GIBCO | 15250-061 | |
| Cell Stripper | Cellgro | 25-056-C1 | |
| Ovation PicoSL WTA | NuGEN | 3310-048 | |
| Agencourt RNAClean XP | Beckman Coulter Genomics | A63987 | |
| pHrodo BioParticles | Invitrogen | P35361 | |
| CCL4 TaqMan qRT-PCR assay | Applied Biosystems | Mm00443111_m1 | |
| CCL5 TaqMan qRT-PCR assay | Applied Biosystems | Mm01302428_m1 | |
| PTGS2 TaqMan qRT-PCR assay | Applied Biosystems | Mm00478374_m1 | |
| TNF TaqMan qRT-PCR assay | Applied Biosystems | Mm00443258_m1 | |
| GAPDH TaqMan qRT-PCR assay | Applied Biosystems | Mm99999915_g1 | |
| Pluronic F127 | Sigma Chemical | 2594628 | |
| Fluorinert FC-40 | Sigma Chemical | 51142-49-5 | |
| Parylene C dimer | Specialty Coating Systems | 28804-46-8 | |
| Teflon-AF | DuPont | AF1600 | |
| Polyimide tape | ULINE | S-11928 | |
| Indium tin oxide (ITO) coated glass substrates | Delta Technologies | CB-40IN-1107 | |
| DMF hub Teflon-coated fused-silica microcapillaries | Polymicro Technologies | TSU100375 | |
| Perfusion chamber microcapillaries | Polymicro Technologies | TSP530700 | |
| Tubing and microcapillary fittings | Sandia National Laboratories | ||
| Polycarbonate tubing | Paradigm Optics | CTPC100-500-5 | |
| 8-port precision syringe pump equipped with 30 mm (500 μl capacity) syringes | Hamilton Company | 54848-01 | |
| Parylene-C vapor deposition instrument | Specialty Coating Systems | PDS 2010 Labcoter 2 | |
| High-voltage function generator | Trek | 615A-1 615-3 | |
| MVX10 microscope | Olympus | Optional (facilitates tracking of droplets on DMF hub) | |
| QIClick digital CCD camera | QImaging | QIClick-F-CLR-12 | Optional (facilitates tracking of droplets on DMF hub) |
Riferimenti
- Young, E. W. K., Beebe, D. J. Fundamentals of microfluidic cell culture in controlled microenvironments. Chemical Society Reviews. 39, 1036-1048 (2010).
- Yeo, L. Y., Chang, H. C., Chan, P. P. Y., Friend, J. R. Microfluidic devices for bioapplications. Small. 7, 12-48 (2011).
- Zhang, B., Kim, M. C., Thorsen, T., Wang, Z. A self-contained microfluidic cell culture system. Biomedical Microdevices. 11, 1233-1237 (2009).
- Skafte-Pedersen, P., et al. A self-contained, programmable microfluidic cell culture system with real-time microscopy access. Biomedical Microdevices. 14, 385-399 (2012).
- Barbulovic-Nad, I., Au, S. H., Wheeler, A. R. A microfluidic platform for complete mammalian cell culture. Lab Chip. 10, 1536-1542 (2010).
- Zhou, Y., Pang, Y., Huang, Y. Openly Accessible Microfluidic Liquid Handlers for Automated High-Throughput Nanoliter Cell Culture. Analytical Chemistry. 84, 2576-2584 (2012).
- Wang, H. Y., Bao, N., Lu, C. A microfluidic cell array with individually addressable culture chambers. Biosensors and Bioelectronics. 24, 613-617 (2008).
- Cimetta, E., Cagnin, S., Volpatti, A., Lanfranchi, G., Elvassore, N. Dynamic culture of droplet-confined cell arrays. Biotechnology Progress. 26, 220-231 (2010).
- Hung, P. J., Lee, P. J., Sabounchi, P., Lin, R., Lee, L. P. Continuous perfusion microfluidic cell culture array for high-throughput cell-based assays. Biotechnology and Bioengineering. 89, 1-8 (2005).
- King, K. R., et al. A high-throughput microfluidic real-time gene expression living cell array. Lab on a Chip. 7, 77-85 (2007).
- Gómez-Sjöberg, R., Leyrat, A. A., Pirone, D. M., Chen, C. S., Quake, S. R. Versatile, fully automated, microfluidic cell culture system. Analytical Chemistry. 79, 8557-8563 (2007).
- Wheeler, A. R. Chemistry – Putting electrowetting to work. Science. 322, 539-540 (2008).
- Gong, J., Kim, C. J. All-electronic droplet generation on-chip with real-time feedback control for EWOD digital microfluidics. Lab Chip. 8, 898-906 (2008).
- Choi, K., et al. Integration of field effect transistor-based biosensors with a digital microfluidic device for a lab-on-a-chip application. Lab Chip. 12, 1533-1536 (2012).
- Barbulovic-Nad, I., Yang, H., Park, P. S., Wheeler, A. R. Digital microfluidics for cell-based assays. Lab Chip. 8, 519-526 (2008).
- Gentilucci, L., de Marco, R., Cerisoli, L. Chemical modifications designed to improve peptide stability: incorporation of non-natural amino acids, pseudo-peptide. 16, 3185-3203 (2010).
- Miller, E. M., Wheeler, A. R. A digital microfluidic approach to homogeneous enzyme assays. Anal. Chem. 80, 1614-1619 (2008).
- Jebrail, M. J., Bartsch, M. S., Patel, K. D. Digital microfluidics: a versatile tool for applications in chemistry, biology. 12, 2452-2463 (2012).
- Ng, A. H. C., Choi, K., Luoma, R. P., Robinson, J. M., Wheeler, A. R. Digital Microfluidic Magnetic Separation for Particle-Based Immunoassays. Analytical Chemistry. 84, 8805-8812 (2012).
- Sista, R. S., et al. Heterogeneous immunoassays using magnetic beads on a digital microfluidic platform. Lab Chip. 8, 2188-2196 (2008).
- Malic, L., Veres, T., Tabrizian, M. Biochip functionalization using electrowetting-on-dielectric digital microfluidics for surface plasmon resonance imaging detection of DNA hybridization. Biosens. Bioelectron. 24, 2218-2224 (2009).
- Malic, L., Veres, T., Tabrizian, M. Nanostructured digital microfluidics for enhanced surface plasmon resonance imaging. Biosens. Bioelectron. 26, 2053-2059 (2011).
- Jebrail, M. J., Wheeler, A. R. Digital microfluidic method for protein extraction by precipitation. Anal. Chem. 81, 330-335 (2009).
- Nelson, W. C., et al. Incubated protein reduction and digestion on an electrowetting-on-dielectric digital microfluidic chip for MALDI-MS. Anal. Chem. 82, 9932-9937 (2010).
- Mousa, N. A., et al. Droplet-scale estrogen assays in breast tissue, blood, and serum. Sci. Transl. Med. 1, 1ra2 (2009).
- Jebrail, M. J., et al. A digital microfluidic method for dried blood spot analysis. Lab Chip. 11, 3218-3224 (2011).
- Choi, K., Ng, A. H. C., Fobel, R., Wheeler, A. R. Digital microfluidics. Annual Review of Analytical Chemistry. 5, 413-440 (2012).
- Jebrail, M. J., et al. Digital Microfluidics for Automated Proteomic Processing. J. Vis. Exp. (33), e1603 (2009).
- Thaitrong, N., et al. Quality Control of Next Generation Sequencing Library through an Integrative Digital Microfluidic Platform. , (2012).
- Shin, Y. J., Lee, J. B. Machine vision for digital microfluidics. Review of Scientific Instruments. 81, (2010).
- Thornton, M., Eward, K. L., Helmstetter, C. E. Production of minimally disturbed synchronous cultures of hematopoietic cells. BioTechniques. 32, 1098-1105 (2002).
- Huang, X., Dai, W., Darzynkiewicz, Z. Enforced adhesion of hematopoietic cells to culture dish induces endomitosis and polyploidy. Cell Cycle. 4, 801-805 (2005).
- Raje, M., et al. Charged nylon membrane substrate for convenient and versatile high resolution microscopic analysis of Escherichia coli & mammalian cells in suspension culture. Cytotechnology. 51, 111-117 (2006).
- Chatterjee, D., Hetayothin, B., Wheeler, A. R., King, D. J., Garrell, R. L. Droplet-based microfluidics with nonaqueous solvents and solutions. Lab Chip. 6, 199-206 (2006).
- Perroud, T. D., et al. Microfluidic-based cell sorting of Francisella tularensis infected macrophages using optical forces. Analytical Chemistry. 80, 6365-6372 (2008).
- Wang, Z., Gerstein, M., Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nature Reviews Genetics. 10, 57-63 (2009).
- Malone, J. H., Oliver, B. Microarrays, deep sequencing and the true measure of the transcriptome. BMC Biology. 9, (2011).
- Wu, M., et al. Microfluidically-unified cell culture, sample preparation, imaging and flow cytometry for measurement of cell signaling pathways with single cell resolution. Lab on a Chip. 12, 2823-2831 (2012).
- Srivastava, N., et al. Fully integrated microfluidic platform enabling automated phosphoprofiling of macrophage response. Analytical Chemistry. 81, 3261-3269 (2009).
- Herr, A. E., et al. Microfluidic immunoassays as rapid saliva-based clinical diagnostics. Proceedings of the National Academy of Sciences of the United States of America. 104, 5268-5273 (2007).

