In Vivo Monitoring of Circadian Clock Gene Expression in the Mouse Suprachiasmatic Nucleus Using Fluorescence Reporters
Summary
This newly developed fluorescence-based technology enables long-term monitoring of the transcription of circadian clock genes in the suprachiasmatic nucleus (SCN) of freely moving mice in real-time and at a high temporal resolution.
Abstract
This technique combines optical fiber mediated fluorescence recordings with the precise delivery of recombinant adeno-associated virus based gene reporters. This new and easy to use in vivo fluorescence monitoring system was developed to record the transcriptional rhythm of the clock gene, Cry1, in the suprachiasmatic nucleus (SCN) of freely moving mice. To do so, a Cry1 transcription fluorescence reporter was designed and packaged into Adeno-associated virus. Purified, concentrated virus was injected into the mouse SCN followed by the insertion of an optic fiber, which was then fixed onto the surface of the brain. The animals were returned to their home cages and allowed a 1-month post-operative recovery period to ensure sufficient reporter expression. Fluorescence was then recorded in freely moving mice via an in vivo monitoring system that was constructed at our institution. For the in vivo recording system, a 488 nm laser was coupled with a 1 × 4 beam-splitter that divided the light into four laser excitation outputs of equal power. This setup enabled us to record from four animals simultaneously. Each of the emitted fluorescence signals was collected via a photomultiplier tube and a data acquisition card. In contrast to the previous bioluminescence in vivo circadian clock recording technique, this fluorescence in vivo recording system allowed the recording of circadian clock gene expression during the light cycle.
Introduction
In mammals, the suprachiasmatic nucleus (SCN) governs the whole body's circadian rhythm to coordinate an individual's response to exogenous environmental changes (e.g., light, temperature, stress, etc.)1. Core clock components consist of Per1-3, Cry1-2, Clock, and Bmal1, and play a central role in regulating the circadian clock of each cell. Each cell in the SCN contains the transcriptional activator, CLOCK/BMAL1, which acts as a heterodimer to induce the expression of PER and CRY. The PER/CRY complex then inhibits the function of CLOCK/BMAL1 to form a transcription-translation feedback loop that takes about 24 h to complete2,3.
Previous studies on the SCN have mainly employed the ex vivo SCN slice culture method4,5,6 and, while this approach has provided valuable information, its limitations have inhibited our ability to obtain data regarding the influence of other brain nuclei on the SCN, as well as the effect of exogenous stimuli (e.g., light) on cells residing in this critical region. In 2001, Hitoshi Okamura's group was the first to use the bioluminescence system to in vivo monitor circadian clock gene expression in the SCN in freely moving mice7. Ken-ichi Honma's group has spent the past few years further developing the bioluminescence in vivo recording system in the SCN8,9,10. Together, these studies have provided researchers with the ability to monitor the circadian clock in constant darkness or after a light pulse. However, because bioluminescence is too dim to allow for continuous monitoring during the light/dark cycle, coupled with the fact that light is the predominant signal required for the SCN-mediated entrainment of circadian clocks11, there is increasing demand for the development of experimental methods which overcome the limitations associated with bioluminescence recording. The current report describes a fluorescence-based system which was constructed to monitor the circadian clock of the SCN in vivo in freely moving mice. This easy-to-use method permits continuous monitoring during the light/dark cycle and allows for the long-term observation of the transcription of circadian clock genes in the SCN in real-time and at high temporal resolution.
Protocol
All procedures in this protocol were conducted with the approval of the Institutional Animal Care and Use Committee (IACUC) of the National Institute of Biological Sciences, Beijing, in accordance with the governmental regulations of China.
1. Construction of the Cry1 Fluorescence Reporter
Note: Previous circadian studies using the bioluminescence system2,12,13 fuse a circadian promoter with a destabilized luciferase (dLuc) to induce rhythmic dLuc expression. To develop a fluorescence reporter, destabilized luciferase was replaced with destabilized fluorescence protein (dVenus) to induce rhythmic dVenus expression. The generation of the P(Cry1)-dVenus reporter is described below, the construct of which is currently being submitted to Addgene and will be available soon for academic use.
- Amplify the functional mouse Cry1 promoter (+328 to -1208 bp region of the Cry1 gene, transcription start site designated as ‘‘-1’’). Use the PCR enzyme (KOD Plus Neo), primers F1 and R1 (see Table 2), mouse genome as a template and the manufacturer’s PCR protocol to amplify the Cry1 promoter DNA fragment.
- Amplify intron 336 of the Cry1 gene, which is important for the circadian clock function of Cry114. Use the PCR enzyme, primers F2 and R2, mouse genome as a template and the manufacturer’s PCR protocol to amplify the intron 336 DNA fragment.
- Amplify Venus. Use the PCR enzyme, primers F3 and R3, pVENUS-N1 plasmid as a template and the manufacturer’s PCR protocol to amplify the Venus DNA fragment.
- Amplify the NLS-D2 DNA fragment. Use the PCR enzyme, primers F4 and R4, pcDNA3.3-d2eGFP plasmid as a template and the manufacturer’s PCR protocol to amplify the NLS-D2 DNA fragment.
- Synthesize DNA fragment 1 as exon 1.
- Synthesize DNA fragment 2 as the linker between intron 336 and the Venus DNA fragment.
- Cut the pAAV-EF1a-double floxed-hChR2(H134R)-mCherry-WPRE-HGHpA vector plasmid with the MluI and EcoRI enzymes. Use the Gel Extraction Kit to purify the larger DNA fragment. Use the manufacturer’s protocol.
- Using a Gibson assembly mix and its standard protocol, assemble all of the DNA fragments together to produce the pAAV-P(Cry1)-intron336-Venus-NLS-D2-WPRE-HGHpA reporter. Use the manufacturer’s protocol.
2. Production of Adeno-associated Virus15
- Prepare the following plasmid DNA stocks: 31.25 µg of pRV1, 31.25 µg of pH21, 125 µg of pFdelta6, and 62.5 µg of pAAV-P(Cry1)-intron336-Venus-NLS- D2-WPRE-HGHpA.
- Prepare five 15 cm dishes of 293T cells and wait until the cells become 90% confluent.
- Transfect the plasmids into the 297T cells following the transfection reagent protocol. For each dish, use 6.25 µg of pRV1, 6.25 µg of pH21, 25 µg of pFdelta6, 12.5 µg of pAAV-P(Cry1)-intron336-Venus-NLS-D2-WPRE-HGHpA, and 120 µL of transfection reagent.
- 72 h after transfection, detach the cells from each plate using a pipette gun; harvest the cells into two 50 mL tubes.
- Pellet the cells at 800 x g for 10 min and discard the supernatant. Wash the cells by placing 20 mL of phosphate-buffered saline (PBS) into each tube; pellet the cells at 800 x g for 10 min, and discard the supernatant.
- Resuspend pellets in 150 mM NaCl and 20 mM Tris solution, pH 8.0; use 25 mL of solution/tube. Add 1.25 mL of freshly prepared 10% sodium deoxycholate (in dH2O) to each tube. Add benzonase nuclease to a final concentration of 50 units per mL, and mix thoroughly. Incubate at 37 °C for 1 h.
- Remove cellular debris by centrifuging at 3000 x g for 15 min. Further ensure that all debris has been removed by pushing the mixture through a 0.45 μm syringe filter; this step will help prevent blocking of heparin columns.
- Set up the heparin columns using a peristaltic pump; set the flow rate to 1 mL/min, ensuring that no air bubbles are introduced into the columns.
- Equilibrate the columns with 10 mL of a 150 mM NaCl, 20 mM Tris solution, pH 8.0. Add the solution containing the virus to each column. Wash the columns with 20 mL of a 100 mM NaCl, 20 mM Tris solution, pH 8.0. Then, using a 5 mL syringe, wash the columns with 1 mL of a 200 mM NaCl and 20 mM Tris solution pH 8.0, followed by 1 mL of a 300 mM NaCl and 20 mM Tris solution, pH 8.0.
- Use a 5 mL syringe to elute the virus with 3 mL of a 400 mM NaCl, 20 mM Tris solution, pH 8.0, followed by 4 mL of a 450 mM NaCl, 20 mM Tris solution, pH 8.0, and finally by a 3 mL of a 500 mM NaCl, 20 mM Tris solution, pH 8.0. Collect the extracted materials.
- For recombinant adeno-associated virus (rAAV) concentration, load the extracted materials, 4 mL at a time, into centrifugal filter units with a 100,000 molecular weight cutoff. Centrifuge each 4-mL sample at 2000 x g for 5 min at room temperature, discarding the flow-through.
- When all extracted materials have been run through the centrifugal filter units, add 4 mL of PBS and centrifuge the mixture into approximately 250 μL. Remove rAAVs (viral titer should be about 2 – 8 x 1012 viral particles per mL) from the centrifugal filter units, aliquot and store at -80 °C until use. At this stage, the virus can be stored for several months.
3. rAAV Injection and Optic Fiber Insertion into the Adult Mouse SCN
- Injection of circadian fluorescence reporter-carrying rAAVs into the SCN
- Load 5 µL of rAAV (the virus titer is about 2 – 8 x 1012 virus particles per mL) into a micro-syringe.
- Anesthetize an adult mouse (2- to 5-month-old, C57BL/6 background) with Nembutal (80 mg/kg) via intraperitoneal injection. Check for sufficient depth of anesthesia by the lack of a toe-pinch response.
- Use scissors to remove fur from the head of the mouse and then mount the mouse into a stereotaxic apparatus. Provide a heat source to anesthetized animals via a circulating hot water blanket or another heating device.
NOTE: Alternatively, use depilatory cream or electric hair clippers to remove the fur. - Clean the surgical site with the appropriate disinfecting agents. For example, this could include sterile water or alcohol and iodine, betadine, or dilute chlorhexidine in alternating scrubs. Repeat at least 3 times. Drape the surgical site.
- Use the scissors to make an incision in the scalp and expose the skull.
- Adjust the stereotaxic apparatus so that bregma and lambda are in the same horizontal plane; make two symmetrical points of bregma in the same horizontal plane.
- Use a sterile cotton swab soaked in 4% H2O2/H2O to corrode the brain periosteum. Then use a clean, sterile cotton swab to clear and dry the surface of the skull.
- Using a micro drill, make a hole about 1 mm in diameter 0.46 mm posterior and 0.25 mm lateral to bregma.
- fiUse a clean cotton sterile swab with physiological saline solution to clear bone fragments from the exposed surface of the brain.
- Insert a micro-syringe into the brain, ensuring that the tip of the micro-syringe is 0.46 mm posterior and 0.25 mm lateral to bregma, and 5.7 mm deep from the surface of the skull.
- Inject 500 nL of rAAV (50 nL min-1) into the SCN, leaving the micro-syringe in place for 10 min after the injection to ensure complete diffusion of rAAV.
- Slowly withdraw the micro-syringe.
- Insert the optical fiber into the SCN area.
- Using an optical fiber knife, cut a fiber 17-mm in length, making sure that each side of the optic fiber is smooth.
- Insert the optic fiber into the ceramic ferrule and fix the optic fiber to a ceramic ferrule with methyl ethyl ketone peroxide (AB glue), ensuring that the surfaces of the optical fiber and ceramic ferrule are flush.
- Clean the fiber with a cold sterilant solution or ethylene oxide.
- After rAAV injection (step 3.1.11), insert the ceramic ferrule-containing optic fiber into the SCN.
- Fix the ceramic ferrule and optical fiber onto the skull using dental resin and allow to dry.
- Smear the surface of the dental resin with back nail polish. Repeat the procedures outlined in 3.1.2 to 3.2.5, omitting the actual injection of rAAV for control.
- Post-surgical care
- Provide mice with post-operative analgesics, such as buprenorphine 2 mg/kg body weight, subcutaneously, twice a day.
- Place mice in a recovery cage while they come out of anesthesia.
- House mice individually under a 12 h light/dark condition (light intensity ~100 lux at the bottom of the cage) with food/water available ad libitum. Eight days should be allotted before further experimentation to allow for sufficient recovery time and to ensure optimal fluorescence reporter expression.
4. In vivo Fluorescence Signal Monitoring and Data Collection
- Eight days after surgery, connect the fiber on the mouse head with the fluorescence signal monitor (e.g., Gene Observer). Check the fluorescence signal in the SCN of control mice to establish the background signal. Conduct the observation and measurement of this signal using fluorescence signal monitor.
- Evaluate the fluorescence signal of experimental mouse with the same operation of the control mouse. Exclude mice which have received the viral vector but which only display the background signal from analysis, as this likely indicates that the tip of the optic fiber has missed the SCN. After these initial measures, return mice to their home cages for another 3 weeks to allow the fluorescence signal to stabilize.
- After 3 weeks, connect the mice with the fluorescence signal monitor. Measure the fluorescence for 15 s every 10 min, at a frequency of 100 Hz. During these measurements, the mice are freely moving in their cages with food/water available ad libitum. Use a light intensity at the bottom of the cage of ~100 lux, and a the laser power at the tip of the optic fiber between 15 – 20 µW.
- After the completion of all recordings, perfuse mice with 4% paraformaldehyde, and remove and slice the brains to check placement of the optic fiber. Exclude mice from analysis if the optic fiber tip is not correctly implanted in the SCN.
5. Data Analysis and Presentation
- Measure fluorescence signal every 10 min for 15 s, at a frequency of 100 Hz, to yield 1500 data points. The average of these 1500 points translates to the fluorescence signal at any given time point. Plotting multiple averages over time produces a signal-to-time curve which corresponds to the transcriptional rhythm of Cry1 under a control light condition in the SCN of freely moving mice.
Representative Results
Fluorescence reporter design of Cry1 was show in Figure 1A. Using the approach detailed above, 500 nL of rAAV-P(Cry1)-intron336-Venus-NLS-D2 was successfully injected into the SCN of an adult mouse, and exhibited robust Venus expression (Figure 1B, 1C). Fluorescence signals recorded under 12h/12h light/dark (LD) and dark/dark (DD) conditions (Figure 2) yielded robust circadian rhythm in both conditions. Finally, fluorescence signal was recorded in long and short photoperiod conditions (Figure 3).
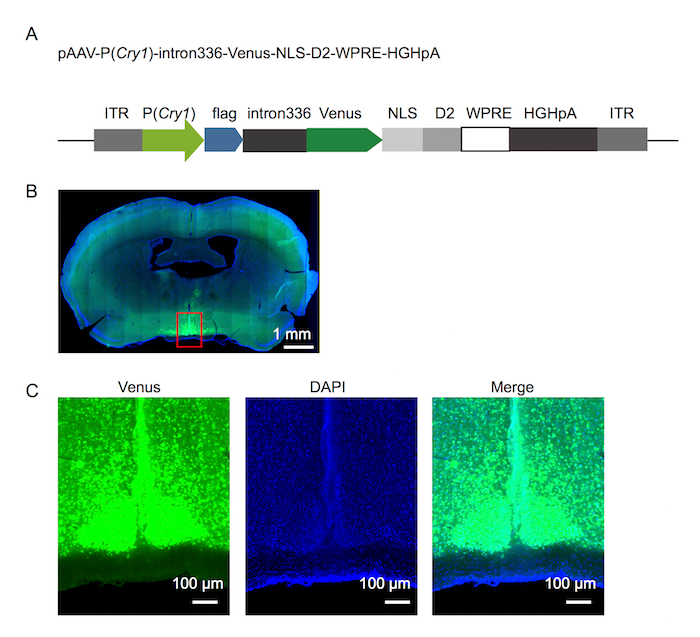
Figure 1. Fluorescence reporter design and it expression in the SCN
(A) The fluorescence reporter design of Cry1. (B) Photomicrograph of the mouse brain showing reporter expression 8 days after the virus injection. (C) Magnification of the area boxed in red on the photomicrograph shown in (B). Please click here to view a larger version of this figure.
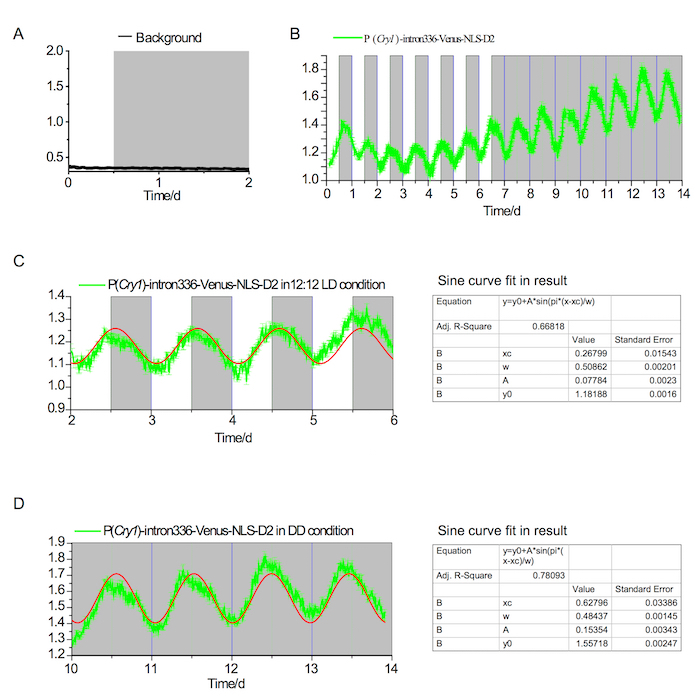
Figure 2. Fluorescence in vivo recording in the SCN under LD and DD conditions
(A) Background fluorescence in the SCN showing no detectable rhythm under both LD and DD conditions. The white and gray areas indicate light-on and light-off periods, respectively. (B) P(Cry1)-intron336-Venus-NLS-D2 expression over the course of 7 days in a 12-12 LD condition and 7 days in a DD condition. (C) Analysis of circadian rhythm in the LD condition (~24.4 h); the red line indicates the converged fit in the Sine curve (the result of the fit is shown in the left panel). (D) Analysis of circadian rhythm in the DD condition (~23.25 h); the red line indicates the converged fit in the Sine curve (the result of the fit is shown in the left panel). Please click here to view a larger version of this figure.
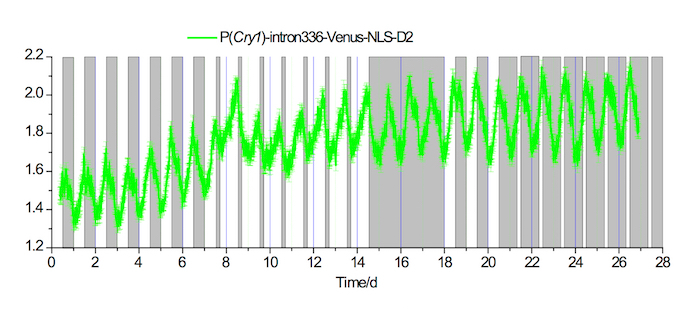
Figure 3. In vivo fluorescence recordings in the SCN under 20-4 long and 4-20 short photoperiod conditions
Example of a fluorescence recording of P(Cry1)-intron336-Venus-NLS-D2 expression in the SCN for 7 days under a 12-12 LD condition, followed by 7 days under a 20-4 long photoperiod condition, 3 days under a DD condition, 3 days under a 12-12 LD condition, and then 7 days under a 4-20 short photoperiod condition. White and gray areas indicate the light-on and light-off periods, respectively. Please click here to view a larger version of this figure.
Table 1. Specific reagents required for protocol (See Table of Materials)
| primer name | sequence | ||
| primer F1 | cactaggggttcctgcggccgcACGCGTGTAAAGATGCACATGTG | ||
| primer R1 | TTGTAACCTTGATACTTACCTACTTAGATCGCAGATCTCGTCCGG | ||
| primer F2 | GTTATGACACAGTGTAGAAACTATGGCATAGGACAGATGACTGTG | ||
| primer R2 | CATGGTCTTTGTAGTCCATGGTGGGTACCtCTTGACAGCTCTACC | ||
| primer F3 | ctgtattttcagggcCCTGCAGGtGTGAGCAAGGGCGAGGAGCTG | ||
| primer R3 | TACCTTTCTCTTCTTTTTTGGAGGCTTGTACAGCTCGTCCATGCC | ||
| primer F4 | GCATGGACGAGCTGTACAAGCCTCCAAAAAAGAAGAGAAAGGTAG | ||
| primer R4 | tgatatcgaattcGGATCCCTACACATTGATCCTAGCAGAAGCAC | ||
Table 2. Primer used in the protocol
Table 3. DNA sequence used in the protocol Please click here to download this file.
Discussion
In contrast to ex vivo methods, such as slice culture4,5, RT-PCR16, and in situ hybridization17, which require that animals be killed, the in vivo recording method allows investigators to study circadian gene expression in a living animal. As such, this technology provides the ability to evaluate the effect of different physical perturbations (e.g., sleep deprivation, stress, food intake, etc.) on the neural circadian clock. In contrast to the in vivo bioluminescence7,8,10 method that can only work in constant dark conditions, the fluorescence in vivo recording method can be applied under light conditions-a key advantage of the technology as light signals re-entrainment of the SCN circadian clock18. Although relatively constant, robust circadian rhythms can be detected with this method, it should be mentioned that the reporter technology does not provide quantitative information and can therefore not be used to measure levels of gene transcription and translation.
There are many critical steps of this protocol. The concentrated rAAV titer should be as high as 2 – 8x 1012 viral particles per mL; lower viral titers would give off dim or even undetectable fluorescence signals. When fixing the head of the mouse into the stereotaxic apparatus, ensure that the skull is horizontal to secure accurate positioning for virus injection and optical fiber implantation. Injection of the virus should be done slowly (≤50 nL min-1) to ensure that the virus remains in the SCN. The ceramic ferrule and optical fiber should be securely fixed on the skull and dental resin should not contact the skin. If the ceramic ferrule and optical fiber are not adequately fixed, they will gradually detach from the mouse head.
This technique can be extended to include other gene reporters (e.g., Per2, Bmal1, c-Fos, Pomc, etc.) by changing the promoter and the corresponding intron19. By reversing the flag-inton336-Venus-NLS-D2 cassette with two reversed lox sequences and combining the reporter with a Cre mouse line, this technology can be used to record activity in specific neurons. For example, combining this approach with a Vip-cre mouse line would allow for the recording of Cry1 transcriptional rhythms in vasoactive intestinal polypeptide-expressing neurons in the SCN. This method can also employ GCaMP6 as a Ca2+ indicator to record the Ca2+ circadian rhythm in the brain.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
We thank members in the Zhang lab for providing stimulating discussions and members in the Zhan lab for providing technical assistance. This research was supported by grants 31500860 (to C.Z.) of the NSFC, 2012CB837700 (to E.E.Z. and C.Z.) of the 973 Program from the M.O.S.T. of China, and by funding from the Beijing Municipal Government. E.E.Z. was supported by the Chinese "Recruitment Program of Global Youth Experts".
Materials
| KOD Plus Neo | TOYOBO | KOD-401 | Reagent |
| pVENUS-N1 | addgene | #61854 | Plasmid |
| pcDNA3.3_d2eGFP | addgene | #26821 | Plasmid |
| pAAV-EF1a-double floxed-hChR2(H134R)-mCherry-WPRE-HGHpA | addgene | #20297 | Plasmid |
| MluI | Thermo Scientific | FD0564 | Reagent |
| EcoRI | Thermo Scientific | FD0274 | Reagent |
| Gibson Assembly Mix | NEB | E2611s | Reagent |
| Lipofectamine 2000 | Thermo Scientific | 12566014 | Reagent |
| Syringe Filter | EMD Millipore | SLHV033RS | 0.45 µm |
| HiTrap heparin columns | gelifesciences | 17-0406-01 | 1 mL |
| Amicon ultra-4 centrifugal filter | EMD Millipore | UFC810024 | 100,000 MWCO |
| Benzonase nuclease | Sigma-Aldrich | E1014 | Reagent |
| Sodium deoxycholate | Sigma-Aldrich | D5670 | Make fresh solution for each batch |
| mouse stereotaxic apparatus | B&E TEKSYSTEMS LTD | #SR-5M/6M | Equipment |
| pentobarbital | SigmaAldrich | #1507002 | Reagent |
| mouse stereotaxic apparatus | B&E TEKSYSTEMS LTD | #SR-5M/6M | Equipment |
| Hydrogen peroxide solution | SigmaAldrich | #216763 | Reagent |
| Optical Fiber | Thorlabs | FT200EMT | 0.39 NA, Ø200 µm |
| microsyringe pump | Nanoliter 2000 Injector, WPI | Equipment | |
| ceramic ferrule | Shanghai Fiblaser | 230 μm I.D., 2.5 mm O.D. | |
| Gene Observer | BiolinkOptics | Equipment |
Riferimenti
- Welsh, D. K., Takahashi, J. S., Kay, S. A. Suprachiasmatic nucleus: cell autonomy and network properties. Annual Review of Physiology. 72, 551-577 (2010).
- Zhang, E. E., Kay, S. A. Clocks not winding down: unravelling circadian networks. Nature Reviews Molecular Cell Biology. 11, 764-776 (2010).
- Takahashi, J. S. Transcriptional architecture of the mammalian circadian clock. Nature Reviews Genetics. 18, 164-179 (2017).
- Yoo, S. H., et al. PERIOD2::LUCIFERASE real-time reporting of circadian dynamics reveals persistent circadian oscillations in mouse peripheral tissues. Proceedings of the National Academy of Sciences U S A. 101, 5339-5346 (2004).
- Davidson, A. J., Castanon-Cervantes, O., Leise, T. L., Molyneux, P. C., Harrington, M. E. Visualizing jet lag in the mouse suprachiasmatic nucleus and peripheral circadian timing system. European Journal of Neuroscience. 29, 171-180 (2009).
- Savelyev, S. A., Larsson, K. C., Johansson, A. S., Lundkvist, G. B. Slice preparation, organotypic tissue culturing and luciferase recording of clock gene activity in the suprachiasmatic nucleus. Journal of Visualized Experiments. (48), (2011).
- Yamaguchi, S., et al. Gene expression: View of a mouse clock gene ticking. Nature. 409, 684-684 (2001).
- Ono, D., Honma, K. I., Honma, S. Circadian and ultradian rhythms of clock gene expression in the suprachiasmatic nucleus of freely moving mice. Science Reports. 5, 12310 (2015).
- Ono, D., Honma, S., Honma, K. Circadian PER2::LUC rhythms in the olfactory bulb of freely moving mice depend on the suprachiasmatic nucleus but not on behaviour rhythms. European Journal of Neuroscience. 42, 3128-3137 (2015).
- Ono, D., et al. Dissociation of Per1 and Bmal1 circadian rhythms in the suprachiasmatic nucleus in parallel with behavioral outputs. Proceedings of the National Academy of Sciences U S A. 114, E3699-E3708 (2017).
- Reppert, S. M., Weaver, D. R. Coordination of circadian timing in mammals. Nature. 418, 935-941 (2002).
- Liu, A. C., et al. Redundant function of REV-ERBalpha and beta and non-essential role for Bmal1 cycling in transcriptional regulation of intracellular circadian rhythms. PLoS Genetics. 4, e1000023 (2008).
- Maywood, E. S., et al. Analysis of core circadian feedback loop in suprachiasmatic nucleus of mCry1-luc transgenic reporter mouse. Proceedings of the National Academy of Sciences U S A. 110, 9547-9552 (2013).
- Ukai-Tadenuma, M., et al. Delay in feedback repression by cryptochrome 1 is required for circadian clock function. Cell. 144, 268-281 (2011).
- McClure, C., Cole, K. L., Wulff, P., Klugmann, M., Murray, A. J. Production and titering of recombinant adeno-associated viral vectors. Journal of Visualized Experiments. 57, e3348 (2011).
- Yamaguchi, Y., et al. Mice genetically deficient in vasopressin V1a and V1b receptors are resistant to jet lag. Science. 342, 85-90 (2013).
- Nagano, M., et al. An abrupt shift in the day/night cycle causes desynchrony in the mammalian circadian center. Journal of Neuroscience. 23, 6141-6151 (2003).
- Golombek, D. A., Rosenstein, R. E. Physiology of Circadian Entrainment. Physiological Reviews. 90, 1063-1102 (2010).
- Mei, L., et al. Long-term in vivo recording of circadian rhythms in brains of freely moving mice. Proceedings of the National Academy of Sciences. 115, 4276-4281 (2018).

