A Human 3D Extracellular Matrix-Adipocyte Culture Model for Studying Matrix-Cell Metabolic Crosstalk
Summary
We describe a 3D human extracellular matrix-adipocyte in vitro culture system that permits dissection of the roles of the matrix and adipocytes in contributing to adipose tissue metabolic phenotype.
Abstract
The extracellular matrix (ECM) plays a central role in regulating tissue homeostasis, engaging in crosstalk with cells and regulating multiple aspects of cellular function. The ECM plays a particularly important role in adipose tissue function in obesity, and alterations in adipose tissue ECM deposition and composition are associated with metabolic disease in mice and humans. Tractable in vitro models that permit dissection of the roles of the ECM and cells in contributing to global tissue phenotype are sparse. We describe a novel 3D in vitro model of human ECM-adipocyte culture that permits study of the specific roles of the ECM and adipocytes in regulating adipose tissue metabolic phenotype. Human adipose tissue is decellularized to isolate ECM, which is subsequently repopulated with preadipocytes that are then differentiated within the ECM into mature adipocytes. This method creates ECM-adipocyte constructs that are metabolically active and retain characteristics of the tissues and patients from which they are derived. We have used this system to demonstrate disease-specific ECM-adipocyte crosstalk in human adipose tissue. This culture model provides a tool for dissecting the roles of the ECM and adipocytes in contributing to global adipose tissue metabolic phenotype and permits study of the role of the ECM in regulating adipose tissue homeostasis.
Introduction
The extracellular matrix (ECM) not only provides a mechanical scaffold for tissues, but also engages in complex crosstalk with cells that reside within it, regulating diverse processes necessary for tissue homeostasis, including cell proliferation, differentiation, signaling, and metabolism1. While healthy ECM plays an essential role the maintenance of normal tissue function, dysfunctional ECM has been implicated in multiple diseases2.
Adipose tissue plays an important role in the pathogenesis of metabolic disease. Obesity is associated with excessive adipocyte hypertrophy and cellular hypoxia, defects in adipocyte cellular metabolism, and adipose tissue endoplasmic reticulum and oxidative stress and inflammation. While poorly understood, these complex processes conspire to impair adipose tissue nutrient buffering capacity, leading to nutrient overflow from adipose tissue, toxicity in multiple tissues, and systemic metabolic disease3,4,5.The sequence of events and specific mechanisms that underlie adipose tissue failure are poorly understood, but alterations in adipose tissue ECM have been implicated. The ECM composition is altered within adipose tissue in human and murine obesity, with increased deposition of ECM protein along with qualitative biochemical and structural differences in the adipose tissue ECM associated with human metabolic disease, including type 2 diabetes and hyperlipidemia6,7,8,9,10,11.
Despite these observations, the role of adipose tissue ECM in mediating adipose tissue dysfunction is not well-defined. This is in part due to a lack of tractable experimental models that permit dissection of the specific roles of ECM and adipocytes in regulating ultimate adipose tissue function. ECM-adipocyte culture better simulates the in vivo environment of native adipose tissue in at least two respects. Firstly, ECM culture provides a molecular environment similar to native adipose tissue, including native collagens, elastins, and other matrix proteins absent in standard 2D culture. Secondly, culture on 2D plastic has been shown to alter adipocyte metabolism via mechanical effects due to decreased elasticity of plastic substrate12, which ECM-culture eliminates.
Methods to engineer biological scaffolds by isolation of ECM from decellularized adipose and other tissues have been studied in the context of regenerative and reconstructive medicine and tissue engineering13,14,15,16,17,18. We have previously published methodology in which we adapted these methods to develop an in vitro 3D model of human ECM-adipocyte culture, using ECM and adipocyte stem cells (preadipocytes) derived from human visceral adipose tissues11. In the present article, we describe these methods in detail. The decellularization procedure for human adipose tissue is a four-day process that involves mechanical and enzymatic treatments to remove cells and lipid, leaving a biological scaffold that maintains characteristics of the tissue from which it is derived. Decellularized ECM supports adipogenic differentiation of human preadipocytes, and when reconstituted with adipocytes, maintains microarchitecture and biochemical and disease-specific characteristics of intact adipose tissue and engages in metabolic functions characteristic of native adipose tissue. This matrix can be studied alone or reseeded with cells, permitting study of interactions and crosstalk between the cellular and extra-cellular components of adipose tissue.
Protocol
Adipose tissues are procured from human subjects undergoing elective bariatric surgery under institutional review board approval.
1. Preadipocyte isolation and culture reagent preparation
- Prepare 2% bovine serum albumin (BSA) in 1x phosphate buffered saline solution (PBS). Filter sterilize, and store at 4 °C.
- Prepare Type II collagenase: 2 mg/mL in 2% BSA in 1x PBS. Prepare immediately before use.
- Prepare Red Blood Cell (RBC) Lysing Solution: 1.5 M NH4Cl, 100 mM NaHCO3, 10 mM disodium EDTA in deionized water (DI/H2O). Store at 4 °C. Prepare 1x RBC Lysing Solution from 10x stock solution in DI/H2O immediately before use.
- Prepare Growth Media: 15% fetal bovine serum (FBS), 1% antibiotic-antimycotic solution (ABAM) in Dulbecco's Modified Eagle Medium:Nutrient Mixture F-12 (DMEM/F12). Filter sterilize, and store at 4 °C.
- Prepare Preadipocyte Freezing Solution: 10% Dimethyl Sulfoxide, 15% FBS in DMEM/F12 media. Filter sterilize, and store at 4 °C.
- Prepare Differentiation Media: 10 mg/L transferrin, 33 µM biotin, 0.5 µM human insulin solution, 17 µM D-pantothenic acid hemicalcium salt, 100 nM dexamethasone, 2 nM 3,3',5-Triiodo-L-thyronine sodium salt (T3), 1 µM ciglitizone, 540 µM 3-Isobutyl-1-methylxanthine (IBMX),1% ABAM in DMEM/F12.Filter sterilize, and store at 4 °C.
2. ECM reagent preparation
- Prepare Freezing Buffer Solution: 10 mM Tris base, 5 mM EDTA, 1% ABAM, 1% phenylmethylsulfonyl fluoride (PMSF) in DI/H2O. Stir solution to dissolve EDTA. Adjust pH to 8.0 with HCl or NaOH.Store at 4 °C for up to 3 months.
- Prepare Enzymatic Solution #1: 1% ABAM in 0.25% trypsin-EDTA. Store at 4 °C for up to 3 months.
- Prepare Rinsing Buffer Solution: 137 mM NaCl, 2.68 mM KCl, 7 mM Na2HPO4, 1.5 mM KH2PO4, 1% ABAM, 1% PMSF in sterilized DI/H2O. Stir to dissolve salts. Adjust pH to 8.0 with HCl or NaOH. Store at 4 °C for up to 3 months.
- Prepare Enzymatic Solution #2: 55 mM Na2HPO4, 17 mM KH2PO4, 4.9 mM MgSO4∙7H2O, 1% ABAM, 1% PMSF in DI/H2O. Store 4 °C for up to 3 months. Stir to dissolve salts. Immediately prior to use, add 80 U/mL lipase from porcine pancreas, type VI-S; 160 U/mL deoxyribonuclease I from bovine pancreas, type II-S; and 100 µg/mL ribonuclease A from bovine pancreas, type III-A.
- Prepare Polar Solvent Extraction Solution: 1% ABAM, 1% PMSF in isopropanol.
CAUTION: Isopropanol is flammable; store in a flammable cabinet at 25 °C and dispose in flammable waste. - Prepare 70% ethanol, 1% ABAM, 1% PMSF in DI/H2O. Add ABAM and PMSF just prior to use.
CAUTION: Ethanol is flammable; store in a flammable cabinet at 25 °C and dispose in flammable waste. - Prepare Storage Solution: 1% ABAM, 1% PMSF in 1x PBS. Store at 4 °C for up to 3 months.
3. Metabolic phenotyping reagent preparation
- Glucose uptake
- Prepare Serum Starvation Media: DMEM/F12, 1% ABAM. Filter sterilize, and store at 4 °C
- Prepare 200 nM human insulin solution in 1x PBS immediately before use.
- Prepare 200 nM human insulin, 0.1 mM 2-Deoxy-D-glucose, 1 µCi/well Deoxy-D-glucose, 2-[1,2-3H(N)]-, in 1x PBS. Prepare immediately before use.
- Lipolysis
- Prepare Isoproterenol diluted in PBS: 3 mM stock solution. Dilute to working concentration of 3 µM for assay.
- Oil Red-O Staining
- Prepare 4% Formalin in DI/H2O. Store at room temperature.
- Prepare Oil Red-O Working Solution. Dilute Oil Red-O Solution (ORO) with DI/H2O in a 3:2 ratio (ORO:DI/H2O). Prepare immediately before use. Filter through filter paper (Table of Materials).
4. Adipose tissue procurement
NOTE: Visceral adipose tissue (VAT) is collected from the greater omentum at the beginning of the operation by the surgeon and transported back to the laboratory on ice for immediate processing. Universal precautions should be used when handling all human tissues and caustic reagents, including performing all work in a laminar flow hood, using complete laboratory safety wear, and no recapping of needles.
- Add 5-10 g of intact VAT to 15-25 mL of Freezing Buffer Solution in a 50 mL conical tube to immerse the tissue sample. Store samples at -80 °C until decellularization, for up to 1 month.
- Use a separate fresh sample of VAT for preadipocyte isolation as described in section 5.
5. Preadipocyte isolation
- Place 2 g of intact VAT in 20 mL of the collagenase, type II, solution in a 50 mL conical tube. Then mince thoroughly by inserting sterile scissors into the conical tube and mincing the tissue within the tube. Once fully minced to a fine slurry, incubate the tissue in the collagenase solution on an orbital shaker at 130 rpm and 37 °C for 60 min.
- Filter the resultant digestate through a 100 µm nylon mesh into a fresh 50 mL conical tube by pouring the digestate from one conical tube through a piece of mesh folded over the top of a fresh conical tube. The digestate at this point should be a yellow-orange liquid with moderate viscosity, with small amounts of residual strands of undigested fibrous tissue. The mesh should capture larger pieces of undigested tissue, which are discarded.
- Centrifuge the sample at 270 x g for 10 min. Remove the supernatant and resuspend the cell pellet in 2 mL of 1x RBC Lysing Solution with a pipette.
- Incubate for 1 min at 25 °C and then add 10 mL of 15% FBS-DMEM/F12. Centrifuge at 270 x g for 10 min.
- Remove the supernatant and resuspend the cell pellet in 10 mL of 15% FBS-DMEM/F12 with a pipette. Transfer the cell suspension to 100 mm Petri dish with a pipette and incubate at 37 °C and 5% CO2, until cells reach 80-100% confluence, typically 2-6 days. Change the media every 2-3 days.
- Detach and wash cells.
- Remove media with a pipette and apply 4 mL of 0.25% trypsin-EDTA to adherent cells. Incubate at 37 °C for 10 min, periodically swirling the plate gently to detach the cells.
- Add 20 mL of 15% FBS-DMEM/F12 and resuspend the detached cells in this media with a pipette. Then transfer to a fresh 50 mL conical tube and centrifuge 270 x g for 10 min.
- Remove the supernatant and discard. Wash the cell pellet once in 1x PBS, and then resuspend the cell pellet in 20 mL of fresh 15% FBS-DMEM/F12 with a pipette. Transfer the cell suspension to a T-150 culture flask.
- Culture cells at 37 °C and 5% CO2. Split and expand cells every 2-3 days as they reach 80-100% confluence by applying 7 mL of 0.25% trypsin-EDTA, expanding from one flask to 8 flasks.
NOTE: This typically requires 3-4 passages, which permits appropriate expansion and retains adipogenic potential and patient- and depot-specific cellular metabolic phenotypes. Passaging preadipocytes in excess of 4-5 passages leads to loss of adipogenic potential. - Detach cells in 8 flasks with 7 mL of 0.25% trypsin-EDTA per flask as described above, and incubate at 37 °C for 10 min.
- Add 8 mL of 15% FBS-DMEM/F12 per flask and resuspend the detached cells with a pipette. Transfer the entire cell suspension divided evenly into three 50 mL conical tubes and centrifuge at 270 x g for 10 min.
- Resuspend the resultant cell pellets in 5 mL of 15% FBS-DMEM/F12 in a 15 mL conical tube and count cells using cell counter and Trypan blue.
- Centrifuge the cell suspension at 270 x g for 10 min. Then resuspend the cell pellet in Preadipocyte Freezing Solution to a final cell concentration of 1 x 106/mL, and aliquot 1 mL of cell suspension per 1.5 mL cryovial tube.
- Store cells in cryovials for 1 days at -80 °C. Then transfer cryovials to liquid nitrogen for long-term storage for 3-6 months.
- When ready for use, thaw one cryovial in a 37 °C water bath for 3-5 min. Resuspend the cells in 20 mL of 15% FBS-DMEM/F12, and centrifuge at 270 x g for 10 min.
- Resuspend the cell pellet in 20 mL of 15% FBS-DMEM/F12, pipette into a single T-150 flask, and then grow to 80% confluence over 2-3 days at 37 °C and 5% CO2.
- Detach cells with 7 mL of 0.25% trypsin-EDTA per flask as described above in step 5.6. Resuspend at 3 million cells per mL (i.e., 6 x 104 cells per 20 µL) in 15% FBS-DMEM/F12, and use as outlined below (section 7, step 7.4).
6. Adipose tissue ECM preparation
- Day 1: Freeze-thaw and enzymatic digestion #1
- Freeze-thaw previously frozen (step 4.2) VAT samples stored in Freezing Buffer Solution in 50 mL conical tubes from -80 °C to 37 °C in a preheated water bath, incubating 20 min with gentle periodic manual agitation. Once thawed, transfer back to -80 °C and incubate 20 min. Repeat freeze-thaw 3x, ending by thawing samples in a 37 °C water bath.
- Using sterile forceps, transfer the VAT samples to fresh 50 mL conical tubes containing 15-25 mL of Enzymatic Solution #1, ensuring that the VAT samples are fully immersed. Then incubate overnight on an orbital shaker (130 rpm, 37 °C).
- Day 2: Enzymatic digestion #2
- Wash samples 3x with 15-25 mL of Rinsing Buffer Solution on an orbital shaker (130 rpm, 37 °C, 20 min each wash). Pour off Rinsing Buffer Solution after each wash.
- Transfer samples to fresh 50 mL conical tubes containing 15-25 mL of Enzymatic Solution #2 and incubate on an orbital shaker (130 rpm, 37 °C, overnight).
- Day 3: Delipidation
- Wash samples 3x with 15-25 mL of Rinsing Buffer Solution on an orbital shaker (130 rpm, 37 °C, 20 min each wash). Pour off Rinsing Buffer Solution after each wash.
- Transfer samples to fresh 50 mL conical tubes containing 15-25 mL of Polar Solvent Extraction Solution and incubate on an orbital shaker (130 rpm, 25 °C, overnight). After this step, a majority of the lipid should be removed, and the samples should be white or translucent in color.
CAUTION: The polar solvent extraction solution is flammable and should be stored and used at 25 °C.
- Day 4: Wash and storage
- Transfer samples to fresh 50 mL conical tubes containing 15-25 mL of Rinsing Buffer Solution. Wash samples 3x on an orbital shaker (130 rpm, 37 °C, 20 min each wash).
- Wash samples 3x with 15-25 mL of 70% ethanol on an orbital shaker (130 rpm, 37 °C, 20 min each wash) pouring off the 70% ethanol solution after each wash.
- Wash samples once with Storage Solution on an orbital shaker (130 rpm, 37 °C, 20 min each wash).
- Using sterile forceps, transfer samples to fresh 50 mL conical tubes containing 15-25 mL of Storage Solution. Ensure enough Storage Solution is used to fully immerse samples. Store at 4 °C for up to 1 month.
7. ECM-adipocyte preparation
- Transfer stored ECM fragments to individual wells of 24-well plate using sterile forceps. Add as many ECM fragments into as many wells as required for the planned downstream assay (e.g., glucose uptake or lipolysis, see below), including duplicates or triplicates. Wash with 500 µL of 70% ethanol 3x on an orbital shaker (130 rpm, 37 °C, 20 min each wash).
- Rehydrate ECM by washing 3x in sterile 1x PBS on an orbital shaker (130 rpm, 37 °C, 20 min each wash).
- Using sterile scissors, cut and weigh ECM into 100 mg fragments. Using sterile forceps, place one 100 mg fragment in each well of a 24-well plate. Incubate at 25 °C for 15 min to allow excess PBS to extrude from fragments. Carefully remove any excess PBS with a pipette.
- Seed each 100 mg ECM fragment with 20 µL of preadipocyte cell suspension (3 million cells per mL, 6 x 104 cells per 20 µL, in 15% FBS-DMEM/F12, from step 5.10). Pipette the cells directly into the ECM by placing the tip of pipette in the ECM and gently expelling the cell suspension into center of the matrix, taking care that the cell suspension does not overflow and end up on the bottom of the well.
- If the cell suspension is overflowing from the ECM where the pipette tip has been placed, remove the tip from that location and insert somewhere else in the ECM. Incubate seeded ECM for 40 min at 37 °C.
NOTE: For RNA extraction for qrtPCR, seed each 500 mg ECM fragments with 3 x 105 cells in 100 µL (3 million cells per mL, i.e., 3 x 105 cells per 100 µL, in 15% FBS-DMEM/F12).
- If the cell suspension is overflowing from the ECM where the pipette tip has been placed, remove the tip from that location and insert somewhere else in the ECM. Incubate seeded ECM for 40 min at 37 °C.
- Fill each well of the 24-well plate with 500 µL of growth media to cover the seeded ECM fragments. Culture at 37 °C and 5% CO2 for 72 h.
- After 72 h, carefully aspirate 15% FBS-DMEM/F12, tilting the plate slightly to allow media to pool below fragment, and placing the pipette tip just adjacent to the ECM fragment without disturbing it. After aspirating, add 500 µL of Differentiation Media, changing media every 2-3 days using a similar technique, for a total culture period of 14 days.
- Check for differentiation using light microscopy: cells will accumulate lipids, turn brown-yellow in color and more spherical in shape.
NOTE: Seeded matrices can be used for metabolic testing (e.g., glucose uptake assay, lipolysis assay, ORO), histology or immunohistochemistry (IHC), or standard tissue imaging. For fixed tissue ORO staining and imaging, freeze ECM-adipocyte samples in liquid nitrogen.
- Check for differentiation using light microscopy: cells will accumulate lipids, turn brown-yellow in color and more spherical in shape.
8. Metabolic phenotyping
- Scanning electron microscopy
- Fix samples in 2.5% glutaraldehyde in Sorensen's phosphate buffer at 25 °C for 12 h. Postfix in 1% osmium tetroxide in Sorensen's phosphate buffer at 4 °C for 1 h.
- Serially dehydrate samples in ethanol. Wash in hexamethyldisalizane, and air-dry. Then mount on a scanning electron microscopy stub with colloidal graphite. Dry, and sputter-coat with gold.
- Capture images with a scanning electron microscope.
- Oil Red-O Staining
- Live Tissue: Oil Red-O Solution
- Carefully aspirate media from wells with a pipette. Then wash samples once with 500 µL of 1x PBS per well.
- Fix samples with 200 µL of 4% formalin in sterile deionized H2O at 25 °C for 15 min. Aspirate formalin with a pipette, wash samples two times with 1x PBS (500 µL each wash).
- Add 200 µL of 60% isopropanol samples at 25 °C for 5 min. Aspirate 60% isopropanol with a pipette.
- Stain samples with Oil Red-O working solution at 25 °C for 5 min. Aspirate Oil Red-O with a pipette and then wash samples 3x with 1x PBS (500 µL each wash). Then image with an optical microscope.
- Fixed Tissue: Oil Red-O Stain Kit
- Flash-freeze ECM-adipocyte samples in optimal cutting temperature (OCT) compound and section (5 µm) on a cryostat.
- Place the slide in 85% propylene glycol in DI/H2O for 2 min. Place the slide in ORO stain at 60 °C for 6 min. Place steh lide in 85% propylene glycol in DI/H2O for 1 min. Rinse the slide twice with DI/H2O.
- Place the slide in Modified Mayer's Hematoxylin from Oil Red-O staining kit for 1 min. Rinse the slide twice with tap water. Rinse slide twice with DI/H2O.
- Mount the coverslip using aqueous mounting medium and image on a microscope.
- RNA extraction from ECM for qrtPCR
NOTE: To maximize RNA yield, use 500 mg ECM fragments seeded with 3 x 105 preadipocytes in 100 µL and differentiate as above in 6-well plates in 3 mL of Differentiation media per well.- Once differentiated, transfer each individual ECM-adipocyte sample into a 50 mL conical tube on ice using sterile forceps.
- Wash empty well with 500 µL of Buffer RLT. Add Buffer RLT to 50 mL conical tube with matching ECM-adipocyte sample.
- Using sterile scissors, finely mince each ECM-adipocyte sample within the 50 mL conical tube, while holding the tube on ice, inserting the scissors into the conical tube to mince the tissue.
- Completely freeze and thaw the conical tubes from -80 °C to 37 °C 3x.
- Centrifuge conical tubes at 500 x g and 4 °C for 10 min.
- Carefully remove the supernatant with a pipette and use for RNA extraction with a Fibrous Tissue RNA extraction Kit (Table of Materials).
- Live Tissue: Oil Red-O Solution
- Glucose uptake assay
- Differentiate 6 x 104 preadipocytes in 100 mg ECM fragments in 0.5 mL of differentiation medium in 24-well plates as described above (section 5).
- After 14 days of differentiation, remove the medium and wash ECM-adipocytes once with 1x PBS. Add 0.5 mL/well Serum Starvation Medium and culture at 37 °C and 5% CO2 for 12 h.
- Remove medium and wash cells twice with 1x PBS. Add 0.5 mL/well 2% BSA in PBS and culture at 37 °C and 5% CO2 for 2 h.
- Wash cells once with 1x PBS, add 0.5 mL/well 1x PBS with or without 200 nM insulin, and incubate at 37 °C for 40 min.
- Aspirate 1x PBS, add 0.5 mL/well 1X PBS with 0.1 mM 2-deoxy-D-glucose, 2 µCi/mL deoxy-D-glucose, 2- [1,2-3H(N)], with or without 200 nM insulin, and incubate at 37 °C and 5% CO2 for 40 min. Use standard precautions for handling and disposal of all radioactive reagents and waste, as mandated by local institutional regulatory statutes.
- Remove medium with a pipette and wash cells 3x with 1x PBS. Add 420 µL of 1% SDS solution in DI/H2O, and lyse cells with vigorous pipetting. Incubate 25 °C for 10 min.
- Collect 5 µL from each well for Bradford protein assay. Transfer 400 µL of remaining cell lysate into 2 mL of scintillation fluid in a scintillation vial. Count 3H-2DG activity on scintillation counter. Analyze data as counts per minute normalized to protein, mg/mL.
- Lipolysis assay
- Differentiate 6 x 104 preadipocytes in 100 mg ECM fragments in 0.5 mL of human differentiation medium in 24-well plates as described above (section 5).
- After 14 days of differentiation, remove medium and wash cells twice with warm 1x PBS. Add 0.5 mL of serum starvation medium (without insulin) with or without 3 µM isoproterenol, and culture adipocytes at 37 °C and 5% CO2 for 72 h.
- Collect culture supernatants, which may be stored at -80 °C until ready for assay. Collect the ECM in microcentrifuge tubes for DNA quantitation for data normalization.
- Pipet 2 µL of each supernatant into a 96-well microplate. Reserve wells for blanks (distilled H2O) and glycerol standard solution provided in Triglyceride Determination Kit.
- Add 270 µL of free glycerol reagent from Triglyceride Determination Kit to each well, pipette to mix. Incubate plate at 37 °C for 5 min.
- Measure absorbance at 540 nm on a microplate spectrophotometer.
- Calculate the concentration of glycerol and normalize with DNA from ECM:

Representative Results
Preparation of adipose tissue ECM, seeding with preadipocytes, and in vitro differentiation into mature adipocytes result in clear sequential morphologic changes in tissue that permits visual assessment of progress throughout the protocol (Figure 1). Preadipocytes used to seed the ECM are isolated using collagenase digestion from separate VAT samples (Figure 2). Scanning electron microscopy of ECM-adipocyte constructs at each stage of processing reveals decellularization of ECM and subsequent recapitulation of lipid-containing adipocytes upon reseeding and differentiation (Figure 3). Collagen 1-specific immunohistochemistry of decellularized ECM reveals maintenance of collagen microarchitecture, while Oil Red-O staining of live and fixed/sectioned 3D ECM-adipocyte constructs reveals lipid-containing adipocytes (Figure 4A). qrtPCR from adipocytes differentiated in ECM demonstrates marked upregulation of adipogenic genes relative to undifferentiated preadipocytes cultured in ECM (Figure 4B). Metabolic phenotyping of ECM-adipocyte constructs studying all combinations of ECM and adipocytes from diabetic (DM) and non-diabetic (NDM) subjects reveals impaired glucose uptake and lipolytic function in ECM-adipocyte constructs from DM relative to NDM tissues, recapitulating DM-specific metabolic defects we have previously reported in human adipocytes in standard 2D culture11. Importantly, NDM ECM rescues insulin-stimulated glucose uptake and lipolytic capacity in DM adipocytes (Figure 4C)11. Together, these data demonstrate disease-specific regulation of adipocyte cellular metabolism by the ECM. Further details are reported in Baker et al.11.
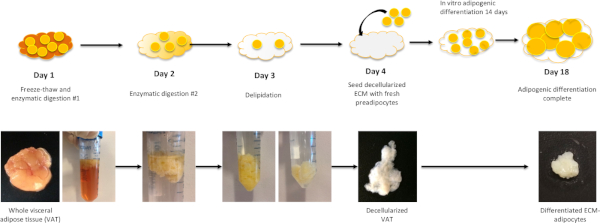
Figure 1: Workflow for ECM-adipocyte preparation. Day 1: Whole visceral adipose tissue samples undergo three freeze-thaw cycles followed by overnight incubation with enzymatic digestion solution #1. Day 2: Following digestion with enzymatic solution #1, samples are digested overnight with enzymatic digestion solution #2. At this point, samples will be partially delipidated. Day 3: Following enzymatic digestion #2, samples undergo overnight incubation with polar solvent extraction solution, which will complete the delipidation. After Day 3, samples should be fully delipidated and white/translucent in color. Samples are washed thoroughly with rinsing buffer solution then 70% ethanol and stored in storage solution in 40 °C until ready for reseeding with preadipocytes. Day 4: Preadipocytes are seeded into decellularized VAT followed by adipogenic differentiation for 14 days. Please click here to view a larger version of this figure.
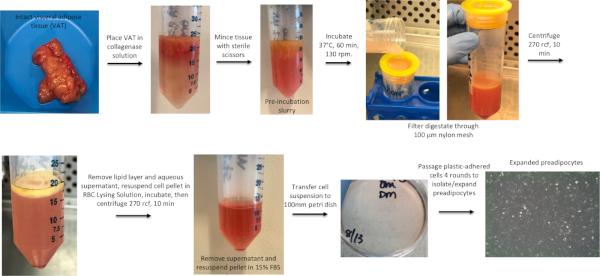

Figure 2: Workflow for preadipocyte isolation. VAT is minced, digested with collagenase, filtered, centrifuged, and the resultant stromovascular cell pellet plated and cultured to expand preadipocytes. See Baker et al. 201721 for further details regarding human preadipocyte isolation and 2D adipocyte culture. Please click here to view a larger version of this figure.
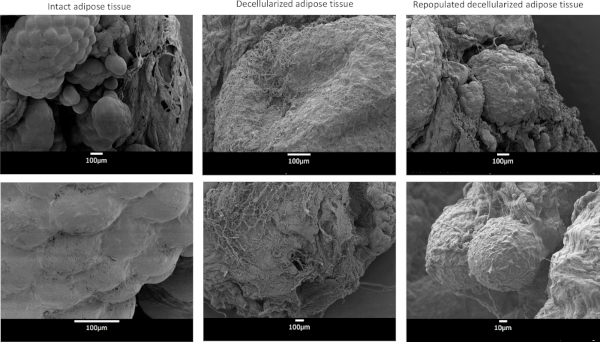

Figure 3: Scanning electron microscopy images. Intact adipose tissue, decellularized adipose tissue, and decellularized adipose tissue repopulated with preadipocytes and differentiated in adipogenic media for 14 days. Please click here to view a larger version of this figure.
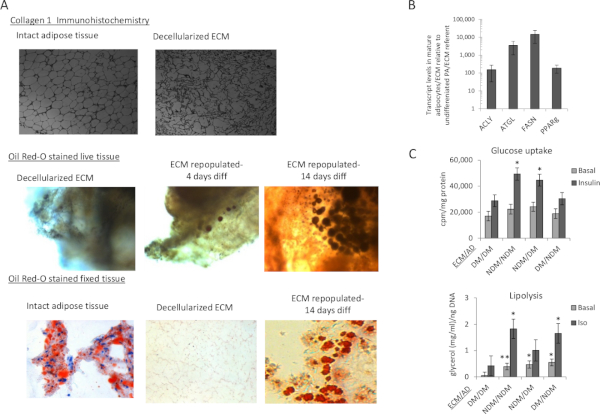
Figure 4: Characterization of adipocytes in ECM. (A) Decellularized adipose tissue ECM maintains microarchitecture and supports adipocyte differentiation: Top: Collagen 1 immunohistochemistry of whole human VAT before and after decellularization demonstrating maintenance of microarchitecture; middle: 3D confocal photomicrographs of live human adipocytes within ECM; intact decellularized human VAT stained with Oil Red-O before reseeding with preadipocytes, 4 days after seeding, and 14 days after seeding with preadipocytes followed by adipogenic differentiation; blue: DAPI staining of cell nuclei; red: Oil Red-O staining of intracellular lipid; bottom: Formalin-fixed, paraffin-embedded, 5 µm sectioned, Oil Red-O-stained human VAT prior to decellularization, immediately after decellularization, and after decellularization, preadipocyte-seeding, and 14 days of adipogenic differentiation, demonstrating cytoplasmic lipid accumulation in adipocytes within ECM. (B) Adipocytes in ECM upregulate adipogenic gene expression: qrtPCR analysis comparing adipogenic gene transcript levels in RNA from human VAT adipocytes differentiated in VAT ECM for 14 days relative to undifferentiated preadipocytes in VAT ECM cultured for 72 h in nonadipogenic media. Data are mean ± SEM. ACLY: ATP citrate lyase, ATGL: adipose triglyceride lipase, FASN: fatty acid synthase, PPARg: peroxisome proliferative activated receptor gamma; ordinate: fold difference in transcript level in mature adipocytes relative to undifferentiated preadipocyte referent=1; all fold differences significant (p<0.001, paired t-test); ECM from 10 subjects, preadipocytes/adipocytes from n=11 subjects. (C) ECM regulates adipocyte metabolism in a DM-specific manner: 3D-ECM from DM or NDM subjects seeded with preadipocytes from DM or NDM subjects, differentiated into adipocytes, and studied with metabolic phenotyping. Data bars labeled with patient source (NDM, DM) of ECM and adipocytes (ECM/AD); e.g., NDM/NDM denotes both ECM and preadipocytes derived from NDM patients, while NDM/DM denotes ECM from NDM patients combined with preadipocytes from DM patients. Data are mean ± SEM. Ordinates: glucose uptake (cpm), normalized to ECM/cell lysate protein concentration(mg/mL); lipolysis: culture supernatant glycerol concentration(mg/mL) normalized to ECM/cell lysate DNA concentration(ng/mL); *p<0.050, **p<0.100 comparing indicated data-point to corresponding data-point (basal or insulin-stimulated for glucose uptake, basal or isoproterenol-stimulated for lipolysis) in DM/DM arm, using mixed model analysis adjusting for repeated measures; ECM from n=9 NDM, 8 DM subjects, preadipocytes from 10 NDM, 9 DM subjects for glucose uptake; ECM from 6 NDM, 6 DM subjects, preadipocytes from 6 NDM, 6 DM subjects for lipolysis. This figure has been adapted from O'Rourke and Lumeng11. Please click here to view a larger version of this figure.
Discussion
The ECM-adipocyte culture model provides a valuable tool for dissecting the individual roles of ECM and cells in dictating ultimate tissue phenotype. The ECM isolation protocol is quite reproducible, but variability in the decellularization process may be observed. The Day 3 delipidation step is a critical point in the protocol. At the completion of the overnight extraction, delipidation of the matrix should be evidenced by the Polar Solvent Solution turning yellow, while the matrix should transition from a yellow-orange color characteristic of intact adipose tissue to translucent/white (Figure 1). If these color changes do not occur, then delipidation is likely not complete. Inter-patient variability in delipidation efficiency is common, but to date we have not observed any correlation of efficiency of delipidation with subject DM status, age, BMI, or other clinical variables. Large samples (greater than 20 g) will not undergo efficient delipidation; processing samples <20 g may prevent this problem. If a sample does not appear completely delipidated after Day 3, divide the sample in half with sterile scissors, then transfer samples to fresh 15-20 mL of Polar Solvent Extraction Solution and place on an orbital shaker for 3-5 h. Some samples may not fully decellularize after repeating the polar solvent extraction step. If a majority of the sample is decellularized, it is possible to cut out the sections containing lipid before using the sample in experiments.
Contamination may occur if sterility is not diligently maintained throughout the process. All work should be performed in a laminar flow hood using standard cell culture precautions. We typically store ECM for up to 3 months at 4 °C, and preadipocytes for up to 6 months in liquid nitrogen. Retention of biofunctionality and disease- and depot-specific properties beyond this time has not been tested.
Confocal microscopy permits visualization of mature adipocytes within the ECM, which may be enhanced with Oil Red-O staining. Scanning electron microscopy (SEM) also provides a non-quantitative method for visualizing adipocytes within ECM. Of note, SEM and Oil Red-O staining suggest unilocular adipocytes in the ECM-adipocyte constructs, a morphology that is similar to in vivo adipocytes and in contrast to the typical multilocular adipocytes observed when preadipocytes undergo adipogenic differentiation on 2D plastic (Figure 3, Figure 4A). This observation suggests that the ECM provides a more physiologic environment for adipogenic differentiation than standard tissue culture on plastic. Fixation and sectioning of ECM-adipocyte constructs may be difficult, as OCT-embedded constructs are fragile and do not section easily. Sectioning embedded tissues immediately after removal from storage from the -80 °C freezer and performing sectioning in a cold-room (4 °C) on a pre-chilled cryostat makes sectioning feasible.
ECM-adipocyte culture provides a physiologic environment that approximates the in vivo adipose tissue environment and provides a sustainable environment for preadipocyte stem cell growth and differentiation, as evidenced by upregulation of adipogenic genes in 3D-ECM-adipocyte culture. ECM-adipocyte cultures also engage in adipocyte metabolic functions, including glucose uptake and lipolysis8. We report glucose uptake as cpm normalized to either total protein extracted from ECM-adipocyte cultures measured with a Bradford assay, or to DNA content measured with a DNA assay kit, and have observed similar results with both methods. Similarly, we report lipolysis as mg/mL of glycerol release normalized to either protein or DNA, also with similar results obtained with either normalization method. Others suggest normalizing adipocyte metabolic data to lipid content, but to date our attempts to reliably extract lipid from ECM constructs has been unsuccessful. Reporting glucose uptake rate as moles of glucose per unit of time and lipolysis as moles of glycerol/free fatty acid per unit of time may permit more accurate comparisons between separate experiments. Isolation of RNA from ECM for qrtPCR analysis is characterized by lower yields compared with cells in 2D culture, but preparation of larger ECM fragments seeded with more cells overcomes this problem, as described in step 8.3.1. We have encountered difficulty in isolating adequate quantities of undegraded protein with phosphomoieties intact for Western blot analysis, which represents a limitation of the model.
Similar methodology for isolating ECM from adipose tissue and seeding with preadipocytes has been previously published, with evidence suggesting that ECM may promote adipogenic differentiation absent classic adipogenic mediators including cAMP agonists, insulin, and PPAR-γ agonists12,14. Our data is the first to demonstrate disease-specific regulation of cellular metabolism by ECM in adipose tissue, using methods in which adipocytes are stimulated with classic adipogenic differentiation factors11; similar studies in the absence of adipogenic stimuli are a target of future research. Others have demonstrated similar disease-specific ECM-cell crosstalk using ECM and cells isolated from lung tissue19,20. Alternative strategies have also been employed, including creation of matrices by mixing homogenized adipose tissue ECM preparations in peptide hydrogels12.
While inter-patient variability in human adipocyte cellular metabolism is observed in both 2D-plastic and 3D-ECM culture, when analyzed in aggregate, these cells manifest disease-, depot- and sex-specific differences in cellular metabolism, as described by our group and others11,21,22,23,24, validating the utility and generalizability of these culture models for the study of adipocyte cellular metabolism. We have demonstrated that adipocytes differentiated in disease-matched ECM (i.e., NDM adipocytes in NDM ECM, DM adipocytes in DM ECM) recapitulate metabolic phenotypes of isolated NDM and DM adipocytes in standard 2D culture, supporting ECM-adipocyte culture as a model for studying disease-specific alterations in adipocyte cellular metabolism in a more physiologic environment. We have not observed differences in the magnitude or direction of metabolic functions, including glucose uptake or lipolysis, when ECM and adipocytes from the same or different donors are studied. In either case, ECM-adipocyte constructs manifest DM-specific metabolic phenotype (e.g., decreased GU in DM/DM relative to NDM/NDM constructs), with no clear difference when comparing constructs comprised of ECM and adipocytes from the same or different donors.
ECM-adipocyte culture allows study of combinations of ECM and preadipocytes from distinct patient populations and tissue depots, permitting analysis of disease- and depot-specific contributions of ECM and adipocytes to ultimate adipose tissue metabolic phenotype. Demonstrating this strength of the ECM-adipocyte culture model, we have shown that ECM from NDM tissue has the capacity to rescue metabolic defects in DM adipocytes (Figure 4C)11. Preliminary data in murine visceral and subcutaneous adipose tissues suggest that murine ECM regulates murine adipocyte metabolism similarly in a depot-specific manner (unpublished data, O'Rourke RW, 2019), suggesting that this model system may be used to study ECM-adipocyte crosstalk in both murine and human systems. Furthermore, this model permits study of isolated manipulation of ECM as a means to regulate adipose tissue phenotype. Preliminary study of ECM treated in conditions that induce glycation targeted specifically to the ECM, for example, suggest that these modifications regulate the cellular metabolic phenotype of cells subsequently seeded into treated ECM (unpublished data, O'Rourke RW, 2019), providing a model for study of the effects of targeted manipulation of the ECM on adipocyte phenotype.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
We thank Danielle Berger, Marilyn Woodruff, Simone Correa, and Retha Geiss for assistance with study coordination. SEM was performed by University of Michigan Microscopy & Image Analysis Laboratory Biomedical Research Core Facility. This project was supported by NIH grants R01DK097449 (RWO), R01DK115190 (RWO, CNL), R01DK090262 (CNL), Veterans Affairs Merit Grant I01CX001811 (RWO), Pilot and Feasibility Grant from the Michigan Diabetes Research Center (NIH Grant P30-DK020572) (RWO), Veterans Administration VISN 10 SPARK Pilot Grant (RWO). Scanning electron microscopy performed by the University of Michigan Microscopy & Image Analysis Laboratory Biomedical Research Core Facility. Figure 4 of this manuscript was originally published in Baker et al., J Clin Endo Metab 2017; Mar 1;102 (3), 1032-1043. doi: 10.1210/jc.2016-2915, and has been reproduced by permission of Oxford University Press [https://academic.oup.com/jcem/article/102/3/1032/2836329]. For permission to reuse this material, please visit http://global.oup.com/academic/rights.
Materials
| 0.25% trypsin-EDTA | Gibco, ThermoFisher Scientific Inc., Waltham, MA, USA | Cat#25200056 | |
| 1.5 mL cryovial tube | Fisher Scientific, ThermoFisher Scientific Inc., Waltham, MA USA | Cat#02-682-557 | |
| 10% Neutral Buffered Formalin | VWR International LLC., Radnor, PA, USA | Cat#89370-094 | |
| 100 µm nylon mesh filter | Corning Inc., Corning, NY, USA | Cat#352360 | |
| 2-Deoxy-D-glucose | Sigma-Aldrich, Inc., St Louis, MO, USA | Cat#D8375 | |
| 2 nM 3,3’-5,Triiodo,L-thyronine sodium salt (T3) | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#T6397 | |
| 24-well tissue culture plates | VWR International LLC., Radnor, PA, USA | Cat#10861-700 | |
| 3-Isobutyl-1-methylxanthine (IBMX) | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#I5879 | |
| 96-well tissue culture plates | VWR International LLC., Radnor, PA, USA | Cat#10861-666 | |
| Antibiotic-Antimycotic Solution (ABAM) | Gibco, ThermoFisher Scientific Inc., Waltham, MA, USA | Cat#15240062 | |
| Biotin | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#B4639 | |
| Bovine Serum Albumin (BSA) | Sigma-Aldrich, Inc., St Louis, MO, USA | Cat#A8806 | |
| Buffer RLT | Qiagen, Hilden, Germany | Cat#79216 | |
| Ciglitizone | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#C3974 | |
| Deoxy-D-glucose, 2-[1,2-3H (N)]- | PerkinElmer Inc., Waltham, MA, USA | Cat#NET328A250UC | |
| Deoxyribonuclease I from bovine pancreas, type II-S | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#D4513 | |
| Dexamethasone | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#D4902 | |
| Dimethyl Sulfoxide | Fisher Scientific, ThermoFisher Scientific Inc., Waltham, MA USA | Cat#BP231 | Flammable, caustic |
| Disodium EDTA | Fisher Scientific, ThermoFisher Scientific Inc., Waltham, MA USA | Cat#BP118 | |
| D-pantothenic acid hemicalcium salt | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#21210 | |
| Dulbecco’s Modified Eagle Medium: Nutrient Mixture F-12 (DMEM/F12 | Gibco, ThermoFisher Scientific Inc., Waltham, MA USA | Cat#11320033 | |
| Ethanol | Decon Labs, Inc., King of Prussia, PA, USA | Cat#DSP-MD.43 | Flammable |
| EVE Cell Counting Slides, NanoEnTek | VWR International LLC., Radnor, PA, USA | Cat#10027-446 | |
| Fetal bovine serum (FBS) | Gibco, ThermoFisher Scientific Inc., Waltham, MA, USA | Cat#10437028 | |
| Glutaraldehyde | Sigma-Aldrich, Inc., St Louis, MO, USA | Cat#G5882 | Caustic |
| Hexamethyldisalizane | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#440191 | Flammable, caustic |
| Human insulin solution | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#I9278 | |
| Isopropanol | Fisher Scientific, ThermoFisher Scientific Inc., Waltham, MA USA | Cat#A415 | Flammable |
| Isoproterenol | Sigma-Aldrich, Inc., St Louis, MO, USA | Cat#I5627 | Flammable |
| KCl | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#S25484 | |
| KH2PO4 | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#P5655 | |
| Lipase from porcine pancreas, type VI-S | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#L0382 | |
| MgSO4*7H2O | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#230391 | |
| Na2HPO4 | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#S5136 | |
| NaCl | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#S3014 | |
| NaHCO3 | Fisher Scientific, ThermoFisher Scientific Inc., Waltham, MA USA | Cat#S233 | |
| NH4Cl | Fisher Scientific, ThermoFisher Scientific Inc., Waltham, MA USA | Cat#A661 | |
| Optimal cutting temperature (OCT) compound | Agar Scientific, Ltd., Stansted, Essex, UK | Cat# AGR1180 | |
| Oil Red-O Solution (ORO) | Sigma-Aldrich, Inc., St Louis, MO, USA | Cat#O1391 | |
| Oil Red-O Stain Kit | American Master Tech Scientific Inc., Lodi, CA, USA | Cat#KTORO-G | |
| Osmium tetroxide | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#201030 | Caustic |
| Phenylmethylsulfonyl fluoride (PMSF) | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#93482 | Caustic |
| Phosphate Buffered Saline Solution (PBS) | Fisher Scientific, ThermoFisher Scientific Inc., Waltham, MA USA | Cat#SH3025601 | |
| Ribonuclease A from bovine pancreas, type III-A | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#R5125 | |
| RNAEasy Fibrous Tissue MiniKit | Qiagen, Hilden, Germany | Cat#74704 | |
| Scintillation Fluid | Fisher Scientific, ThermoFisher Scientific Inc., Waltham, MA USA | Cat#SX18 | |
| Scintillation Counter | |||
| Scissors, forceps, sterile | |||
| Sorensen's phosphate buffer | Thomas Scientific, Inc., Swedesboro, NJ | CAS #: 10049-21-5 | |
| T-150 culture flask | VWR International LLC., Radnor, PA, USA | Cat#10062-864 | |
| TaqMan Gene Expression Master Mix | ThermoFisher Scientific Inc., Waltham, MA USA | Cat#4369016 | |
| Temperature-controlled orbital shaker | |||
| Tissue Homogenizer, BeadBug Microtube Homogenizer | Benchmark Scientific | Cat#D1030 | |
| Transferrin | Sigma-Aldrich, Inc. St Louis, MO, USA | Cat#T3309 | |
| Triglyceride Determination Kit | Sigma-Aldrich, Inc., St Louis, MO, USA | Cat#TR0100 | |
| Trypan blue stain, 0.4% | VWR International LLC., Radnor, PA, USA | Cat#10027-446 | |
| Type II collagenase | Gibco, ThermoFisher Scientific Inc., Waltham, MA, USA | Cat#17101015 | |
| Whatman Reeve Angel filter paper, Grade 201, 150mm | Sigma-Aldrich, Inc., St Louis, MO, USA | Cat#WHA5201150 |
Riferimenti
- Frantz, C., Stewart, K. M., Weaver, V. M. The extracellular matrix at a glance. Journal of Cell Science. 123, 4195-4200 (2010).
- Berrier, A. L., Yamada, K. M. Cell-matrix adhesion. Journal of Cell Physiology. 213 (3), 565-573 (2007).
- Trayhurn, P. Hypoxia and adipose tissue function and dysfunction in obesity. Physiology Reviews. 93 (1), 1-21 (2014).
- O’Rourke, R. W., Lumeng, C. N. Obesity heats up adipose tissue lymphocytes. Gastroenterology. 145 (2), 282-285 (2013).
- Engin, A. The Pathogenesis of Obesity-Associated Adipose Tissue Inflammation. Advances in Experimental Medicine and Biology. 960. 960, 221-245 (2017).
- Dankel, S. N., et al. COL6A3 expression in adipocytes associates with insulin resistance and depends on PPARγ and adipocyte size. Obesity (Silver Spring). 22 (8), 1807-1813 (2014).
- Divoux, A., et al. Fibrosis in human adipose tissue: composition, distribution, and link with lipid metabolism and fat mass loss. Diabetes. 59, 2817-2825 (2010).
- Lackey, D. E., et al. Contributions of adipose tissue architectural and tensile properties toward defining healthy and unhealthy obesity. American Journal of Physiology, Endocrinology, and Metabolism. 306 (3), E233-E246 (2014).
- Muir, L. A., et al. Adipose tissue fibrosis, hypertrophy, and hyperplasia: correlations with diabetes in human obesity. Obesity (Silver Spring). 24 (3), 597-605 (2016).
- Spencer, M., et al. Adipose tissue macrophages in insulin-resistant subjects are associated with collagen VI and fibrosis and demonstrate alternative activation. American Journal of Physiology, Endocrinology, and Metabolism. 299 (6), E1016-E1027 (2010).
- Baker, N. A., et al. Diabetes-specific regulation of adipocyte metabolism by the adipose tissue extracellular matrix. Journal of Clinical Endocrinology and Metabolism. 102 (3), 1-12 (2017).
- Pellegrinelli, V., et al. Human adipocyte function is impacted by mechanical cues. Journal of Patholology. 233 (2), 183-195 (2014).
- Flynn, L. E. The use of decellularized adipose tissue to provide an inductive microenvironment for the adipogenic differentiation of human adipose-derived stem cells. Biomaterials. 31 (17), 4715-4724 (2010).
- Perea-Gil, I., et al. In vitro comparative study of two decellularization protocols in search of an optimal myocardial scaffold for recellularization. American Journal Translational Research. 7 (3), 558-573 (2015).
- Porzionato, A., et al. Decellularized omentum as novel biologic scaffold for reconstructive surgery and regenerative medicine. European Journal of Histochemistry. 57 (1), e4 (2013).
- Tebyanian, H., et al. A Comparative Study of Rat Lung Decellularization by Chemical Detergents for Lung Tissue Engineering. Open Access Macedonian Journal of Medical Sciences. 5 (7), 859-865 (2017).
- Crapo, P. M., Gilbert, T. W., Badylak, S. F. An overview of tissue and whole organ decellularization processes. Biomaterials. 32 (12), 3233-3243 (2011).
- Wang, L., Johnson, J. A., Zhang, Q., Beahm, E. K. Combining decellularized human adipose tissue extracellular matrix and adipose-derived stem cells for adipose tissue engineering. Acta Biomaterials. 9 (11), 8921-8931 (2013).
- Booth, A. J., et al. Acellular normal and fibrotic human lung matrices as a culture system for in vitro investigation. American Journal of Respiratory and Critical Care Medicine. 186 (9), 866-876 (2012).
- Parker, M. W., et al. Fibrotic extracellular matrix activates a profibrotic positive feedback loop. Journal of Clinical Investigation. 124 (4), 1622-1635 (2014).
- Baker, N. A., Muir, L. A., Lumeng, C. N., O’Rourke, R. W. Differentiation and Metabolic Interrogation of Human Adipocytes. Methods in Molecular Biology. 1566, 61-76 (2017).
- O’Rourke, R. W., et al. Hexosamine biosynthesis is a possible mechanism underlying hypoxia’s effects on lipid metabolism in human adipocytes. PLoS One. 8 (8), e71165 (2013).
- Tchkonia, T., et al. Fat depot-specific characteristics are retained in strains derived from single human preadipocytes. Diabetes. 55 (9), 2571-2578 (2006).
- Tchoukalova, Y. D., et al. Sex- and depot-dependent differences in adipogenesis in normal-weight humans. Obesity (Silver Spring). 18 (10), 1875-1880 (2010).

