In Utero Electroporation of Multiaddressable Genome-Integrating Color (MAGIC) Markers to Individualize Cortical Mouse Astrocytes
Summary
Astrocytes tile the cerebral cortex uniformly, making the analysis of their complex morphology challenging at the cellular level. The protocol provided here uses multicolor labeling based on in utero electroporation to single out cortical astrocytes and analyze their volume and morphology with a user-friendly image analysis pipeline.
Abstract
Protoplasmic astrocytes (PrA) located in the mouse cerebral cortex are tightly juxtaposed, forming an apparently continuous three-dimensional matrix at adult stages. Thus far, no immunostaining strategy can single them out and segment their morphology in mature animals and over the course of corticogenesis. Cortical PrA originate from progenitors located in the dorsal pallium and can easily be targeted using in utero electroporation of integrative vectors. A protocol is presented here to label these cells with the multiaddressable genome-integrating color (MAGIC) Markers strategy, which relies on piggyBac/Tol2 transposition and Cre/lox recombination to stochastically express distinct fluorescent proteins (blue, cyan, yellow, and red) addressed to specific subcellular compartments. This multicolor fate mapping strategy enables to mark in situ nearby cortical progenitors with combinations of color markers prior to the start of gliogenesis and to track their descendants, including astrocytes, from embryonic to adult stages at the individual cell level. Semi-sparse labeling achieved by adjusting the concentration of electroporated vectors and color contrasts provided by the Multiaddressable Genome-Integrating Color Markers (MAGIC Markers or MM) enable to individualize astrocytes and single out their territory and complex morphology despite their dense anatomical arrangement. Presented here is a comprehensive experimental workflow including the details of the electroporation procedure, multichannel image stacks acquisition by confocal microscopy, and computer-assisted three-dimensional segmentation that will enable the experimenter to assess individual PrA volume and morphology. In summary, electroporation of MAGIC Markers provides a convenient method to individually label numerous astrocytes and gain access to their anatomical features at different developmental stages. This technique will be useful to analyze cortical astrocyte morphological properties in various mouse models without resorting to complex crosses with transgenic reporter lines.
Introduction
Astrocytes play numerous vital functions in brain development and physiology1. Beside their role at the blood-brain barrier where they regulate nutrient uptake and blood flow, they actively contribute to synapse formation and function while producing neuromodulators that can alter neuronal activity and behavior2. Furthermore, astrocyte dysfunction contributes to a variety of neurological disorders3. Astrocytes located in the cerebral cortex display an elaborate morphology enabling extensive contact with neuronal processes. These contacts, essential for circuit function, also control astrocyte morphogenesis and synaptogenesis through cell-adhesion proteins4. Neuroscientists need convenient and robust tools to investigate astrocyte development and morphogenesis in their neurological models of interest. However, due to the close apposition of astrocytes to their neighbors and their uniform three-dimensional tiling, it is challenging to single out cortical astrocytes and comprehensively assess their morphology using immunomarkers.
Currently, two main genetic engineering strategies enable labeling and individualization of cortical astrocytes in situ: sparse reporter activation in transgenic mouse lines or somatic transgenesis using electroporation of reporter plasmids. The first strategy relies on breeding a floxed reporter mouse line with mice expressing an inducible form of Cre recombinase activated specifically in astrocytes upon tamoxifen delivery (e.g., Aldh1l1-CreERT25). Several disadvantages are associated with this strategy. First, breeding transgenic mice requires a large number of animals and multiple assays are typically needed to determine the proper dose of tamoxifen to provide adequately sparse labeling of cortical astrocytes. Analyzing cortical astrocyte phenotypes in a genetic mouse model of interest will require even more breeding and mouse consumption. Furthermore, in utero tamoxifen injection is known to interfere with parturition, making this strategy difficult to apply to the study of the earliest stages of astrocyte development. In vivo DNA electroporation is an alternative tamoxifen-free strategy that relies on a minimum number of animals6. Performed either at embryonic or postnatal stages, this approach consists of injecting reporter plasmids in the lateral ventricles of rodents followed by electric pulses that create pores in the cell membrane, hence allowing DNA to enter progenitor cells lining the ventricle. Subsequently, the reporter transgenes carried by the electroporated plasmids are processed by the targeted cell machinery and expressed7. Two electroporation methods have been previously described to label mouse cortical astrocytes: 1) Postnatal Astrocyte Labeling by Electroporation (PALE), which relies on the electroporation of 1–2 single-color episomal reporter plasmids at early postnatal stages4; 2) The StarTrack strategy based on in utero electroporation (IUE) of multiple single-color integrative reporter plasmids8,9,10. Although these two techniques efficiently label PrA in the cerebral cortex, they also present some limitations. In their initial version, both methods rely on a glial fibrillary acidic protein (GFAP) promoter to drive expression in astrocytes, which may bias the labelling toward radial glia as well as pial and reactive astrocytes that express GFAP more strongly than normal resting PrA11,12. Regarding PALE, other disadvantages are the late stage of electroporation, which prevents labeling of early-born PrA (or those originating from early delaminating progenitors) and analysis of early stages of astroglia development, and the use of episomal vectors that become diluted through successive divisions during the massive proliferation that PrA undergo during the first postnatal week13,14. In contrast to PALE, StarTrack is based on the embryonic electroporation of integrative reporter plasmids that allow tracking the contribution of both embryonic and postnatal progenitors to PrA. An updated StarTrack scheme relying on the ubiquitin C promoter (UbC-StarTrack) achieves broader expression of fluorescent reporters in both the neuronal and glial descent (astrocytes included) of neural progenitors15,16,17. However, in its current version, implementation of this approach is complex, as it relies on an equimolar mixture of 12 distinct plasmids expressing six fluorescent proteins (FP) with partial excitation and emission spectra overlap.
Presented here is a straightforward in utero electroporation-based multicolor labeling method using integrative reporter constructs driven by a strong and broadly active promoter to single out cortical astrocytes14. In addition, an easy image analysis pipeline using both licensed (e.g., Imaris) and open access (Vaa3D18,19,20) image analysis software is provided to segment astrocyte territorial volume and arborization, respectively. Compared to the previously described methods, this strategy relies solely on 1–2 multicolor integrative transgenes Multiaddressable Genome-Integrating Color Markers (MAGIC Markers or MM21) directed to the cytoplasmic and (optionally) nuclear cell compartment whose expression is driven by a synthetic CAG promoter comprised of a cytomegalovirus enhancer, chicken β-actin promoter, and rabbit β-globin splice acceptor site22. This enables labeling and tracking of cortical astrocytes, from embryonic to late postnatal stages, independent of GFAP expression14,23. Each of these transgenes bears the following four distinct FP: eBFP, mTurquoise2/mCerulean, EYFP, and tdTomato/mCherry, which display minimal spectral overlap that can be easily circumvented with 1) Sequential channel acquisition; 2) Optimized excitation power and collection gain; and 3) Specific dichroic filters to collect narrow FP emission windows. The MM strategy uses Cre/lox recombination with a self-excisable Cre recombinase (seCre) to drive stochastic expression of FP in a cellular population. A single copy of MM transgene expresses FP in a mutually exclusive manner, while multiple transgenes give rise to FP combinations, creating dozens of distinct hues. Genomic integration of the transgenes is driven by the piggyBac (PB) or Tol2 transposition system24,25,26. Therefore, through in utero electroporation, the MM toolkit and the multicolor ‘mosaic’ that it generates enable simultaneous marking of multiple adjacent cortical progenitors and the tracking of their glial descent, including cortical astrocytes, over long periods. Color contrasts resulting from the expression of distinct FP permit delineation of the contour of PrA and subsequently extract key information about their territorial volume (using IMARIS) and complex morphology (using Vaa3D). The multicolor strategy presented in detail here is a convenient and robust method that gives quick and easy access to the cortical astrocyte surface and morphology in wild type mice at various developmental stages, and is easily adaptable to investigate astrocyte anatomical features in mouse models of neurological diseases without using transgenic reporter lines.
Protocol
All animal procedures described here were carried out in accordance with institutional guidelines. Animal protocols have been approved by the Charles Darwin animal experimentation ethical board (CEEACD/N°5).
1. Preparation of endotoxin-free plasmids for MAGIC Markers in utero electroporation
- Bacterial transformation
- On ice, thaw DH5 alpha competent cells stored at -70 °C.
- Warm up the agar plates containing the appropriate antibiotic (100 µg/mL ampicillin or 50 µg/mL kanamycin) at 37 °C.
- Add 1 µL of 5–50 ng of MAGIC Markers plasmid DNA in 10 µL of thawed DH5 alpha competent cells and incubate on ice for 10 min without mixing.
- For the heat shock transformation, place the aliquot in a 42 °C water bath for 45 s, then place immediately on ice and wait 3–5 min.
- Under sterile conditions, add 230 µL of SOC medium and incubate for 1 h at 37 °C.
- Spread the content of the aliquot over the agar plate and incubate overnight at 37 °C.
- Plasmid culture
- The following morning, under sterile conditions, pick up a colony from the agar plate and put it in a 14 mL tube containing 2 mL of LB medium with appropriate antibiotic. Let it incubate for the day at 37 °C in a shaking incubator at 300 rpm.
- At the end of the day, seed 300 mL of LB with antibiotic using the 2 mL starter culture from step 1.2.1 and incubate overnight at 37 °C in a shaking incubator at 300 rpm.
- Plasmid DNA preparation
- The following morning proceed with the purification of MAGIC Markers plasmid DNA (i.e., PB-CAG-Cytbow and Tol2-CAG-Nucbow as well as CAG-driven plasmids expressing PB and Tol2 transposases and Cre recombinase) using an endotoxin-free maxiprep kit following the manufacturer’s protocol.
- Elute the DNA in 200 µL of sterile water and estimate its concentration using a spectrophotometer prior to storage at -20 °C.
2. Preparation for MAGIC Markers in utero electroporation (MM IUE)
- Solution preparation
- Warm 30 mL of 0.9% saline solution at 37 °C in a water bath and keep it warm during the entire duration of the surgery.
- Prepare a plasmid mix containing PB-CAG-Cytbow and Tol2-CAG-Nucbow (final concentration, 0.8 µg/µL each), PB and Tol2 transposases (final concentration, 0.4 µg/µL each), CAG-seCre (final concentration, 0.16 µg/µL), and 0.01% Fast Green dye in PBS without Ca2+ and Mg2+.
NOTE: For astrocyte anatomical reconstruction purposes, the Tol2-CAG-Nucbow construct may be omitted. However, this plasmid is useful to distinguish doublets of closely juxtaposed astrocytes and when using MAGIC Markers to probe clonal relationships among astrocytes14. - Prepare anesthetic solution containing 100 µL of ketamine (100 mg/mL) and 100 µL of xylazine (20 mg/mL) diluted in 2 mL of saline solution.
- Prepare analgesic solution by diluting 0.3 mg/mL buprenorphine stock solution 1:10 in saline solution.
- Preparation of the surgery material
- Sterilize the surgical tools (see Table of Materials) at high temperature in a glass bead sterilizer or equivalent.
- Place a drop of electrode gel in a 35 mm dish.
- Insert the micropipette in the microinjector holder, break the tip of the micropipette, and aspire the DNA solution.
- Place a 1 µL drop of Fast Green solution (0.01%) in the lid of a 3 cm dish and, using it as a reference, adjust the diameter of the micropipette’s tip by breaking its tip with fine forceps. Adjust the pressure parameter so that equivalent size drops are produced by the microinjector, thereby enabling delivery of approximately 1 µL of DNA solution per injection.
- Preparation of the pregnant female mouse
- Weigh the RjOrl:SWISS pregnant mouse.
- Perform an intraperitoneal injection with 12.5 µL per gram of body weight (BW) of anesthetic solution. Wait for 5 min and verify that the mouse is asleep by pinching its toe.
- Once the animal is unresponsive to pinch, inject it subcutaneously with 1.6 µL/g BW of analgesic solution.
- Add a drop of ocular gel on each eye to prevent drying during the surgery and place the animal belly up on the warming pad.
- Gently shave its abdomen, clean it with a pad soaked with iodine, and sanitize the shaved area with an alcohol pad.
- Arrange an operating field by placing sterile compresses around the shaved, cleaned, and sanitized area.
3. In utero electroporation (IUE)
- Intraventricular injection
- Cut a 2 cm-vertical incision along the midline starting in the lower part of the abdomen, through the skin, and then through the underlying muscle. Expose the uterine horns by gently manipulating the embryonic bags and assess the location of the cervix and the number of embryonic bags on each side of the cervix.
- Orient the brain of the E15.5 embryo to be electroporated in order to see the bregma, easily recognized as its location matches with the junction of the three main blood vessels that run along the cerebral fissures.
- Imagine a virtual line between the bregma (visible through the skull) and the eye; introduce the micropipette between the virtual line and the longitudinal fissure, then press the injector’s foot pedal to deliver 1 µL of DNA solution in the lateral ventricle of the targeted hemisphere.
NOTE: When injected at the proper location, the filled lateral ventricle appears blue, indicating that it has been filled with the DNA solution.
- Electroporation
- Apply the electrodes (see Table of Materials), previously dipped in electrode gel, on both sides of the injected embryo with the anode covering the injected hemisphere. Press the foot pedal of the electroporator to deliver a series of four 50 ms pulses of 35 V, each separated by a 950 ms interval.
- Dampen the embryo with warmed saline solution.
NOTE: The embryos should be kept humid using warmed saline solution during the entire surgical operation and the uterus should not dry out. - Repeat steps 3.1.2–3.2.2 for each embryo.
- Once all targeted embryos have been electroporated, replace the uterine horns in the abdomen by gently pushing them with forceps back to their original position. Fill the abdominal cavity with warmed saline solution to prevent the uterus from drying while sutures are made.
- First close the abdominal muscle with a continuous absorbable suture, and then the skin with multiple individual stitches (~10) using 4-0 suture.
- Put the animal in a clean cage, lying on its side on a clean paper towel and turn the animal to the other side every 5–10 min until it wakes up and starts moving on its own.
- Assess the state of the animal the following day, especially if a nest has been made.
NOTE: No postsurgery treatment is required. In the absence of a nest, any sign of pain (e.g., prostration, shaggy fur), and/or heavy bleeding, the animal should be euthanized without delay.
4. Tissue harvesting and sectioning
- Tissue collection
- Inject phenobarbital (100 mg/kg BW) intraperitoneally for terminal anesthesia at the desired harvesting time. In this case, the harvesting times were at postnatal (P) days P4, P7, and P21.
- Perform intracardiac perfusion using cold premade paraformaldehyde-based solution.
- Dissect out the brain and place it overnight at 4 °C in the paraformaldehyde-based solution for postfixation.
- Histology
- The following morning rinse the brain 3x for 10 min with 1x PBS.
- Embed the brain in 3% agarose dissolved in 1x PBS.
- Cut 80 µm sections using a vibrating-blade microtome.
- Collect sections in a 24 well plate prefilled with 1x PBS.
- Mount sections in the mounting medium (see Table of Materials) between the slide and coverslip. Keep the mounted slides at -20 °C for optimal fluorescent protein preservation and long-term storage.
5. Multichannel confocal imaging
- Microscope settings
- Set up the configuration of the confocal microscope to separately excite mCerulean/mTurquoise2, EYFP, tdTomato/mCherry using 440, 515, and 559 nm laser lines, respectively.
- Use a 20x 0.8 NA (or higher NA) objective and adjust XY sampling and Z-step size according to Nyquist criteria.
NOTE: Images acquired with higher resolution (e.g., 60x 1.4 NA oil objective) and deconvolution algorithms will enable reconstruction of the finer details of astrocyte arbors. However, the experimenter should keep in mind that the finest astrocyte processes may not be resolved by conventional optical imaging.
- Image acquisition
- Find the brightest astrocytes in the electroporated area.
- Because labeled cells located close to the surface might appear brighter than those deeper in the slice, adjust the acquisition settings on the surface cells to avoid saturating the images.
- Adjust acquisition settings separately for each of the three channels while making sure to avoid pixel saturation using HiLo LUT, which displays zero values as blue and maximum pixel values as red.
NOTE: An adequately balanced image should display only a few blue and almost no red pixels. - Acquire tiled 1,024 x 1,024 pixel Z-stacks using the motorized stage of the microscope, with a 10% overlap between adjacent stacks to subsequently enable mosaic reconstructions of the electroporated area or the zone of interest.
6. Astrocyte territorial volume segmentation
NOTE: This is performed using a commercial software program (e.g., IMARIS).
- Preparation of the dataset
- Crop the labeled astrocytes within the 3D reconstruction (250 x 250 pixels) by selecting cells entirely enclosed in the imaged tissue section.
NOTE: It is also possible to work directly on larger volumes if the computer used can handle them. - Click on the Blue Surface button in the Object toolbar of the Surpass view to create a new Surface for each astrocyte.
- To obtain better visualization contrast, first adjust the minimal and maximal color contrast values by clicking on Edit | Show display Adjustment option and dragging both handles. If needed, change the channel color by clicking directly on the channel name in the Display Adjustment window in order to select a new color.
NOTE: Preferentially always work with the same color display to prevent visual bias. - Click on Edit | Image Properties to manually indicate the voxel size in micrometers. If the voxel size is not accurate, the volume calculations will be incorrect.
- Save the new settings.
- Crop the labeled astrocytes within the 3D reconstruction (250 x 250 pixels) by selecting cells entirely enclosed in the imaged tissue section.
- Surface segmentation
- Use the Blue Icon to introduce a surface. A Surface icon and box will appear on the Scene list. A Volume box allows to see/hide the dataset.
- Select the surface line and click on Skip Automatic Creation | Edit Manually.
- Click on Contour | Visibility and click None. Then move to the Select view and click on the Draw button.
- Click on Mode to select the drawing mode.
- Use the fast and efficient Isoline semiautomated drawing tool. Define the cell outline by moving the mouse pointer on it. If the isoline preview does not match the astrocyte outline, adjust the mouse pointer position. To validate the preview, left click on the selection. To correct potential mistakes, use the Board window to Delete the current Z-plane contour or press Ctrl + Z.
- Using Slice | Position to navigate through the Z-planes, move to the next plane by changing the Z-plane number and start a new contour. Preferentially start from the middle of the cell and then move to the extremities.
- When contours have been drawn in all the Z-planes containing the astrocyte, click on Create Surface. Before creating a surface, check that all the components are connected. If not, select the largest or connect relevant ones before exporting volume data.
- Check the quantitative data displayed in the Data Panel and export statistics in spreadsheet format, saving volume value and unit.
- Save the surface under a new name to access it later.
7. Tracing astrocyte arborization
NOTE: This is done using open access software program Vaa3D.
- Preparation of the dataset
- Load the image stack and search for isolated astrocytes or nearby astrocytes displaying distinct colors.
NOTE: The resolution of the reconstruction can be improved by increasing the objective NA and sampling. Ultimately, however, due to the resolution limit of confocal microscopy, the finest astrocyte processes cannot be accurately traced. Care should be taken to avoid oversegmenting beyond the resolution of the acquired images. - Using Fiji, crop 250 x 250 pixel image stacks around each astrocyte.
- As Vaa3D tracing is performed on only one color, select one channel and deactivate the other channels.
- Convert the image to RGB and save it in .tiff format.
- Load the image stack and search for isolated astrocytes or nearby astrocytes displaying distinct colors.
- Arbor tracing
- Open the RGB image in Vaa3D. Go to Image | Data | Geometry then image resampling, and adjust X/Y and Z voxel size values.
NOTE: While reconstructing astrocyte arbors, care should be taken not to trace details finer than the resolution provided by the images. - Open a 3D window by clicking on Visualize | 3D viewer for the entire image. Right-click on a 3D image and choose 1-right-click to define marker. Point with the cursor the center of the astrocyte and then right-click to set a marker.
NOTE: Click on Escape to have access to the 3D view again. - Go to Advanced | 3D tracing | Vaa3D-Neuron2-auto-tracing.
- On the newly opened window, select the channel on which to apply the plug-in.
- Choose a background threshold (often between 30 and 90). Adapt this value for each astrocyte if needed.
- Uncheck Radius from 2D and keep Auto-down sample and auto-resample checked.
- Set cnn_type at 3, length_thresh at 1, and SR_ratio at 0.1.
- After clicking ok, ensure that the plug-in automatically generates two files based on the original name. The suffix –ini.swc and -coordinateX–coordinateY-coordinateZ-app2.swc will be added. Manually add a distinctive label to these name files in order not to overwrite them while continuing to run the plug-in.
- Open Object Manager, go to Neuron | Line structure, select the traced astrocyte, click on Display Mode, and select Always line mode.
- Go back to the 3D window, where a skeleton of the astrocyte appears.
NOTE: If segmentation and signal do not match, repeat steps 7.2.3–7.2.10 and adjust the threshold until they do. Make sure to re-enter the other parameters as they are reset when the plug-in is restarted. - Right-click on skeleton and select first choice neuron | line#1…APP2_Tracing to access quantitative measurements that will appear on a new window Surface| Object Annotation.
NOTE: Due to the limited resolution of light microscopy, measurements displayed under the listed items Number of branches and Number of bifurcations provide only a rough estimate of astrocyte arbor complexity.
- Open the RGB image in Vaa3D. Go to Image | Data | Geometry then image resampling, and adjust X/Y and Z voxel size values.
Representative Results
Electroporation of MAGIC Markers in embryonic cortical progenitors allows for the labeling of astrocytes from early to late stages of cerebral cortex development (Figure 1). These astrocytes were found in all cortical layers at various postnatal stages (P4, P7, P21) as they dispersed widely into the entire cerebral cortex. They were assessed with tiled confocal images acquired with a 20x 0.8 NA (or higher NA) objective and assembled as Z-stack reconstructions (Figure 2). MAGIC Markers combinatorial labeling enabled individualization of cortical astrocytes and extraction of information regarding their volume and morphology. Using the commercial image analysis software, the contour of individual astrocytes was delineated on each individual optical section of confocal image stacks to segment and reconstruct the territorial domain occupied by each astrocyte (Figure 3). From the same Z-image stacks, the branched morphology of singled-out cortical astrocytes was segmented using the open access software that allows extraction of the skeleton of the main astrocyte processes (Figure 4). These segmentation and tracing tools provided a semiquantitative assessment of the increase of territorial volume (Figure 3) and morphological complexity (Figure 4) that occurred for individual cortical astrocytes from early to late postnatal stages14. These approaches also revealed the heterogeneity of volume and morphology displayed by distinct cortical astrocytes at the same stage of development, as illustrated in Figure 2, Figure 3, and Figure 4.
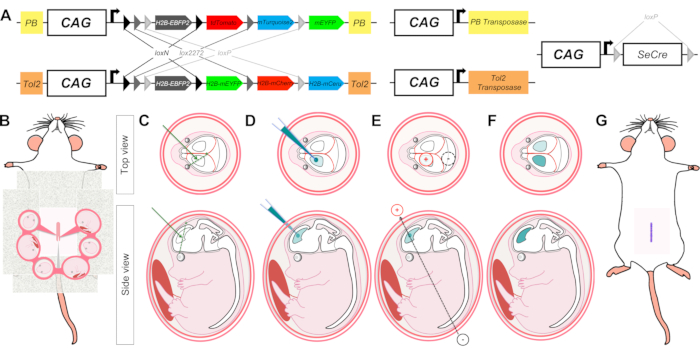
Figure 1: Schematic representation of MAGIC Markers (MM) in utero electroporation (IUE) to label cortical progenitors and their descent during brain development. (A) The MM toolkit comprises several plasmids encoding PB-Cytbow, Tol2-Nucbow, PB and Tol2 transposases, and self-excisable Cre recombinase (seCre). In MM constructs, three pairs of incompatible lox sites (loxN, lox2272, and loxP) flank four distinct FP coding sequences (EBFP2, mTurquoise2/mCerulean, EYFP, tdTomato/mCherry) and create mutually exclusive possibilities of excision upon Cre recombination. Before Cre action, only the first gene (EBFP2) is expressed. After Cre-mediated excision induced by seCre, either tdTomato/mCherry (red FP), EYFP (green FP), or mTurquoise2/mCerulean (cyan FP) is expressed. Co-expression of FP from multiple MAGIC Markers copies yields color combinations in the cytoplasm (PB-CAG-Cytbow) or nucleus (Tol2-CAG-Nucbow) of labeled cells. PB and Tol2 transposition endfeet framing the MM cassettes permit their integration into the genome of cortical progenitors when MM constructs are coelectroporated along with PB and Tol2 transposases coding plasmids. (B–G) Graphic illustration of IUE successive steps including the laparotomy of anesthetized pregnant mice (B), injection of the MM plasmid mix (C) in the lateral ventricles of the embryos (D), delivery of electric pulses through carefully positioned tweezertrodes (E) to target cortical progenitors in one of the two brain hemispheres (F), and suturing of pregnant mouse abdomen (G). Please click here to view a larger version of this figure.
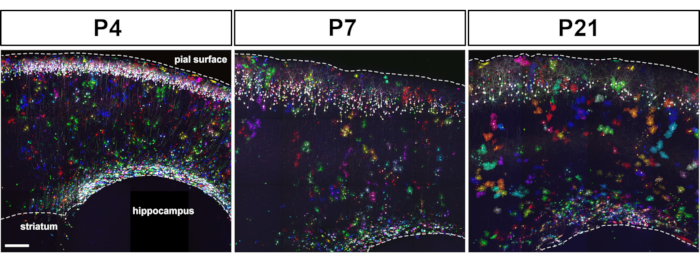
Figure 2: Following in utero electroporation of MAGIC Markers toolkit, multicolor astrocytes were found scattered in the entire cerebral cortex at postnatal stages. IUE of plasmids driving MM, seCre, PB, and Tol2 transposases expression in E15.5 mouse cortical progenitors resulted in labeling of layer 2–3 neurons and astrocytes at P4, P7, and P21. Expression level and color palette depended on the number of MM transgenes integrated in the genome of cortical progenitors. Montage of maximal intensity projections from tiled confocal image stacks acquired on 80 µm sagittal brain sections. Scale bar: 200 µm. Please click here to view a larger version of this figure.
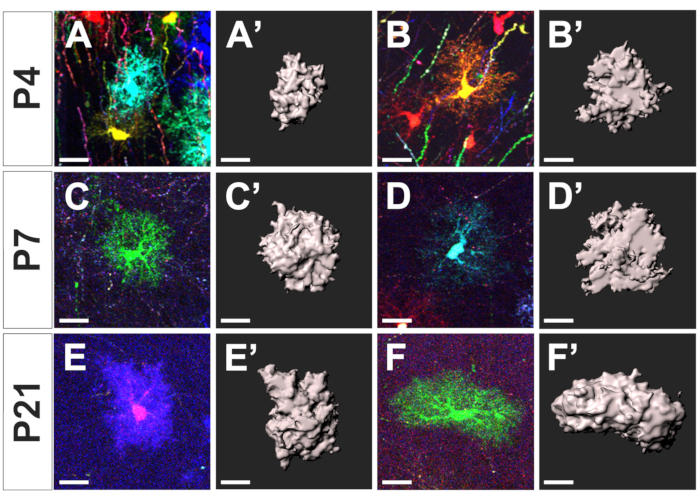
Figure 3: Segmentation of cortical astrocyte territorial domain at distinct developmental stages. Maximal intensity projections of cropped confocal Z-stack framing individual astrocytes (A–F) and their associated territorial domain segmented with the commercial software (A’-F’) at three distinct developmental stages: P4 (A-B, A’-B’), P7 (C-D, C’-D’), and P21 (E-F, E’-F’), respectively. Scale bar: 20 µm. Please click here to view a larger version of this figure.
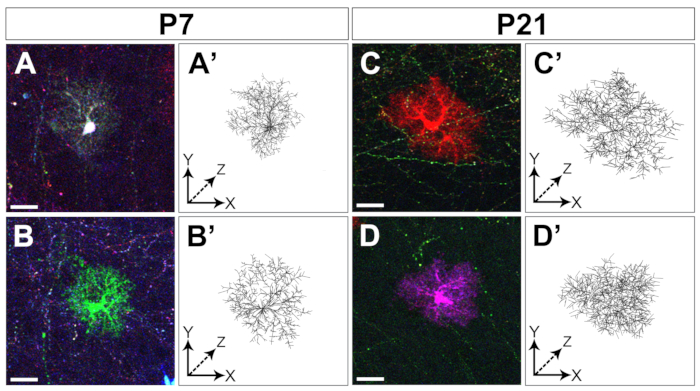
Figure 4: Tracing of cortical astrocyte arborization. Examples of two distinct astrocytes cropped from confocal Z-stacks (A-D) and reconstruction of their arborization coarsely segmented with Vaa3D (A’-D’) collected at two distinct developmental stages: P7 (A-B, A’-B’) and P21 (C-D, C’-D’), respectively. Scale bar: 20 µm. Please click here to view a larger version of this figure.
Discussion
In utero electroporation (IUE) of MAGIC Markers in cortical progenitors (Figure 1) enabled labeling of astrocytes throughout the postnatal cerebral cortex at different postnatal stages (P4-P7-P21, Figure 2). Interestingly, the stage of IUE is not critical, as electroporation performed from E13.5 to E15.5 yields similar labeling patterns concerning cortical astrocytes14. However, the location of labeled pyramidal neurons in the cortical parenchyma varies with the electroporation stage. Indeed, IUE performed at E15 marks layer 2–3 neurons whereas IUE performed at E13 labels pyramidal neurons in all cortical layers, from layer 5 to layer 2–36,27. This joint labeling of cortical pyramidal neurons following MM IUE is the main limitation of this method, as it prevents semiautomated segmentation of astrocyte morphology in layers where dense neuronal labeling interferes with astrocyte processes recognition. Should that be a problem, MM could be electroporated at postnatal stages as in PALE. At P4, labeling of radial glia fibers still present at that stage may also interfere with Vaa3D arbor segmentation. While cumbersome, a solution if one wishes to proceed with astrocyte arbor reconstruction at this stage is to manually remove radial glial fibers by progressively replacing the radial fiber signal with black pixels around the astrocyte of interest using Adobe Photoshop.
Despite this limitation, MM IUE is a powerful technique when adequately performed. A few critical steps need to be handled with care: 1) embryos must be kept humid during the entire surgery procedure and carefully manipulated to increase their survival; 2) using glass capillaries with a large diameter or squeezing embryonic bags too tightly can lead to bag rupture and therefore embryos’ death; 3) during DNA injection, blood vessels must be avoided to prevent bleeding; 4) the whole procedure should not last more than 40 minutes from anesthesia to suture in order to maximize embryo survival; 5) stress plays a critical role in IUE success and therefore extra sources of stress such as change of cage, transportation, noises, and vibration must be avoided from 5 days prior surgery to 7 days after birth in order to prevent abortion and cannibalism.
Of note, experimenters wishing to specifically target cells born at a given stage can use the MM toolkit without adding the piggyBac and Tol2 transposases such that only the cells born at the time of the electroporation express the combinatorial labels. Another advantage of the method is the flexibility that it confers in terms of the density of labeled cells and their location in various brain regions. Indeed, denser labeling of cortical astrocytes can be achieved by increasing the total concentration of MM transgenes while keeping plasmids ratio constant (1:10 ratio for Cre recombinase/MM constructs and 1:2 ratio for transposases/MM constructs). Contrary to monochrome approaches, such as electroporation of single-color transposons or Cre electroporation in Ai9 mice, where the ability to single out individual astrocytes requires sparse labeling, color contrasts offered by the MAGIC Markers strategy enable the individualization of astrocytes over a wide range of labeling densities. In addition, positioning the electrode probes in distinct orientations permits targeting distinct brain regions such as the prospective striatum (anode in the ventral position, opposite to the dorsal position required to achieve electroporation in the cerebral cortex), or hippocampus (anode in medial position)28. Finally, IUE can be performed in different mice strains such as outbred (OF1, Swiss) and inbred (C57BL/6J or N) mice, which opens the way for use of the MM toolkit in transgenic animal disease models. However, to successfully achieve IUE in inbred mice, one should adapt the number of pulses (three pulses for C57BL/6 mice versus five pulses in Swiss), voltage (30 V instead of 35 V), and analgesic dose (0.15 mg/mL buprenorphine stock solution and an injected volume of 0.8 µL/g BW).
In comparison with transgenic animal breeding or the PALE and StarTrack approaches, this method offers several advantages. To begin with, in contrast to the breeding strategy, it uses few animals. It also allows labeling of cortical astrocytes since the earliest stages of their development, including embryonic stages, unlike PALE4, which relies on postnatal electroporation. Furthermore, in comparison to the twelve reporter constructs used in the StarTrack approach8, this strategy relies on only two multicolor transgenes, thus making DNA mix preparation and imaging simpler. Moreover, the balance between the different colors stochastically expressed by MM is intrinsically determined by the structure of the transgenes and does not depend on the mixing of different components by the experimenter. In addition, this strategy can extend beyond simple anatomical consideration and can be successfully applied for multiclonal analysis of astrocyte development, as demonstrated in previously published work14. This work, using rare color combinations of cytoplasmic and nuclear markers to define cortical astrocyte clones, demonstrated that they display extensive variability in terms of spatial distribution, structural organization, number, and subtype of generated cells.
Cortical astrocytes born from MM-labeled cortical progenitors displayed significant color contrast and dispersed widely across the entire cerebral cortex (Figure 2). Simple multichannel Z-stack acquisitions using confocal microscopy were used to access key astrocyte features such as their territorial volume (Figure 3) and their morphological complexity (Figure 4) at several postnatal stages. Beyond astrocytes, this methodology may be adapted to study the morphology of other glial cells such as oligodendrocytes. However, it should be kept in mind that the limited resolution offered by confocal microscopy can only provide a partial rendering of astrocyte morphological complexity. While images obtained with higher resolution (e.g., 63x 1.4 NA oil objective) and deconvolution algorithms can be used to reconstruct finer details of astrocyte arbors14, the finest processes cannot be resolved with conventional optical imaging. Nevertheless, the strategy presented here will be of interest to screen efficiently for a potential phenotype affecting cortical astrocyte volume or morphology in mouse models of neurological diseases.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
We thank S. Fouquet and the imaging and animal core facilities of Institut de la Vision and Institut des Neurosciences de Montpellier (MRI and RAM) for technical assistance. This work was supported by fellowships from Région Ile-de-France and Fondation ARC pour la Recherche sur le Cancer to S.C, and from Université Paris-Saclay (Initiatives Doctorales Interdisciplinaires) to L.A., by funding from European Research Council (ERC-SG 336331, PI J. Valette) to E.H., by Agence Nationale de la Recherche under contracts ANR-10-LABX-65 (LabEx LifeSenses), ANR-11-EQPX-0029 (Equipex Morphoscope2), ANR-10-INBS-04 (France BioImaging), by Fondation pour la Recherche Médicale (Ref. DBI20141231328), by the European Research Council (ERC-CoG 649117, PI J. Livet) and by ATIP-Avenir program (PI K. Loulier).
Materials
| 1.1 Bacteria transformation | |||
| Ampicillin | Euromedex | EU0400-C | |
| DH5 alpha competent cells | Fisher Scientitic | 11563117 | |
| Ice box | Dutscher | 139959 | |
| Kanamycin | Sigma | 60615 | |
| LB Agar | Sigma | L2897 | |
| SOC medium | Fisher Scientitic | 11563117 | |
| Sterile petri dish- 10 cm | Thermo Fisher | 150350 | |
| Water bath | VWR | 462-0556H | |
| 1.2 Plasmid culture | |||
| 14 ml culture tube | Dutscher | 187262 | |
| Glass erlenmeyer- 2L | Fisher Scientitic | 11383454 | |
| LB medium | Sigma | L3522 | |
| 1.3 Plasmid DNA preparation | |||
| NucleoBond Xtra Maxi Plus EF | Macherey-Nagel | 740426.10 | |
| 2.1 Preparation of the solutions | |||
| 26 G x 1/2 needle | Terumo | 8AN2613R1 | |
| 30 G x 1/2 needle | Terumo | 8AN3013R1 | |
| Fast Green | Sigma Aldrich | F7272 | |
| NaCl | VWR | 27810.295 | |
| Single-use polypropylene syringe, 1 mL | Dutscher | 50002 | |
| 2.2 Preparation of the surgery material | |||
| Adson Forceps – DeBakey Pattern- 12.5 cm | FST | 11617-12 | |
| Arched tip Forceps- 10 cm | FST | 11071-10 | |
| Glass bead sterilizer Steri 250 | Sigma | Z378569 | |
| Glass micropipette 1 mm diameter | FHC | 10-10-L | |
| Graefe Forceps – Titanium 1 mm Tips Slight Curve- 10 cm | FST | 11651-10 | |
| Graefe Forceps – Titanium 1 mm Tips Straight- 10 cm | FST | 11650-10 | |
| Iris Scissors – Delicate Straight- 9 cm | FST | 14060-09 | |
| Laboratory tape | Fisher Scientitic | 11730454 | |
| Microinjector | INJECT+MATIC | No catalog number | |
| Olsen-Hegar Needle Holder – 12 cm | FST | 12002-12 | |
| Optical fiber | VWR | 631-1806 | |
| Plastic-coated white paper | Distrimed | 700103 | |
| Signagel electrode gel | Free-Med | 15-60 | |
| Sterile Petri dish- 35 mm | Dutscher | 056714 | |
| Sterilizer, glass dry bead, Steri 250 | Sigma | Z378569 | |
| 2.3 Preparation of the pregnant female mouse | |||
| Alcohol pad | Alcomed | 1731000 | |
| Buprecare | Axience | 0.3 mg/ml | |
| Compress | tRAFFIN | 70189 | |
| Ketamine | Merial | Imalgene 1000 | |
| Ocular gel | tvm lab | Ocry-gel | |
| RjOrl:SWISS mice | Janvier Labs | ||
| Vetadine, 10% solution | Vetoquinol | 4337400113B | |
| Warming pad | Harvard Apparatus | 72-0493 | |
| Xylazine | Bayer | Rompun 2% | |
| 3.2 Electroporation | |||
| Absorbable suture Size 4-0 45 cm Suture 1-Needle 19 mm Length 3/8 Circle Reverse | Novosyn | C0068220 | |
| Electroporateur Sonidel | Sonidel | NEPA 21 | |
| Sterile transfer pipets (individually wrapped) | Dutscher | 043202S | |
| Tweezers with 3 mm platinium disk electrodes | Sonidel | CUY650P3 | |
| 4.1 Tissue harvesting and sectioning | |||
| 24-well plate | Falcon | 353047 | |
| Agarose | Lonza | 50004 | |
| Antigenfix | Microm Microtech | U/P0014 | |
| Coverslip | Dutscher | 100266 | |
| Dolethal | Vetoquinol | DOL202 | |
| DPBS (10X), no calcium, no magnesium | Fisher Scientific | 11540486 | |
| Nail polish | EMS | 72180 | |
| Slide | Dutscher | 100001 | |
| Vectashield | Vectorlabs | H-1000 | |
| Vibratome | Leica | VT1000S | |
| 5. Multichannel confocal imaging | |||
| 20X oil NA 0.85 | Olympus | ||
| Confocal Laser Scanning Microscope | Carl Zeiss | LSM880 | |
| Confocal Laser Scanning Microscope | Olympus | FV1000 | |
| Plan Apochromat 20x/0.8 M27 | Carl Zeiss | ||
| 6. Astrocyte territorial volume segmentation | |||
| IMARIS 8.3 and later versions | Bitplane | ||
| 7. Astrocyte arborization tracing | |||
| 3D Visualization-Assisted Analysis software suite (Vaa3D) | HHMI – Janelia Research Campus /Allen Institute for Brain Science |
Riferimenti
- Clarke, L. E., Barres, B. A. Emerging roles of astrocytes in neural circuit development. Nature Reviews Neuroscience. 14 (5), 311-321 (2013).
- Ma, Z., Stork, T., Bergles, D. E., Freeman, M. R. Neuromodulators signal through astrocytes to alter neural circuit activity and behaviour. Nature. 539 (7629), 428-432 (2016).
- Blanco-Suárez, E., Caldwell, A. L. M., Allen, N. J. Role of astrocyte-synapse interactions in CNS disorders: Astrocyte-synapse disease. The Journal of Physiology. 595 (6), 1903-1916 (2017).
- Stogsdill, J. A., et al. Astrocytic neuroligins control astrocyte morphogenesis and synaptogenesis. Nature. 551 (7679), 192-197 (2017).
- Srinivasan, R., et al. New Transgenic Mouse Lines for Selectively Targeting Astrocytes and Studying Calcium Signals in Astrocyte Processes in Situ and In Vivo. Neuron. 92 (6), 1181-1195 (2016).
- Tabata, H., Nakajima, K. Efficient in utero gene transfer system to the developing mouse brain using electroporation: visualization of neuronal migration in the developing cortex. Neuroscienze. 103 (4), 865-872 (2001).
- Shimogori, T., Ogawa, M. Gene application with in utero electroporation in mouse embryonic brain. Development, Growth & Differentiation. 50 (6), 499-506 (2008).
- García-Marqués, J., López-Mascaraque, L. Clonal Identity Determines Astrocyte Cortical Heterogeneity. Cerebral Cortex. 23 (6), 1463-1472 (2013).
- Martín-López, E., García-Marques, J., Núñez-Llaves, R., López-Mascaraque, L. Clonal Astrocytic Response to Cortical Injury. PLoS ONE. 8 (9), 74039 (2013).
- Gutiérrez, Y., et al. Sibling astrocytes share preferential coupling via gap junctions. Glia. 67 (10), 1852-1858 (2019).
- Lee, Y., Messing, A., Su, M., Brenner, M. GFAP promoter elements required for region-specific and astrocyte-specific expression. Glia. 56 (5), 481-493 (2008).
- Yoon, H., Walters, G., Paulsen, A. R., Scarisbrick, I. A. Astrocyte heterogeneity across the brain and spinal cord occurs developmentally, in adulthood and in response to demyelination. PloS One. 12 (7), 0180697 (2017).
- Ge, W. P., Miyawaki, A., Gage, F. H., Jan, Y. N., Jan, L. Y. Local generation of glia is a major astrocyte source in postnatal cortex. Nature. 484 (7394), 376-380 (2012).
- Clavreul, S., et al. Cortical astrocytes develop in a plastic manner at both clonal and cellular levels. Nature Communications. 10 (1), 4884 (2019).
- Figueres-Oñate, M., García-Marqués, J., López-Mascaraque, L. UbC-StarTrack, a clonal method to target the entire progeny of individual progenitors. Scientific Reports. 6 (1), 33896 (2016).
- Figueres-Oñate, M., García-Marqués, J., Pedraza, M., De Carlos, J. A., López-Mascaraque, L. Spatiotemporal analyses of neural lineages after embryonic and postnatal progenitor targeting combining different reporters. Frontiers in Neuroscience. 9, 87 (2015).
- Figueres-Oñate, M., Sánchez-Villalón, M., Sánchez-González, R., López-Mascaraque, L. Lineage Tracing and Cell Potential of Postnatal Single Progenitor Cells In Vivo. Stem Cell Reports. 13 (4), 700-712 (2019).
- Peng, H., Ruan, Z., Long, F., Simpson, J. H., Myers, E. W. V3D enables real-time 3D visualization and quantitative analysis of large-scale biological image data sets. Nature Biotechnology. 28 (4), 348-353 (2010).
- Peng, H., Bria, A., Zhou, Z., Iannello, G., Long, F. Extensible visualization and analysis for multidimensional images using Vaa3D. Nature Protocols. 9 (1), 193-208 (2014).
- Peng, H., et al. Virtual finger boosts three-dimensional imaging and microsurgery as well as terabyte volume image visualization and analysis. Nature Communications. 5 (1), 4342 (2014).
- Loulier, K., et al. Multiplex Cell and Lineage Tracking with Combinatorial Labels. Neuron. 81 (3), 505-520 (2014).
- Niwa, H., Yamamura, K., Miyazaki, J. Efficient selection for high-expression transfectants with a novel eukaryotic vector. Gene. 108 (2), 193-199 (1991).
- Abdeladim, L., et al. Multicolor multiscale brain imaging with chromatic multiphoton serial microscopy. Nature Communications. 10 (1), 1662 (2019).
- Chen, F., LoTurco, J. A method for stable transgenesis of radial glia lineage in rat neocortex by piggyBac mediated transposition. Journal of Neuroscience Methods. 207 (2), 172-180 (2012).
- Siddiqi, F., et al. Fate Mapping by PiggyBac Transposase Reveals That Neocortical GLAST+ Progenitors Generate More Astrocytes Than Nestin+ Progenitors in Rat Neocortex. Cerebral Cortex. 24 (2), 508-520 (2014).
- Yoshida, A., et al. Simultaneous expression of different transgenes in neurons and glia by combining in utero electroporation with the Tol2 transposon-mediated gene transfer system. Genes to Cells. 15 (5), 501-502 (2010).
- LoTurco, J., Manent, J. B., Sidiqi, F. New and Improved Tools for In Utero Electroporation Studies of Developing Cerebral Cortex. Cerebral Cortex. 19, 120-125 (2009).
- Pacary, E., et al. Visualization and Genetic Manipulation of Dendrites and Spines in the Mouse Cerebral Cortex and Hippocampus using in utero Electroporation. Journal of Visualized Experiments. (65), e4163 (2012).

