Generation and Assembly of Virus-Specific Nucleocapsids of the Respiratory Syncytial Virus
Summary
For in-depth mechanistic analysis of the respiratory syncytial virus (RSV) RNA synthesis, we report a protocol of utilizing the chaperone phosphoprotein (P) for coexpression of the RNA-free nucleoprotein (N0) for subsequent in vitro assembly of the virus-specific nucleocapsids (NCs).
Abstract
The use of an authentic RNA template is critical to advance the fundamental knowledge of viral RNA synthesis that can guide both mechanistic discovery and assay development in virology. The RNA template of nonsegmented negative-sense (NNS) RNA viruses, such as the respiratory syncytial virus (RSV), is not an RNA molecule alone but rather a nucleoprotein (N) encapsidated ribonucleoprotein complex. Despite the importance of the authentic RNA template, the generation and assembly of such a ribonucleoprotein complex remain sophisticated and require in-depth elucidation. The main challenge is that the overexpressed RSV N binds non-specifically to cellular RNAs to form random nucleocapsid-like particles (NCLPs). Here, we established a protocol to obtain RNA-free N (N0) first by co-expressing N with a chaperone phosphoprotein (P), then assembling N0 with RNA oligos with the RSV-specific RNA sequence to obtain virus-specific nucleocapsids (NCs). This protocol shows how to overcome the difficulty in the preparation of this traditionally challenging viral ribonucleoprotein complex.
Introduction
Nonsegmented negative-sense (NNS) RNA viruses include many significant human pathogens, such as rabies, Ebola, and respiratory syncytial virus (RSV)1,2. RSV is the leading cause of respiratory illness such as bronchiolitis and pneumonia in young children and older adults worldwide3. Currently, no effective vaccines or antiviral therapies are available to prevent or treat RSV4. As part of the life cycle, the RSV genome serves as the template for replication by the RSV RNA dependent RNA polymerase to produce an antigenome, which in turn acts as the template to generate a progeny genome. Both genome and antigenome RNAs are entirely encapsidated by the nucleoprotein (N) to form the nucleocapsids (NCs)3. Because the NCs serve as the templates for both replication and transcription by the RSV polymerase, proper NC assembly is crucial for the polymerase to gain access to the templates for RNA synthesis5. Interestingly, based on the structural analyses of the NNS viral polymerases, it is hypothesized that several N proteins transiently dissociate from the NCs to allow the access of the polymerase and rebind to RNA after the RNA synthesis6,7,8,9,10,11,12.
Currently, the RSV RNA polymerization assay has been established using purified RSV polymerase on short naked RNA templates13,14. However, the activities of the RSV polymerase do not reach optimal, as observed in the non-processive and abortive products generated by the RSV polymerase when using naked RNA templates. The lack of NC with virus-specific RNA is a primary barrier for the further mechanistic understanding of the RSV RNA synthesis. Therefore, using an authentic RNA template becomes a critical need to advance the fundamental knowledge of RSV RNA synthesis. The known structures of the nucleocapsid-like particles (NCLPs) from RSV and other NNS RNA viruses reveal that the RNAs in the NCLPs are either random cellular RNAs or average viral genomic RNAs15,16,17,18,19. Together, the main hurdle is that N binds non-specifically to cellular RNAs to form NCLPs when N is overexpressed in the host cells.
To overcome this hurdle, we established a protocol to obtain RNA-free (N0) first and assemble N0 with authentic viral genomic RNA into NCLPs20. The principle of this protocol is to obtain a large quantity of recombinant RNA-free N (N0) by co-expressing N with a chaperone, the N-terminal domain of RSV phosphoprotein (PNTD). The purified N0P could be stimulated and assembled into NCLPs by adding RSV-specific RNA oligos, and during the assembly process, the chaperone PNTD is displaced upon the addition of RNA oligos.
Here, we detail a protocol for the generation and assembly of RSV RNA-specific NCs. In this protocol, we describe the molecular cloning, protein preparation, in vitro assembly, and validation of the complex assembly. We highlight the cloning strategy to generate bi-cistronic constructs for protein coexpression for molecular cloning. For protein preparation, we describe the procedures of cell culture, protein extraction, and the purification of the protein complex. Then we discuss the method for in vitro assembly of the RSV RNA-specific NCs. Finally, we use size exclusion chromatography (SEC) and negative stain electron microscopy (EM) to characterize and visualize the assembled NCLPs.
Protocol
1. Molecular cloning
NOTE: Ligase Independent Cloning (LIC) was used to make an RSV bi-cistronic coexpression construct plasmid. LIC is a method developed in the early 1990s, which uses the 3’-5’ Exo activity of the T4 DNA polymerase to create overhangs with complementarity between the vector and the DNA insert21,22. The constructs were made using the 2BT-10 vector DNA, which consists of a 10x His tag at the N-terminal of the Open Reading Frame (ORF) (Figure 1).
- Perform linearization of LIC vectors using SSPI digestion.
- Combine 10 μL of SSPI 10X buffer, 4 μL of SSPI enzyme at a concentration of 5 U/μL, the equivalent volume of 5 μg of vector miniprep DNA, and sterile ddH2O to 100 μL.
- Incubate the digest for 3 hours at 37 °C.
- Run the digest on a 1.0% agarose gel for the extraction of vector DNA.
- Use a gel extraction kit to perform extraction and purification. Suspend the final volume of vector DNA in 30 μL of ddH2O and store it at -20 °C.
- Prepare the DNA inserts for N1-391 and P1-126 using the N1-391 Forward Primer, N1-391 Reverse Primer, P1-126 Forward Primer, and P1-126 Reverse Primer (Table 1).
NOTE: There is sufficient overlap with the linearized vector to ensure a melting temperature of between 55 °C and 60 °C. For the reverse primer, there is sufficient overlap with the reverse complementary strand of the linearized vector for the same reason.- Perform PCR amplification of the DNA insert using the conditions in Table 2 and Table 3.
- Extract the amplified DNA insert. Run the PCR products from the previous step on a 1.0% agarose gel, then extract and purify the bands by gel extraction. Suspend the final volume of extracted DNA in 15 μL of ddH2O.
- T4 DNA polymerase treatment of vector and insert DNA (Table 4).
NOTE: Treatment must be performed separately for the vector DNA and the insert DNA.- Incubate the mixture for 40 minutes at room temperature. Then, heat-inactivate the polymerase at 75 °C for 20 minutes. Store the reaction mixture at -20 °C.
- Anneal the LIC vector and the insert vector.
- Set up a negative control with 2 μL of LIC vector DNA and 2 μL of sterile ddH2O.
- Combine 2 μL of insert DNA and 2 μL of LIC vector DNA from the previous T4 DNA polymerase reactions in a 0.2 mL tube.
- Perform the annealing reaction at room temperature for 10 minutes.
- Quench the reaction with 1.3 μL of EDTA at a concentration of 25 mM.
- Transform the reaction into 100 μL of E. coli Top10 competent cells and plate them on an ampicillin selection plate23,24.
- Identify the positive constructs.
- Prepare the plasmid miniprep solution. This can be done through colony picking and inoculation in LB media. Usually, 3 colonies are sufficient.
- Incubate the mixture overnight at 37 °C.
- Centrifuge the mixture at 4,560 x g for 10 minutes and discard the supernatant.
- Resuspend the pellets in 250 μL of P1 buffer and prepare plasmid minipreps using a spin miniprep kit.
- Conduct a digestion analysis of the miniprep product using AseI or other restriction enzymes.
- Load samples on 0.8% agarose gel and run the digested plasmid. Analyze the gel under a UV lamp.
- Use a sequencing service to validate the sequence of the positive product.
- Obtain the coexpression DNA insert.
- Perform PCR to obtain N1-391 and P1-126 using the previously constructed 2BT-10 N1-391 and 2BT-10 P1-126 as templates.
- Perform the 1st PCR to obtain N1-391 from the 2BT-10 N1-391 construct using the PCR conditions in Table 2 and Table 3 with the N1-391 Forward primer and Reverse Primer 5 ́-GTGAAGATCCTGGCTGATGCAATGCGGCGGCG
CGCCGCGATCGCGGATCC-3 ́. - Perform the 2nd PCR to obtain P1-126 from the 2BT-10 P1-126 construct using the Forward primer: 5’-CCGCCGCATTGCATCAGCCAGGATCTTCACTGC
AGGACTCGAGTTCTAGA-3 ́and the P1-126 Reverse primer (use the PCR conditions in Table 2 and Table 3). - Finally, perform overlap PCR on the mixed products of the previous 2 PCR reactions to merge N1-391 and P1-126. Use the N1-391 Forward primer and P1-126 Reverse primer. Use the PCR conditions in Table 5 and Table 6.
- Join the vector and the DNA insert.
- Treat the overlap PCR product with T4 DNA polymerase following the protocol in step 1.3.
- Anneal the LIC vector and PCR product following the protocol in step 1.4.
- Identify the positive constructs following the protocol in step 1.5.
2. Protein expression and purification
NOTE: Use E. coli for the bi-cistronic construct of the coexpression of both N and P. Culture the cells at 37 °C, but carry out the expression at a reduced temperature (16 °C) overnight. Purify the protein complexes through a combination of cobalt column, ion exchange, and size exclusion chromatography (Figure 2).
- Use the E. coli BL21(DE3) strain for protein production. Grow 4 L cell cultures at 37 °C in LB (Luria Broth) medium until OD600 reaches 0.6.
- Lower the temperature to 16 °C. An hour later, induce the expression with 0.5 mM isopropyl 1-thio–D-galactopyranoside (IPTG) overnight.
- Centrifuge the cells at 4,104 x g for 25 min and then discard the supernatant.
- Resuspend the cell pellets in 200 mL of lysis buffer A (50 mM sodium phosphate, pH 7.4, 500 mM NaCl, 5 mM imidazole, 10% glycerol, and 0.2% NP40). Use 50 mL of lysis buffer to resuspend the cell pellets from 1 L of cell culture.
- Lyse the cells by sonication for 15 min, 3 seconds on, and 3 seconds off. Then centrifuge cells at 37,888 x g for 40 min.
NOTE: The protocol can be paused by freezing the cells before sonication in a -80 °C freezer. - Load the supernatant into a cobalt gravity column (diameter x length: 2.5 cm x 10 cm) with ~10mL of beads pre-equilibrated with 5-10 column volumes (CV) of lysis buffer.
- Wash the column with 5 CV of buffer B (50 mM Tris-HCl pH 7.4, 1 M NaCl, 10% glycerol, and 5 mM imidazole) and 5 CV of buffer C (50 mM Tris-HCl pH 7.4, 500 mM NaCl, 10% glycerol, and 5 mM imidazole).
- Elute the protein from the beads using 2 CV of buffer D (50 mM Tris-HCl pH 7.4, 500 mM NaCl, and 250 mM imidazole).
- Dilute the eluted protein 5x with QA buffer (50 mM Tris-HCl pH 8.0, 5% Glycerol) for the Q column.
- Wash the 5 mL of Q column with QA buffer to equilibrate the column, then load the diluted sample into the Q column using the peristaltic pump (e.g., Rabbit).
- Load the Q column into the HPLC machine along with QA buffer and QB buffer(50 mM Tris-HCl pH7.4, 1.5 M NaCl, 5% glycerol). Set up the flow rate as 1 mL/min.
- Run the “pump wash” program to wash the machine with QB buffer followed by QA buffer (1-2 CVs/each). Set the system flow to 3 mL/min.
- Set the UV1 to 280 nm and UV2 to 260 nm. Use a 96 deep-well plate to collect the fractions.
- Elute proteins using a stepwise gradient of elution agent (QB Buffer) applying 3-4 CV of each concentration, increasing the percentage by 5% each time starting at 0% QB. N0P protein complex will come out at 15% QB Buffer.
- Once all of the protein is eluted, wash the column with 100% QB Buffer (2 CV).
- Isolate the protein by gel filtration Superdex 200 Increase 10/300 GL column (diameter x length: 1.0 cm x 30 cm) and equilibrate with buffer E (20 mM HEPES pH 7.4 and 200 mM NaCl).
- Analyze protein-containing fractions by SDS-PAGE.
3. In vitro assembly of the virus-specific NC
NOTE: The in vitro assembly of the RSV-specific NC (N:RNA) was performed by incubating the prepared N0P complex with RNA oligos. Then, SEC chromatography was used to separate the assembly complex from the N0P and excess RNA (Figure 2).
- Mix and incubate the purified N0P complex with RNA oligo with the molecular ratio of 1:1.5 at room temperature for 1 hour, usually 1 mL of the protein N0P with a concentration of 1 mg/mL is enough for the next step. Set up the control sample, which only contains the same amount of N0P protein.
- Pre-equilibrate the gel filtration Superdex 200 Increase 10/300 GL column with the buffer E (20 mM HEPES, pH 7.4, 200 mM NaCl).
- Centrifuge the sample with 21,130 x g for 15 min, remove any precipitation and load the supernatant to the SEC column.
- Compare the SEC chromatography images of N:RNA assembly sample and N0P control sample, combine the A260/A280 ratio to identify which peaks are the assembled N-RNA, N0P, and free RNA.
- Collect the peak fractions, run the SDS-PAGE gel, or make grids.
- For the assembly N-RNA complex, collect all the fractions of N-RNA peak, do the RNA extraction and run the Urea-PAGE gel to double-check the length of specific RNA, which is the same as mixed and incubated at the first step.
4. Making negative stain grids
NOTE: Negative stain electron microscopy (EM) is a method in which the molecules are adsorbed to a carbon film and then embedded in a layer of heavy metal atoms. Negative stain EM produces a high image contrast, making it easy to see and computationally align the particles. Another advantage is that the adsorption of the particles to a carbon film usually induces the molecules to adhere to the grid with few preferred orientations. When the molecules are in a similar orientation, it is easy to separate them into structurally distinct classes. Negative stain EM is thus the appropriate technique to guide sample preparation25,26 (Figure 3).
- Microwave or heat 5 mL of ddH2O in a small glass beaker using a Tungsten heater until it is boiling.
- Weigh 37.5 mg of uranyl formate and add to 5 mL of heated ddH2O to make a 0.75% uranyl formate staining solution. Stir under an aluminum foil covered beaker to protect from light.
- Add 4 μL of 10 M NaOH to the staining solution and continue to stir for 15 min, protected from the light.
- Filter the solution through a 0.22 mm filter into a test tube.
- Use glow discharge to make the continuous carbon-coated EM grids hydrophilic27.
NOTE: The grids are placed inside a chamber connected to a power supply. When high voltage is applied, the gas within the chamber ionizes, and the negatively charged ions deposit on the carbon grids to make them hydrophilic. - Cut and fold a parafilm strip. Pipet 2 drops of 40 μL of the buffer on one side of the parafilm, and pipet another 2 drops of 40 μL of staining solution to the other side.
- To make the EM carbon grids, apply 3 μL of protein sample for 1 minute.
- Blot the grids against a blotting paper.
NOTE: The grids are washed 2x with buffer, and 1x with the 0.75% uranyl formate staining solution blotted after each step. - Hold the grid surface in the second drop of 0.75% uranyl formate staining solution for 30 seconds.
- Blot the grid against a blotting paper to remove excess stain solution and allow the grid to air-dry.
- Store the grids in the grid box before imaging.
Representative Results
Purification of RNA-free N0P protein
With this protocol, a large-scale soluble heterodimeric RSV N0P complex can be obtained. The full length of N and N terminal part of P proteins were co-expressed with 10X His-Tag on the N protein in E. coli. N0P was purified using a cobalt column, ion exchange, and size exclusion chromatography. N0P contains both the full-length N and N terminal P but did not contain cellular RNA based on the UV absorbance A260/A280 ratio20 (Figure 4).
Assembly of N-RNA and checking with negative stain
We then demonstrated that the purified N0P could be stimulated and assembled into Nuceloplasmid-like particles (NCLPs) by incubating with specific RNA oligos. The NCLPs were assembled by incubating the N0P with RNA oligos with the ratio of 1:1.5 at room temperature for 1 hour and then run through the gel filtration column. When the N:RNA complex forms, it shows three peaks: the 1st peak is N:RNA, the 2nd peak is N0P, and the 3rd peak is excess free RNA. The highest fraction of the N:RNA peak creates the negative stain grids for checking with EM20 (Figure 5).
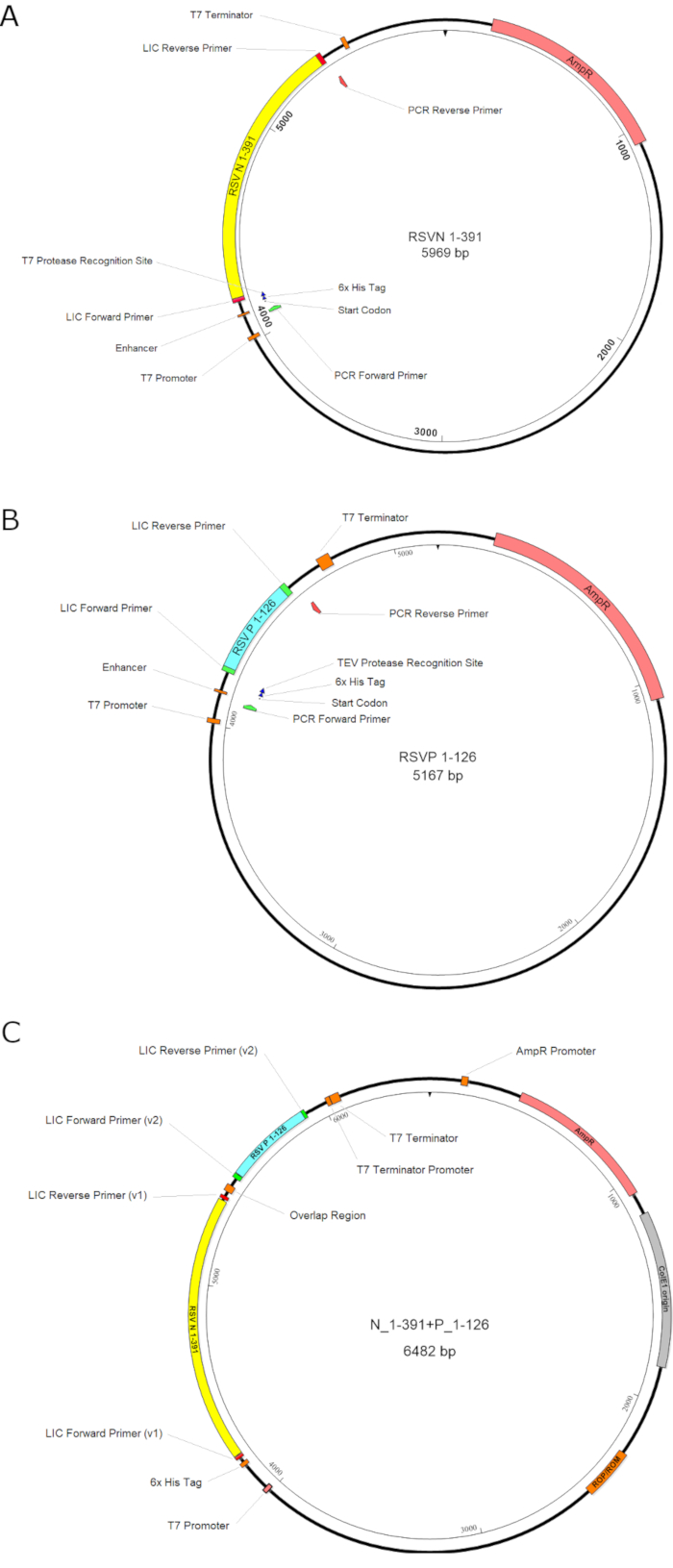
Figure 1. The illustration of the plasmid constructions. A. The construct of the RSV N1-391; B. The construct of the RSV P1-126; C. The bi-cistronic construct for the coexpression of N1-391 and P1-126. The first gene RSV N1-391, the second gene RSV P1-126, the antibiotic-resistant gene (AmpR), and the promoters are highlighted in yellow, cyan, pink, and orange boxes, respectively. In summary, the gene inserts of the RSV N1-391 and the RSV P1-126 are constructed separately and assembled. Please click here to view a larger version of this figure.
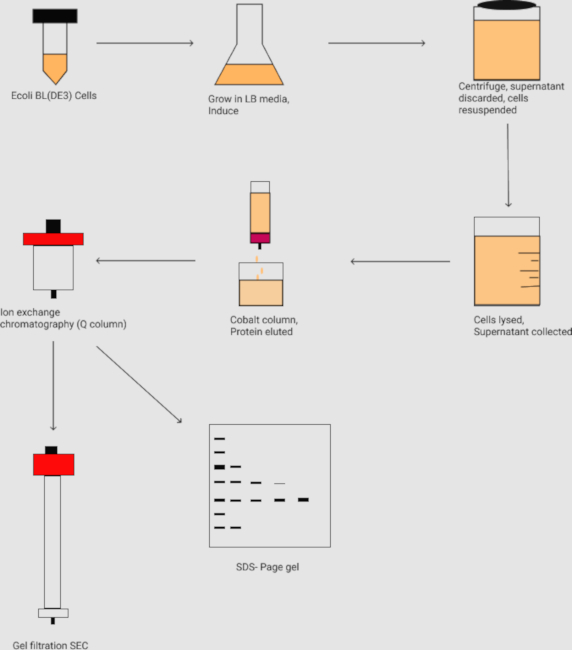
Figure 2. The flowchart of the purification of the protein complex N1-391P1-126. It outlines the inoculation and large scale grow-ups of the E. coli cell culture and harvesting the cell by centrifugation. Followed by cell lysis, the protein samples are purified by the affinity chromatography (i.e., Co2+ column), ion-exchange chromatography (i.e., Q column), and gel filtration size exclusion (SEC) chromatography. The protein samples are further analyzed by the SDS-PAGE gel. Please click here to view a larger version of this figure.
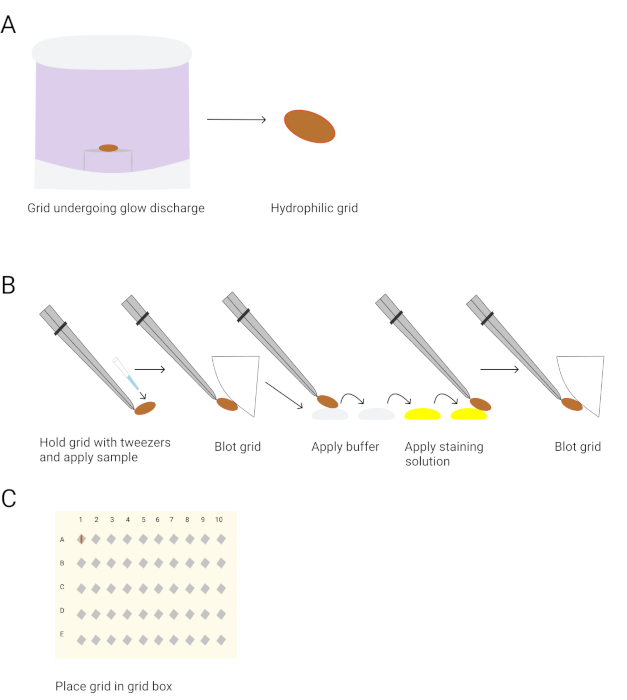
Figure 3. Preparation of negative stain EM grids for imaging. A. Glow discharges the grids. B. The procedure for negative staining grids is shown. The tweezers are used to pick up a grid, followed by applying the protein sample for 1 minute. The grids are blotted with blotting paper. The grid is washed twice with buffer and twice with the 0.75% uranyl formate staining solution. The grids are held in the second staining solution drop for 30 seconds. The grids are blotted after each wash and air-dried after the final blotting. C. The grids are stored in the grid box for imaging. Please click here to view a larger version of this figure.
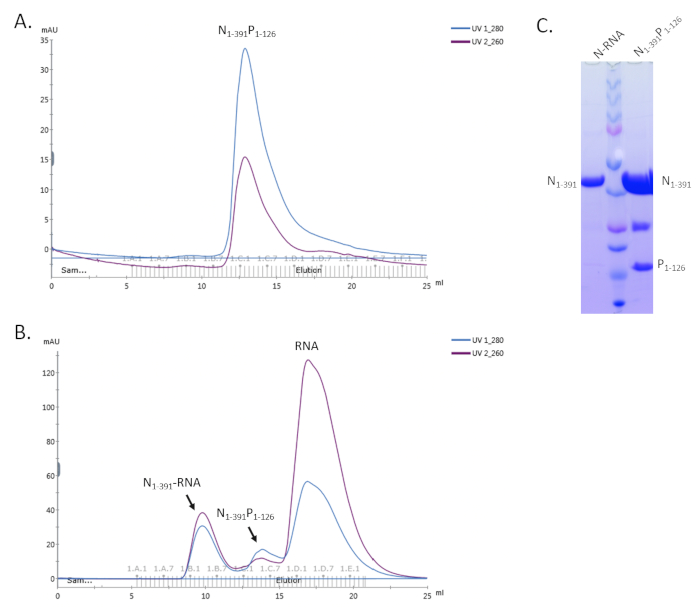
Figure 4. Representative results of the copurification of the N1-391P1-126 complex. A. The SEC profile of N1-391P1-126. B. The SEC profile of the assembly N1-391P1-126 with RNA. C. The SDS-PAGE gel shows the N protein only for the N-RNA complex and the bands for both N and P proteins of the N1-391P1-126 complex. Please click here to view a larger version of this figure.
Figure 5. Representative images of N-RNA. A and B are representative negative stain EM images of N-RNA from the N1-391-RNA peak in Figure 4. The N-RNA complexes are stained using the procedure described in Figure 3. Please click here to view a larger version of this figure.
| Primers | |||
| N1-391 Forward | 5’-TACTTCCAATCCAATGCAATGGCCCTGAGCAAAGTGAAG-3’ | ||
| N1-391 Reverse | 5’-TTATCCACTTCCAATGTTATTACAGTTCCACGTCGTTGTCCTTGG-3' | ||
| P1-126 Forward | 5’-TACTTCCAATCCAATGCAATGGAAAAGTTCGCCCCCGAG-3' | ||
| P1-126 Reverse | 5’-TTATCCACTTCCAATGTTATTACTGGTCGTTGATTTCCTCGTAGC-3’ | ||
Table 1. Primer sequences.
| PCR amplification of DNA insert | |
| 15 μL | 10x Pfu polymerase reaction buffer |
| 3 μL | Forward primer (100 μM concentration) |
| 3 μL | Reverse primer (100 μM concentration) |
| 15 μL | dNTP mix at 2.5 mM concentration |
| 6 μL | Plasmid DNA contains the gene of N or P (100ng/ μL) |
| 7 μL | DMSO |
| 3 μL | Pfu polymerase at 2.5U/μL |
| Volume to fill to 150 μL | Sterile ddH2O |
Table 2. PCR amplification of DNA insert reagents.
| PCR amplication of DNA insert | |||
| Step | Time | Temperature | Cycles |
| Denaturation | 4 min. | 95 ºC | 1 |
| Denaturation | 45 sec. | 95 ºC | 30 |
| Annealing | 30 sec. | 62 ºC | |
| Extension* | 90 sec. | 72 ºC | |
| Extension | 10 min. | 72 ºC | 1 |
| Hold | ∞ | 4 ºC | 1 |
| The 150 μL mixture can be run in three separate PCR reactions (3 x 50μL). | |||
| *For the Pfu DNA polymerase, 1 kb/min is the recommended speed for the extension phase. Here, both the lengths of the N1-391 gene or the P1-126 gene are shorter than 1.5 Kb. | |||
| Thus, 90 seconds was used for the extension step. | |||
Table 3. PCR amplification of DNA insert thermocycling program.
| T4 DNA polymerase treatment | |
| 10 x Buffer | 2 μL |
| Vector/Insert DNA (0.1 pmol vector or 0.2 pmol insert) | 5 μL |
| dNTP* at 25 mM | 2 μL |
| DTT at 100 mM | 1 μL |
| T4 DNA polymerase (LIC qualified) | 0.4 μL (1.25 U) |
| Sterile ddH2O | 9.6 μL |
| *dGTP was used for the vector, and dCTP was used for the insert DNA. | |
Table 4. T4 DNA polymerase treatment.
| Overlap PCR | |
| 15 μL | 10 X Pfu polymerase reaction buffer |
| 3 μL | Forward primer (100 μM) |
| 3 μL | Reverse primer (100 μM) |
| 15 μL | dNTP Mix (2.5 mM) |
| 3 μL | DNA from 1st round PCR which contain the gene N1-391 (100 ng/μL ) |
| 3 μL | DNA from 1st round PCR which contain the gene P1-126 (100 ng/μL ) |
| 7 μL | DMSO |
| 3 μL | Pfu polymerase (2.5 U/μL) |
| Volume to fill to 150 μL | Sterile ddH2O |
Table 5. Overlap PCR reagents.
| Overlap PCR | |||
| Step | Time | Temperature | Cycles |
| Denaturation | 4 min. | 95 °C | 1 |
| Denaturation | 45 sec. | 95 °C | 30 |
| Annealing | 30 sec. | 62 °C | |
| Extension* | 2 min. | 72 °C | |
| Extension | 10 min. | 72 °C | 1 |
| Hold | ∞ | 4 °C | 1 |
| The 150 μL mixture can be run in three separate PCR reactions (3 x 50 μL). | |||
| *For the Pfu DNA polymerase, 1 kb/min is the recommended speed for the extension phase. Here, the total length of the gene N1-391 and P1-126 is shorter than 2.0 Kb. Thus, 2 minutes were used for the extension step. | |||
Table 6. Overlap PCR thermocycling program.
Discussion
The known nucleocapsid-like particle (NCLP) structures of the nonsegmented negative-sense (NNS) RNA viruses show that the assembled NCLPs are the complex N with host cellular RNAs when overexpressed in bacterial or eukaryotic expression systems15,16,17,18,19. Previous studies have attempted to get the RNA free N with a variety of methods, such as the RNase A digestion, high salt washing, or adjusting different pH buffers to remove the nonspecific cellular RNAs28,29. However, none of the above methods can be successfully used for the assembly of the virus-specific NCs. To obtain the RNA-free RSV N, we also tried a combination of methods, including RNase A digestion, high salt (1.5M NaCl) wash, adjusting the buffer pH from pH 5.0 to pH 9.0, protein denaturation and renaturation. After many failed trials, we still could not get RNA-free N with the above methods. We will briefly discuss the attempts and potential reasons.
One method to obtain RNA-free N is to digest host cellular RNA in assembled NCLPs with RNase A. In VSV, the incubation of the purified NCLP with RNase A at a final concentration of 1mg/ml at 37 °C for 1 h completely removed RNA from the NC28. Purified empty oligomeric N was then incubated with poly-A (250-nt or longer) in a molar ratio of 1:5 in the presence of RNase inhibitors. Analysis of the RNA isolated from reconstituted N:poly-A showed that the RNA was approximately 90 nt in length. This suggested that the RNA outside the nucleoprotein is susceptible to nonspecific digestion by the contaminated RNase A from the previous step. The strategy of RNase A digestion to remove RNA was not successful when applied to RSV. This may be due to two reasons. First, contaminated RNase A will digest the RSV-specific RNA, which will subsequently be incubated and assembled with N. Second, the efficiency of the RNase A digestion is much lower in RSV. This is because the RNAs assemble differently in different NNS viruses. The known crystal structures of N:RNA show that RNA binds outside of the NC in VSV, but inside of the NC in RSV30,31. The configuration of RNA binding inwards of the NC may lead to low efficiency for RNase A to access and digest.
Another method to get RNA-free N is to make the truncations that cut both the N-terminal motif (N-arm) and the C-terminal motif (C-arm) of N. However, this truncated RNA-free N cannot be used to assemble with RNA into a stable NC because the N-arm of N is folded into its neighboring subunits by interacting with the C-terminal domain (CTD) of the precedent N subunit. The extended C-arm is positioned to the CTD of the next N subunit31.
An additional method to get RNA-free N is to prepare mutant N. For example, the result obtained by Galloux et al. showed that RSV RNA free N0P complex could be obtained by coexpression of a K170A/R185A double N mutant with the N-terminus of P in bacteria32. However, it has two potential issues for further structural characterizations. One issue is the low stability of this mutant complex at high concentrations. The other issue is that the mutant complex lost the RNA binding ability, which cannot be used in the next step of assembly.
Despite the tremendous challenges, we have established and optimized the protocol to obtain virus-specific NCs using the coexpression of N with a chaperone P. Recently, another successful method is to make a chimeric fusion construction encoding for P1-50-TEV-N1-405-8xHis for MeV33,34. N0P can be obtained after the TEV protease cleavage. The purity of N0P depends on the efficiency of TEV cleavages, which cut the chimeric fusion between N and P protein.
Instead of using the chimeric fusion method, we designed a bi-cistronic coexpression construct. Specifically, the coexpression constructs of the N0P complex have been designed and engineered in two open reading frames, comprising the full length of N (1-391) with a 10x His-tag at N-terminal in the first ORF, and the N-terminal peptides (1-126) from P in the second ORF20. Briefly, the overall procedure of the N0P purification is to purify His-tagged N0P and N-RNA from cellular lysis samples with cobalt beads, remove the nonspecific RNA and N:cellular RNA complex with Q column, and get a pure N0P complex with the SEC column. In the SEC step, the ratio of A260/A280 can be monitored and double-checked with RNA extraction from the N0P peak fractions.
Collectively, in this protocol, the most critical steps are the strategy to design the construction of the coexpression of N0P complex and using a series affinity and ion-exchange column to separate the RNA free N0P complex from the other N-cellular RNA complexes. The efficiency of the coexpression strategy to get N0P complex is relatively low; around 50% N protein is still N-cellular RNA complex. The protocol may also be applied for getting RNA free N0P and assembly with specific RNA with N to get N:RNA complex of other NNS viruses.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
The research programs in the Liang laboratory at Emory are supported by the US National Institute of General Medical Sciences (NIGMS), National Institutes of Health (NIH) under award number R01GM130950, and the Research Start-Up Fund at Emory University School of Medicine. The author acknowledges the members of the Liang laboratory for helpful support and critical discussion.
Materials
| Agarose | SIgma | A9539-500G | making construct using LIC method |
| Amicon Ultra-15 Centrifugal Filter Unit | Millipore | UFC901024 | concentrate the protein sample |
| Ampicillin sodium | GOLD BIOTECHNOLOGY | 5118.111317A | antibiotic for cell culture |
| AseI | NEB | R0526S | making construct using LIC method |
| Cobalt (High Density) Agarose Beads | Gold Bio | H-310-500 | For purification of His-tag protein |
| Corning LSE Digital Dry Bath Heater | CORNING | 6885-DB | Heate the sample |
| dCTP | Invitrogen | 10217016 | making construct using LIC method |
| dGTP | Invitrogen | 10218014 | making construct using LIC method |
| Glycerol | Sigma | G5516-4L | making solution |
| HEPES | Sigma | H3375-100G | making solution |
| HiTrap Q HP | Sigma | GE29-0513-25 | Protein purification |
| Imidazole | Sigma | I5513-100G | making solution |
| IPTG (Isopropyl-beta-D-thiogalactopyranoside) | GOLD BIOTECHNOLOGY | 1116.071717A | induce the expression of protein |
| Microcentrifuge Tubes | VWR | 47730-598 | for PCR |
| Misonix Sonicator XL2020 Ultrasonic Liquid Processor | SpectraLab | MSX-XL-2020 | sonicator for lysing cell |
| Negative stain grids | Electron Microscopy Sciences | CF400-Cu-TH | For making negative stain grids |
| New Brunswick Innova 44/44R | eppendorf | M1282-0000 | Shaker for culturing the cell |
| Nonidet P 40 Substitute | Sigma | 74385-1L | making solution |
| OneTaq DNA Polymerase | NEB | M0480L | PCR |
| QIAquick Gel Extraction Kit | QIAGEN | 28706 | Purify DNA |
| SSPI-HF | NEB | R3132S | making construct using LIC method |
| Superose 6 Increase 10/300 GL | Sigma | GE29-0915-96 | Protein purification |
| T4 DNA polymerase | Sigma | 70099-3 | making construct using LIC method |
| Thermo Scientific Sorvall RC 6 Plus Centrifuge | Fisher Scientific | 36-101-0816 | Centrifuge, highest speed 20,000 rpm |
| Trizma hydrochloride | Sigma | T3253-250G | making solution |
| Uranyl Formate | Electron Microscopy Sciences | 22451 | making negative stain solution |
Riferimenti
- Whelan, S. P., Barr, J. N., Wertz, G. W. Transcription and replication of nonsegmented negative-strand RNA viruses. Current Topics in Microbiology and Immunology. 283, 61-119 (2004).
- Lamb, R. A., Knipe, D. M., Howley, P. M. . Fields virology. , (2013).
- Collins, P. L., Fearns, R., Graham, B. S. Respiratory syncytial virus: virology, reverse genetics, and pathogenesis of disease. Current Topics in Microbiology and Immunology. 372, 3-38 (2013).
- Fearns, R., Deval, J. New antiviral approaches for respiratory syncytial virus and other mononegaviruses: Inhibiting the RNA polymerase. Antiviral Research. 134, 63-76 (2016).
- Grosfeld, H., Hill, M. G., Collins, P. L. RNA replication by respiratory syncytial virus (RSV) is directed by the N, P, and L proteins; transcription also occurs under these conditions but requires RSV superinfection for efficient synthesis of full-length mRNA. Journal of Virology. 69 (9), 5677-5686 (1995).
- Pan, J., et al. Structure of the human metapneumovirus polymerase phosphoprotein complex. Nature. 577 (7789), 275-279 (2020).
- Horwitz, J. A., Jenni, S., Harrison, S. C., Whelan, S. P. J. Structure of a rabies virus polymerase complex from electron cryo-microscopy. Proceedings of the National Academy of Sciences of the United States of America. 117 (4), 2099-2107 (2020).
- Cao, D., et al. Cryo-EM structure of the respiratory syncytial virus RNA polymerase. Nature Communications. 11 (1), 368 (2020).
- Abdella, R., Aggarwal, M., Okura, T., Lamb, R. A., He, Y. Structure of a paramyxovirus polymerase complex reveals a unique methyltransferase-CTD conformation. Proceedings of the National Academy of Sciences of the United States of America. 117 (9), 4931-4941 (2020).
- Gilman, M. S. A., et al. Structure of the Respiratory Syncytial Virus Polymerase Complex. Cell. 179 (1), 193-204 (2019).
- Liang, B., et al. Structure of the L Protein of Vesicular Stomatitis Virus from Electron Cryomicroscopy. Cell. 162 (2), 314-327 (2015).
- Jenni, S., et al. Structure of the Vesicular Stomatitis Virus L Protein in Complex with Its Phosphoprotein Cofactor. Cell Reports. 30 (1), 53-60 (2020).
- Renner, M., et al. Nucleocapsid assembly in pneumoviruses is regulated by conformational switching of the N protein. Elife. 5, 12627 (2016).
- Cox, R. M., Plemper, R. K. Structure and organization of paramyxovirus particles. Current Opinion in Virology. 24, 105-114 (2017).
- Desfosses, A., et al. Self-organization of the vesicular stomatitis virus nucleocapsid into a bullet shape. Nature Communications. 4, 1429 (2013).
- Green, T. J., et al. Common mechanism for RNA encapsidation by negative-strand RNA viruses. Journal of Virology. 88 (7), 3766-3775 (2014).
- Jamin, M., Yabukarski, F. Nonsegmented Negative-Sense RNA Viruses-Structural Data Bring New Insights Into Nucleocapsid Assembly. Advances in Virus Research. 97, 143-185 (2017).
- Wan, W., et al. Structure and assembly of the Ebola virus nucleocapsid. Nature. 551 (7680), 394-397 (2017).
- Mendes, A., Kuhn, R. J. Alphavirus Nucleocapsid Packaging and Assembly. Viruses. 10 (3), (2018).
- Gao, Y., et al. In vitro trackable assembly of RNA-specific nucleocapsids of the respiratory syncytial virus. Journal of Biological Chemistry. 295 (3), 883-895 (2020).
- Aslanidis, C., de Jong, P. J. Ligation-independent cloning of PCR products (LIC-PCR). Nucleic Acids Research. 18 (20), 6069-6074 (1990).
- Li, C., Evans, R. M. Ligation independent cloning irrespective of restriction site compatibility. Nucleic Acids Research. 25 (20), 4165-4166 (1997).
- Hanahan, D. Studies on transformation of Escherichia coli with plasmids. Journal of Molecular Biology. 166 (4), 557-580 (1983).
- Green, R., Rogers, E. J. Transformation of chemically competent E. coli. Methods in Enzymology. 529, 329-336 (2013).
- De Carlo, S., Harris, J. R. Negative staining and cryo-negative staining of macromolecules and viruses for TEM. Micron. 42 (2), 117-131 (2011).
- Ohi, M., Li, Y., Cheng, Y., Walz, T. Negative Staining and Image Classification – Powerful Tools in Modern Electron Microscopy. Biological Procedures Online. 6, 23-34 (2004).
- Aebi, U., Pollard, T. D. A glow discharge unit to render electron microscope grids and other surfaces hydrophilic. Journal of Electron Microscopy Technique. 7 (1), 29-33 (1987).
- Green, T. J., et al. Access to RNA encapsidated in the nucleocapsid of vesicular stomatitis virus. Journal of Virology. 85 (6), 2714-2722 (2011).
- Alvarez Paggi, D., et al. A conformational switch balances viral RNA accessibility and protection in a nucleocapsid ring model. Archives of Biochemistry and Biophysics. 671, 77-86 (2019).
- Green, T. J., Zhang, X., Wertz, G. W., Luo, M. Structure of the vesicular stomatitis virus nucleoprotein-RNA complex. Science. 313 (5785), 357-360 (2006).
- Tawar, R. G., et al. Crystal structure of a nucleocapsid-like nucleoprotein-RNA complex of respiratory syncytial virus. Science. 326 (5957), 1279-1283 (2009).
- Galloux, M., et al. Characterization of a viral phosphoprotein binding site on the surface of the respiratory syncytial nucleoprotein. Journal of Virology. 86 (16), 8375-8387 (2012).
- Milles, S., et al. Self-Assembly of Measles Virus Nucleocapsid-like Particles: Kinetics and RNA Sequence Dependence. Angewandte Chemie Interntional Edition English. 55 (32), 9356-9360 (2016).
- Desfosses, A., et al. Assembly and cryo-EM structures of RNA-specific measles virus nucleocapsids provide mechanistic insight into paramyxoviral replication. Proceedings of the National Academy of Sciences of the United States of America. 116 (10), 4256-4264 (2019).

