Single-Molecule Dwell-Time Analysis of Restriction Endonuclease-Mediated DNA Cleavage
Summary
Using quantum-dot-labeled DNA and total internal reflection fluorescence microscopy, we can investigate the reaction mechanism of restriction endonucleases while using unlabeled protein. This single-molecule technique allows for massively multiplexed observation of individual protein-DNA interactions, and data can be pooled to generate well-populated dwell-time distributions.
Abstract
This novel total internal reflection fluorescence microscopy-based assay facilitates the simultaneous measurement of the length of the catalytic cycle for hundreds of individual restriction endonuclease (REase) molecules in one experiment. This assay does not require protein labeling and can be carried out with a single imaging channel. In addition, the results of multiple individual experiments can be pooled to generate well-populated dwell-time distributions. Analysis of the resulting dwell-time distributions can help elucidate the DNA cleavage mechanism by revealing the presence of kinetic steps that cannot be directly observed. Example data collected using this assay with the well-studied REase, EcoRV – a dimeric Type IIP restriction endonuclease that cleaves the palindromic sequence GAT↓ATC (where ↓ is the cut site) – are in agreement with prior studies. These results suggest that there are at least three steps in the pathway to DNA cleavage that is initiated by introducing magnesium after EcoRV binds DNA in its absence, with an average rate of 0.17 s-1 for each step.
Introduction
Restriction endonucleases (REases) are enzymes that effect sequence-specific double-strand breaks in DNA. The discovery of REases in the 1970s led to the development of recombinant DNA technology, and these enzymes are now indispensable laboratory tools for genetic modification and manipulation1. Type II REases are the most widely used enzymes in this class as they cleave DNA at a fixed location either within or near their recognition sequence. However, there is a great deal of variation among the Type II REases, and they are divided into several subtypes based on particular enzymatic properties rather than being classified according to their evolutionary relationships. Among each subtype, there are frequent exceptions to the classification scheme, and many enzymes belong to multiple subtypes2. Thousands of Type II REases have been identified, and hundreds of them are commercially available.
However, in spite of the diversity among the Type II REases, very few REases have been studied in detail. According to REBASE, the restriction enzyme database established by Sir Richard Roberts in 19753, published kinetics data are available for fewer than 20 of these enzymes. Furthermore, while some REases have been directly observed at the single-molecule level while diffusing along the DNA prior to encountering and binding to their recognition sequence4,5,6,7, there are very few single-molecule studies of their cleavage reaction kinetics. The existing studies either do not report adequate statistics to undertake detailed analysis of the variation in the times at which single cleavage events take place8,9,10 or are not capable of capturing the full distribution of cleavage times11. This type of analysis can reveal the presence of relatively long-lived kinetic intermediates and could lead to better understanding of the mechanisms of REase-mediated DNA cleavage.
At the single-molecule level, biochemical processes are stochastic-the waiting time for a single instance of the process to occur, τ, is variable. However, many measurements of τ can be expected to obey a probability distribution, p(τ), that is indicative of the type of process taking place. For instance, a single-step process, such as the release of a product molecule from an enzyme, will obey Poisson statistics, and p(τ) will take the form of a negative exponential distribution:
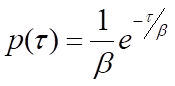
where β is the mean waiting time. Note that the rate of the process, k, will be equal to 1/β, the inverse of the mean waiting time. For processes that require more than one step, p(τ) will be the convolution of the single-exponential distributions for each of the individual steps. A general solution for the convolution of N single-exponential decay functions with identical mean waiting times, β, is the gamma probability distribution:
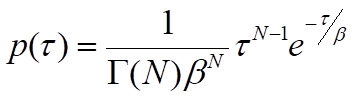
where Γ(N) is the gamma function, which describes the interpolation of the factorial of N-1 to non-integer values of N. Although this general solution can be used as an approximation when the mean waiting times of individual steps are similar, it must be understood that the presence of relatively fast steps will be masked by steps with significantly longer waiting times. In other words, the value of N represents a lower limit on the number of steps12. With an adequate number of waiting-time measurements, the parameters β and N can be estimated by binning the events and fitting the gamma distribution to the resulting histogram or by using a maximum-likelihood estimation approach. This type of analysis can therefore reveal the presence of kinetic steps that cannot be easily resolved in ensemble assays and requires a large number of observations to estimate parameters accurately12,13.
This paper describes a method to use quantum-dot-labeled DNA and total internal reflection fluorescence (TIRF) microscopy to observe hundreds of individual REase-mediated DNA cleavage events in parallel. The design of the assay makes it possible to pool the results of several experiments and can create dwell-time distributions containing thousands of events. The high photostability and brightness of quantum dots permit a 10 Hz time resolution without sacrificing the ability to observe cleavage events occurring even many minutes after the start of the experiment. Good temporal resolution and a broad dynamic range, combined with the ability to collect a large data set, allow accurate characterization of the dwell-time distributions to uncover the presence of multiple kinetic steps in the cleavage pathways of REases, which have turnover rates in the 1 min-1 range. In the case of EcoRV, three kinetic steps can be resolved, all of which have been identified through other means, confirming that the assay is sensitive to the presence of such steps.
Duplex DNA substrates containing the recognition sequence of interest are produced by annealing a biotinylated oligonucleotide to a complementary strand labeled with a single, covalently attached semiconductor nanocrystal quantum dot. These substrates are introduced into a flow channel built on top of a glass coverslip with a lawn of high molecular weight polyethylene glycol (PEG) molecules covalently attached to its surface. The DNA substrates are captured via a biotin-streptavidin-biotin linkage by a fraction of the PEG molecules that have a biotin at their free end. In TIRF microscopy, an evanescent wave that decays exponentially with distance from the glass-liquid interface provides illumination; the penetration depth is on the order of the wavelength of the light used. Under these conditions, only quantum dots that are tethered to the surface by a DNA molecule that has been captured on the functionalized glass surface will be excited. Quantum dots that are free in solution will not be constrained within the illuminated region and therefore will not luminesce. If the DNA tethering a quantum dot to the surface is cleaved, that quantum dot will be free to diffuse away from the surface, and it will disappear from the fluorescence image.
Although many Type II REases are known to bind DNA in the absence of magnesium14, all require magnesium to mediate DNA cleavage15. These REases can bind the surface-immobilized DNA in the absence of magnesium. When magnesium-containing buffer is flowed through a channel with REase prebound to the DNA, cleavage begins immediately, as indicated by the disappearance of quantum dots. The synchronization achieved by prebinding the REase molecules, and then initiating DNA cleavage by introducing magnesium, facilitates measurement of the lag time to the completion of DNA cleavage independently for each molecule in the population of enzymes observed in an experiment. Fluorescein is included as a tracer dye in the magnesium-containing buffer to indicate the arrival of magnesium into the field of view. As no enzyme is included in the magnesium-containing buffer, the lag time from the arrival of magnesium-containing buffer to the disappearance of each quantum dot indicates the time it takes for an REase that is already bound to the DNA to cleave the DNA and release the quantum dot from the glass surface. Quantum dot disappearance happens quickly and results in a sharp decrease in the intensity trajectory, providing a clear indication of the time at which a given DNA molecule is cleaved. The determination of event times is accomplished by mathematical analysis of intensity trajectories, and a typical experiment results in hundreds of identifiable cleavage events. The results of multiple experiments can be pooled to provide adequate statistics to allow estimation of the parameters, N and β, by either nonlinear least-squares or maximum-likelihood analysis.
Protocol
Representative Results
Discussion
The DNA substrate for this assay is labeled with a quantum dot using a two-step reaction scheme using sulfo-SMCC. This bifunctional crosslinker consists of an NHS ester moiety that can react with a primary amine, and a maleimide moiety that can react with a sulfhydryl group20. The thiolated oligonucleotides used to prepare the substrate are shipped in their oxidized form. It is important to reduce and purify them, as described, before proceeding with the coupling procedure, or the efficiency of th…
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This work was supported by Award Number K12GM074869 to CME from the National Institute of General Medical Sciences. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institute of General Medical Sciences or the National Institutes of Health.
Materials
| 3-aminopropyltriethoxysilane (APTS) | Sigma Aldrich | 440140-100ML | Store in dessicator box |
| 5 Minute Epoxy | Devcon | 20845 | use for sealing the microfluidic device |
| acetone | Pharmco | 329000ACS | use for cleaning coverslips |
| bath sonicator | Fisher Scientific | CPXH Model 2800 | catalog number 15-337-410 |
| Beaker, glass, 100 mL | |||
| Benchtop centrifuge | |||
| Biotin-PEG-Succinimidyl Valerate (MW 5,000) | Laysan Bio | BIO-SVA-5K | Succinimidyl valerate has a longer half-life than succinimidyl carbonate |
| biotinylated oligonucleotide | Integrated DNA Technologies | custom – see protocol for design considerations | Request 5' Biotin modification and HPLC purification |
| Bovine Serum Albumin (BSA) | VWR | 0903-5G | prepare a 10 mg/mL solution (aq) and heat to 95 °C before using |
| Centri-Spin-10 Size Exclusion Spin Columns | Princeton Separations | CS-100 or CS-101 | used to purify thiolated oligonucleotides after reducing the disulfide bond |
| Centrifuge tubes 1.5. mL | |||
| Coverslips, 1-inch square glass | |||
| coverslip holders | |||
| diamond point wheel | Dremel | 7134 | use for drilling holes in quartz flow cell topper |
| dithiothreitol (DTT) | Thermo Scientific | A39255 | No-Weigh Format, 7.7 mg/vial |
| drill press rotary tool workstation stand | Dremel | 220-01 | facilitates quartz drilling |
| EcoRV (REase used to generate example data) | New England Biolabs | R0195T or R0195M | Use 100,000 units/mL stock to avoid adding excess glycerol Check REBASE for suppliers of other REases |
| ethanol | various | CAS 64-17-5 | denatured or 95% are acceptable, use for cleaning coverslips |
| Ethylenediaminetetraacetic acid (EDTA) | Sigma Aldrich | EDS | BioUltra, anhydrous, store in dessicator box |
| Flea Micro Spinbar | Fisherbrand | 14-513-65 | 3 mm x 10 mm size to fit beneath coverslip rack |
| fluorescein | Acros Organics | 17324 | use to make experimental buffers |
| gravity convection oven | Binder | 9010-0131 | |
| handheld rotary multitool | Dremel | 8220 | use for drilling holes in quartz flow cell topper |
| ImagEM X2 EM-CCD Camera | Hamamatsu | C9100-23B | air cooling is adequate for this experiment, use HCImage software or similar to control |
| Imaging spacer, double-sided, adhesive | |||
| Jar, glass with screw cap, (approximately 50 mm diameter by 50 mm high) | |||
| magnesium chloride hexahydrate | Fisher Bioreagents | BP214-500 | use to make experimental buffer with magnesium |
| MATLAB software | Data analysis | ||
| metal tweezers | Fisher Brand | 16-100-110 | |
| methoxy-PEG-Succinimidyl Valerate (MW 5,000) | Laysan Bio | M-SVA-5K | Both PEGs should have the same NHS ester so that the rate of reaction is consistent |
| microcentrifuge | Eppendorf | 5424 | |
| multiposition magnetic stirrer | VWR | 12621-022 | |
| N-cyclohexyl-2-aminoethanesulfonic acid (CHES) | Acros Organics | AC20818 | CAS 103-47-9, use to make CHES buffer |
| orbital shaker and heater for microcentrifuge tubes | Q Instruments | 1808-0506 | with 1808-1061 adaptor for 24 x 2.0 mL or 15 x 0.5 mL tubes |
| Parafilm | |||
| PE60 Polyethylene tubing (inner diameter 0.76 mm, outer diameter 1.22 mm) | Intramedic | 6258917 | 22 G blunt needles are a good fit for this tubing size |
| Phosphate-Buffered Saline (PBS) 10x | Sigma Aldrich | P7059 | Use at 1x strength |
| potassium hydroxide | VWR Chemicals BDH | BDH9262 | use a 1 M solution to clean coverslips |
| Qdot 655 ITK Amino (PEG) Quantum Dots | Invitrogen | Q21521MP | |
| Quartz Slide, 1 inch square, 1 mm thick | Electron Microscopy Sciences | 72250-10 | holes must be drilled in the corners for inlet and outlet tubing insertion |
| reinforced plastic tweezers | Rubis | K35a | use for handling coverslips and building microfluidic device |
| SecureSeal Adhesive Sheets | Grace Biolabs | SA-S-1L | cut to form spacer for microfluidic device |
| Single channel syringe pump for microfluidics | New Era Pump Systems | NE-1002X-US | fitted with a 50 mL syringe and a 22 G blunt needle |
| Slide-a-Lyzer MINI Dialysis Devices, 10 kDa MWCO, 0.1 mL | Thermo Scientific | 69570 or 69572 | used for buffer exchange during quantum dot coupling to DNA |
| sodium bicarbonate | EMD Millipore | SX0320 | use to make buffer for surface functionalization; 100 mM, pH 8 |
| sodium chloride | Macron | 7581-12 | use to make experimental buffers |
| Sodium phosphate dibasic solution (BioUltra, 0.5 M in water) | Sigma Aldrich | 94046 | use to make 100 mM sodium phosphate buffer |
| Sodium phosphate monobasic solution (BioUltra, 5M in water) | Sigma Aldrich | 74092 | use to adjust pH of 100 mM sodium phosphate buffer |
| Streptavidin from Streptomyces avidinii | Sigma Aldrich | S4762 | dissolve at 1 mg/mL and store 25 mL aliqouts at -20 ℃ |
| Sulfosuccinimidyl-4-(N-maleimidomethyl) cyclohexane-1-carboxylate (sulfo-SMCC) | Thermo Scientific | A39268 | No-Weigh Format, 2 mg/vial |
| Syringe fitted with blunt 21 G needle | |||
| Syringe pump | |||
| thiolated oligonucleotide | Integrated DNA Technologies | custom – see protocol for design considerations | Request 5' Thiol Modifier C6 S-S and HPLC purificaiton |
| TIRF imaging system with 488 nm laser illumination | various | custom built | |
| Tris -HCl | Research Products International | T60050 | use to make experimental buffers |
| Tris base | JT Baker | 4101 | use to make experimental buffers |
| Tween-20 | Sigma | P7949 | use to make blocking buffer |
| Ultrapure water | |||
| vortex mixer | VWR | 10153-842 | |
| Wash-N-Dry Coverslip Rack | Electron Microscopy Sciences | 70366-16 | used for surface functionalization of coverslips |
Riferimenti
- Roberts, R. J. How restriction enzymes became the workhorses of molecular biology. Proceedings of the National Academy of Sciences of the United States of America. 102 (17), 5905-5908 (2005).
- Loenen, W. A., Dryden, D. T., Raleigh, E. A., Wilson, G. G., Murray, N. E. Highlights of the DNA cutters: a short history of the restriction enzymes. Nucleic Acids Research. 42 (1), 3-19 (2014).
- Roberts, R. J., Vincze, T., Posfai, J., Macelis, D. REBASE–a database for DNA restriction and modification: enzymes, genes and genomes. Nucleic Acids Research. 43, 298-299 (2015).
- Bonnet, I., et al. Sliding and jumping of single EcoRV restriction enzymes on non-cognate DNA. Nucleic Acids Research. 36 (12), 4118-4127 (2008).
- Biebricher, A., Wende, W., Escude, C., Pingoud, A., Desbiolles, P. Tracking of single quantum dot labeled EcoRV sliding along DNA manipulated by double optical tweezers. Biophysical Journal. 96 (8), 50-52 (2009).
- Blainey, P. C., et al. Nonspecifically bound proteins spin while diffusing along DNA. Nature Structural & Molecular Biology. 16 (12), 1224-1229 (2009).
- Huang, C. -. F., et al. Direct visualization of DNA recognition by restriction endonuclease EcoRI. Journal of Experimental & Clinical Medicine. 5 (1), 25-29 (2013).
- Palma, M., et al. Selective biomolecular nanoarrays for parallel single-molecule investigations. Journal of American Chemical Society. 133 (20), 7656-7659 (2011).
- Reinhard, B. M., Sheikholeslami, S., Mastroianni, A., Alivisatos, A. P., Liphardt, J. Use of plasmon coupling to reveal the dynamics of DNA bending and cleavage by single EcoRV restriction enzymes. Proceedings of the National Academy of Sciences of the United States of America. 104 (8), 2667-2672 (2007).
- van den Broek, B., Noom, M. C., Wuite, G. J. DNA-tension dependence of restriction enzyme activity reveals mechanochemical properties of the reaction pathway. Nucleic Acids Research. 33 (8), 2676-2684 (2005).
- Gambino, S., et al. A single molecule assay for measuring site-specific DNA cleavage. Analytical Biochemistry. 495, 3-5 (2016).
- Floyd, D. L., Harrison, S. C., van Oijen, A. M. Analysis of kinetic intermediates in single-particle dwell-time distributions. Biophysical Journal. 99 (2), 360-366 (2010).
- Loparo, J. J., van Oijen, A., Hinterdorfer, P., Oijen, A. Single-molecule enzymology. Handbook of Single-Molecule Biophysics. , 165-182 (2009).
- Taylor, J. D., Badcoe, I. G., Clarke, A. R., Halford, S. E. EcoRV restriction endonuclease binds all DNA sequences with equal affinity. Biochimica. 30 (36), 8743-8753 (1991).
- Pingoud, A., Jeltsch, A. Structure and function of type II restriction endonucleases. Nucleic Acids Research. 29 (18), 3705-3727 (2001).
- Tanner, N. A., Loparo, J. J., van Oijen, A. M. Visualizing single-molecule DNA replication with fluorescence microscopy. Journal of Visualized Experiments: JoVE. (32), e1529 (2009).
- Kulczyk, A. W., Tanner, N. A., Loparo, J. J., Richardson, C. C., van Oijen, A. M. Direct observation of enzymes replicating DNA using a single-molecule DNA stretching assay. Journal of Visualized Experiments: JoVE. (37), e1689 (2010).
- Baldwin, G. S., Vipond, I. B., Halford, S. E. Rapid reaction analysis of the catalytic cycle of the EcoRV restriction endonuclease. Biochimica. 34 (2), 705-714 (1995).
- Winkler, F. K., et al. The crystal-structure of Ecorv endonuclease and of Its complexes with cognate and non-cognate DNA fragments. EMBO Journal. 12 (5), 1781-1795 (1993).
- Sioss, J. A., Stoermer, R. L., Sha, M. Y., Keating, C. D. Silica-coated, Au/Ag striped nanowires for bioanalysis. Langmuir. 23 (22), 11334-11341 (2007).

