Rapid, Affordable, and Uncomplicated Production of Bacterial Cell-free Lysate
Summary
This protocol describes a rapid and simple method to produce bacterial lysate for cell-free gene expression, using an engineered strain of Escherichia coli and requiring only standard laboratory equipment.
Abstract
Cell-free gene expression offers the power of biology without the complications of a living organism. Although many such gene expression systems exist, most are quite expensive to buy and/or require special equipment and finely honed expertise to produce effectively. This protocol describes a method to produce bacterial cell-free lysate that supports high levels of gene expression, using only standard laboratory equipment and requiring minimal processing. The method uses an Escherichia coli strain producing an endolysin that does not affect growth, but which efficiently lyses a harvested cell pellet following a simple freeze-thaw cycle. The only further processing required is a brief incubation followed by centrifugation to clear the autolysate of cellular debris. Dynamic gene circuits can be achieved through heterologous expression of the ClpX protease in the cells before harvesting. An E. coli strain lacking the lacZ gene can be used for high-sensitivity, cell-free biosensing applications using a colorimetric or fluorescent readout. The entire protocol requires as few as 8-9 hours, with only 1-2 hours of hands-on labor from inoculation to completion. By reducing the cost and time to obtain cell-free lysate, this method should increase the affordability of cell-free gene expression for various applications.
Introduction
Gene expression in cell-free lysates has several advantages over using live cells1,2,3,4. Lysates can be easily modified biochemically and used in conditions that could be detrimental to or impossible to achieve in live cells. Gene expression circuits do not have to contend or compete with host biological processes, and testing new genetic circuits is as simple as adding DNA. For these reasons, cell-free gene expression has found various applications, from biosensors5,6 to rapidly prototyping synthetic gene circuits7,8 to developing artificial cells9. Most cell-free gene expression utilizes cellular lysates that have been highly processed, generally requiring long and complex protocols, specialized equipment, and/or sensitive steps that can lead to significant variation between users and batches10,11.
This paper describes a simple, efficient method for producing cell-free lysate that requires minimal processing and expertise (Figure 1A)12. The method relies on E. coli cells that are engineered to lyse following a simple freeze-thaw cycle. The cells express an endolysin from phage lambda that degrades the cell wall. As the cells are growing, this endolysin remains in the cytoplasm, sequestered from the cell wall. However, a simple freeze-thaw cycle disrupts the cytoplasmic membrane, releasing the endolysin into the periplasm, where it degrades the cell wall, resulting in rapid cell lysis. The protocol can be completed with only a few hours of hands-on work and requires only a freezer, a centrifuge capable of 30,000 × g (for optimal results; lower speeds can be used with more care not to disturb the pellet), a vortex mixer, and a simple buffer solution. Functional lysate can even be produced by freeze-drying the cells and rehydrating them in situ. However, this method produces lysates with lower activity, presumably due to the remaining cell debris.
The lysates are highly active for cell-free gene expression, and they can be enhanced in various ways depending on the end use. The rate of protein synthesis can be further increased by concentrating the lysate using standard spin concentrators. Linear DNA can be protected from degradation by adding purified GamS protein. Protein degradation, necessary for more complex circuit dynamics such as oscillation, can be achieved by co-expressing a ClpX hexamer in the autolysate-producing strain13. Finally, LacZ-based visual readouts are enabled by using an autolysate strain lacking lacZ. Overall, this method produces highly active cell-free lysate that is suitable for a wide range of applications.
Protocol
1. Prepare media and buffers.
- Prepare 2xYTPG medium.
- Mix 62 g 2xYT powder, 5.99 g potassium phosphate monobasic, 13.93 g potassium phosphate dibasic, and deionized water to 2 L.
- Autoclave on liquid cycle with an exposure time of 30 min14.
- To 400 mL of 2xYTP media from 1.1.2, add 7.2 g D-glucose (dextrose) and mix until dissolved.
- Filter-sterilize through a 0.2 µm filter.
- Prepare S30A buffer.
- Mix Tris-HCl (pH 7.7, 50 mM final concentration), potassium glutamate (60 mM final), and magnesium glutamate (14 mM final).
- Adjust the pH to 7.7 using 10 M KOH.
- Prepare Solution 1 (see Table 1).
- Resuspend 4-(2-hydroxyethyl)-1- piperazineethanesulfonic acid (HEPES) in 2 mL of water.
- Adjust the pH to 8.0 using KOH.
- Add all other components from Table 1.
- Adjust the pH to 7.6 using 10 M KOH. Filter-sterilize.
- Prepare 2.5x premix solution (see Table 2).
- Mix all the components in Table 2.
- Adjust the pH to 7.5 using KOH.
- Aliquot and freeze at -80 °C.
NOTE: A 20 µL reaction uses 8.9 µL of premix.
2. Prepare cells.
- Streak the autolysate cells onto LB agar plates containing 50 µg/mL ampicillin using an inoculating loop and grow at 37 °C (see Note 1).
- Pick a single colony into a starter culture of LB/ampicillin medium using a pipet tip and grow at 37 °C overnight.
- Inoculate 400 mL of 2xYTPG medium containing 50 µg/mL ampicillin with 400 µL of starter culture, and grow at 37 °C in a 1 L Erlenmeyer flask, shaking at 300 rpm.
- Periodically measure the culture's optical density at 600 nm (OD600) using a spectrophotometer to read an optical cuvette with a 1 cm path length. When the OD600 exceeds 1, begin diluting the culture 5-fold before measurements to ensure that the measurements remain within the linear range of a typical laboratory spectrophotometer. Continue growing the cells until the 5-fold diluted culture reaches OD600 of 0.3 (corresponding to a culture OD600 of 1.5).
3. Prepare the lysate.
- Prepare S30A buffer supplemented with 2 mM dithiothreitol (DTT). Mix 3 mL of S30A buffer with 6 µL of DTT stock solution at 1 M. Place on ice for use in step 3.7.
- Harvest the cells by centrifuging at 1800 × g for 15 min at room temperature.
- Discard the supernatant by pouring it off and using a pipet to remove any remaining liquid.
- Resuspend the pellet in 45 mL of cold (4-10 °C) S30A buffer using a vortex mixer.
- Weigh an empty 50 mL centrifuge tube, transfer the cells into it, and repeat steps 3.2-3.3 to wash the cells.
- Weigh the pellet, subtracting the weight of an empty 50 mL tube. Make sure to carefully aspirate any remaining supernatant to ensure an accurate measurement of pellet weight.
NOTE: A typical yield is ~1.3 g of cell pellet from 400 mL of production culture. - Add 2 volumes of cold S30A buffer supplemented with 2 mM dithiothreitol, i.e., 2 mL of buffer for every 1 g of cell pellet, and resuspend the cells by vigorously vortex mixing.
- Freeze the cells. Place the 50 mL tube containing the cells in a -20 °C or -80 °C freezer until the pellet is thoroughly frozen.
NOTE: The freezing step is a good stopping point for the day. - Thaw the cells in a room temperature water bath.
- Vortex vigorously for 2-3 min.
- Incubate at 37 °C for 45 min with shaking at 300 rpm.
- Clear the sample of heavy cellular debris by centrifuging in transparent centrifuge tubes at 30,000 × g for 45 min at 4 °C.
NOTE: If a centrifuge capable of 30,000 x g is not available, centrifuge for 45 min at 21,000 × g, and use additional caution in step 3.13, as the pellet will be less compact. - Carefully transfer the supernatant to a new tube with a pipet, avoiding disturbing the pellet as much as possible. If the transferred supernatant is contaminated with material from the pellet, repeat the previous step.
- Transfer the supernatant to 1.5 mL centrifuge tubes and centrifuge once more at 21,000 × g (or the maximum speed of a tabletop centrifuge) for 5 min.
- Aliquot the cleared autolysate into the desired volumes, carefully avoiding any remaining pellet, and freeze at -80 °C or use immediately.
NOTE: A single 20 µL reaction uses 8 µL of autolysate.
4. Cell-free gene expression
NOTE: The autolysate is now ready for any desired end-use. The following is an example standard protocol for cell-free gene expression.
- For a 20 µL reaction, mix on ice 8 µL of autolysate and 8.9 µL of premix. See NOTE at the end of the protocol section regarding the optimization of magnesium glutamate and PEG 8000 concentrations.
- Add DNA (e.g., pBEST-OR2-OR1-Pr-UTR1-deGFP-T500 to a final concentration of 8 nM), any other reagents, and water to 20 µL.
- Place the reaction in a 384-well microplate and measure the fluorescence time course and/or endpoints using a plate reader. For green fluorescent protein (GFP), use an excitation wavelength of 485 nm and an emission wavelength of 520 nm.
5. Protocol modifications
NOTE: The following modifications of the protocol enable it to serve other applications.
- Cell-free gene expression using linear DNA templates
- Perform the steps in section 4, supplementing the reaction with 2.2 µM purified GamS protein (expressed and purified as described12) before the addition of the linear DNA.
- Cell-free gene expression incorporating protein degradation
- In step 2.1, use autolysate cells containing the plasmid pACYC-FLAG-dN6-His (see the Table of Materials). In all growth media, additionally include 34 µg/mL chloramphenicol.
- In step 2.3, include 40 µM isopropyl β-D-1-thiogalactopyranoside (IPTG) in the growth medium to induce expression from the plasmid.
- Repeat steps 3.2-3.4 (washing) two additional times (for a total of three washes) to ensure the complete removal of chloramphenicol, which is a translation inhibitor. For the first two washes (step 3.4), substitute S30A buffer with phosphate-buffered saline (pH 7.4).
- In step 4.2, supplement with an additional 3 mM ATP (added from a stock solution of 100 mM ATP in water, pH 7.2) and 4.5 mM magnesium glutamate (using a 1 M stock solution in water) (final concentrations) to compensate for high ATP use by ClpXP, as well as chelation of magnesium by the additional ATP.
- Cell-free gene expression using LacZ-based readouts (including colorimetric)
- In step 2.1, use autolysate cells that do not natively express LacZ (see the Table of Materials).
- Alternatively, prepare the lysate directly from freeze-dried cells.
- Perform all steps from 1.1 to 3.7.
- Mix 8 µL of cell suspension with 8.9 µL of premix.
- Add plasmid DNA (if desired), other custom reagents, and water to reach a final volume of 20 µL.
- Transfer the reaction to a 384-well microplate and freeze-dry it.
NOTE: Freeze-dried samples can be stored for up to a week and possibly longer. - To begin the reaction, rehydrate it with 18 µL of deionized water supplemented with any desired DNA or other reagents.
- Follow the fluorescence dynamics in a plate reader.
NOTE 1: Autolysate cells are designed to lyse upon a freeze-thaw cycle, so it is particularly important to use cryoprotectant when making frozen stocks. We froze stocks in 24% wt/vol glycerol and stored them at -80 °C.
NOTE 2: Magnesium ions and PEG 8000 are critical for lysate performance. The base 2.5x premix, based on previously published data, contains 6 mM magnesium glutamate and 4.8% wt/vol PEG 8000, which become 2.4 mM and 1.9%, respectively, in the final reaction. The autolysate prepared with the protocol here typically performs best with an additional 5 mM Mg-glutamate and 1.5% PEG 8000 in the final reaction. However, this can be optimized in the range of an additional 0-10 mM Mg glutamate and an additional 0-3% PEG 8000 (compared to the base premix). To prepare the premix with the recommended additional 5 mM Mg-glutamate and 1.5% PEG 8000 (final concentrations), mix 380 µL of premix with 4.75 µL of magnesium glutamate at 1 M and 36.1 µL of PEG 8000 at 40% weight/volume.
Representative Results
Representative results can be observed by using autolysate to express GFP from a constitutively expressing plasmid, here pBEST-OR2-OR1-Pr-UTR1-deGFP-T500, and recording a time course of GFP fluorescence in a plate reader (Figure 1B). A dilution series of plasmid DNA found strong expression even at 1 nM DNA. Compared to a commercially available lysate, the autolysate can produce a greater total yield and achieved a greater maximum production rate, calculated as the time derivative of the GFP time course (Figure 1C,D). For additional results using this method, see Didovyk et al.12. This protocol has relatively few failure points; however, suboptimal results could be obtained if the autolysate is not sufficiently cleared of cell debris. Optimal results can also require optimizing the concentrations of PEG-8000 and Mg2+ for each batch of lysate (see Note 2).
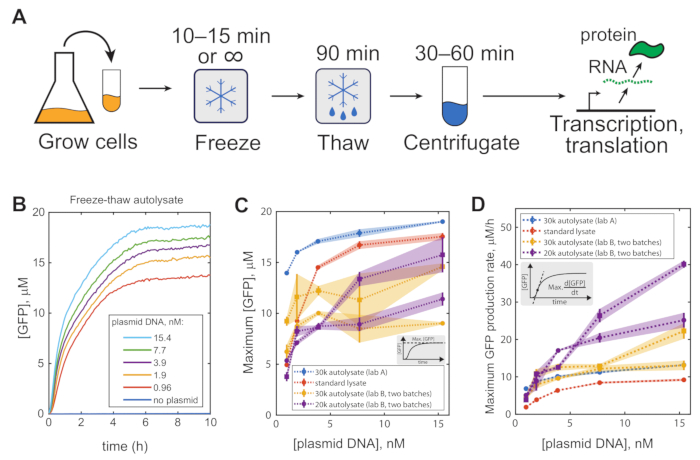
Figure 1: Representative results. (A) Visual representation of the protocol. (B) Example time course of GFP expression from pBEST-OR2-OR1-Pr-UTR1-deGFP-T500 in a freeze-thaw autolysate. Maximum GFP production (C) and maximum production rate (D) of autolysate prepared in 2 different labs by 2 different researchers, using a 30,000 x g centrifuge (blue and orange) or a 20,000 x g centrifuge (purple), as compared to a commercial reference lysate (orange, MYtxtl-70-960M from MYcroarray). All error bars are mean absolute error of two technical replicates. This figure has been modified from 12. Copyright 2017 American Chemical Society. Abbreviation: GFP = green fluorescent protein. Please click here to view a larger version of this figure.
| Solution 1: Add water to 4 mL total | |
| Name | Weight (mg) |
| 3-PGA | 386.4 |
| ATP | 52 |
| cAMP | 13.8 |
| CoA | 11.2 |
| CTP | 28.4 |
| Folinic acid | 1.9 |
| GTP | 47.6 |
| HEPES | 667.2 |
| NAD | 12.3 |
| Spermidine | 8.1 |
| tRNAs | 11.2 |
| UTP | 29.5 |
Table 1: Solution 1. Components of solution 1.
| 2.5x Premix | |
| Reagent | Volume (μL) |
| Amino acid mix containing 24 mM each, except for leucine which is at 20 mM | 2500 |
| dithiothreitol (DTT) at 100 mM | 100 |
| Magnesium glutamate at 1 M | 25 |
| PEG-8000 at 40% wt/vol | 500 |
| Potassium glutamate at 2 M | 303 |
| Solution 1 (see Table 1) | 714.3 |
Table 2: Premix. Components of the premix solution.
Discussion
The protocol described here yields highly active bacterial lysate for cell-free gene expression. The key is to use cells carrying the plasmid pAD-LyseR, which expresses the lambda phage endolysin cytosolically. These cells are potentiated to lyse themselves upon permeabilization of the inner membrane, allowing the endolysin access to the cell wall, which the method achieves through a simple freeze-thaw cycle. Because the cells effectively lyse themselves, the product is referred to as autolysate. After the cells have lysed, the only remaining steps are incubation and centrifugation to clear the autolysate of cellular debris.
Compared to other methods for producing bacterial lysate, this approach is notably simple and rapid, yet it does not sacrifice the quality of the lysate. The protocol can be completed in 8-9 h after inoculating the production culture, with only 1-2 hours of hands-on labor. The only recommended piece of equipment that is not entirely standard for molecular biology laboratories is a centrifuge capable of achieving 30,000 × g. However, autolysate of comparable quality can be produced even with a lower-speed centrifuge (Figure 1C, D); the user would just have to be more careful removing the lysate from the pellet, perhaps leaving behind slightly more liquid to ensure clean samples. This simplicity is not merely a matter of convenience; less complicated protocols tend to yield more reproducible results, with less variation when performed with different hands. The modification presented in step 5.4, in which cells are freeze-dried along with all other reagents, presents an even simpler protocol, although with reduced protein production yields. Notably, in this modification, the centrifugation steps to clear the lysate of cellular debris are skipped, which further reduces processing labor; however, the remaining debris reduces expression from the lysate12.
In recent years, many approaches to producing cell-free lysate have been published, summarized recently by Cole et al.15. These studies have explored various strategies for cell type, growth conditions, lysis methodologies, and post-processing. Most other methods for lysis have required specialized equipment such as a French press, homogenizer, bead beater, or sonicator. Fujiwara and Doi omitted this equipment in favor of a freeze-thaw cycle similar to the one described here, except that they rendered the cells susceptible to lysis by treating them with lysozyme rather than expressing an endolysin16. Although this is roughly as simple a protocol as the one described here, the lysozyme-treated cells must be washed while in their fragile state without disrupting them prematurely, which could require experimental finesse and introduce a source of variability.
In addition to lysate, cell-free gene expression requires a premix solution containing energy sources, RNA and protein monomers, and other small molecules. The premix recipe was described and optimized previously17, with a few modifications. The premix used here contained approximately 4 times higher concentrations of amino acids, as well as additional magnesium glutamate and PEG 8000 corresponding to final reaction concentrations of 7.5 mM and 3.5% weight/volume, respectively. Optimal results may require adjusting the supplemental magnesium glutamate and PEG 8000 concentrations for each new batch of lysate, although the above concentrations consistently produced good results (see Note 2). Unique applications may require reoptimizing these concentrations, for example, when using ClpX-supplemented lysate13.
A standard E. coli strain for producing cell-free lysate is BL21-Gold (DE3). A derivative of these cells containing the autolysis plasmid pAD-LyseR has been deposited in a strain and plasmid repository (see the Table of Materials). Also available are a derivative lacking genomic lacZ to dramatically reduce the background for circuits that use LacZ-based output and an expression plasmid pAD-GamS to be used for the purification of the GamS protein that can protect linear DNA from degradation. These cell strains and plasmids should be useful for a variety of applications in cell-free gene expression.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
The authors thank Zachary Sun and Richard Murray (California Institute of Technology) for kindly providing the plasmid P_araBAD-gamS, and Kaeko Kamei (Kyoto Institute of Technology) for kindly providing a high-speed cooling centrifuge. This work was supported by grants from the National Institutes of Health and from the ARO MURI program and was partly supported by the Leading Initiative for Excellent Young Researchers, MEXT, Japan.
Materials
| 2xYT media | EMD Millipore | 4.85008 | or equivalent |
| 3-PGA | Sigma Aldrich | P8877 | or equivalent |
| Amicon Ultra-15 centrifugal filter unit, 3 kDa cutoff | Millipore Sigma | UFC900308 | optional, can be used to concentrate lysate, select concentrator capacity appropriate for the volume to be concentrated |
| ampicillin | Sigma Aldrich | A0166-5G | or carbenicillin, a more stable variant |
| ATP | Sigma Aldrich | A8937 | or equivalent |
| cAMP | Sigma Aldrich | A9501 | or equivalent |
| CoA | Sigma Aldrich | C4282 | or equivalent |
| CTP | United States Biosciences | 14121 | or equivalent |
| D-glucose (dextrose) | Fisher Scientific | AAA1749603 | or equivalent |
| dithiothreitol (DTT) | Sigma Aldrich | D0632-1G | or equivalent |
| E. coli BL21-Gold (DE3) carrying pAD-LyseR | Addgene | 99244 | |
| E. coli BL21-Gold (DE3) ΔlacZ carrying pAD-LyseR | Addgene | 99245 | |
| Folinic acid | Sigma Aldrich | F7878 | or equivalent |
| GTP | United States Biosciences | 16800 | or equivalent |
| HEPES | Sigma Aldrich | H3375-25G | or equivalent |
| LB media | Fisher Scientific | DF0446075 | or equivalent |
| magnesium glutamate | Sigma Aldrich | 49605-250G | or equivalent |
| NAD | Sigma Aldrich | N6522 | or equivalent |
| potassium glutamate | Sigma Aldrich | G1501-100G | or equivalent |
| potassium hydroxide (KOH) | Sigma Aldrich | 221473-25G | for adjusting pH |
| potassium phosphate dibasic | Fisher Scientific | BP363-500 | or equivalent |
| potassium phosphate monobasic | Fisher Scientific | BP362-500 | or equivalent |
| Spermidine | Sigma Aldrich | 85558 | or equivalent |
| Tris-HCl | Fisher Scientific | 9310500GM | or equivalent |
| tRNA mix | Roche | 10109541001 | or equivalent |
| UTP | United States Biosciences | 23160 | or equivalent |
Riferimenti
- Dudley, Q. M., Karim, A. S., Jewett, M. C. Cell-free metabolic engineering: biomanufacturing beyond the cell. Biotechnology Journal. 10 (1), 69-82 (2015).
- Smith, M. T., Wilding, K. M., Hunt, J. M., Bennett, A. M., Bundy, B. C. The emerging age of cell-free synthetic biology. FEBS Letters. 588 (17), 2755-2761 (2014).
- Casteleijn, M. G., Urtti, A., Sarkhel, S. Expression without boundaries: cell-free protein synthesis in pharmaceutical research. International Journal of Pharmaceutics. 440 (1), 39-47 (2013).
- He, M. Cell-free protein synthesis: applications in proteomics and biotechnology. New Biotechnology. 25 (2-3), 126-132 (2008).
- Pardee, K., et al. Paper-based synthetic gene networks. Cell. 159 (4), 940-954 (2014).
- Pardee, K., et al. low-cost detection of Zika virus using programmable biomolecular components. Cell. 165 (5), 1255-1266 (2016).
- Takahashi, M. K., et al. Characterizing and prototyping genetic networks with cell-free transcription-translation reactions. Methods. 86, 60-72 (2015).
- Siegal-Gaskins, D., Tuza, Z. A., Kim, J., Noireaux, V., Murray, R. M. Gene circuit performance characterization and resource usage in a cell-free “breadboard”. ACS Synthetic Biology. 3 (6), 416-425 (2014).
- Niederholtmeyer, H., Chaggan, C., Devaraj, N. K. Communication and quorum sensing in non-living mimics of eukaryotic cells. Nature Communications. 9, 5027 (2018).
- Sun, Z. Z., et al. Protocols for implementing an Escherichia coli based TX-TL cell-free expression system for synthetic biology. Journal of Visualized Experiments: JoVE. (79), e50762 (2013).
- Dopp, J., Jo, Y., Reuel, N. Methods to reduce variability in E. coli-based cell-free protein expression systems. Synthetic and Systems Biotechnology. 4 (4), 204-211 (2019).
- Didovyk, A., Tonooka, T., Tsimring, L., Hasty, J. Rapid and scalable preparation of bacterial lysates for cell-free gene expression. ACS Synthetic Biology. 6 (12), 2198-2208 (2017).
- Tonooka, T., Niederholtmeyer, H., Tsimring, L., Hasty, J. Artificial cell on a chip integrated with protein degradation. 2019 IEEE 32nd International Conference on Micro Electro Mechanical Systems (MEMS). , 107-109 (2019).
- Garibaldi, B., et al. Validation of autoclave protocols for successful decontamination of category A medical waste generated from care of patients with serious communicable diseases. Journal of Clinical Microbiology. 55 (2), 545-551 (2017).
- Cole, S., Miklos, A., Chiao, A., Sun, Z., Lux, M. Methodologies for preparation of prokaryotic extracts for cell-free expression systems. Synthetic and Systems Biotechnology. 5 (4), 252-267 (2020).
- Fujiwara, K., Doi, N. Biochemical preparation of cell extract for cell-free protein synthesis without physical disruption. PLoS ONE. 11 (4), 0154614 (2016).
- Shin, J., Noireaux, V. Efficient cell-free expression with the endogenous E. coli RNA polymerase and sigma factor 70. Journal of Biological Engineering. 4, 8 (2010).

