Isolation and Characterization of the Immune Cells from Micro-dissected Mouse Choroid Plexuses
Summary
This study uses flow cytometry and two different gating strategies on isolated perfused mice brain choroid plexuses; this protocol identifies the main immune cell subsets that populate this brain structure.
Abstract
The brain is no longer considered as an organ functioning in isolation; accumulating evidence suggests that changes in the peripheral immune system can indirectly shape brain function. At the interface between the brain and the systemic circulation, the choroid plexuses (CP), which constitute the blood-cerebrospinal fluid barrier, have been highlighted as a key site of periphery-to-brain communication. CP produce the cerebrospinal fluid, neurotrophic factors, and signaling molecules that can shape brain homeostasis. CP are also an active immunological niche. In contrast to the brain parenchyma, which is populated mainly by microglia under physiological conditions, the heterogeneity of CP immune cells recapitulates the diversity found in other peripheral organs. The CP immune cell diversity and activity change with aging, stress, and disease and modulate the activity of the CP epithelium, thereby indirectly shaping brain function. The goal of this protocol is to isolate murine CP and identify about 90% of the main immune subsets that populate them. This method is a tool to characterize CP immune cells and understand their function in orchestrating periphery-to-brain communication. The proposed protocol may help decipher how CP immune cells indirectly modulate brain function in health and across various disease conditions.
Introduction
Since the discovery of the blood-brain barrier by Paul Erhlich in the late 19th century, the brain has been considered virtually separated from the other organs and the bloodstream. Yet, this last decade has seen the emergence of the concept that brain function is shaped by various biological factors, such as gut microbiota and systemic immune cells and signals1,2,3,4. In parallel, other brain borders such as meninges and choroid plexuses (CP) have been identified as interfaces of active immune-brain cross talk rather than inert barrier tissues5,6,7,8.
The CP constitute the blood-cerebrospinal fluid barrier, one of the borders separating the brain and the periphery. They are located in each of the four ventricles of the brain, i.e., the third, the fourth, and both lateral ventricles, and are adjacent to areas involved in neurogenesis such as the subventricular zone and subgranular zone of the hippocampus3. Structurally, the CP are composed of a network of fenestrated blood capillaries enclosed by a monolayer of epithelial cells, which are interconnected by tight and adherens junctions9,10. Major physiological roles of the CP epithelium involve the production of cerebrospinal fluid, which flushes the brain from waste metabolites and protein aggregates, and the production and controlled blood-to-brain passage of various signaling molecules including hormones and neurotrophic factors11,12,13. Secreted molecules from CP shape brain's activity, i.e., by modulating neurogenesis and microglial function14,15,16,17,18,19, which makes CP crucial for brain homeostasis. CP also engage in various immune activities; whereas the main immune cell type in the brain parenchyma under non-pathological conditions is microglia, the diversity of CP immune cell populations is as broad as in peripheral organs3,7, suggesting that various channels of immune regulation and signaling are at work at the CP.
The space between the endothelial and epithelial cells, the CP stroma, is mainly populated by border-associated macrophages (BAM), which express pro-inflammatory cytokines and molecules related to antigen presentation in response to inflammatory signals3. Another subtype of macrophages, Kolmer's epiplexus cells, are present on the apical surface of the CP epithelium20. CP stroma is also a niche for dendritic cells, B cells, mast cells, basophils, neutrophils, innate lymphoid cells, and T cells which are mostly effector memory T cells able to recognize central nervous system antigens7,21,22,23,24. In addition, the composition and activity of immune cell populations at the CP changes upon systemic or brain perturbation, for example, during aging10,14,15,21,25, microbiota perturbation7, stress26, and disease27,28. Notably, these changes were suggested to indirectly shape brain function, i.e., a shift of CP CD4+ T cells towards Th2 inflammation occurs in brain aging and triggers immune signaling from the CP that may shape aging-associated cognitive decline14,15,21,25,29. Illuminating the properties of the CP immune cells would thus be crucial to better understand their regulatory function on CP epithelium physiology and secretion and thereby decipher their indirect impact on brain function under healthy and disease conditions.
CP are small structures that contain only a few immune cells. Their isolation requires microdissection after a preliminary step of perfusion; immune cells in the bloodstream would otherwise constitute major contaminants. This protocol aims to characterize the myeloid and T cell subsets of the CP using flow cytometry. This method identifies about 90% of the immune cell populations that compose mouse CP under non-inflammatory conditions, in accordance with recently published works using other methods to dissect immune CP heterogeneity7,10,28. This protocol could be applied to characterize changes in the CP immune cell compartment with disease and other experimental paradigms in vivo.
Protocol
All the procedures agreed with the guidelines of the European Commission for the handling of laboratory animals, directive 86/609/EEC. They were approved by the ethical committees No. 59, by the CETEA/CEEA No. 089, under the number dap210067 and APAFIS #32382-2021070917055505 v1.
1. Preparation of the materials
- Store all antibodies (Table of Materials) at 4 °C, protected from light exposure.
- DAPI stock solution (1 mg/mL): Resuspend the powder in PBS-/- (Table of Materials), aliquot, and store at -20 °C.
- DAPI working solution (0.1 mg/mL): Dilute DAPI stock solution with PBS-/- at a ratio of 1:10.
- Magnetic-activated cell sorting (MACS) buffer: Prepare 2 mM ethylene diamine tetraacetic acid (EDTA) and 0.5% bovine serum albumin (BSA) (Table of Materials) in PBS-/-.
- Collagenase IV stock solution (20 U/μL): Resuspend the powder in PBS+/+ (Table of Materials), aliquot, and store at -20 °C.
NOTE: Collagenase IV requires MgCl2 to be fully active; do not freeze-thaw Collagenase IV solution. - Anesthetic-analgesic solution: Mix 150 μL of ketamine (150 mg/kg), 25 μl of xylazine (5 mg/kg), and 330 μl of buprecare (0.1 mg/kg) in 1 mL of PBS-/-.
NOTE: Prepare anesthetic-analgesic solution extemporaneously and do not keep it longer than a day. - Heparin solution (100 U/mL): Resuspend the powder in PBS-/-.
- Prepare the infusion inset system connecting a tube to a decanter Erlenmeyer filled with PBS-/-. Connect a 23 G needle to the extremity of the infusion tube. Open the infusion inset and let the PBS run until there are no bubbles in the tube.
- Prepare the binocular loupe equipped with a light.
2. Housing of C57BL/6 mice
- For the analysis of CP myeloid or T cells, use four mice for each and pool the four CP (two from each lateral ventricle, the third ventricle and the fourth ventricle CP; for eight mice in total). Not pooling CP carries the risk of failing to detect the rare immune populations in CP due to their low abundance.
- Let the mice acclimatize for at least 7 days prior to any experimentation. Keep the mice under pathogen-free conditions at constant temperature and humidity, in a 12/12 h or 14/10 h light/dark cycle, with water and standard pellet food ad libitum.
3. PBS perfusion and brain dissection
- Weight each mouse and inject 10 μL/g of the anesthetic-analgesic mix solution (step 1.6) intraperitoneally.
- Wait around 30 min for efficient analgesia (check for the depth of anesthesia by pinching the mouse's digits).
- Position the anesthetized mouse flat on its back, on the dissection support, and tape its palms to the dissection support.
- Pinching the skin of the animal with forceps and using scissors, open the abdomen, the diaphragm, and the thorax to expose the heart.
NOTE: When the thorax is opened, the brain becomes anoxic. One must proceed precisely and swiftly through the next steps of the perfusion. - Inject 20 μL of 100 U/mL heparin solution directly into the left ventricle.
- With fine scissors, make an incision of at least 3 mm in the right atrium to allow the blood to flow out of the body.
- Immediately after, insert the 23 G needle placed at the extremity of the infusion tube through the tip of the left ventricle.
- Open the infusion system at the maximum and wait for complete perfusion: Using a 23 G needle, at a flow rate of about 6 mL/min, the perfusion is complete in 3 min.
NOTE: The even discoloration of organs such as the liver attests for perfusion efficacy. - Close the infusion system and remove the needle from the heart ventricle.
- Remove the tape from the mouse's palms. Place the mouse in a ventral position.
- Pinching the skin of the animal with forceps and using scissors, remove the skin of the top of the head from the eyes to the ears.
- With the help of scissors, cut the skull first between the eyes and then laterally from each eye to the spinal cord just above the masseter muscles. For this, use the extremity of scissors and proceed gently to avoid damaging the brain with the scissors.
- Open the skull with forceps by pinching it from the extremity between the eyes.
- Use scissors to cut the spinal cord and extract the brain with forceps, placing their two points on the lateral side of the brain to tilt it and place it in a Petri dish filled with ice-cold PBS+/+. The CP are then immediately collected.
NOTE: Check for discoloration of the brain to verify perfusion efficacy.
4. Dissection of the Choroid Plexus from the brain
- Position the brain dorsal side up in the Petri dish and below the objectives of the binocular loupes.
- With the help of forceps to maintain the brain in place, insert the two ends of another forceps down through the midline between the hemispheres.
- Use the forceps to pull the cortex with the callosum and hippocampus away from the septum, exposing the lateral ventricle and a part of the third ventricle.
- Identify the lateral CP as a long veil lining the lateral ventricle that is flaring at both ends. Use the two ends of a thin forceps to catch the lateral CP. Be careful to collect the triangular posterior part that may be hidden by the posterior fold of the hippocampus.
- Pull the cortex with the corpus callosum and hippocampus of the contralateral hemisphere away from the septum to expose the entire third ventricle and the opposite lateral ventricle.
- Collect with fine forceps the third CP, which can be identified as a short structure with a granular surface aspect.
- Collect the other lateral CP.
- Insert the two ends of a forceps down between the cerebellum and midbrain. Detach the cerebellum from the pons and medulla to expose the fourth ventricle.
- Identify the fourth CP as a long globular structure with a granular surface aspect that lines the fourth ventricle from the lateral right end to the left end between the cerebellum and the medulla. Collect the fourth CP with fine forceps.
NOTE: Silane-base coating of forceps may prevent stickiness of CP on forceps. - Repeat steps 3.1-4.9 for each of the other seven mice. In the meantime, the collected CP can be kept temporally in a tube placed on ice.
5. Preparation of samples for flow cytometry analysis
- Fill the tube containing all dissected CP to 750 μL of PBS+/+.
NOTE: The deposition of dissected CP with thin forceps in the tube brings a small volume of PBS+/+. Rather complete the existing volume to 750 μL than remove and replace it with fresh 750 μL of PBS+/+ to avoid a possible loss of the CP tissue. - Add 15 μL of 20 U/μL Collagenase IV stock solution (see step 1.5) for a 400 U/mL final concentration.
NOTE: DNAse I (150 μg/mL) may prevent excessive cell clumping. - Incubate CP with Collagenase IV at 37 °C under mild agitation (300 RPM) for 45 min.
- Gently pipette up and down around 10 times to finalize the CP dissociation.
- Fill CP tube to 1.5 mL with MACS buffer (see recipe step 1.4) to stop collagenase IV activity.
NOTE: Collagenase IV activity is stopped at low temperature and inhibited by serum albumin. - Centrifuge the cells at 500 x g, for 5 min, at 4 °C. Discard the supernatant and wash the cells with 1.5 mL of MACS buffer.
- Centrifuge the cells at 500 x g for 5 min, at 4 °C. Discard the supernatant and resuspend cells in 220 μL of MACS buffer.
- Separate an aliquot of 10 μL from the cell suspension to be used as unstained control (next step 5.18).
- Separate an aliquot of 10 μL from the cell suspension to be used as DAPI-mono-stained control (next step 5.12).
- Preincubate the remaining 200 μL with anti-mouse CD16/CD32 blocking antibody (1:100) for 20 min, at 4 °C, to block nonspecific Fc-mediated interactions.
- Split the cells into two 100 μL tubes (one for myeloid cells analysis, the other for T cells analysis) (next step 5.14).
- Take the sample with 10 μL of cells for the DAPI-mono-stained control (step 5.9) and fill it up to 100 μL with MACS buffer (next step 5.14).
- Prepare eleven new tubes containing 100 μL of MACS for the antibody mono-stained (step 5.14.4) and all-stained controls (step 5.14.5) and add a drop of compensation beads into each.
NOTE: The compensation controls will allow setting up the voltage of fluorescent detection laser and assess the potential unspecific signal detection of fluorescent dye in the other channels that will be further compensated. - Antibody incubation of samples:
- Myeloid sample: Add 0.1 mg/mL DAPI (1:100) (see step 1.3), FITC anti-mouse CD45 (1:100), PE-Dazzle 594 anti-mouse CD11b (1:100), APC anti-mouse CX3CR1 (1:100), BV711 anti-mouse Ly6C (1:100), PE anti-mouse F4/80 (1:100), and APC-Cy7 anti-mouse IA-IE (1:100).
- T cells sample: Add 0.1 mg/mL DAPI (1:100), FITC anti-mouse CD45 (1:100), PE-Dazzle 594 anti-mouse CD11b (1:100), APC anti-mouse TCRβ (1:100), PE anti-mouse CD8a (1:100), and APC-Cy7 anti-mouse CD4 (1:50).
- DAPI-mono-stained control (from step 5.12): Add 0.1 mg/mL DAPI (1:100).
- Antibody mono-stained compensation beads: Add each of the nine previously used antibodies for myeloid and T cells samples (same dilution), separately (one for each tube) to the nine tubes containing the beads.
NOTE: The mono-stained compensation controls will allow the determination of positive peaks of fluorescence for each of the used fluorescent markers during the compensation step. The analyzer will compare them to the negative signal observed in the unstained control CP cells. - Antibody all-stained compensation beads: Add all the antibodies used for myeloid staining to one tube and those used for T cells staining to another.
NOTE: The all-stained compensation controls will allow setting up the voltage of each fluorescent laser detection and assess the potential unspecific signal detection of fluorescent dye in the other channels. - Incubate for 30-45 min on ice, protected from light exposure.
- Fill all tubes up to 1.5 mL with MACS buffer. Centrifuge the cells and compensation beads at 500 x g, for 5 min, at 4 °C.
- Discard the supernatant and wash the cells and compensation beads in 1.5 mL of MACS buffer.
- Centrifuge the cells and compensation beads at 500 x g, for 5 min, at 4 °C. Discard the supernatant.
- Resuspend the cells and compensation beads in 500 μL of MACS buffer. Fill unstained control (step 5.8) up to 500 μL with MACS buffer.
- Filter each tube with CP cells through a 70 μm strainer (unstained CP, DAPI mono-stained control, and myeloid and T cells samples).
6. Flow cytometry
NOTE: The flow cytometer used in this protocol is equipped with the following 5 lasers: a 355 nm UV laser, a 405 nm Violet laser, a 488 nm Blue laser, a 561 nm Yellow-Green laser and a 637 nm Red laser.
- Perform the daily quality control checks on the cytometer using the cytometer setup, tracking (CST) interface, and the CST control beads to ensure consistency between analyses, instrument quality, and consistent target fluorescence intensities.
- To set up the flow cytometry experiment, create a new experiment with bivariate plots and histograms. Ensure inclusion of a forward scatter area (FSC-A) and side scatter area (SSC-A) plots as well as histogram plots for each color to monitor the acquisition.
- Define the cell morphology by setting up the PMT voltage for FSC-A and SSC-A parameters on unstained control CP cells.
- Define the voltages of each fluorescent marker by setting up the negative peaks of the fluorochromes on unstained control CP cells and the positive peaks of the fluorochromes using compensation beads stained with all antibodies and ensuring that the detector signals are not off the scale and are not too strongly cross-detected in other channels.
- Create a compensation matrix to analyze each of the single-color compensation controls. After gating on appropriate FSC-A/SSC-A populations, check single-stained controls on histogram plots and record compensation controls. After recording all single-stained samples, calculate the compensation matrix.
- Record all events detected for both myeloid and T cell populations.
7. CP myeloid cells gating
- Select FSC-A vs. SSC-A and gate for cells (based on size).
- Create a daughter gate for single cells with FSC-A vs. forward scatter height (FSC-H).
- Create a daughter gate for live cells with DAPI vs. FSC-A to exclude DAPI+ cells.
- Create a daughter gate for CD45+ immune cells with CD45 vs. FSC-A.
- Create a daughter gate from the CD45+ cells and select CD11b vs. F4/80, identify the following two groups: the CD11b+ F4/80+ and the CD11b+ F4/80– populations (next step 7.8).
- Create a daughter gate for CD11b+ F4/80+ cells and select CX3CR1 vs. IA-IE. Identify both CX3CR1+ IA-IE+ BAM that are further denominated as CD11b+ F4/80high CX3CR1+ IA-IE+ BAM, and CX3CR1+ IA-IE– macrophages.
- Create a daughter gate for CD11b+ F4/80+ CX3CR1+ IA-IE– macrophages and select F4/80 vs. CD11b. Identify both the CD11bhigh F4/80intermediate and the CD11b+ F4/80high populations that are further called CD11bhigh F4/80intermediate CX3CR1+ IA-IE– Kolmer's epiplexus macrophages and CD11b+ F4/80high CX3CR1+ IA-IE– BAM, respectively.
NOTE: CD11bhigh F4/80intermediate CX3CR1+ IA-IE– and CD11b+ F4/80high CX3CR1+ IA-IE– cells appeared a bit intermixed between F4/80intermediate-F4/80high and CD11bhigh-CD11b+ levels. - From the CD11b+ F4/80– population (step 7.5), gate for Ly6C vs. IA-IE. Select both IA-IE+ Ly6C– population and IA-IE– Ly6C+ cells that mainly correspond to monocytes/neutrophils.
NOTE: A high presence of IA-IE– Ly6C+ cells likely reflects problems in the perfusion efficacy of animals without an inflammatory condition; their abundance should be low. - Create a daughter gate for CD11b+ F4/80– IA-IE+ Ly6C– population and select CD11b vs. CX3CR1, identify both CD11bhigh CX3CR1low and CD11b+ CX3CR1– populations.
8. CP T cells gating
- Select FSC-A vs. SSC-A and gate for cells (based on size).
- Create a daughter gate for single cells with FSC-A vs. forward scatter height (FSC-H).
- Create a daughter gate for live cells with DAPI vs. FSC-A to exclude DAPI+ cells.
- Create a daughter gate for CD45+ immune cells with CD45 vs. FSC-A.
- Create a daughter gate from the CD45+ cells and select TCRβ vs. CD11b. Exclude CD11b+ myeloid cells and select TCRβ+ CD11b– T cells population.
- Create a daughter gate for TCRβ+ CD11b– population and select CD4 vs. CD8a. Identify both CD8– CD4+ T cells and CD8+ CD4– T cells.
Representative Results
The flow cytometry analyses presented here successfully revealed the major subsets of myeloid and T cells (Figure 1 and Figure 2, respectively), and their relative total number per mouse in a highly reproducible manner (Figure 3).
The flow cytometry analysis of myeloid cells showed that CP are populated by CD11b+ CX3CR1+ F4/80high BAM, representing almost 80% of the CD45+ immune cells at the CP. These BAM were divided into two different populations according to their expression of MHC-II: the IA-IE+ BAM that constituted the major group (72% of the CD45+ immune cells), in line with previously published data7, and the IA-IE– BAM. The CD11bhigh F4/80intermediate CX3CR1+ IA-IE– population of Kolmer's epiplexus macrophages7,2 was also identified and only represented about 1.2% of the CD45+ CP immune cells, consistently with the literature7. Among the CD11b+ F4/80– immune cells, two different populations of Ly6C– IA-IE+ cells were characterized that likely correspond to dendritic cells, and that can be divided into CD11bhigh CX3CR1low and CD11b+ CX3CR1– populations which constituted around 3.1% and 3.7% of the CD45+ CP immune cells, respectively. CD45+ CD11b+ F4/80– IA-IE– Ly6C+ cell population including both monocytes and neutrophils constituted only 0.5% of CD45+ immune cells in accordance with previous reports and attesting for the high efficacy of the PBS perfusion3. These results are largely consistent with previously published data assessing CP immune populations by single-cell RNA sequencing7.
The analysis and gating strategy of T cells highlighted the presence of the minor population of CD45+ CD11b– TCRβ+ cells. Among them, about 30-50 CD8– CD4+ and CD8+ CD4– T cells per mouse populated CP under physiological conditions, representing 1.3% and 0.9% of the CD45+ CP immune cells, respectively. CD4+ cells were slightly more abundant than CD8+ T cells, which is consistent with previous results in terms of both number and proportion21,30. This gating strategy also revealed a population of CD45+ CD11b– TCRβ+ CD4– CD8– cells, which may include NKT cells, and a recently described regulatory T cell subset10,28,31. The proposed flow cytometry analysis and gating strategies allowed the cell type annotation of almost 90% of the immune cells (CD45+) that compose the mouse CP.
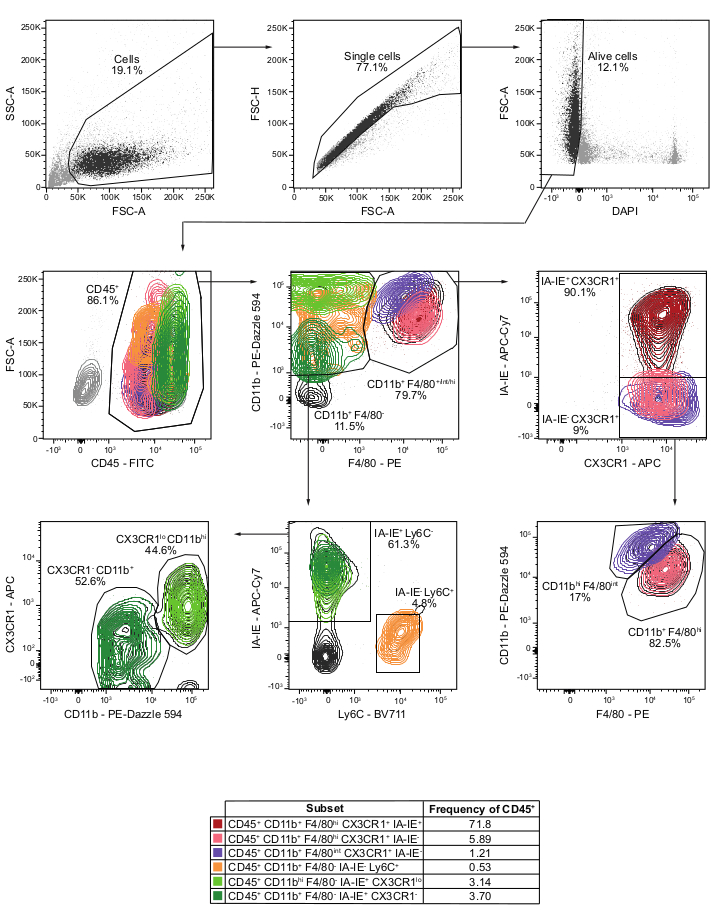
Figure 1: Flow cytometry analysis and gating strategy of myeloid cells from CP of perfused mice. Cells are gated based on size using FSC-A vs. SSC-A. Singlet cells are selected using FSC-A vs. FSC-H. Live cells are gated by excluding DAPI+ cells on DAPI vs. FSC-A plot. CD45+ immune cells are then identified on a CD45 vs. FSC-A plot. Gating CD11b vs. F4/80, the CD11b+ F4/80+ and the CD11b+ F4/80– populations are selected. Gating CD11b+ F4/80+ population using CX3CR1 vs. IA-IE, both the CD11b+ F4/80hi CX3CR1+ IA-IE+ BAM and the CX3CR1+ IA-IE– populations are determined. This latter population is then gated using F4/80 vs. CD11b to better separate CD11b+ F4/80high CX3CR1+ IA-IE– BAM and CD11bhigh F4/80intermediate CX3CR1+ IA-IE– Kolmer's epiplexus macrophages. Gating Ly6C vs. IA-IE on CD11b+ F4/80–, both CD11b+ F4/80– IA-IE– Ly6C+ cells that mainly correspond to monocytes and neutrophils and CD11b+ F4/80– IA-IE+ Ly6C– cells that likely represent dendritic cells population are selected. CD11b+ F4/80– IA-IE+ Ly6C– cells are gated using CD11b vs. CX3CR1 allowing identification of CD11bhigh CX3CR1low and CD11b+ CX3CR1– cells. If not specified, values below each gated population represent the frequency of the parent population. hi: high; int: intermediate; lo: low. Please click here to view a larger version of this figure.
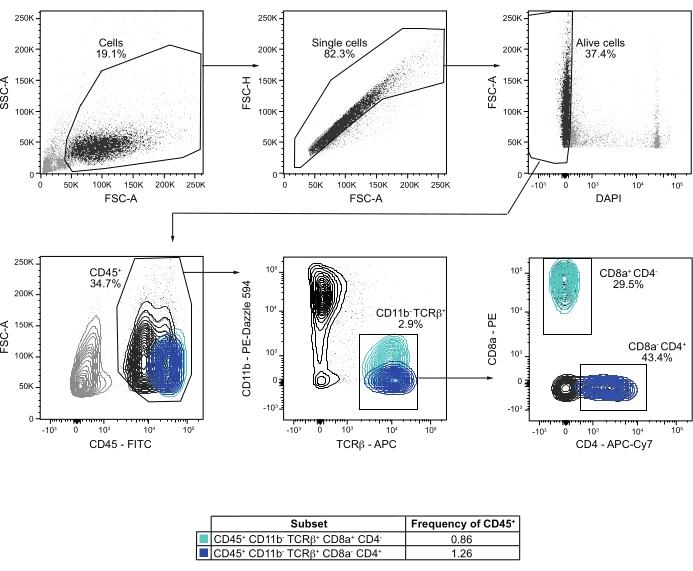
Figure 2: Flow cytometry analysis and gating strategy of T cells from CP of perfused mice. Cells are gated based on size using FSC-A vs. SSC-A. Single cells are selected using FSC-A vs. FSC-H. Live cells are identified excluding DAPI+ cells on DAPI vs. FSC-A plot. CD45+ immune cells are then gated on a CD45 vs. FSC-A plot. Gating CD11b vs. TCRβ, CD11b– TCRβ+ are selected. Gating CD4 vs. CD8, the proportions of CD8– CD4+ and CD4– CD8+ T cells are determined. If not specified, values below each gated population represent the frequency of the parent population. Please click here to view a larger version of this figure.
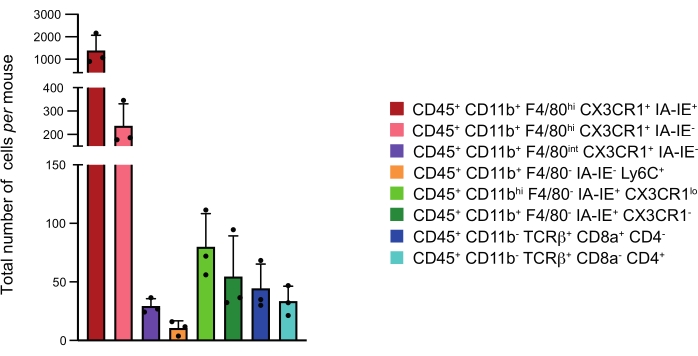
Figure 3: Numbers of the major immune cell subsets at the CP per mouse. Mean of the total number of cells per mouse of the different immune subsets identified in the four CP pooled together. Representation: Mean + standard deviation. Please click here to view a larger version of this figure.
Discussion
Studies aiming to understand the immunological contributions to brain homeostasis and disease have mainly focused on cells residing within the brain parenchyma, neglecting brain borders such as CP, which are nevertheless crucial contributors to brain function2,3. The analysis of immune cell populations at CP is challenging due to the small size of CP, low numbers of resident immune cells, and complicated access to this tissue. Flow cytometry performed on total brain immune cells (CD45+) does not allow the characterization of the rare immune populations that reside in the CP. To characterize the immune composition of the CP with high precision, flow cytometry analyses were conducted on CP dissected from PBS-perfused mice. This approach allows the exclusion of circulating CD45+ cells and CP CD45– cells such as pericytes, endothelial and epithelial cells32. The gating strategies focusing on myeloid and T cell populations identify the main CP immune subsets.
A critical step of the current protocol is the PBS perfusion. As the CP immune cells are rare, blood contamination could completely mask their local heterogeneity. Liver and brain discoloration is always checked as control, and therefore blood contamination is minimal. Consistently, CD11b+ F4/80– Ly6C+, which includes both monocytes and neutrophils present in the blood rather than the brain in non-inflammatory conditions3, accounted for only 0.5% of the CD45+ cells, demonstrating a high efficacy of PBS perfusion. Another helpful control for PBS perfusion efficacy might be the incorporation of antibodies against Ter119, a marker specific for erythrocytes. Finally, an additional analysis of the CP from PBS-perfused mouse brain sections by immunostaining and microscopy may also be useful to verify whether any blood immune cells that may be strongly adherent to the endothelium have resisted the PBS perfusion and remain attached at the lumen of the vasculature.
The myeloid gating strategy revealed two populations of macrophages: CD11b+ F4/80high CX3CR1+ IA-IE+/-BAM and CD11b+ F4/80intermediate CX3CR1+ IA-IE– Kolmer's epiplexus cells, which express high levels of CD11b and low levels of F4/807. The gates of Kolmer's epiplexus cells7,20 and IA-IE– BAM were not clearly separated using CD11b and F4/80 markers. Incorporation of the CD206 marker into this gating strategy may help to better differentiate CD206+ IA-IE– BAM from CD206low Kolmer's epiplexus cells7.
Almost 7% of the CP immune cells appeared to be CD11b+ F4/80– Ly6C– IA-IE+. This subset was composed of two populations according to their levels of CD11b and CX3CR1 expression. These cells likely correspond to dendritic cells, but this remains to be confirmed by analyzing their CD11c expression either by FACS3 or immunofluorescence33. The CD11b+ F4/80– Ly6C– IA-IE+ population also likely includes mast cells. Incorporating antibodies for the detection of CD117 and c-kit into the gating strategy would help to distinguish them from CD11c+ dendritic cells3,34. The proportion of CD45+ CD11b+ F4/80+ Ly6C+ cells among CP immune cells was determined. However, they include both monocytes and neutrophils, and the addition of the Ly6G marker to the gating strategy of CD45+ CD11b+ F4/80+ Ly6C+ cells would determine the ratio of Ly6G– monocytes to Ly6G+ neutrophils. Beyond the described CD45+ CD11b– TCRβ+ population, the gating strategy on CD45+ CD11b– populations may be enriched using NK1.1 and CD11c markers to identify NK cells, TCRγδ to analyze γδ T cells, CD19 and/or B220 to characterize B cells, and CD138 to identify plasma cells3. Expression of reporter genes can also be included in this analysis. For example, using Foxp3-GFP transgenic mice could be helpful in identifying CP-resident T regulatory (Treg) cells3,30. Lastly, this protocol can also incorporate intracellular staining of transcription factors or cytokines21.
Whereas the immune cells of the brain parenchyma have been extensively studied, much less is known about immune cells populating brains barriers, such as the CP3,35,36. CP in mice are small tissues located in four different areas of the brain, and their isolation requires a quite complex microdissection. Because of this, the CP were mostly studied using imaging. Staining of CP immune cells from whole CP dissected tissue or from brain sections followed by microscopic analysis provided major advances in the understanding of CP immune mechanisms14,15,21,22,26,27,29,30,33. However, this method does not allow for extensive analysis of the CP immune cell diversity as only four to six markers can be analyzed at a time. In addition, some subpopulations such as CD4+ or CD8+ T cells are quite rare at the CP, and quantitative analysis of their phenotypes with microscopy would be challenging. The flow cytometry approach allows for the analysis of tens of markers in parallel for every immune cell, providing a better-suited tool for quantitative assessment of the CP immune composition.
More recently, single nuclei transcriptomic technology (snRNA-seq) came out as a powerful method to study CP heterogeneity10,28. However, it is not well-suited for analyzing CP immune cells as they constitute only a small fraction of the CP cells. Since snRNA-seq samples the cells at the proportion they are present in the tissue, the immune cells are underrepresented in the snRNA-seq, and their low numbers pose an obstacle for in-depth analysis10,28.
Single-cell transcriptomics (scRNA-seq) has been recently applied to precisely map the immune cell heterogeneity of isolated CP immune cells7. While scRNA-seq is quite complex and costly to perform, flow cytometry may serve as a more accessible way to survey immune subpopulations at the CP. Lastly, this protocol can serve as part of a cell sorting protocol, where the selected subtypes of cells are sorted using FACS for analysis with molecular methods, such as scRNA-seq or bulk RNA analysis.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
We thank the Institut Pasteur Animalerie Centrale and the CB-UTechS facility members for their help. This work was supported financially by Institut Pasteur.
Materials
| anti-mouse CD16/CD32 | BD Biosciences | 553142 | Flow cytometry antibody |
| Albumin, bovine | MP Biomedicals | 160069 | Blocking reagent |
| APC anti-mouse CX3CR1 | BioLegend | 149008 | Flow cytometry antibody |
| APC anti-mouse TCRb | BioLegend | 109212 | Flow cytometry antibody |
| APC-Cy7 anti-mouse CD4 | BioLegend | 100414 | Flow cytometry antibody |
| APC-Cy7 anti-mouse IA-IE | BioLegend | 107628 | Flow cytometry antibody |
| BD FACSymphony A5 Cell Analyzer | BD Biosciences | Flow cytometry analyzer | |
| BV711 anti-mouse Ly6C | BioLegend | 128037 | Flow cytometry antibody |
| Collagenase IV | Gibco | 17104-019 | Enzyme to dissociate CP tissue |
| DAPI | Thermo Scientific | 62248 | Live/dead marker |
| EDTA | Ion chelator | ||
| fine scissors | FST | 14058-11 | Dissection tool |
| FITC anti-mouse CD45 | BioLegend | 103108 | Flow cytometry antibody |
| Flow controller infusion inset | CareFusion | RG-3-C | Blood perfusion inset |
| FlowJo software | BD Biosciences | Analysis software | |
| forceps | FST | 11018-12 | Dissection tool |
| Heparin | Sigma-Aldrich | H3149-10KU | Anticoagulant |
| Imalgene | Boehringer Ingelheim | Ketamine, anesthesic | |
| OneComp eBeads | Invitrogen | 01-1111-42 | Control beads to realize compensation |
| PBS-/- | Gibco | 14190-094 | Buffer |
| PBS+/+ | Gibco | 14040-091 | Buffer |
| PE anti-mouse CD8a | BioLegend | 100708 | Flow cytometry antibody |
| PE anti-mouse F4/80 | BioLegend | 123110 | Flow cytometry antibody |
| PE-Dazzle 594 anti-mouse CD11b | BioLegend | 101256 | Flow cytometry antibody |
| Rompun | Bayer | Xylazine, anesthesic | |
| thin forceps | Dumoxel Biology | 11242-40 | Dissection tool |
| Vetergesic | Ceva | Buprenorphin, analgesic |
Riferimenti
- Morais, L. H., Schreiber, H. L., Mazmanian, S. K. The gut microbiota-brain axis in behaviour and brain disorders. Nature Reviews Microbiology. 19 (4), 241-255 (2021).
- Deczkowska, A., Schwartz, M. Targeting neuro-immune communication in neurodegeneration: Challenges and opportunities. Journal of Experimental Medicine. 215 (11), 2702-2704 (2018).
- Croese, T., Castellani, G., Schwartz, M. Immune cell compartmentalization for brain surveillance and protection. Nature Immunology. 22 (9), 1083-1092 (2021).
- Erny, D., et al. Host microbiota constantly control maturation and function of microglia in the CNS. Nature Neuroscience. 18 (7), 965-977 (2015).
- Mrdjen, D., et al. High-dimensional single-cell mapping of central nervous system immune cells reveals distinct myeloid subsets in health, aging, and disease. Immunity. 48 (2), 380-395 (2018).
- Korin, B., et al. single-cell characterization of the brain’s immune compartment. Nature Neuroscience. 20 (9), 1300-1309 (2017).
- van Hove, H., et al. A single-cell atlas of mouse brain macrophages reveals unique transcriptional identities shaped by ontogeny and tissue environment. Nature Neuroscience. 22 (6), 1021-1035 (2019).
- Ajami, B., et al. Single-cell mass cytometry reveals distinct populations of brain myeloid cells in mouse neuroinflammation and neurodegeneration models. Nature Neuroscience. 21 (4), 541-551 (2018).
- Wolburg, H., Paulus, W. Choroid plexus: Biology and pathology. Acta Neuropathologica. 119 (1), 75-88 (2010).
- Dani, N., et al. A cellular and spatial map of the choroid plexus across brain ventricles and ages. Cell. 184 (11), 3056-3074 (2021).
- Falcão, A. M., Marques, F., Novais, A., Sousa, N., Palha, J. A., Sousa, J. C. The path from the choroid plexus to the subventricular zone: Go with the flow. Frontiers in Cellular Neuroscience. 6, (2012).
- Shipley, F. B., et al. Tracking calcium dynamics and immune surveillance at the choroid plexus blood-cerebrospinal fluid interface. Neuron. 108 (4), 623-639 (2020).
- Mazucanti, C. H., et al. Release of insulin produced by the choroids plexis is regulated by serotonergic signaling. JCI Insight. 4 (23), (2019).
- Baruch, K., et al. Aging-induced type I interferon response at the choroid plexus negatively affects brain function. Science. 346 (6205), 89-93 (2014).
- Deczkowska, A., et al. Mef2C restrains microglial inflammatory response and is lost in brain ageing in an IFN-I-dependent manner. Nature Communications. 8 (1), (2017).
- Silva-Vargas, V., Maldonado-Soto, A. R., Mizrak, D., Codega, P., Doetsch, F. Age-dependent niche signals from the choroid plexus regulate adult neural stem cells. Cell Stem Cell. 19 (5), 643-652 (2016).
- Iliff, J. J., et al. Impairment of glymphatic pathway function promotes tau pathology after traumatic brain injury. Journal of Neuroscience. 34 (49), 16180-16193 (2014).
- Redzic, Z. B., Preston, J. E., Duncan, J. A., Chodobski, A., Szmydynger-Chodobska, J. The choroid plexus-cerebrospinal fluid system: From development to aging. Current Topics in Developmental Biology. 71, 1-52 (2005).
- da Mesquita, S., et al. Functional aspects of meningeal lymphatics in ageing and Alzheimer’s disease. Nature. 560 (7717), 185-191 (2018).
- Schwarze, E. -. W. The origin of (Kolmer’s) epiplexus cells. Histochemistry. 44 (1), 103-104 (1975).
- Baruch, K., et al. CNS-specific immunity at the choroid plexus shifts toward destructive Th2 inflammation in brain aging. Proceedings of the National Academy of Sciences of the United States of America. 110 (6), 2264-2269 (2013).
- Kunis, G., et al. IFN-γ-dependent activation of the brain’s choroid plexus for CNS immune surveillance and repair. Brain. 136 (11), 3427-3440 (2013).
- Prinz, M., Priller, J. Microglia and brain macrophages in the molecular age: From origin to neuropsychiatric disease. Nature Reviews Neuroscience. 15 (5), 300-312 (2014).
- Goldmann, T., et al. fate and dynamics of macrophages at central nervous system interfaces. Nature Immunology. 17 (7), 797-805 (2016).
- Fung, I. T. H., et al. Activation of group 2 innate lymphoid cells alleviates aging-associated cognitive decline. Journal of Experimental Medicine. 217 (4), (2020).
- Kertser, A., et al. Corticosteroid signaling at the brain-immune interface impedes coping with severe psychological stress. Science Advances. 5, 4111 (2019).
- Shechter, R., et al. Recruitment of beneficial M2 macrophages to injured spinal cord is orchestrated by remote brain choroid plexus. Immunity. 38 (3), 555-569 (2013).
- Yang, A. C., et al. Dysregulation of brain and choroid plexus cell types in severe COVID-19. Nature. 595 (7868), 565-571 (2021).
- Baruch, K., et al. PD-1 immune checkpoint blockade reduces pathology and improves memory in mouse models of Alzheimer’s disease. Nature Medicine. 22 (2), 135-137 (2016).
- Baruch, K., et al. Breaking immune tolerance by targeting Foxp3+ regulatory T cells mitigates Alzheimer’s disease pathology. Nature Communications. 6, 7967 (2015).
- Rodríguez-Rodríguez, N., Flores-Mendoza, G., Apostolidis, S. A., Rosetti, F., Tsokos, G. C., Crispín, J. C. TCR-α/β CD4 − CD8 − double negative T cells arise from CD8 + T cells. Journal of Leukocyte Biology. 108 (3), 851-857 (2020).
- Schafflick, D., et al. Single-cell profiling of CNS border compartment leukocytes reveals that B cells and their progenitors reside in non-diseased meninges. Nature Neuroscience. 24 (9), 1225-1234 (2021).
- Quintana, E., et al. DNGR-1+ dendritic cells are located in meningeal membrane and choroid plexus of the noninjured brain. GLIA. 63 (12), 2231-2248 (2015).
- Kabashima, K., et al. Biomarkers for evaluation of mast cell and basophil activation. Immunological Reviews. 282 (1), 114-120 (2018).
- Li, Q., Barres, B. A. Microglia and macrophages in brain homeostasis and disease. Nature Reviews Immunology. 18 (4), 225-242 (2018).
- Borst, K., Dumas, A. A., Prinz, M. Microglia: Immune and non-immune functions. Immunity. 54 (10), 2194-2208 (2021).

