DNA Virus Detection System Based on RPA-CRISPR/Cas12a-SPM and Deep Learning
Summary
We present a protocol that combines recombinase polymerase amplification with a CRISPR/Cas12a system for trace detection of DNA viruses and builds portable smartphone microscopy with an artificial intelligence-assisted classification for point-of-care DNA virus detection.
Abstract
We report a fast, easy-to-implement, highly sensitive, sequence-specific, and point-of-care (POC) DNA virus detection system, which combines recombinase polymerase amplification (RPA) and CRISPR/Cas12a system for trace detection of DNA viruses. Target DNA is amplified and recognized by RPA and CRISPR/Cas12a separately, which triggers the collateral cleavage activity of Cas12a that cleaves a fluorophore-quencher labeled DNA reporter and generalizes fluorescence. For POC detection, portable smartphone microscopy is built to take fluorescent images. Besides, deep learning models for binary classification of positive or negative samples, achieving high accuracy, are deployed within the system. Frog virus 3 (FV3, genera Ranavirus, family Iridoviridae) was tested as an example for this DNA virus POC detection system, and the limits of detection (LoD) can achieve 10 aM within 40 min. Without skilled operators and bulky instruments, the portable and miniature RPA-CRISPR/Cas12a-SPM with artificial intelligence (AI) assisted classification shows great potential for POC DNA virus detection and can help prevent the spread of such viruses.
Introduction
In recent years, epidemics of infectious diseases caused by different viruses have occurred frequently, including the Ebola virus disease (EVD) epidemic in 20141 and 20182, the Middle East Respiratory Syndrome (MERS) in 20153, the Zika virus disease epidemic in 20154, the Coronavirus disease 2019 (COVID-19) caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)5 and the continuing Monkeypox caused by Monkeypox virus (MKPV) in 20226. These sudden outbreaks of epidemic infectious diseases cause a large number of deaths and bring huge economic losses and social unrest. A rapid and accurate detection system is urgently required to quickly diagnose the infection and prevent the virus's further spread.
Recently, clustered regularly interspaced short palindromic repeats (CRISPR) and CRISPR-associated (Cas) proteins have gained worldwide attention and have shown promising results in nucleic acid detection7,8,9,10,11,12,13,14,15. The CRISPR/Cas12a protein, guided by CRISPR RNA (crRNA), binds to and cleaves the target DNA. This activity leads to the release of nonspecific single-stranded DNA (ssDNA), known as trans-cleavage, and can be utilized to enhance the detection signal for nucleic acid detection. Some traditional detection methods like polymerase chain reaction (PCR), quantitative real-time PCR (qPCR), and enzyme-linked immunosorbent assay (ELISA) are complicated, time-consuming, and costly for point-of-care (POC) detection. Our previous work successfully developed an automated, integrated, and cost-effective detection system for the African swine fever virus (ASFV) based on CRISPR/Cas12a technology. In this system, we achieved a detection limit of 1 pM within a 2-h time frame without the need for amplification. The CRISPR/Cas12a system and recombinase polymerase amplification (RPA) are combined to improve the sensitivity and specificity for trace DNA detection. Compared with other isothermal amplification techniques, RPA is simple in design and convenient in operation since it has a shorter reaction time without sophisticated temperature control equipment.
For POC detection of pathogens, instruments such as smartphone microscopy (SPM), handheld fluorimeter, or lateral flow strips are developed for the results readouts16,17,18. SPM captures images through a camera and uploads them to some mobile applications for fast data analysis. Such microscopy makes a portable, cheap, and miniatured signal acquisition system with high sensitivity and has shown advantages in detecting pathogens such as H5N1, the Zika virus, and SARS-CoV-219,20. Therefore, we build a portable SPM to catch the fluorescence signals triggered by RPA-CRISPR/Cas12a detection of the target DNA virus. The ssDNA reporter probe linking a fluorophore and a quencher will be cleaved when CRISPR/Cas12a recognizes the target DNA virus, and the fluorescence emitted by the fluorophore can be captured by SPM.
Compared to the professional software usually used to obtain the results information from the fluorescence images from SPM21, some experts use machine learning and deep learning to quantify the concentrations of virus DNA after attaining fluorescence images22, which is more time-consuming. When it comes to classifying medical images, conventional neural networks (CNNs) are often used to learn features from the raw pixelated images in an end-to-end manner23,24,25,26. Popular CNN-based deep learning models like AlexNet, DenseNet-121, and EfficientNet-B7 have been successfully applied in this field27,28. However, obtaining large data sets in specific domains can be challenging, necessitating transfer learning29,30. This approach pre-trains a deep learning model with a large dataset, and the pre-trained model is used as the starting point for a new task with a small dataset. This technique can reduce the need for large data sets, combat overfitting, and reduce training time31. Herein, we use deep learning models with transfer learning for the binary classification of the positive and negative samples' fluorescence images.
In this method, we combine RPA and the CRISPR/Cas12a system for the trace detection of DNA viruses. Target DNA is amplified and recognized by RPA and CRISPR/Cas12a separately, which triggers the collateral cleavage activity of Cas12a that cleaves a fluorophore-quencher labeled DNA reporter and generalizes fluorescence. We build a portable SPM to take the fluorescent images for POC detection and develop deep learning models for binary classification. The schematic of the built POC detection system is shown in Figure 1. Without skilled operators and bulky instruments, the RPA-CRISPR/Cas12a-SPM with artificial intelligence (AI) assisted classification shows great potential for POC DNA virus detection.
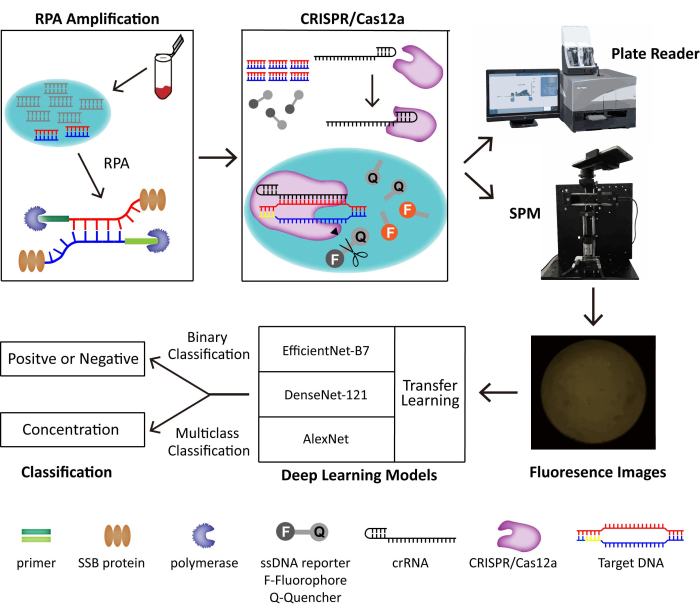
Figure 1: The schematic of the RPA-CRISPR/Cas12-SPM detection system along with AI classification for collected images. The nucleic acids of animal-derived samples are released by PINDBK. The Target DNA of the virus is amplified and recognized specifically by the RPA-CRISPR/Cas12a system. CRISPR/Cas12a bonds with crRNA and the Cas12a-crRNA complex bonds with target DNA, which triggers the collateral cleavage of CRISPR/Cas12a on the ssDNA reporter probes. The fluorophore on the reporter is released, and the fluorescence is detected by a commercialized plate reader or the SPM we build. Three different deep learning models, including AlexNet, DenseNet-121, and EfficientNet-B7 with transfer learning, are used to classify the fluorescence images. This figure is reused with permission from Lei et al.35. Please click here to view a larger version of this figure.
Protocol
1. Processing of samples
- Take Frog Virus 3 (FV3, genera Ranavirus, family Iridoviridae), a double-stranded DNA virus. Select the major capsid (mcp) gene as the target for the detection of FV3 since it is highly conserved and usually regarded as the target for ranavirus detection. The target sequence selected is shown in Table 1.
NOTE: Frog Virus 3 is taken as an example in this protocol. - For target DNA fragments preparation, use DNA fragments of the mcp gene from FV3 and Infectious Spleen and Kidney Necrosis Virus (ISKNV, another Ranavirus).
NOTE: In this study, the target DNA fragments were commercially obtained from a company listed in the Table of Materials. They are regarded as the target and the control of subsequent detection.
| Name | Sequence | ||
| FV3 MCP | NTS: 5’ …gtaacccggctttcGGGCAGCAGTTTCGGTCGGCGTtcccaggtcg… 3’ (240 bp) | ||
| TS: 5’ …ccgacctgggaACGCCGACCGAAACTGCTGCCCtgctgcccgaaagc… 3’ (240 bp) | |||
| ISKNV MCP | NTS: 5’ …ggccatgccaatttTGGGCAGGAGTTTAGTGTGACGgtggcgaggg… 3’ (231 bp) | ||
| TS: 5’ …ccctcgccaccgtcACACTAAACTCCTGCCCAAAATtggcatggcc… 3’ (231 bp) | |||
Table 1: Target sequence selected in this method.
2. RPA reaction
- Design and synthesize the RPA primer pairs for the target sequence. The sequences of RPA primer pairs are described in Table 2.
- Prepare the 5x RPA reaction buffer (Table 3) and MgCl2 (100 mM).
- Mix the four key RPA enzymes (UvsX, UvsY, GP32, Bsu protein) in the 1x RPA reaction buffer, along with the primers designed in advance, as detailed in Table 4.
- Vortex the mixture thoroughly.
- Add 1 µL of the target obtained from step 1 for each RPA reaction, and mix it thoroughly by vortex again.
- Add 7 µL of MgCl2 (100 mM) to initiate the reaction. The final volume of each RPA reaction is 50 µL.
- Perform the assay at 37 °C for 30 min.
- The RPA product can be stored at 4 °C for a few days. As RPA products degrade, use for further detection as soon as possible to obtain better diagnosis results.
- [Optional] Conduct DNA gel electrophoresis.
- Take out 5 µL of RPA products, add an appropriate volume of 6x DNA loading buffer, and conduct DNA gel electrophoresis in the Tris-acetate EDTA (TAE) buffer.
- Run the electrophoresis under 120 V for about 20 min until the bands of loading buffer reach the bottom of the gel.
- Determine whether the target sequence is successfully amplified from the sample by comparing the size of the bands of the sample with the bands of the marker.
| Name | Sequence |
| RPA primer F | ATGTCTTCTGTAACTGGTTCAGGTATCACA |
| RPA primer R | GGCGTTGAGGATGTAATCCCCCGACCTGGG |
Table 2: RPA primers used in this method.
| Component | Original concentration | Addition |
| PEG 20,000 | – | 114 mg |
| ATP | 100 mM | 125 μL |
| dNTPs | 25 mM | 48 μL |
| Tris-HCl | 1 M | 125 μL |
| DTT | 1 M | 125 μL |
| Phosphocreatine | 1 M | 250 μL |
| Creatine kinase | 10 μg/μL | 50 μL |
| ddH2O | – | 277 μL |
| Total Volume | – | 1 mL |
Table 3: The composition of the 5x RPA reaction buffer (pH 7.5).
| Component | Original concentration | Addition |
| 5x RPA reaction buffer | – | 10 μL |
| UvsX protein | 5 mg/mL | 2.6 μL |
| UvsY protein | 5 mg/mL | 0.9 μL |
| GP32 protein | 5 mg/mL | 2.54 μL |
| Bsu protein | 5 mg/mL | 0.88 μL |
| forward primer | 100 μM | 0.25 μL |
| reversed primer | 100 μM | 0.25 μL |
| ddH2O | – | 24.58 μL |
| Target | – | 1 μL |
| *MgCl2 | 100 mM | 7 μL |
| Total Volume | 50 μL | |
| *MgCl2 needs to be lastly added to initiate the RPA reaction. | ||
Table 4: The composition of the RPA reaction.
3. CRISPR/Cas12a detection without SPM
- Use the crRNAs (Table of Materials) of the target sequence and the ssDNA reporter probe linking a fluorophore and a quencher for CRISPR/Cas12a. Here, carboxy tetramethylrhodamine (TAMRA) is linked as the fluorophore at the 5' ends of ssDNA reporter probes and Black Hole Quencher-2 (BHQ2) as the quencher at the 3' ends. The detailed sequence of crRNA and ssDNA reporter are described in Table 5.
- Prepare the Lachnospiraceae bacterium Cas12a (LbCas12a) protein with 10x CRISPR/Cas12a reaction buffer.
- Dissolve 1 µL of RPA reaction product from section 2 in 1x CRISPR/Cas12a reaction buffer with LbCas12a-crRNA complexes and 500 nM ssDNA reporter probe in a 100 µL reaction volume.
- After mixing LbCas12a and crRNA, let the mixture sit for at least 5 min to form a functional complex. After incubation, add other components to the reaction mixture and carry out the whole reaction at 37 °C. The final volume of each CRISPR/Cas12a reaction is 100 µL. The detailed concentrations of each component in each CRISPR/Cas12a reaction are described in Table 6.
- Perform the 100 µL CRISPR/Cas12a detection reaction at 37 °C for 30 min.
- Examine the fluorescence signals constantly by a microplate reader at an excitation wavelength of 535 nm and an emission wavelength of 595 nm with a gain of 60.
NOTE: The detection of different wavelengths of excitation and emission light depends on the choice of the fluorophore and quencher in the previously designed ssDNA reporter probes. - For these collected data, divide the control value by the positive samples' measurement to normalize all the data, then integrate them for a two-sample t-test analysis.
| Name | Sequence | |
| LbCas12a crRNA for FV3 | uaauuucuacuaaguguagauGGGCAGCAGTTTTCGGTCGGCGT | |
| ssDNA reporter | /5TAMRA/TTATT/3BHQ2 | |
Table 5: Sequences of CRISPR/Cas12a crRNA and ssDNA reporter used in this method.
| Component | Original concentration | Addition |
| NEBuffer r2.1 | – | 10 μL |
| Lba Cas12a (Cpf1) | 10 μM | 0.5 μL |
| crRNA | 10 μM | 0.625 μL |
| Wait for at least 5 min to let the LbCas12a/crRNA complex combine. | ||
| DNA reporter | 100 μM | 0.5 μL |
| ddH2O | – | 87.375 μL |
| Target | – | 1 μL |
| Total Volume | – | 100 μL |
Table 6: The composition of the CRISPR/Cas12a Reaction.
4. SPM setup
- For the excitation path, set a laser beam to pass through the neutral density (ND) filters to attenuate the laser intensity.
- Generate a collimated beam from an aspherical lens (hereinafter referred to as L1) and reflect it by the dichroic mirror (DM).
- Direct the light onto the glass slide where the sample is placed through the objective (20x) to illuminate and excite the fluorescence of the sample. The sample stage enables precise focal plane adjustment, directing the beam to the objective's back focal plane. The steps mentioned above form the excitation path of the SPM instrument.
- For the emission path, position an external lens (hereinafter referred to as L2) to form an intermediate image on the other side of the objective. The objective simultaneously illuminates the sample and collects the emission signal.
- Record the fluorescence signal from the sample using the smartphone placed at the end of the emission path. Use a stable bracket to avoid shaking.
- Set a bandpass filter between L1 and the smartphone's camera to filter out the excited light while allowing only the emitted light from the sample to reach the camera, which can optimize the detection.
- Immobilize the SPM setup on a breadboard for portable deployment.
NOTE: The schematic and the physical appearance of the SPM device for fluorescence detection based on the RPA-CRISPR/Cas12a reaction are shown in Figure 2. The CRISPR/Cas12a detection takes place on the pre-treated glass slide described in the next step.
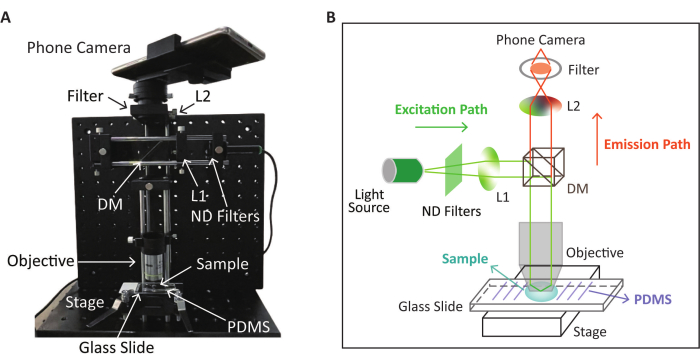
Figure 2: Schematic and physical appearance of the SPM device used for fluorescence detection. (A) The physical appearance of the SPM device for fluorescence image collection after the RPA-CRISPR/Cas12a reaction. (B) Schematic of the SPM device for fluorescence detection based on the RPA-CRISPR/Cas12a reaction. This figure is modified (adjusted image position and color) with permission from Lei et al.35. Please click here to view a larger version of this figure.
5. Treatment of glass slide for detection with SPM
- Prepare polydimethylsiloxane (PDMS) by mixing the base and curing agent with a ratio of 10:1, followed by baking on a hotplate at 80 °C for 2 h.
- Treat both PDMS and the glass slide (Length: 75 mm; Height: 50 mm) with oxygen plasma treatment for 120 s, then press them together.
- Bake the glass/PDMS at 95 °C for 2 h; it is sealed permanently by the Si-O-Si bond. PDMS has high transparency and no auto-fluorescence, which is conducive to SPM detection. The treatment of the glass slide and PMDS is performed as per He et al.32
6. CRISPR/Cas12a detection with SPM
- Use the crRNAs (Table of Materials) of the target sequence and the ssDNA reporter probe linking a fluorophore and a quencher for CRISPR/Cas12a. In this method, carboxy tetramethylrhodamine (TAMRA) is linked as the fluorophore at the 5' ends of ssDNA reporter probes and Black Hole Quencher-2 (BHQ2) as the quencher at the 3' ends. The detailed sequence of crRNA and ssDNA reporter are described in Table 5.
- Prepare the Lachnospiraceae bacterium Cas12a (LbCas12a) protein with 10x CRISPR/Cas12a Reaction Buffer.
- Dissolve 1 µL of RPA reaction product from section 2 in 1x CRISPR/Cas12a reaction buffer with LbCas12a-crRNA complexes and 500 nM ssDNA reporter probe in a 100 µL reaction volume.
- After mixing LbCas12a and crRNA, let the mixture sit for at least 5 min to form a functional complex. After incubation, add other components to the reaction mixture and complete the reaction at 37 °C. The final volume of each CRISPR/Cas12a reaction is 100 µL. The detailed concentrations of each component in each CRISPR/Cas12a reaction are described in Table 6.
- Perform the 100 µL CRISPR/Cas12a detection reaction on the pre-treated glass slide and cover it with a coverslip. Incubate the glass slide with reaction at RT for 10 min.
- Measure the fluorescence signals by the SPM. Put the glass slide with the detection reaction on the stage of the SPM, maintain an appropriate distance, adjust the focal length and clarity, then look for the field of view of the reaction and focus it to capture an image.
NOTE: A standard curve should be obtained first so that the data from the samples can be scaled to the approximate concentration range. Various concentrations of purified targets are used, including 10 nM, 1 nM, 100 pM, and 10 pM, another virus as negative control (before RPA).
7. Dataset and data augmentation
- Collect the fluorescence images from the detection assay in section 6 as the datasets. Repeat at least three parallel detections for each sample to ensure data parallelism.
- Some appropriate ways to achieve higher parallelism can be approved. For example, when collecting the fluorescence images from each sample, manually focus and look for a relatively brighter field to take images. At the same time, photograph each sample to attain the fluorescence signal in more than five different locations.
- Measure the mean grey value of each image and the standard deviation of the mean gray value in a concentration group by ImageJ.
- Set an intensities range [median – standard deviation, median + standard deviation] for data cleaning.
NOTE: In the images obtained in the early steps, there may be images with large differences, so it is necessary to screen the images. If the images' intensities are out of the set threshold, they should be considered outliers and excluded. - Label the images for the purified target at increasing concentrations with 0-6 in ascending order, respectively.
- To enhance the system's robustness and prevent over-fitting, implement image augmentation techniques such as horizontal flipping, vertical flipping, and random noise through transform functions in Python. This helps to introduce variations in the dataset.
8. Transfer learning
- As the backbone network, adopt the deep learning model AlexNet33 for classification.
- To satisfy the constraint of the pre-trained model in section 7, reshape the input images to 224 pixels x 224 pixels x 3 channels (height and width of 224 pixels and a depth of 3 channels for the red, green, and blue color channels) through transform functions in Python.
NOTE: This step is a common preprocessing for heterogeneous data in transfer learning, including transform. - Use a pre-trained backbone network with the ImageNet dataset to extract features while leveraging the weights of intermediate hidden layers that were learned.
- In the fluorescence classification job context, substitute the final fully connected layer of the neural network, which originally included 1000 neurons for the ImageNet task, with a fully connected layer with either 2 or 7 neurons.
- Evaluate the performance of the setup training model by using a series of metrics, including confusion matrix, accuracy, precision, recall, and F1-score based on Lawton and Viriri34.
Representative Results
This method focuses on a fast, easy-to-implement, highly sensitive, and point-of-care (POC) detection system for DNA viruses. Primer pairs design for the RPA reaction and crRNA design for CRISPR/Cas12a reaction are two of the essential parts since they will affect the efficiency of the RPA-CRISPR/Cas12a reaction and influence the subsequent detection and classification.
In this method, FV3 is regarded as an example of DNA virus detection. Some RPA primer pairs for FV3 are designed, and the most efficient one is picked and made sure that it shows good amplification efficiency, which can be confirmed by ensuring the brightness and size of bands through gel electrophoresis of the RPA products as mentioned in step 2.8. At the same time, we have designed CRISPR/Cas12a crRNAs and chose the most efficient one based on the stronger fluorescence signal triggered in CRISPR/Cas12a reaction without RPA. The sequence of the three forward and three backward primers for RPA and three crRNA for Cas12a are shown in Table 7. Based on the agarose gel electrophoresis results shown in Figure 3, the 6th pair of primers gives better amplification efficiency (F2 and R1), which is selected for the method. As shown in Figure 4, crRNA-3 is the most efficient for collateral cleavage, which is selected in the method.
| RPA primer F1 | ATGTCTTCTGTAACTGGTTCAGGTATCACAAG | |
| RPA primer F2 | ATGTCTTCTGTAACTGGTTCAGGTATCACA | |
| RPA primer F3 | TCTTCTGTAACTGGTTCAGGTATCACAAGTGGT | |
| RPA primer R1 | GGCGTTGAGGATGTAATCCCCCGACCTGGG | |
| RPA primer R2 | GGCGTTGAGGATGTAATCCCCCGACCTGGGAA | |
| RPA primer R3 | CGTTGAGGATGTAATCCCCCGACCTGGGAACG | |
| LbCas12a crRNA-1 | uaauuucuacuaaguguagauATCGACTTGGCCACTTATGACAA | |
| LbCas12a crRNA-2 | uaauuucuacuaaguguagauTCAAGGAGCACTACCCCGTGGGG | |
| LbCas12a crRNA-3 | uaauuucuacuaaguguagauGGGCAGCAGTTTTCGGTCGGCGT | |
Table 7: The RPA primers and crRNA sequences for reaction optimization.
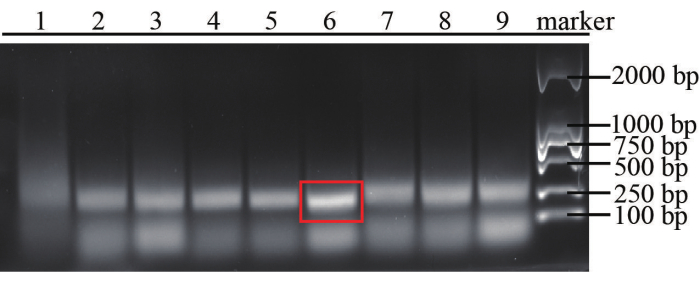
Figure 3: Results of agarose gel electrophoresis for assessment of RPA efficiency. The RPA reactions were carried out using different combinations of primers under the same condition with a 1 nM purified target. Number 1 was regarded as the combination of RPA-F3 and RPA-R3, 2 as RPA-F3 and RPA-R2, 3 as RPA-F3 and RPA-R1, 4 as RPA-F2 and RPA-R3, 5 as RPA-F2 and RPA-R2, 6 represented RPA-F2 and RPA-R1, 7 as RPA-F1 and RPA-R3, 8 as RPA-F1 and RPA-R2, 9 as RPA-F1 and RPA-R2. Please click here to view a larger version of this figure.
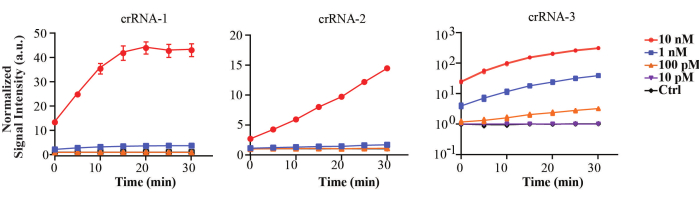
Figure 4: Detection of the FV3 target fragments with three different crRNAs by the CRISPR/Cas12a. The concentrations of purified fragments applied in the detection were from 10 nM to 100 pM target DNA versus 10 nM control DNA with 30 min incubation. This figure is reused with permission from Lei et al.35. Please click here to view a larger version of this figure.
To generate the data in Figure 5, we applied optimized RPA primers, crRNA, and the SPM setup. To get a better sense of the concentration in the samples, we amplified 240 bp DNA fragments of the highly conserved major capsid (MCP) gene from FV3 and Infectious spleen and kidney necrosis virus (ISKNV, Iridoviridae) as target and control through a PCR amplification kit, which is run with the procedures shown in Table 8. The size of the amplified DNA is confirmed through agarose gel electrophoresis, and we extracted the purified dsDNA target and control from the gel. The purified DNA fragments are quantified by using a NanoDrop 2000 spectrophotometer.
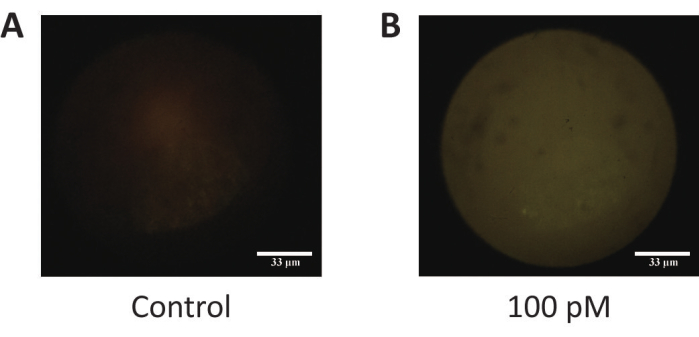
Figure 5: The fluorescence images of purified target DNA and control DNA. (A) The negative control. (B) Purified target DNA (100 pM). This figure is reused with permission from Lei et al.35. Please click here to view a larger version of this figure.
| Stage | Temperature (°C) | Time | Cycle(s) |
| Hot start | 95 | 5 min | 1 |
| Denaturation | 95 | 30 s | 35 |
| Annealing | 60 | 45 s | |
| Extension | 72 | 30 s | |
| Extension | 72 | 5 min | 1 |
| Storage | 4 | – | – |
Table 8: The procedure of the PCR amplification.
For reference only, a dataset including 162 fluorescence images is obtained after data cleaning. The number of fluorescence images for further analysis needs to be expanded, and we trained the dataset by using 337 fluorescence images for binary classification and 398 for multiclass classification.
We achieved detection within around 40 min, including 30 min for RPA amplification and 10 min for LbCas12a detection after RPA. When using the developed detection system with selected RPA primers and crRNA, the signal intensity of 10 aM FV3 and control shows significant differences, as shown in Supplementary Figure 1.
Supplementary Figure 1: The statistics of the signal intensity for fluorescence images collected in RPA-LbCas12a reactions with various concentrations of purified fragments. *: p < 0.05; **: p < 0.01; ***: p < 0.001; ****: p < 0.0001; F: FV3, I: ISKNV; a.u.: absorbance unit. This figure is reused with permission from Lei et al.35. Please click here to download this File.
Discussion
In this method, we develop a fast, easy-to-implement, highly sensitive, sequence-specific, and POC DNA virus detection system with AI assistance. After obtaining samples, RPA is applied to amplify the target sequence, and then CRISPR/Cas12a can recognize the target DNA and release fluorescence, which enlarges the detection signal. Portable smartphone microscopy is built to take fluorescence images, and deep learning models with transfer learning are used for binary classification of the positive and negative samples’ images. FV3 is taken as the positive sample in this method and then investigated from multiple perspectives to improve the specificity and sensitivity of the proposed system.
The first strategy is designing specific RPA primers and screening out the most efficient primers. The amplification efficiency of the RPA reaction is influenced by different primers36. The specificity of the primers used to amplify the target gene of the DNA virus strains can be confirmed by blasting against the standard database of nucleotide sequencing. Since there is no guide for designing efficient RPA primer pairs, experimental validation is required37.
Another approach is to design the crRNA of the target virus for CRISPR/Cas12a reaction since the collateral cleavage efficiency significantly contributes to the sensitivity of detection38,39. The strength of the triggered fluorescence signal in the CRISPR/Cas12a reaction without RPA needs to be considered in selecting the better crRNA. Besides, the collateral activity of the LbCas12a protein is constantly activated for 24 h32, which can benefit detection.
For the purpose of identifying viruses and visualizing viral particles, fluorescence microscopy has been applied40,41. However, conventional microscopy is cumbersome and pricey for quick POC diagnosis. Thus, in the proposed system, a portable SPM with less cost is built to attain the fluorescence signal by capturing the images for POC detection. Such portable SPM costs much less than a commercialized plate reader or real-time PCR machine, which is cost-friendly for detection system setup.
This proposed platform also needs to be further improved in some aspects. When applying this detection platform to other DNA virus detection, both RPA primer and Cas12a-crRNA sequence need to be redesigned. Since there is no guide for their design, experiments need to be carried out to evaluate their efficiency. Although the proposed system can meet the advantages of portability, the size of the whole system still needs to be further reduced to obtain a lighter one. Since the mobile phone camera itself is not easy to focus on for close-range shooting, it takes time to adjust during the process of obtaining fluorescent images, including the field of view, for images with better quality.
With advancements in related technology and new applications thereof, efficiency in POC detection will continue to improve. The method presented here is one example of an improvement that may help with POC detection based on the CRISPR/Cas12a system and SPM platform, with novel AI assistance. Additionally, the integrated system might be utilized to identify any DNA target due to the crRNA’s reprogrammability. The RPA-CRISPR/Cas12a-AI system will have a significant position in the identification of DNA pathogens in terms of detection sensitivity, specificity, time, and reliability.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This work is supported by the National Natural Science Foundation of China 31970752, Science, Technology, Innovation Commission of Shenzhen Municipality JCYJ20190809180003689, JSGG20200225150707332, JSGG20191129110812708, WDZC20200820173710001; Shenzhen Bay Laboratory Open Funding, SZBL2020090501004; China Postdoctoral Science Foundation 2020M680023; and General Administration of Customs of the People's Republic of China 2021HK007.
Materials
| 20x amplification | OLYMPUS | OPLN20X | |
| 532 nm green laser | Thorlabs | PL201 | with 0.9 mW output power |
| 535 nm cutoff wavelength | chrome | AT535 | |
| 6x DNA loading buffer | Thermo scientific | R0611 | |
| 96-well black microplate | Corning Incorporated | 3603 | Black with flat clear bottom |
| Aspherical lens | Lubang | N/A | |
| Bandpass filter | SEMROCK | FF01-542/27-25 | |
| Bsu DNA Polymerase | ATG Biotechnology | M103 | Large Fragment |
| crRNA | Sangon Biotech | N/A | |
| DNA fragments | Sangon Biotech | N/A | |
| Dichroic holders | Ruicage | N/A | |
| Dichroic mirror | SEMROCK | FF555-Di03-25×36 | with a cutoff wavelength of 535 nm |
| E.Z.N.A Gel Extraction Kit | Omega Biotek | D2500-02 | |
| EnGen Lba Cas12a (Cpf1) | New England Biolabs (Beijing) LTD | M0653T | |
| Filter holders | Ruicage | N/A | |
| Fluorophore-ssDNA-Quencher reporter probes | Sangon Biotech | N/A | TAMRA (carboxy tetramethylrhodamine) as the fluorophore at the 5 ends; BHQ2 (Black Hole Quencher-2) as the quencher at the 3 ends |
| GP32 | ATG Biotechnology | M104 | |
| ImageJ | Open-source | Version 1.53t 24 | Downloaded from https://imagej.nih.gov/ij/ |
| Microplate reader | SPARK, TECAN | N/A | |
| Multi-Block thermal Cycler PCR instrument | LongGene | N/A | |
| NanoDrop 2000/2000c Spectrophotometers | Thermo Scientific | ND-2000 | |
| NEBuffer r2.1 | New England Biolabs (Beijing) LTD | B6002S | 10x CRISPR/Cas12a Reaction buffer |
| Oxygen plasma treatment | Electro-Technic Products | N/A | |
| Pathogen Inactivate, Nucleic acid extraction-free, Direct-to-PCR Buffer with Proteinase K (PINDBK) | Ebio | PINDBK -25mL | |
| PCR primer pairs | Sangon Biotech | N/A | |
| PDMS | Dow Corning | Sylgard 184 | |
| RPA primer pairs | Sangon Biotech | N/A | |
| Smartphone | Huawei | Mate10 | |
| Translation stages | Ruicage | N/A | |
| Transmitted neutral density filters | Thorlabs | ND40A | |
| Triplet achromatic lenses | Thorlabs | TRH127-020-A | |
| UvsX | ATG Biotechnology | M105 | |
| UvsY | ATG Biotechnology | M106 |
Riferimenti
- Gire, S. K., et al. Genomic surveillance elucidates Ebola virus origin and transmission during the 2014 outbreak. Science. 345 (6202), 1369-1372 (2014).
- The Ebola Outbreak Epidemiology Team. Outbreak of Ebola virus disease in the Democratic Republic of the Congo, April-May 2018: an epidemiological study. Lancet. 392 (10143), 213-221 (2018).
- Zumla, A., Hui, D. S., Perlman, S. Middle East respiratory syndrome. Lancet. 386 (9997), 995-1007 (2015).
- Plourde, A. R., Bloch, E. M. A literature review of Zika virus. Emerg Infect Dis. 22 (7), 1185-1192 (2016).
- Yuan, X., et al. Current and perspective diagnostic techniques for COVID-19. ACS Infect Dis. 6 (8), 1998-2016 (2020).
- Minhaj, F. S., et al. Monkeypox outbreak – nine states, May 2022. MMWR Morb Mortal Wkly Rep. 71 (23), 764-769 (2022).
- Bao, M., et al. Challenges and opportunities for clustered regularly interspaced short palindromic repeats based molecular biosensing. ACS Sens. 6 (7), 2497-2522 (2021).
- Broughton, J. P., et al. CRISPR-Cas12-based detection of SARS-CoV-2. Nat Biotechnol. 38 (7), 870-874 (2020).
- Chen, J. S., et al. CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science. 360 (6387), 436-439 (2018).
- Gootenberg, J. S., et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science. 356 (6336), 438-442 (2017).
- Kellner, M. J., Koob, J. G., Gootenberg, J. S., Abudayyeh, O. O., Zhang, F. SHERLOCK: nucleic acid detection with CRISPR nucleases. Nat Protoc. 14 (10), 2986-3012 (2019).
- Mukama, O., et al. An ultrasensitive and specific point-of-care CRISPR/Cas12 based lateral flow biosensor for the rapid detection of nucleic acids. Biosens Bioelectron. 159, 112143 (2020).
- Schwank, G., et al. Functional repair of CFTR by CRISPR/Cas9 in intestinal stem cell organoids of cystic fibrosis patients. Cell Stem Cell. 13 (6), 653-658 (2013).
- Yin, L., Man, S., Ye, S., Liu, G., Ma, L. CRISPR-Cas based virus detection: Recent advances and perspectives. Biosens Bioelectron. 193, 113541 (2021).
- Dronina, J., Bubniene, U. S., Ramanavicius, A. The application of DNA polymerases and Cas9 as representative of DNA-modifying enzymes group in DNA sensor design (review). Biosens Bioelectron. 175, 112867 (2021).
- Fozouni, P., et al. Amplification-free detection of SARS-CoV-2 with CRISPR-Cas13a and mobile phone microscopy. Cell. 184 (2), 323-333.e9 (2021).
- Kumar, M., et al. FnCas9-based CRISPR diagnostic for rapid and accurate detection of major SARS-CoV-2 variants on a paper strip. eLife. 10, e67130 (2021).
- Lee, R. A., et al. Ultrasensitive CRISPR-based diagnostic for field-applicable detection of Plasmodium species in symptomatic and asymptomatic malaria. Proc Natl Acad Sci U S A. 117 (41), 25722-25731 (2020).
- Ganguli, A., et al. Hands-free smartphone-based diagnostics for simultaneous detection of Zika, Chikungunya, and Dengue at point-of-care. Biomed Microdevices. 19 (4), 73 (2017).
- Yeo, S. J., et al. Smartphone-based fluorescent diagnostic system for highly pathogenic H5N1 viruses. Theranostics. 6 (2), 231-242 (2016).
- von Chamier, L., et al. Democratising deep learning for microscopy with ZeroCostDL4Mic. Nat Commun. 12 (1), 2276 (2021).
- Shiaelis, N., et al. Virus detection and identification in minutes using single-particle imaging and deep learning. ACS Nano. 17 (1), 697-710 (2020).
- Liu, Y., et al. Mixed-UNet: Refined class activation mapping for weakly-supervised semantic segmentation with multi-scale inference. Front. Comput. Sci. 4, 1036934 (2022).
- Lawrimore, J., Doshi, A., Walker, B., Bloom, K. AI-assisted forward modeling of biological structures. Front Cell Dev Biol. 7, 279 (2019).
- Yang, Y., Hu, Y., Zhang, X., Wang, S. Two-stage selective ensemble of CNN via deep tree training for medical image classification. IEEE Trans Cybern. 52 (9), 9194-9207 (2022).
- Zhang, R., et al. RCMNet: A deep learning model assists CAR-T therapy for leukemia. Comput Biol Med. 150, 106084 (2022).
- Xie, Y., et al. Stroke prediction from electrocardiograms by deep neural network. Multimed Tools Appl. 80, 17291-17297 (2021).
- Wang, J., Zhu, H., Wang, S., Zhang, Y. -. D. A review of deep learning on medical image analysis. Mobile Netw Appl. 26, 351-380 (2021).
- Artoni, P., et al. Deep learning of spontaneous arousal fluctuations detects early cholinergic defects across neurodevelopmental mouse models and patients. Proc Natl Acad Sci U S A. 117 (38), 23298-23303 (2020).
- Li, J., et al. DeepLearnMOR: a deep-learning framework for fluorescence image-based classification of organelle morphology. Plant Physiol. 186 (4), 1786-1799 (2021).
- Yosinski, J., Clune, J., Bengio, Y., Lipson, H. How transferable are features in deep neural networks. Proceedings of the 27th International Conference on Neural Information Processing Systems. 2, 3320-3328 (2014).
- He, Q., et al. High-throughput and all-solution phase African Swine Fever Virus (ASFV) detection using CRISPR-Cas12a and fluorescence based point-of-care system. Biosens Bioelectron. 154, 112068 (2020).
- Krizhevsky, A., Sutskever, I., Hinton, G. E. ImageNet classification with deep convolutional neural networks. Commun. ACM. 60 (6), 84-90 (2017).
- Lawton, S., Viriri, S. Detection of COVID-19 from CT lung scans using transfer learning. Comput Intell Neurosci. 2021, 5527923 (2021).
- Lei, Z., et al. Detection of frog virus 3 by integrating RPA-CRISPR/Cas12a-SPM with deep learning. ACS Omega. 8 (36), 32555-32564 (2023).
- Chen, Z., Huang, J., Zhang, F., Zhou, Y., Huang, H. Detection of shrimp hemocyte iridescent virus by recombinase polymerase amplification assay. Mol Cell Probes. 49, 101475 (2020).
- Fu, X., Sun, J., Ye, Y., Zhang, Y., Sun, X. A rapid and ultrasensitive dual detection platform based on Cas12a for simultaneous detection of virulence and resistance genes of drug-resistant Salmonella. Biosens Bioelectron. 195, 113682 (2022).
- Habimana, J. D., et al. Mechanistic insights of CRISPR/Cas nucleases for programmable targeting and early-stage diagnosis: A review. Biosens Bioelectron. 203, 114033 (2022).
- Liang, Y., Lin, H., Zou, L., Deng, X., Tang, S. Rapid detection and tracking of Omicron variant of SARS-CoV-2 using CRISPR-Cas12a-based assay. Biosens Bioelectron. 205, 114098 (2022).
- Sivaraman, D., Biswas, P., Cella, L. N., Yates, M. V., Chen, W. Detecting RNA viruses in living mammalian cells by fluorescence microscopy. Trends Biotechnol. 29 (7), 307-313 (2011).
- Wang, I. H., Burckhardt, C. J., Yakimovich, A., Greber, U. F. Imaging, tracking and computational analyses of virus entry and egress with the cytoskeleton. Viruses. 10 (4), 166 (2018).

