Investigation of Beige Fat Biology and Metabolism Using the CRISPR SunTag-p65-HSF1 Activation System
Summary
This protocol presents the use of CRISPR SunTag-p65-HSF1 (SPH) in adipocytes (AdipoSPH) as an alternative strategy to adeno-associated virus (AAV) for investigating beige fat biology. In vivo injection of AAV-carrying sgRNA targeting the endogenous Prdm16 gene is sufficient to induce beige fat development and enhance the thermogenic gene program.
Abstract
Clustered regularly interspaced short palindromic repeats (CRISPR) technology has prompted a revolution in biology, and recent tools have been applied far beyond the originally described gene editing. The CRISPR activation (CRISPRa) system combines the catalytically inactive Cas9 (dCas9) protein with distinct transcription modules to induce endogenous gene expression. SunTag-p65-HSF1 (SPH) is a recently developed CRISPRa technology that combines components of synergistic activation mediators (SAMs) with the SunTag activators. This system allows the overexpression of single or multiple genes by designing a customized single-guide RNA (sgRNA). In this study, a previously developed SPH mouse was used to generate a conditional mouse expressing SPH in adipocytes (adiponectin Cre lineage), named AdipoSPH. To induce a white-to-beige fat (browning) phenotype, an adeno-associated virus (AAV) carrying sgRNA targeting the endogenous Prdm16 gene (a well-established transcription factor related to brown and beige fat development) was injected into the inguinal white adipose tissue (iWAT). This mouse model induced the expression of endogenous Prdm16 and activated the thermogenic gene program. Moreover, in vitro SPH-induced Prdm16 overexpression enhanced the oxygen consumption of beige adipocytes, phenocopying the results of a previous Prdm16 transgenic mouse model. Thus, this protocol describes a versatile, cost-effective, and time-effective mouse model for investigating adipose tissue biology.
Introduction
Beige (or brite) adipocytes are uncoupling protein 1 (UCP1)-expressing and mitochondrial-rich adipocytes that reside within white adipose tissue (WAT) depots. Beige fat emerges from a subset of adipocyte progenitors or mature white adipocytes in response to cold exposure and other stimuli1,2. Beige adipocytes can convert energy into heat in a UCP1-dependent or independent manner3. Regardless of its thermogenic function, beige fat can also improve metabolic health by other means, such as the secretion of adipokines and anti-inflammatory and anti-fibrotic activities. Studies in mice and humans have shown that the induction of beige fat improves whole-body glucose and lipid homeostasis3. However, although our knowledge of beige fat biology has evolved rapidly in recent years, most of its metabolic benefits and related mechanisms are still not fully understood.
Clustered regularly interspaced short palindromic repeats (CRISPR) were first described in eukaryotic cells as a tool capable of generating a double-strand break (DSB) at a specific site in the genome through the nuclease activity of the Cas9 protein4,5. Cas9 is guided by a synthetic single-guide RNA (sgRNA) to target a specific genomic region, leading to a DNA DSB. In addition to using the nuclease Cas9 for editing purposes, CRISPR-Cas9 technology has evolved to be used as a sequence-specific gene regulation tool6. The development of a catalytically inactive Cas9 protein (dCas9) and the association of transcriptional modules capable of enhancing gene expression has given rise to CRISPR activation (CRISPRa) tools. Several CRISPRa systems have emerged, such as VP647,8, synergistic activation mediator (SAM)9, SunTag10,11, VPR12,13, and SunTag-p65-HSF1 (SPH)14, which combines the components of SAM and SunTag activators. It has recently been demonstrated that the induced expression of neurogenic genes in N2a neuroblasts and primary astrocytes is higher using SPH compared to other CRISPRa systems14, demonstrating SPH as a promising CRISPRa tool.
Here, we took advantage of a previously developed SPH mouse14 to generate a conditional mouse model expressing SPH specifically in adipocytes using the adiponectin Cre lineage (AdipoSPH). Using an adeno-associated virus (AAV) carrying the gRNA targeting the endogenous Prdm16 gene, browning (white to beige conversion) of inguinal WAT (iWAT) was induced to increase the expression of the thermogenic gene program. Moreover, in vitro Prdm16 overexpression enhanced oxygen consumption. Therefore, this protocol provides a versatile SPH mouse model for exploring the mechanisms of beige fat development within adipose tissue.
Protocol
Animal studies were performed in accordance with the University of Campinas Guide for the Care and Use of Laboratory Animals (protocol CEUA #5810-1/2021).
1. Molecular cloning
- Design of single guide RNAs (sgRNAs)
- Design sgRNAs for CRISPR activation using CHOPCHOP, available at https://chopchop.cbu.uib.no/, or any other suitable tool. Use the following parameters to design sgRNA targeting the Prdm16 gene: Target: Prdm16; In: Mus musculus; Using: Crispr/Cas9; For: Activation.
NOTE: Design sgRNAs for each region of interest spread over a 200 bp upstream transcription start site (TSS) region. For instance, the sgRNA targeting Prdm16 used in this study binds 154 bp upstream of TSS. - Add overhangs to the sgRNA to match the SacI restriction site in the vector backbone pAAV-U6-gRNA-CBh-mCherry (see Table of Materials). Include: 5'- (N20)AGCT-3' (N = nucleotides). For example, the sequence targeting the Prdm16 gene is 5'- CGAGCTGCGCTGAAAAGGGG-3', and with overhangs is 5'- CGAGCTGCGCTGAAAAGGGGAGCT-3'.
- Obtain the 3' sgRNA reverse complement sequence using the tool available at https://arep.med.harvard.edu/labgc/adnan/projects/Utilities/revcomp.html. For example, the 3’ sgRNA sequence targeting the Prdm16 gene is 3’- TCGAGCTCGACGCGACTTTTCCCC-5’.
- Design sgRNAs for CRISPR activation using CHOPCHOP, available at https://chopchop.cbu.uib.no/, or any other suitable tool. Use the following parameters to design sgRNA targeting the Prdm16 gene: Target: Prdm16; In: Mus musculus; Using: Crispr/Cas9; For: Activation.
- Annealing of single-stranded complementary oligonucleotides
- Add 1 µL of each 5' and 3' single-stranded oligonucleotide (stock concentration: 100 µM), 1 µL of T4 ligase buffer, 0.5 µL of T4 polynucleotide kinase (PNK) (1 × 104 units/mL), and 6.5 µL of H2O to a final reaction volume of 10 µL. Anneal the complementary single-stranded oligonucleotides using a thermocycler under the following conditions: 37 °C for 30 min and 95 °C for 5 min, followed by a ramp-down rate of 5 °C/min.
NOTE: The PNK enzyme is supplied with PNK buffer and does not contain enough ATP required for the phosphorylation reaction (see Table of Materials). To simplify the reaction, use the T4 ligase buffer (instead of PNK buffer). T4 ligase buffer provides the appropriate amount (1 mM ATP) of phosphate for the phosphorylation reaction. The PNK enzyme provides 5' end phosphorylation of oligonucleotides for the subsequent ligation reaction.
- Add 1 µL of each 5' and 3' single-stranded oligonucleotide (stock concentration: 100 µM), 1 µL of T4 ligase buffer, 0.5 µL of T4 polynucleotide kinase (PNK) (1 × 104 units/mL), and 6.5 µL of H2O to a final reaction volume of 10 µL. Anneal the complementary single-stranded oligonucleotides using a thermocycler under the following conditions: 37 °C for 30 min and 95 °C for 5 min, followed by a ramp-down rate of 5 °C/min.
- Ligation of annealed sgRNA oligonucleotides
- Add 25 ng of plasmid pAAV-U6-gRNA-CBh-mCherry to 2 µL of annealed sgRNA oligonucleotides, 1 µL of SacI enzyme, 2 µL of 10x T4 DNA ligase buffer, 1 µL of T4 DNA ligase (1-3 u/µL) (see Table of Materials), and H2O to a final reaction volume of 10 µL.
- Perform ligation by incubating the reaction mixture using a thermocycler under the following conditions: 15 cycles of 37 °C for 5 min and 25 °C for 5 min, followed by holding at 4 °C.
- Transformation followed by colony polymerase chain reaction (PCR)
- Transform the competent E. coli DH10B cells (see Table of Materials) with 4 µL of the ligation product using heat shock (42 °C for 45 s) and spread on an agar plate containing 100 µg/mL ampicillin.
- Confirm transformed colonies by colony PCR using the PCR master mix (see Table of Materials). Pick the colony and mix with 5 µL of master mix, 0.1 µL of universal primer (stock concentration: 100 µM), 0.1 µL of sgRNA reverse primer (stock concentration: 100 µM) (Table 1), and 5 µL of H2O. Run the PCR using a thermocycler under the following conditions: initial denaturation (94 °C for 2 min), followed by 35 cycles of denaturation (94 °C for 20 s), annealing (60 °C for 30 s), extension (72 °C for 30 s), and a final elongation step (72 °C for 5 min). Resolve the DNA using agarose (1.5%) gel electrophoresis in 0.5x TAE buffer at 90 V for 30 min.
NOTE: Positive clones give a band of ~280 bp. - Submit positive samples for Sanger sequencing using universal primer (Table 1, Supplemental File 1).
- Plasmid purification
- Purify the plasmid from a positive clone using a plasmid purification kit (see Table of Materials), following the manufacturer's instructions.
NOTE: Purify the plasmid using anion-exchange resin or another kit suitable for use in cell transfection. - Incubate the positive clone (12 h at 37 °C with shaking at 200 rpm) using a standard bacterial growth medium containing 100 µg/mL ampicillin.
- Centrifuge the bacterial cells at 6,000 × g for 10 min at 4 °C. Follow the next steps according to the manufacturer's kit instructions.
- Purify the plasmid from a positive clone using a plasmid purification kit (see Table of Materials), following the manufacturer's instructions.
2. AAV packaging
NOTE: AAV packaging was performed according to previous publications15,16 with minor modifications.
- Plate 293T cells (see Table of Materials) in a 175 cm2 flask (seeding 5 × 106 cells per flask) using 25 mL of Dulbecco's Modified Eagle's Medium (DMEM) containing 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin (P/S). Incubate at 37 °C, 5% CO2, and 95% humidity until the cells reach 50%-70% confluence.
- Mix 14 µL of polyethylenimine (1 µg/µL) with 1 mL of 150 mM NaCl. Mix the three plasmids required for AAV production in a 1:1:1 molar ratio (i.e., 17.7 µg of pAdDeltaF6, 7.9 µg of AAV2/8 (see Table of Materials), and 5.9 µg of cloned sgRNA from step 1 in 1 mL of 150 mM NaCl. Transfer the polyethylenimine:NaCl mixture drop by drop to the tube containing the DNA (mixture of plasmids) and incubate for 20 min at 25 °C.
NOTE: pAdDeltaF6 is a Helper plasmid and pCapsid pAAV2/8 is a packaging plasmid expressing Replication (Rep) and Capsid (cap) genes. Both plasmids are necessary to produce AAV. - Before transfection, replace the cell growth medium with 18 mL of DMEM containing 1% FBS and L-alanyl-L-glutamine (0.5 g/L). Add 2 mL of the polyethylenimine:DNA mixture to each culture flask and incubate the cells in a CO2 incubator (37 °C, 5% CO2, 95% humidity).
- After 5 h, add 5 mL of DMEM supplemented with 10% FBS and L-alanyl-L-glutamine (0.5g/L).
- After 3 days of incubation, detach the cells from the culture flask using a cell scraper. Collect the 293T cells from 10 (175 cm2) cell culture flasks into 50 mL conical tubes and add DMEM (see Table of Materials) up to 30 mL.
- Add 3 mL of chloroform to each 50 mL conical tube containing the 293T cells and mix using a vortex mixer at high speed for 5 min. Resuspend the cells by adding 7.6 mL of 5 M NaCl and vortex briefly. Centrifuge at 3,000 × g and 4 °C for 5 min.
- Transfer the aqueous phase into a new conical tube and add 9.4 mL of 50% (v/v) polyethylene glycol (PEG) 8000. Mix with a vortex mixer at high speed for 10 s and put the samples on ice for 1 h.
- Centrifuge at 3,000 × g at 4 °C for 30 min. Remove the supernatant and allow the pellet to dry for 10 min.
- Add 1.4 mL of HEPES (50 mM, pH 8) and mix for 5 min using a vortex mixer at high speed. Add 3.5 µL of 1 M MgCl2, 14 µL of DNase I (20 units/µL), and 1.4 µL of RNase A (10 µg/µL). Incubate in a 37 °C water bath for 20 min, and then transfer the samples into new 1.5 mL tubes (700 µL in each tube).
- Add 700 µL of chloroform to each 1.5 mL tube and mix using a vortex mixer at high speed for 10 s. Centrifuge at 3,000 × g at 4 °C for 5 min and transfer the aqueous phase to a new tube. Repeat this step 3x.
- Evaporate the chloroform for 30 min in a biosafety cabinet. Then, transfer 300 µL of the aqueous phase into a 0.5 mL ultra-centrifugation tube (see Table of Materials). Spin the filter at 14,000 × g at 25 °C for 5 min.Remove the flow-through and spin the filter again until the entire volume has passed through the filter.
- Wash the filter by adding 300 µL of Dulbecco's phosphate-buffered saline (DPBS) and mix the solutions by pipetting. Centrifuge the filter for 5 min at 14,000 × g at 25 °C. Repeat this washing step 4x.
- Centrifuge the filter for 8 min at 14,000 × g at 25 °C. Place the filter upside down into a new tube, and spin for 2 min at 1,000 × g at 25 °C.
3. Titration of the AAV by qPCR
- Prepare a standard curve for AAV titration and determine the AAV titer according to the original study by Fripont et al.15.
4. In vivo injection of AAV into the inguinal white adipose tissue (iWAT)
- Separate the surgical tools and supplies needed for surgery and sterilize them as recommended for each specific material.
- Anesthetize the mouse with 100 mg/kg ketamine and 10 mg/kg xylazine by intraperitoneal injection. Confirm anesthesia by applying vigorous pressure on the paw and tail and checking for a reflex. Apply ointment to the mouse's eyes to avoid eye dryness during the surgery.
- Place the anesthetized mouse in the supine position and shave a small area on the flanks, proximal to the hip joints for iWAT injections, with a shaver. Apply depilatory cream for 5 min. Remove residual cream with water to avoid skin burn.
- Disinfect the skin using three alternating rounds of applying antiseptic solution (povidone-iodine) on the skin with a clean gauze and 70% alcohol. Discard the gauze after each use.
- Perform a final application of an antiseptic solution on the skin. Then, make a 1-2 cm incision with sterilized scissors in the proximal area of the joints, and hold the skin open using forceps to expose the fat depot. The WAT can be found attached to the skin on both sides, extending from the beginning on the back and down toward the testis.
NOTE: Use drapes to avoid contamination during surgery or suture procedures. - Using forceps, through the incision, gently pull the fat depot upward to ensure the injection is at the correct location and depth.
NOTE: Be careful not to remove the tissue from its original location. - Fill the microliter syringe (gauge: 33; point style: 4; angle: 12; length: 10) (see Table of Materials) with 2.5 µL (5.6 × 1010 viral genomes [VG]/µL) of the AAV (containing the sgRNA targeting the endogenous Prdm16 gene). Carefully insert the needle at a 30°-45° angle into the iWAT. Repeat the injection 5x into different locations of the tissue to homogeneously infect the whole iWAT fat pad. A total volume of 15 µL is recommended to infect the iWAT.
NOTE: The depth of insertion of the syringe depends on the thickness of the fat pad deposit. Use the no-touch technique to draw up the injection. - Close the shaved skin incision using 4/0 monofilament sutures. Place the mouse on a heat pad until consciousness is regained. Monitor the animal every 10-15 min until it fully recovers. After the animal regains consciousness, observe the locomotor profiling, which should be linear and have no signs of distress, or pain.
- Perform postoperative pain control within 48 h after surgery by administering tramadol hydrochloride (5 mg/kg) intraperitoneally for 3 days (2x/day).
NOTE: Watch for signs of distress and discomfort and monitor water and food intake. Give an effective dose of analgesic in a preemptive manner, preferably before or at the beginning of the surgery. - Keep the mouse in a cage with free access to food and water during the healing period. After the healing period, proceed to euthanize the mouse. In this study, euthanasia was performed by an overdose of injectable anesthetics (intraperitoneal) from 3x the inducing dose (300-360 mg/kg ketamine hydrochloride + 30-40 mg/kg xylazine hydrochloride) followed by decapitation.
NOTE: It is strongly recommended to keep the animals housed in individual cages until fully recovered from anesthesia.
5. In vitro differentiation of stromal vascular cells (SVFs) into beige adipocytes
- Perform isolation and plating of primary SVFs from the iWAT of AdipoSPH mice according to Aune et al.17. Seed SVFs derived from AdipoSPH mice iWAT into a 6-well plate containing complete medium (DMEM containing 3.1 g/L glucose, 0.5 g/L L-alanyl-L-glutamine, 10% FBS, and 2.5% P/S) for 1-2 h.
NOTE: The SVF fraction contains a mixture of different cell types. At this step, it is not possible to define the number of seeded progenitor cells that give rise to adipocytes. - Aspirate the medium, wash the well 2x using phosphate-buffered saline (1x PBS), and replace with fresh complete medium. Incubate the cells at 37 °C, 5% CO2, 95% humidity until the cells reach 70%-80% confluency.
- Induce differentiation (day 0) by treating the cells with the induction medium (Table 2).
NOTE: The drug cocktail of the induction medium is necessary to enhance beige adipocyte differentiation and for expression of the thermogenic gene program17. - After 2 days (day 2), replace the induction medium with the maintenance medium (Table 2).
- After 2 days (day 4), replace the maintenance medium with fresh maintenance medium (Table 2) for 2 to 3 days.
- Change the maintenance medium every 48 h until the preadipocytes are fully differentiated into adipocytes (typically 6 days after the addition of induction medium). Mature adipocytes can be observed using light microscopy, as the differentiated cells appear to be loaded with lipid droplets.
6. In vitro AAV infection of SVFs
NOTE: SVFs derived from AdipoSPH mice iWAT were infected with AAV-carrying sgRNA-Prdm16 as previously described by Wang et al.18 with a few modifications.
- Grow the cells on a 6-well culture plate with complete medium until the cells reach 70%-80% confluence, as previously described in steps 5.1-5.3.
- Mix 5.6 × 1010 VG/µL of AAV-carrying sgRNA-Prdm16 with 2 mL of complete medium and hexadimethrine bromide (8 µg/mL) (see Table of Materials). Transduce the cells by replacing the complete medium and adding the complete medium containing AAV. Incubate the transduced cells for 12 h at 37 °C, 95% humidity, and 5% CO2.
- Split and seed the cells as described in step 5 for cell proliferation and differentiation into beige adipocytes.
NOTE: For the oxygen consumption assay, seed 4.0 × 104 cells (from step 6.2) per well with induction medium in a 24-well cell culture plate. The subsequent steps of cell proliferation and differentiation are performed as described in step 5. The oxygen consumption assay is performed when the cells reach 80%-100% confluence and are fully differentiated, as previously reported19.
Representative Results
AdipoSPH mice were developed by breeding SPH and Adipoq-Cre mouse strains. Both mouse strains were in a hybrid C57BL6J-DBA/2J background (according to the commercial supplier; see Table of Materials). The SPH mouse lineage was originally described by Zhou et al.14.
In vivo beige adipocyte development through AdipoSPH-mediated Prdm16 overexpression
To evaluate the capacity of the model described in this study to develop beige adipocytes in vivo, the AAV carrying the sgRNA targeting Prdm16 gene was injected into the iWAT of AdipoSPH mice. Prdm16 is a well-established transcription factor that determines beige adipocyte development and function20,21. It is important to mention that here we chose a previously tested and validated sgRNA targeting the endogenous Prdm16 gene14. AAV carrying an empty sgRNA was used as a control. At 10 days after AAV injection, iWAT was harvested for histological and gene expression analyses (Figure 1A). As expected, immunofluorescence images of iWAT from AdipoSPH mice demonstrated the expression of dCas9 in both the control and Prdm16 groups (Figure 1B). Additionally, mCherry expression confirmed the success of AAV infection of iWAT in both the control and Prdm16 groups (Figure 1B). SPH-induced Prdm16 expression clearly induced a widespread accumulation of multilocular beige adipocytes into iWAT (Figure 1B). Moreover, quantitative PCR (qPCR) revealed an increased expression of Prdm16 and the thermogenic gene program (Ucp1, Cox8b, Ppargc1a, and Cidea) in the Prdm16 group compared to the control (Figure 1C).
Promotion of beige adipocyte development and enhancement of oxygen consumption in vitro by AdipoSPH-mediated Prdm16 overexpression
Next, the suitability of this model for studying beige adipocyte biology in vitro was investigated. To this end, preadipocytes derived from the iWAT of AdipoSPH mice were transduced with AAV carrying the sgRNA sequence targeting Prdm16. At seven days after differentiation, beige adipocytes were used for the analysis of gene expression and oxygen consumption (Figure 2A). Empty sgRNA was used as a control. It is worth noting that the removal of STOP codon by CRE recombinase and activation of SPH machinery was under the control of the adiponectin promoter/enhancer, resulting in the activation of the endogenous Prdm16 gene in mature adipocytes. Gene expression analysis confirmed the increased expression of the endogenous Prdm16 gene and thermogenic genes in the Prdm16 overexpression group compared to the control (Figure 2B). To confirm that the SPH-induced beige adipocytes were functionally thermogenic, a high-resolution respirometry assay was performed on the primary adipocytes. The respirometry data analysis and interpretation was performed as recently described22. SPH-induced Prdm16 expression resulted in higher basal and maximal oxygen consumption than that in control cells (Figure 2C). Importantly, the data indicated enhanced uncoupled respiration (oligomycin-insensitive) in the Prdm16 group compared with that in control group (Figure 2C, Table 3). Taken together, these results revealed that SPH-induced expression of endogenous Prdm16 recapitulates the browning phenotype and enhances oxygen consumption rates observed in conventional Prdm16 Tg mice.
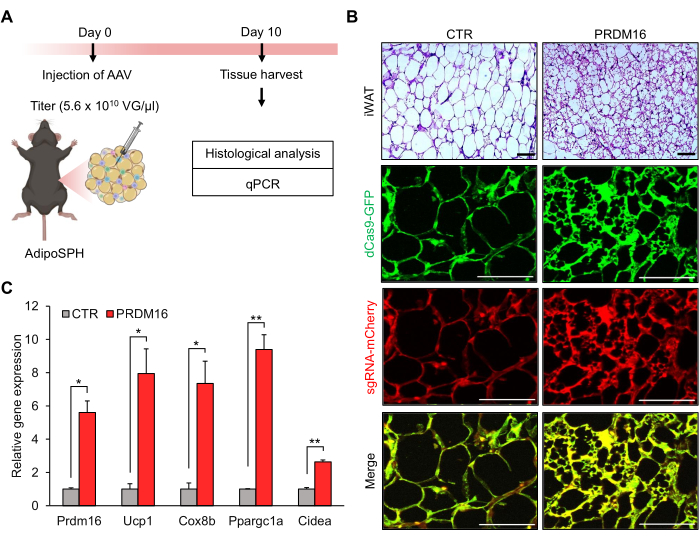
Figure 1: Beige fat development by injection of adeno-associated virus (AAV) targeting endogenous Prdm16 gene into inguinal white adipose tissue (iWAT) of AdipoSPH mice. (A) Schematic illustration of the in vivo beige adipocyte experimental design (created using Biorender.com). (B) Histological (hematoxylin and eosin [H&E] staining) images of iWAT. Scale bar = 50 µm. (C) Quantitative PCR (qPCR) of Prdm16 and thermogenic genes (Ucp1, Cox8b, Ppargc1α, and Cidea; n = 3). Data are presented as mean ± SEM. *p < 0.05; **p < 0.01 for Prdm16 versus control (CTR) by unpaired Student's t-test. Please click here to view a larger version of this figure.
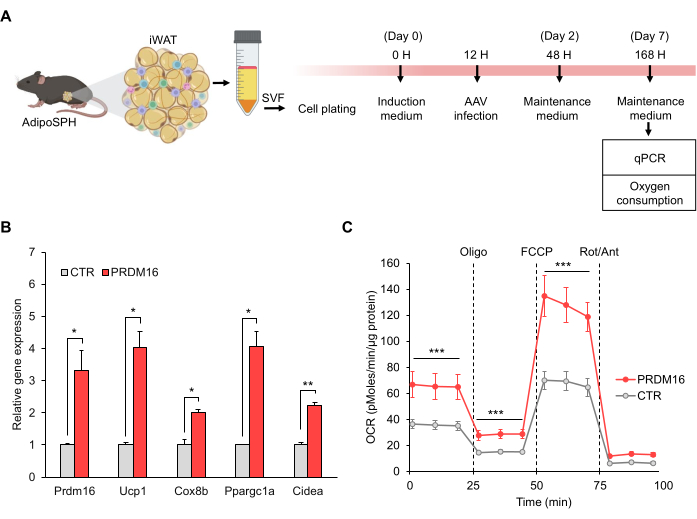
Figure 2: SPH-induced beige adipocyte differentiation by Prdm16 overexpression. (A) Schematic illustration of the in vitro beige adipocyte experimental design (created using Biorender.com). (B) Relative gene expression of Prdm16 and thermogenic genes (Ucp1, Cox8b, Ppargc1α, and Cidea; n = 3). (C) Oxygen consumption rate (OCR), n = 6. Data are presented as mean ± SEM. *p < 0.05; **p < 0.01; ***p < 0.001 for Prdm16 versus control (CTR) by (B) unpaired Student's t-test and (C) two-way repeated-measures ANOVA, followed by Tukey's test. Please click here to view a larger version of this figure.
Supplemental Figure 1: Sanger sequencing of Prdm16 sgRNA. (A) Schematic illustration demonstrating the alignment of single-stranded complementary Prdm16 oligonucleotides. (B) Sanger sequencing of Prdm16 sgRNA using universal primer. Please click here to download this File.
| Gene | Species | Forward | Reverse | |||||
| Gene expression | Prdm16 | mouse | CAGCACGGTGAAGCCATTC | GCGTGCATCCGCTTGTG | ||||
| Ucp1 | mouse | TCTCAGCCGGCTTAATGACTG | GGCTTGCATTCTGACCTTCAC | |||||
| Ppargc1a | mouse | AGCCGTGACCACTGACAACGAG | GCTGCATGGTTCTGAGTGCTAAG | |||||
| Cox8b | mouse | GAACCATGAAGCCAACGACT | GCGAAGTTCACAGTGGTTCC | |||||
| Cidea | mouse | ATCACAACTGGCCTGGTTACG | TACTACCCGGTGTCCATTTCT | |||||
| 36B4 | mouse | TCCAGGCTTTGGGCATCA | CTTTATCAGCTGCACATCACTCAGA | |||||
| Molecular cloning | sgRNA Prdm16 sequence | mouse | CGAGCTGCGCTGAAAAGGGG | CCCCTTTTCAGCGCAGCTCG | ||||
| universal primer | mouse | GAGGGCCTATTTCCC ATGATTCCTTCATAT |
||||||
Table 1: Primer sequences used in the study.
| Medium | Composition | |||
| Complete medium | Dulbecco's Modified Eagle Medium (DMEM) with L-Glutamine | |||
| 10% of fetal bovine serum (FBS) | ||||
| 2.5% of penicillin/streptomycin | ||||
| Induction medium | Complete medium | |||
| Indomethacin, final concentration 125 μM (0.125 M stock in ethanol). NOTE: Indomethacin must be heated to 90 °C for 10 seconds to be dissolved. | ||||
| Insulin, final concentration 20 nM (1 mM stock, Add 1 μL of HCl to solubilize (5.73 mg/mL). Store stock at -20 °C. | ||||
| Dexamethasone, final concentration 2 μg/mL (2 mg/mL stock in ethanol) | ||||
| 3-Isobutyl-1-methylxanthine (IBMX), final concentration 500 μM (0.25 M stock in 0.5 M KOH ) | ||||
| 3,3',5-Triiodo-L-thyronine (T3), final concentration 1 nM (10 μM stock, dissolve T3 in 1 M HCl and EtOH 1:4 to stock). | ||||
| Rosiglitazone, final concentration 1 μg/mL (1 mg/mL stock in ethanol) | ||||
| Maintenance medium | Complete medium | |||
| Insulin, final concentration 20 nM (1 mM stock). Add 1 μL of HCl to solubilize (5.73 mg/mL). | ||||
| Rosiglitazone, final concentration 1 μg/mL (1 mg/mL stock in ethanol) | ||||
Table 2: Composition of media used in the study.
| Parameters | CTR | PRDM16 | p-value | ||
| No mitochondrial oxygen consumption (pMoles/min/µg protein) | 8.19 ± 1.40 | 10.80 ± 1.83 | 0.01 | ||
| Basal respiration (pMoles/min/µg protein) | 28.82 ± 5.20 | 52.58 ± 13.73 | 0.001 | ||
| Maximal respiration (pMoles/min/µg protein) | 63.81 ± 9.80 | 122.94 ± 22.31 | < 0.001 | ||
| Spare respiratory capacity (pMoles/min/µg protein) | 34.98 ± 11.09 | 70.36 ± 26.06 | 0.006 | ||
| Oligo insensitive (pMoles/min/µg protein) | 8.27 ± 2.29 | 15.85 ± 5.48 | 0.005 | ||
| Oligo sensitive (pMoles/min/µg protein) | 20.54 ± 5.68 | 36.72 ± 14.79 | 0.016 | ||
| Coupling efficiency | 71.28 ± 1.51 | 69.84 ± 3.05 | 0.163 | ||
| Cell RCR | 7.70 ± 1.46 | 7.75 ± 2.43 | 0.485 | ||
Table 3: Respiratory parameters of SPH-induced beige adipocytes.
Discussion
One of the most useful non-editing applications of CRISPR technology is the interrogation of gene function through the activation of endogenous genes using CRISPRa systems6. SPH is a powerful CRISPRa that was originally described to induce the conversion of astrocytes into active neurons by targeting several neurogenic genes14. In this study, AdipoSPH was demonstrated to be a suitable tool for investigating beige fat biology by activating the expression of endogenous Prdm16 in adipocytes. SPH-induced Prdm16 overexpression in adipocytes leads to activation of the thermogenic gene program and induces the browning phenotype, phenocopying a previous transgenic (Tg) Prdm16 mice model23. Prdm16 was chosen in this study as the target for inducing beige adipocytes based on the central role of this transcription factor in regulating brown and beige fat differentiation, as well as the regulation of the thermogenic gene program (e.g., enhancing Ucp1 expression) in differentiated adipocytes. Nevertheless, the current model allows for overexpression of any endogenous gene at the discretion of the researchers and the aim of the study.
Traditional transgenic models investigating adipose tissue biology rely on the development of genetically modified rodents overexpressing a transgene under the control of a promoter/enhancer restricted to adipocytes (for example, Fabp4 or adiponectin)24. Although current technology has accelerated the pace of the development of these models, the costs and time to develop them are still a common issue experienced by the scientific community. In contrast, AdipoSPH is a cheaper and more versatile model for gain-of-function approaches than traditional Tg mice. Moreover, AdipoSPH is well suited for studying long non-coding RNA and transcript isoforms, which are not particularly suitable for Tg mice models.
AdipoSPH also presents several advantages in investigating adipose tissue biology compared to the simple administration of AAV, a current alternative approach for gain-of-function experiments25. First, AAV delivered to adipose tissue promotes overexpression by introducing several copies of a transgene. In contrast, the SPH activation of endogenous genes represents a more natural mechanism of action. Second, transgenes delivered by AAV (using constitutive promoters) normally result in their overexpression in “any cell type” within the adipose tissue microenvironment. In contrast, AdipoSPH-induced expression of endogenous genes is restricted to adipocytes. Third, the packaging size (~4.5kb) is a typical limitation of AAV for some transgenes26. However, AAV-carrying sgRNA requires a simple and easy cloning strategy for customized 20 nucleotides. Finally, AAV administration typically reaches systemic circulation and infects other tissues and organs. Although some AAVs contain microRNAs to mitigate the expression of transgenes in particular tissues, such as the liver and heart25, this strategy cannot prevent AAV infection in other tissues and organs. In contrast, AdipoSPH-induced expression of endogenous genes is limited to adipocytes, even considering some leakage of AAV into systemic circulation. Thus, AdipoSPH is also a suitable model for the systemic administration of AAV-carrying sgRNA and the investigation of adipose tissue regulation of whole-body energy homeostasis.
Although the AdipoSPH model offers some advantages, it has limitations that are important to consider. AAV is currently the most common platform for in vivo delivery of sgRNA. However, some challenges in AAV manufacturing and immunological issues remain unresolved26,27. Moreover, different amounts and distributions of injected AAV throughout adipose tissue result in intra- and inter-tissue variability of gene expression. We consider that sgRNA vector cloning, AAV production, and the homogeneous administration of AAV into the iWAT are critical steps for the success of this protocol. Finally, different sgRNAs distinctively activate the expression of endogenous genes in the SPH system14. The main recommendation is to design and test three to five sgRNA sequences within 200 bp upstream of the transcription start site (TSS) of the gene of interest. Therefore, defining the best sgRNA sequence for a target gene typically requires laborious in vitro assays prior to in vivo experiments.
AdipoSPH is also an attractive model for investigating other aspects of adipose-tissue biology. For instance, adipocytes promote the secretion of numerous adipokines28,29; however, the whole-body effects of most of these molecules remain poorly understood. AdipoSPH is a suitable model to address this issue by overexpressing single or multiple endogenous genes in mature adipocytes and investigating their local or systemic effects. Thus, AdipoSPH is a unique tool for studying the biology and physiology of adipose tissue.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
The authors thank the support received from Centro Multidisciplinar para Investigação Biológica na Área da Ciência em Animais de Laboratório (Cemib), Unicamp, for the generation of AdipoSPH mice, the Inmmunometabolism and Cell Signaling Laboratory, and National Institute of Science and Technology on Photonics Applied to Cell Biology (INFABIC) for all experimental support. We thank the financial support from Sao Paulo Research Foundation (FAPESP): 2019/15025-5; 2020/09308-1; 2020/14725-0; 2021/11841-2.
Materials
| 3,3',5-Triiodo-L-thyronine | Sigma-Aldrich | T2877 | |
| 3-Isobutyl-1-methylxanthine | Sigma-Aldrich | I5879 | |
| AAVpro 293T Cell Line | Takarabio | 632273 | |
| Amicon Ultra Centrifugal Filter | Merckmillipore | UFC510008 | 100 KDa |
| Dexamethasone | Sigma-Aldrich | D1756 | |
| Dulbecco's Modification of Eagles Medium (DMEM) | Corning | 10-017-CV | |
| Dulbecco's Modified Eagle Medium (DMEM) F-12, GlutaMAX™ supplement | Gibco | 10565-018 | high concentrations of glucose, amino acids, and vitamins |
| Dulbecco's phosphate buffered saline (DPBS) | Sigma-Aldrich | D8662 | |
| Excelta Self-Opening Micro Scissors | Fisher Scientific | 17-467-496 | |
| Fetal bovine serum | Sigma-Aldrich | F2442 | |
| Fisherbrand Cell Scrapers (100 pk) | Fisher Scientific | 08-100-241 | |
| Fisherbrand High Precision Straight Tapered Ultra Fine Point Tweezers/Forceps | Fisher Scientific | 12-000-122 | |
| Fisherbrand Sharp-Pointed Dissecting Scissors | Fisher Scientific | 08-940 | |
| Glycerol | Sigma-Aldrich | G5516 | |
| HEPES | Sigma-Aldrich | H3375-25G | |
| Hexadimethrine bromide (Polybrene) | Sigma-Aldrich | H9268 | |
| Indomethacin | Sigma-Aldrich | I7378 | |
| Insulin | Sigma-Aldrich | I9278 | |
| LigaFast Rapid DNA Ligation System | Promega | M8225 | |
| Maxiprep purification kit | Qiagen | 12162 | |
| Microliter syringe | Hamilton | 80308 | Model 701 |
| NEB 10-beta/Stable | New England Biolabs | C3019H | E. coli competent cells |
| pAAV2/8 | Addgene | 112864 | |
| pAAV-U6-gRNA-CBh-mCherry | Addgene | 91947 | |
| pAdDeltaF6 | Addgene | 112867 | |
| PEG 8000 | Sigma-Aldrich | 89510 | |
| Penicillin/streptomycin | Gibco | 15140-122 | |
| Polyethylenimine | Sigma-Aldrich | 23966 | Linear, MW 25000 |
| Povidone-iodine | Rioquímica | 510101303 | Antiseptic |
| Rosiglitazone | Sigma-Aldrich | R2408 | |
| SacI enzyme | New England Biolabs | R0156 | |
| Surgical Design Premier Adson Forceps | Fisher Scientific | 22-079-741 | |
| Syringe | Hamilton | 475-40417 | |
| T4 DNA Ligase | Promega | M180B | |
| T4 DNA ligase buffer | New England Biolabs | B0202S | |
| T4 PNK enzyme kit | New England Biolabs | M0201S | |
| Tramadol Hydrochloride | SEM | 43930 | |
| Vidisic Gel | Bausch + Lomb | 99620 |
Riferimenti
- Wang, W., Seale, P. Control of brown and beige fat development. Nature Reviews Molecular Cell Biology. 17 (11), 691-702 (2016).
- Wu, J., et al. Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell. 150 (2), 366-376 (2012).
- Cohen, P., Kajimura, S. The cellular and functional complexity of thermogenic fat. Nature Reviews Molecular Cell Biology. 22 (6), 393-409 (2021).
- Cong, L., et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 339 (6121), 819-823 (2013).
- Mali, P., et al. RNA-guided human genome engineering via Cas9. Science. 339 (6121), 823-826 (2013).
- Dominguez, A. A., Lim, W. A., Qi, L. S. Beyond editing: Repurposing CRISPR-Cas9 for precision genome regulation and interrogation. Nature Reviews Molecular Cell Biology. 17 (1), 5-15 (2016).
- Gilbert, L. A., et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell. 154 (2), 442-451 (2013).
- Perez-Pinera, P., et al. RNA-guided gene activation by CRISPR-Cas9-based transcription factors. Nature Methods. 10 (10), 973-976 (2013).
- Konermann, S., et al. Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature. 517 (7536), 583-588 (2015).
- Gilbert, L. A., et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell. 159 (3), 647-661 (2014).
- Tanenbaum, M. E., Gilbert, L. A., Qi, L. S., Weissman, J. S., Vale, R. D. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell. 159 (3), 635-646 (2014).
- Chavez, A., et al. Highly efficient Cas9-mediated transcriptional programming. Nature Methods. 12 (4), 326-328 (2015).
- Chavez, A., et al. Comparison of Cas9 activators in multiple species. Nature Methods. 13 (7), 563-567 (2016).
- Zhou, H., et al. In vivo simultaneous transcriptional activation of multiple genes in the brain using CRISPR-dCas9-activator transgenic mice. Nature Neuroscience. 21 (3), 440-446 (2018).
- Fripont, S., Marneffe, C., Marino, M., Rincon, M. Y., Holt, M. G. Production, purification, and quality control for adeno-associated virus-based vectors. Journal of Visualized Experiments. (143), e58960 (2019).
- Negrini, M., Wang, G., Heuer, A., Björklund, T., Davidsson, M. AAV production everywhere: a simple, fast, and reliable protocol for in-house aav vector production based on chloroform extraction. Current Protocols in Neuroscience. 93 (1), 103 (2020).
- Aune, U. L., Ruiz, L., Kajimura, S. Isolation and differentiation of stromal vascular cells to beige/brite cells. Journal of Visualized Experiments. (73), e50191 (2013).
- Wang, Q., et al. Post-translational control of beige fat biogenesis by PRDM16 stabilization. Nature. 609 (7925), 151-158 (2022).
- Oeckl, J., Bast-Habersbrunner, A., Fromme, T., Klingenspor, M., Li, Y. Isolation, culture, and functional analysis of murine thermogenic adipocytes. STAR Protocols. 1 (3), 100118 (2020).
- Cohen, P., et al. Ablation of PRDM16 and beige adipose causes metabolic dysfunction and a subcutaneous to visceral fat switch. Cell. 156 (1-2), 304-316 (2014).
- Harms, M., Seale, P. Brown and beige fat: development, function and therapeutic potential. Nature Medicine. 19 (10), 1252-1263 (2013).
- Divakaruni, A. S., Jastroch, M. A practical guide for the analysis, standardization and interpretation of oxygen consumption measurements. Nature Metabolism. 4 (8), 978-994 (2022).
- Seale, P., et al. Prdm16 determines the thermogenic program of subcutaneous white adipose tissue in mice. The Journal of Clinical Investigation. 121 (1), 96-105 (2011).
- Valet, P., Tavernier, G., Castan-Laurell, I., Saulnier-Blache, J. S., Langin, D. Understanding adipose tissue development from transgenic animal models. Journal of Lipid Research. 43 (6), 835-860 (2002).
- Bates, R., Huang, W., Cao, L. Adipose tissue: an emerging target for adeno-associated viral vectors. Molecular Therapy. Methods & Clinical Development. 19, 236-249 (2020).
- Wang, D., Tai, P. W. L., Guangping, G. Adeno-associated virus vector as a platform for gene therapy delivery. Nature Reviews Drug Discovery. 18 (5), 358-378 (2019).
- Colella, P., Ronzitti, G., Mingozzi, F. Emerging issues in AAV-mediated in vivo gene therapy. Molecular Therapy. Methods & Clinical Development. 8, 87-104 (2018).
- Deshmukh, A. S., et al. Proteomics-based comparative mapping of the secretomes of human brown and white adipocytes reveals EPDR1 as a novel batokine. Cell Metabolism. 30 (5), 963-975 (2019).
- Sponton, C. H., et al. The regulation of glucose and lipid homeostasis via PLTP as a mediator of BAT-liver communication. EMBO reports. 21 (9), 49828 (2020).

