Design and Development of a Model to Study the Effect of Supplemental Oxygen on the Cystic Fibrosis Airway Microbiome
PREPARAZIONE ISTRUTTORI
CONCETTI
Student Protocol
This study received approval from the Partners Institutional Review Board (Protocol # 2018P002934). Inclusion criterion included adult patients with cystic fibrosis who provided written informed consent for the study. There was no exclusion criterion. According to protocol guidelines, all sputum samples were collected from patients with cystic fibrosis during a scheduled outpatient visit with their clinical provider.
1. Artificial Sputum Medium Preparation
NOTE: Quantities listed here are for the production of 1 L of final artificial sputum medium, and assume the specific reagents listed in the Materials Table. Numbers must be adjusted for other volumes or for the use of different reagents to ensure the same final product. See Table 1 for target concentrations.
- Artificial sputum chemical mix (ASCM)
NOTE: ASCM makes up 25% of the final medium volume. It is shelf-stable and can be prepared in bulk or in advance. If being prepared for later use, autoclave the chemical mix and safely store it at room temperature.- Mix the constituent chemical stock solutions.
- Prepare 1 M NaCl stock: Add 58.44 g of NaCl per liter of sterile water.
- Prepare 1 M KCl stock: Add 74.55 g of KCl per liter of sterile water.
- Prepare 1 M MgSO4 stock: Add 246.47 g of MgSO4·7H2O per liter of sterile water, or 120.37 g of anhydrous MgSO4 per liter of sterile water.
- Prepare 1 M glucose stock: Add 180.16 g of glucose per liter of sterile water.
- Autoclave sterilize the chemical stock solutions, as well as an empty 250 mL bottle. Perform the autoclaving steps to at least standard values of 121 °C and 15 PSI for 30 min.
- Add 80.59 mL of sterile water to the empty 250 mL bottle.
- Add 152.30 mL of 1 M NaCl stock to the mix.
- Add 15.8 mL of 1 M KCl stock to the mix.
- Add 610 µL of 1 M MgSO4 stock to the mix.
- Add 700 µL of 1 M glucose stock to the mix.
- Mix the constituent chemical stock solutions.
- Artificial sputum mucin mix (ASMM)
NOTE: ASMM makes up 50% of the final medium volume. Ensure that it is prepared on the same day as the final medium batch.- Add 450 mL of sterile water to an empty 1 L bottle.
- Add 50 mL of 10x Phosphate-Buffered Saline (PBS) to the bottle.
- Add a disposable magnetic stir bar to the bottle.
- Autoclave the bottle containing PBS and the stir bar.
- Measure out 10 g of mucin powder and add it to the PBS.
- Shake the bottle vigorously for preliminary mixing.
- Place the bottle onto a hot plate with a magnetic stirrer. Set heat to medium-high targeting 50 °C and stirring speed to 1100 rpm. Ramp up the speed gradually so that the bar does not fly off the magnet.
- Allow to heat and stir for 15 min.
- Pick up the bottle with heat-resistant gloves. Observe to check if mucin powder settles out of the solution.
- If mucin powder is not fully dissolved, return the bottle to heat/stirrer for 5-min intervals until it is completely dissolved.
- Allow the mucin mix to cool to room temperature.
- Artificial sputum biological mix (ASBM)
NOTE: ASBM is 25% of the final medium volume. Prepare it on the same day as the final medium batch, and unlike the other mixes, do not expose its components to any heat.- Thaw the 100x vitamin stock in a 4 °C fridge or on ice.
NOTE: Pre-portion the vitamin stock into 10 mL aliquots to minimize the number of freeze/thaw cycles. - Add 124.24 mL of sterile water to the empty autoclaved 250 mL bottle.
- Add 25.76 mL of 50x essential amino acid stock to the mix.
- Add 80.14 mL of 100x non-essential amino acid stock to the mix.
- Add 10 mL of (thawed) 100x vitamin stock to the mix.
- Add 1 mL of 1000x trace metals stock to the mix.
- Add 8.33 mL of 30% egg yolk emulsion to the mix.
- Add 400 µL of 10 g/L ferritin stock to the mix.
- Mix the solution well via manual shaking.
- Thaw the 100x vitamin stock in a 4 °C fridge or on ice.
- Artificial sputum medium (ASM)
- Add 250 mL of ASCM to the 1 L bottle containing ASMM.
- Add 250 mL of ASBM to the medium bottle.
- Titrate the medium with basic MOPS buffer (1 M) to reach a pH of 6.3 on a narrow range pH paper. Prior to titration, the medium mix will be too acidic.
- Refrigerate the resulting artificial sputum medium at 4 °C until it is ready for filtration.
- To start the filtration process, transfer 200 mL of unfiltered artificial sputum medium to a vacuum filtration system with a 0.22 µm pore size filter.
- Connect the filtration system to the vacuum pump, turn on the vacuum pump, set it to 70 mbar, and then place the chamber on an orbital shaker shaking at 90 rpm in a cold room at 4 °C.
- Top off with an additional 150 mL of the medium as an appreciable amount is filtered. It takes 1-2 days to filter 1 L of medium completely.
- Repeat with additional chambers until all the media is filtered.
NOTE: Try not to filter more than 350 mL of the medium through the same 0.22 µm filter since mucin will plug the filter over time.
- Refrigerate filtered artificial sputum medium at 4 °C until ready for use. Use ASM within one month of preparation for best results.
2. Oxygen Sparging
- Sparging station setup
NOTE: This protocol should only need to be done in full once, after which point the setup can be maintained through simple maintenance as necessary. See Figure 1 for a visual schematic of the oxygen sparging system.- Obtain and properly secure the compressed air and oxygen tanks.
CAUTION: High pressure makes the tanks extremely dangerous when mishandled. Ensure that the tanks are completely sealed and secured, there are no leaks when the tank is closed, and that all handling personnel is fully trained in their use. - Attach an air regulator to the compressed air tank with a wrench. For optimal flow reading on the regulator, attach the regulator as close as possible to an upright position.
- Attach an oxygen regulator to the compressed oxygen tank, attaching as close as possible to an upright position. Depending on the oxygen tank, one may need to invert the direction to tighten.
- Connect the tubing from the regulators to a Y-connector to combine the gas flow from the two tanks.
- Connect the output of the Y-connector to the central T-junction valve.
- Connect one side of the central T-junction valve to a gas pressure gauge.
- Connect the other side of the gas pressure gauge to a 25 mm diameter sterile syringe filter with a 0.22 µm pore size.
- Attach a second 25 mm diameter syringe filter to a syringe without a plunger to be used as a gas release during sparging.
- Connect the final side of the central T-junction valve to a second T-junction valve for the oxygen monitor.
- Connect a 25 mm diameter syringe filter to one side of this second T-junction valve, along with tubing to attach 18 G needles.
- Connect the final side of the second T-junction valve to the oxygen monitoring apparatus.
- Connect a cut-off tube to the other side of the oxygen monitoring apparatus to be used as a gas release during monitoring.
CAUTION: When testing/using the oxygen sparging system, take careful note of the position of the T-junctions and ensure it matches the intended path through the system. Failure to do so will result in pressure buildup inside the system and cause components to fail and come apart . - For the maintenance of the system and to keep it working at optimal performance, the following practices are beneficial.
- Reinforce the connections with liberal amounts of Teflon tape to greatly improve their seal and reduce the chance of components coming apart from the internal pressure.
- Keep combined flow rate under 10 L/min to mitigate maximum pressure and prevent failures.
- If a leak is suspected, use a detergent solution such as commercially available liquid leak detectors to identify its location easily, as it will bubble above any gas leaks. Patch leaks using a Polytetrafluoroethylene tape (e.g., Teflon).
- Replace the 25 mm diameter syringe filters in the oxygen sparging system bi-weekly, but this varies with use frequency. Over time, particles caught in the filter reduce the gas flow rate and cause pressure buildups.
- Calibrate the oxygen monitor to 21% oxygen compressed air prior to carrying out measurements.
- Upon completion of the system use, turn off the tanks and bleed the excess gas from the regulators until the flow completely stops.
- Obtain and properly secure the compressed air and oxygen tanks.
- Serum bottle culture sparging
- Label 500 mL autoclaved serum bottles with sample identifiers, date/time of inoculation, and target oxygen percentage.
- In a biological hood, add 24 mL of the artificial sputum medium to each serum bottle being set up.
- Add 1 mL of the patient sputum homogenized with an 18-G needle (diluted with sterile saline if necessary to obtain sufficient volume of sample for each culture condition) to each serum bottle.
- Using sterile tweezers, place the autoclaved rubber stoppers onto the top of each serum bottle.
- Press down the rubber stoppers, take care not to touch the underside of the stopper with hands.
- Remove the bottles from the hood, apply and crimp the aluminum seals. Remove the center piece from the seals.
- Wipe down the top of bottles with an alcohol wipe and pass them through a Bunsen burner flame.
- Affix a sterile 18-G needle to a plunger-less syringe with a filter. Insert this gas release into the bottle first.
- Affix a sterile 18-G needle to the gas output from the system and insert the gas output needle into the bottle as well.
- Route the T-junctions from the tanks through the oxygen monitor. Verify that the target oxygen concentration is flowing through the system. Target approximately 5 L/min of gas flow.
- Reroute the T-junctions from the tanks through to the gas output. The gas starts to flow through the serum bottle.
CAUTION: Pay close attention to the pressure gauge during oxygen sparging. If pressure increases unexpectedly, shut the system off immediately. - Run oxygen sparge through the serum bottle for 1 min. At 5 L/min, this allows for 10 air exchanges and ensures the internal atmosphere reaches the desired partial pressure.
- Remove the gas release 18-G needle.
- Allow the pressure in the serum bottle to build to +1 atmosphere (2 atmospheres at sea level) and then immediately remove the gas output needle.
NOTE: Maintaining pressure aids retention of hyperoxic conditions over time. - Place the serum bottle into a 37 °C incubator shaker at 150 RPM. Incubate the samples for three 24-h intervals. At each 24-h interval, take an aliquot for downstream analysis, re-sparge the samples and return them to incubation for a total incubation time of 72 h.
- Outflow oxygen measurement
- Calibrate the oxygen meter to 21% compressed air, and then turn off the tank.
- Route the serum bottle intake through the oxygen monitor and affix a sterile needle to the end.
- Insert the needle through the rubber stopper into the serum bottle.
- Wait for the outflow reading to stabilize. A low flow rate out of the serum bottles means this may take up to 2 min. Report the peak difference from room air (number furthest from 21%).
- If performing multiple readings, flush the system with compressed air between readings.
Design and Development of a Model to Study the Effect of Supplemental Oxygen on the Cystic Fibrosis Airway Microbiome
Learning Objectives
These protocols were applied to 50 expectorated sputum samples from pwCF presenting for routine care to an outpatient cystic fibrosis clinic at Massachusetts General Hospital in Boston, Massachusetts. Each patient's sputum was cultured under 21%, 50%, and 100% oxygen conditions using the artificial sputum medium, with 0.5 mL aliquots taken from each culture at 24 h, 48 h, and 72 h of culture time for testing. Cultures were photographed when extractions were made to track visual changes. In addition, a 0.5 mL aliquot of each primary sputum sample was taken prior to culturing. This resulted in 10 discrete samples per patient and a final N of 500 samples. Of these, sputum from 11 patients (11 uncultured sputa, 11 cultured sputa from 21% oxygen at 48 h of incubation) underwent nucleic acid extraction17, sequencing libraries were generated using a commercial DNA library preparation kit, and metagenomic sequencing was performed on a whole genome sequencing platform targeting ~ 5 Gb of sequence per sample with 150 base pair, paired-end reads. Raw reads were processed using the bioBakery suite of tools18, which includes quality control and removal of human "contaminant" sequences and taxonomic profiling with the MetaPhlAn3 profiler19. At the time of nucleic acid extraction, 10 million cells of Imtechella halotolerans, a halotolerant species normally found in estuary ecosystems and not in human microbial communities, were spiked into each sample, allowing the quantification of absolute microbial load for each sample20.
Figure 2 shows individual and average outflow oxygen measurements and pH levels over the course of the culture process for 50 sputum samples cultured under each oxygen condition and an example of a visual differential culture phenotype. Cultures were maintained at 37 °C except during brief periods when sparging and removal of sample aliquots was performed. With both sparging intervals of 12 h and 24 h, elevated oxygen concentrations were maintained, although a drop over time was observed for all three oxygen conditions, with 100% oxygen falling to approximately 85%, 50% oxygen falling to 40%, and 21% oxygen falling to 18%. Oxygen conditions remained distinct, and importantly, elevated oxygen concentrations were maintained throughout the process for hyperoxic samples. pH measurements showed a greater degree of variability but stayed well within a physiologically normal range with no statistically significant changes over time. These measurements indicate that these methods maintain discrete differential oxygen conditions throughout the culture process. Lastly, an example of one of many visual culture phenotypes that differentiated across oxygen concentration is shown. This sample had marked turbidity differences after 72 h of culture, with higher oxygen associated with lower visual turbidity. Differential culture phenotypes support the presence of hyperoxia-induced effects on culture communities.
Figure 3 compares microbial load, microbial diversity, and microbial community composition between uncultured sputum and cultured sputum (21% oxygen condition for a period of 48 h). Measurements reveal the only major difference introduced by the act of culturing to be an approximately 20-fold increase in microbial load as compared to uncultured sputum. The immune system and the typical mechanical sputum clearance mechanisms such as coughing normally serve as a regulatory process limiting the microbial load in the lung, even in cases of dysfunction and infection like those seen in pwCF. Ex vivo culture has no such regulatory mechanisms, and microbial communities are instead free to proceed toward cellular saturation. Alpha and beta diversity metrics indicate that despite this difference in microbial load, the underlying community composition remains well-preserved, with minimal global differences introduced by the culture process.
Figure 4 expands on the comparison between uncultured and cultured sputum samples, looking at the binary presence/absence of the 120 microbial species conclusively identified by shotgun metagenomics sequencing from cultured and uncultured sputum obtained from 11 patients. Microbes are clustered based on phylogenetic similarities. 46 (38.3%) of these species were identified in both uncultured and cultured samples (cyan color), while 35 (29.2%) were exclusively identified in uncultured samples (yellow) and 39 (32.5%) were exclusively identified in cultured samples (blue). It is likely that there is greater parity than what we identified using sequencing in terms of what is present and what is absent, but some taxa fall below the sequencing detection threshold in some cases. The differences do indicate that the culture process introduces some bias in cultured compared to uncultured sputum. Most notably, culturing increases the presence of fungi such as Candida and Aspergillus, as well as Enterobacterales members including Escherichia, Serratia, and Streptococcus members. Conversely, Bacteroidetes members such as Prevotella and Clostridiales, which are anaerobes, were present in uncultured samples but not present in cultured samples. This may be attributed to the lack of an anaerobic condition in our experimental model.
Figure 5 shows absorbance-based growth curves of common CF lung pathogens isolated from sputum obtained from 50 different pwCF. These isolates represent phenotypically different clinical isolates obtained using enrichment culture procedures from the Massachusetts General Hospital Clinical Microbiology Laboratory, and include Pseudomonas aeruginosa (N = 53), Staphylococcus aureus (N = 37), Stenotrophomonas maltophilia (N = 12), Klebsiella pneumoniae (N = 3), and Achromobacter sp (N = 7). Growth curves were obtained by culturing each isolate in artificial sputum media at 37 °C in the dark, with ASM sans bacterial inoculation serving as a negative control. The transparent quality of ASM (which is due to filter rather than heat sterilization) allows for conducting optical measures to estimate growth curves. Optical readings at 600 nm (OD600) were taken every 10 min, and the first 24 h of each curve are shown. The absence of changes in optical readings in the ASM-only negative control indicates cultures free of contamination. The demonstrative curves shown here follow typical growth curve patterns indicating the viability of this ASM recipe as a medium for the absorbance-based generation of growth curves.
| Column 1 | Column 2 | Column 3 | Column 4 | Column 5 | Column 6 | Column 7 | Column 8 | Column 9 | |
| Value | Comstock | Kirchner | Sriramulu | Palmer | Flynn | Gallagher | Lai | Source | |
| Mucin | 2% w/v | 0.5% w/v | 0.5% w/v | – | 1% w/v | 2% w/v | 1% w/v | Flynn | |
| Sodium Chloride | 85.5 mM | 85.5 mM | 85.5 mM | 66.6 mM | 89.8 mM | 85.5 mM | 152.3 mM | Lapierre | |
| Potassium Chloride | 29.5 mM | 29.5 mM | 29.5 mM | 15.8 mM | – | 2.95 mM | 15.8 mM | Palmer | |
| Magnesium Sulfate | – | – | – | 0.6 mM | 1 mM | 1 mM | 0.61 mM | Palmer | |
| Iron Sulfate | – | – | – | 0.0036 mM | – | – | – | – | |
| Ammonium Chloride | – | – | – | 2.3 mM | 60 mM | – | – | – | |
| Monopotassium Phosphate | – | – | – | 2.5 mM | 60 mM | – | – | – | |
| Glucose | – | – | – | 3.2 mM | 13 mM | 40 mM | 0.7 mM | Sambeek | |
| Lactate | – | – | – | 9 mM | – | – | – | – | |
| Essential Amino Acids | 14.45x | 0.25 g/L | 0.25 g/L | Per Acid | 0.5x | 0.375x | 1.29x | Palmer | |
| Non-Essential Amino Acids | 28.9x | 0.25 g/L | 0.25 g/L | Per Acid | 0.25x | 0.5x | 8.01x | Palmer | |
| Vitamins | – | – | – | – | – | 1x | 1x | Gallagher | |
| Trace Metals | – | – | – | – | 1x | 1x | 1x | Flynn | |
| Egg Yolk | 0.25% | 0.25% | 0.25% | – | – | 0.25% | 0.25% | Kirchner | |
| Ferritin | 0.0003 g/L | – | – | – | – | 0.0004 g/L | 0.0004 g/L | Gallagher | |
| Salmon Sperm DNA | 1.4 g/L | 4 g/L | 4 g/L | – | – | 1.4 g/L | – | – | |
| DPTA | – | – | 0.0059 g/L | – | – | – | – | – | |
| pH | – | 6.9 | – | 6.8 | – | – | 6.3 | Lapierre | |
| Storage | 0 | 4° | – | – | 4° | 4° | 4° | Kirchner | |
| Sterilization | Autoclave | Filter | Autoclave | – | Autoclave | Autoclave | Filter | Kirchner | |
Table 1: Artificial sputum medium recipe derived from review of literature. (Column 1) Reagents and key values in the formulation of artificial sputum medium. (Columns 2–7) Comparison of recipes from extant literature8,9,10,12,14,15. (Columns 8–9) Artificial sputum medium recipe detailed in this protocol and the corresponding sources that informed each selected value10,11,12,13,14,15.
| Column 1 | Column 2 | Column 3 |
| CF Sputum | ASM | |
| Total Amino Acids | 10.25 mM | 10.76 mM |
| Alanine | 0.96 mM | 0.80 mM |
| Arginine | 0.17 mM | 0.94 mM |
| Asparagine | 0.91 mM | |
| Aspartic Acid | 0.45 mM | 0.80 mM |
| Cysteine | 0.09 mM | 0.33 mM |
| Glutamic Acid | 0.84 mM | 0.80 mM |
| Glycine | 0.65 mM | 0.80 mM |
| Histidine | 0.28 mM | 0.35 mM |
| Isoleucine | 0.60 mM | 0.52 mM |
| Leucine | 0.87 mM | 0.52 mM |
| Lysine | 1.15 mM | 0.64 mM |
| Methionine | 0.34 mM | 0.13 mM |
| Ornithine | 0.36 mM | |
| Phenylalanine | 0.29 mM | 0.26 mM |
| Proline | 0.90 mM | 0.80 mM |
| Serine | 0.78 mM | 0.80 mM |
| Threonine | 0.58 mM | 0.52 mM |
| Tryptophan | 0.07 mM | 0.06 mM |
| Tyrosine | 0.43 mM | 0.26 mM |
| Valine | 0.60 mM | 0.52 mM |
Table 2: Amino acid concentrations previously described in cystic fibrosis sputum and in artificial sputum medium recipe detailed in this protocol. (Column 1) Key amino acids. (Column 2) Amino acid concentrations of sputum from people with cystic fibrosis12. (Column 3) Amino acid concentrations in artificial sputum medium detailed in this protocol
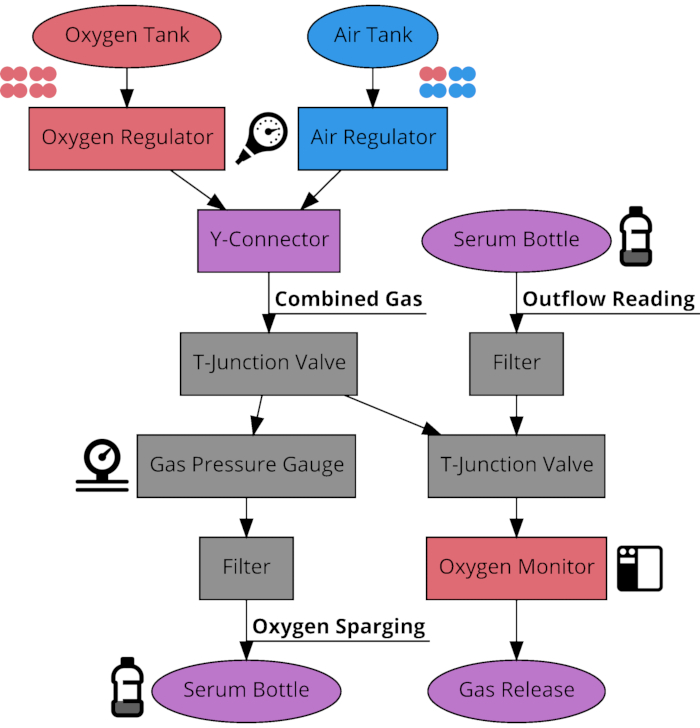
Figure 1: Connection schematic of oxygen sparging system components. Flow diagram of the connections between components of the system used to sparge serum bottles to desired oxygen concentrations between 21% and 100%. The system has 3 modes of use determined by the position of the two T-junction valves. The system can route gas from the tanks through the gas output or through the oxygen percentage monitor, as well as route outflow gas from previously sparged serum bottles through the monitor to check concentration after time has elapsed. Please click here to view a larger version of this figure.
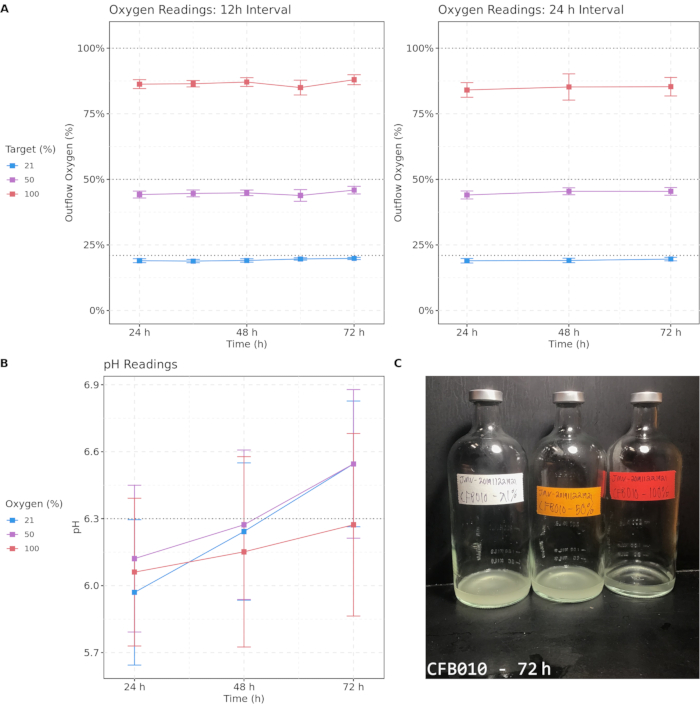
Figure 2: Target oxygen concentrations are approximately maintained with both 12 h and 24 h sparging intervals, and pH remains in the physiological range during culture. (A) Outflow oxygen readings from 12 h and 24 h oxygen sparging intervals over a 72-h period. (B) pH readings for samples measured every 24 h. (C) An example image of cultured sample CFB010 after 72 h, displaying differential turbidity across oxygen concentrations. The color indicates target oxygen percentage; error bars denote 95% confidence intervals. Critical thresholds are emphasized with dashed lines. Please click here to view a larger version of this figure.
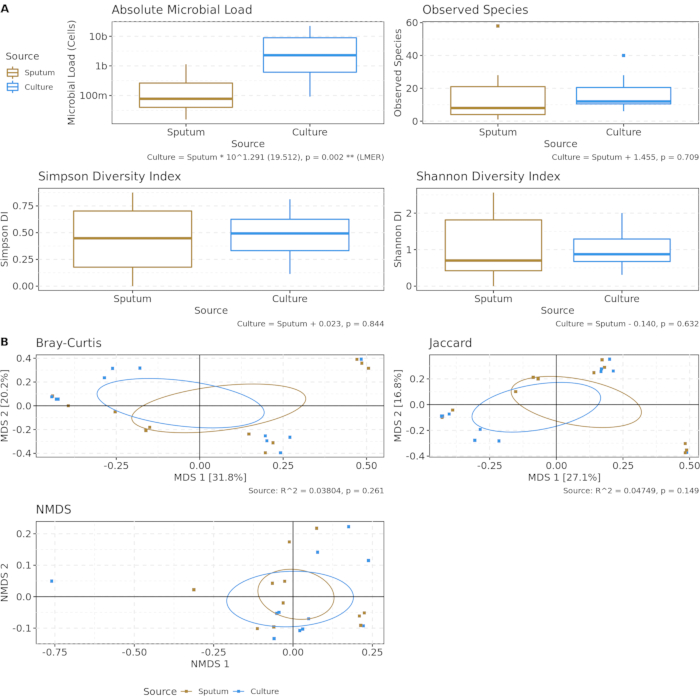
Figure 3: Culturing increases microbial load, but underlying community composition is preserved. Uncultured sputum (yellow) and cultured sputum (blue) using artificial sputum medium at 21% oxygen for 48 h. Aliquots underwent nucleic acid extraction and shotgun metagenomics sequencing to detect possible bias introduced from culture conditions. (A) Absolute microbial load (determined by spike-in controls) and alpha diversity metrics. Using linear mixed effects models, uncultured vs. cultured sputum predicted microbial load but not alpha diversity. (B) Ordination of first two components of beta diversity metrics, controlling for difference in microbial load. No significant difference in either metric after PERMANOVA. Please click here to view a larger version of this figure.
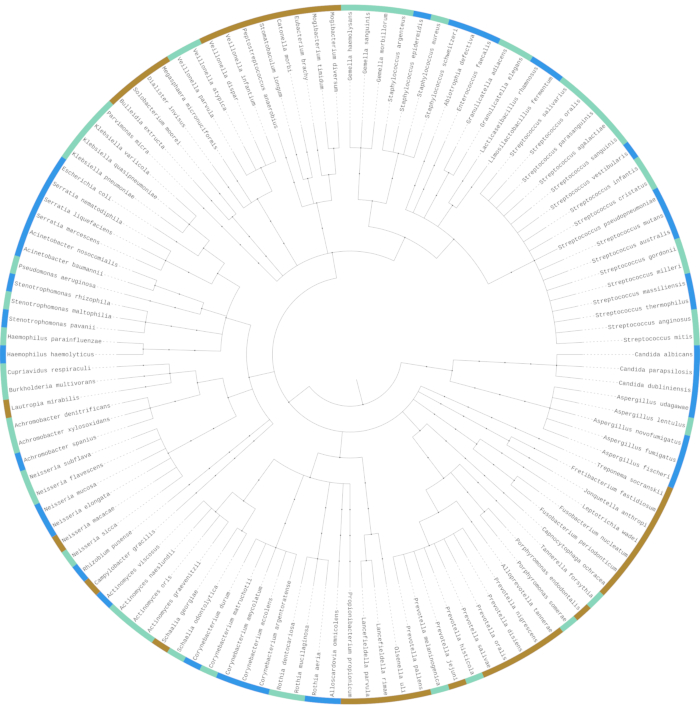
Figure 4: The majority of identified taxa are present in both source sputum and culture, while others only appear in source sputum or culture. Shotgun metagenomics sequencing used to compare differences in microbial community composition between uncultured and cultured sputum samples. Phylogenetic tree of all identified microbial species in sequenced samples (N = 120). Species marked with yellow (N = 35, 29.2%) were only seen in uncultured sputum samples. Species marked with blue (N = 39, 32.5%) were only seen in artificial sputum medium culture samples. Species marked with cyan (N = 46, 38.3%) were seen in both uncultured and cultured samples. Please click here to view a larger version of this figure.
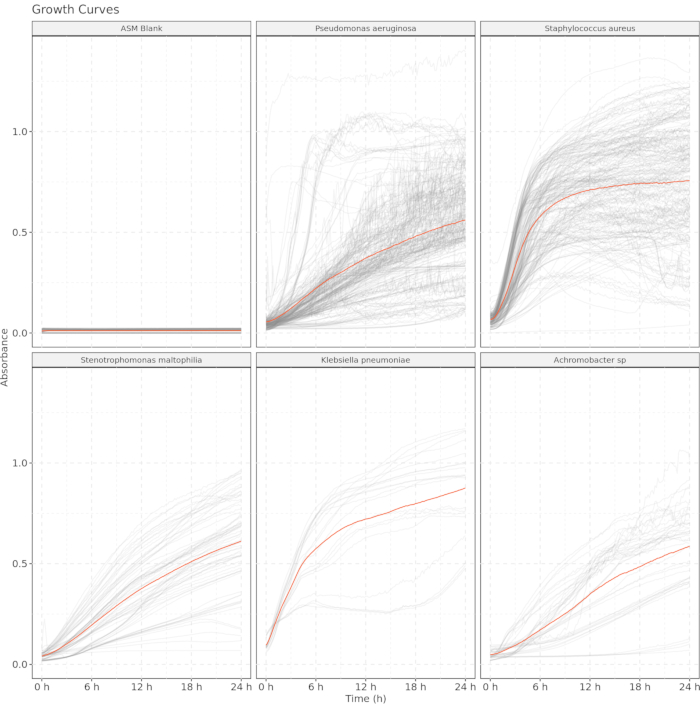
Figure 5: Artificial sputum medium is sufficiently transparent to be used as a growth curve medium for culturing clinical isolates. Optimal density readings at 600 nm were taken every 10 min, and the first 24 h of each curve are shown. Gray lines represent individual readings, and orange lines represent the mean absorbance for each taxon. Artificial sputum medium blank included as a control. Please click here to view a larger version of this figure.
List of Materials
| BME Vitamins (100x) Solution | MilliporeSigma | B6891 | Concentrated solution of supplemental vitamins. |
| Crimper, 30 mm | DWK Life Sciences | 224307 | Crimper for attaching aluminum seals to serum bottles. |
| D-(+)-Glucose | MilliporeSigma | G7021 | Solid glucose powder (dextrorotatory isomer). |
| Diaphragm Pump ME 2 NT | VACUUBRAND | 20730003 | Vacuum pump for vacuum filtration. |
| Egg Yolk Emulsion | HiMedia | FD045 | Sterile emulsion of 30% egg yolk in saline. |
| Ferritin, Cationized from Horse Spleen | MilliporeSigma | F7879 | Ferritin (iron-storage protein) solution. |
| FIREBOY plus Safety Bunsen Burner | Integra Biosciences | 144000 | Bunsen burner with user interface and safety features. |
| Hydrion pH Paper (1.0–14.0) | Micro Essential Laboratory | 94 | pH testing paper for the range of 1.0–14.0. |
| Hydrion pH Paper (4.0–9.0) | Micro Essential Laboratory | 55 | pH testing paper for the range of 4.0–9.0. |
| Hydrion pH Paper (6.0–8.0) | Micro Essential Laboratory | 345 | pH testing paper for the range of 6.0–8.0. |
| Hypodermic Needle-Pro EDGE Safety Device, 18 G | Smiths Medical | 401815 | 18 G needles with safety caps. |
| In-Line Pressure Gauge | MilliporeSigma | 20469 | Gas pressure gauge for monitoring bottle pressure. |
| Innova 42 Incubated Shaker | Eppendorf | 2231000756 | Combination incubator/orbital shaker. |
| Luer-Lok Syringe with Attached Needle | Becton Dickinson | 309580 | Combination 3 mL syringe and 18 G needle. |
| Luer Valve Assortment | World Precision Instruments | 14011 | Valves for gas flow tubing. |
| LSE Orbital Shaker | ThermoFisher Scientific | 6780-NP | Orbital shaker to agitate media during filtration. |
| Magnesium Sulfate Heptahydrate | MilliporeSigma | M2773 | Solid epsom salt (magnesium sulfate heptahydrate). |
| Medical Air Single Stage Regulator with Flowmeter | Western Enterprises | M1-346-15FM | Air flow rate regulator with 15 L/min meter. |
| MEM Amino Acids (50x) Solution | MilliporeSigma | M5550 | Concentrated solution of essential amino acids. |
| MEM Non-Essential Amino Acids (100x) Solution | MilliporeSigma | M7145 | Concentrated solution of non-essential amino acids. |
| Millex-GP Filter, 0.22 µm | MilliporeSigma | SLMP25SS | 0.22 µm polyethersulfone membrane sterile syringe filter. |
| Milli-Q Academic | MilliporeSigma | ZMQS60E01 | Milli-Q sterile water filtration system. |
| MiniOX 3000 Oxygen Monitor | MSA | 814365 | Gas flow oxygen percentage monitor. |
| MOPS Buffer (1 M, pH 9.0) | Boston BioProducts | BBM-90 | MOPS buffer for adjusting media pH. |
| Mucin from Porcine Stomach | MilliporeSigma | M2378 | Mucin (glycosylated gel-forming protein) powder. |
| Natural Polypropylene Barbed Fitting Kit | Harvard Apparatus | 72-1413 | Connectors for gas flow tubing. |
| Nextera XT DNA Library Preparation Kit | Illumina | FC-131-1096 | Library preparation for identification during sequencing. |
| NovaSeq 6000 Sequencing System | Illumina | 770-2016-025-N | Shotgun sequencing platform for generating sample reads. |
| Oxygen Single Stage Regulator with Flowmeter | Western Enterprises | M1-540-15FM | Oxygen flow rate regulator with 15 L/min meter. |
| Oxygen Tubing with 2 Standard Connectors | SunMed | 2001-01 | Tubing for connecting gas system components. |
| Phosphate buffered saline, 10x, pH 7.4 | Molecular Biologicals International | MRGF-6235 | Concentrated phosphate-buffered saline solution. |
| PC 420 Hot Plate/Stirrer | Marshall Scientific | CO-PC420 | Combination hot plate/stirrer. |
| Potassium Chloride | MilliporeSigma | P9541 | Solid potassium chloride salt. |
| PTFE Disposable Stir Bars | ThermoFisher Scientific | 14-513-95 | Disposable magnetic stir bars. |
| PTFE Thread Seal Teflon Tape | VWR | 470042-938 | Teflon tape for reinforcing gas system connections. |
| Q-Gard 2 Purification Cartridge | MilliporeSigma | QGARD00D2 | Purification cartridge for Milli-Q system. |
| Reusable Media Storage Bottles | ThermoFisher Scientific | 06-423A | Bottles for mixing and storing culture media. |
| Rubber Stopper, 30 mm, Gray Bromobutyl | DWK Life Sciences | 224100-331 | Rubber stoppers for serum bottles. |
| Serum Bottle with Molded Graduations, 500 mL | DWK Life Sciences | 223952 | Glass serum bottles for sealed culturing. |
| Small Bore Extension Set | Braun Medical | 471960 | Tubing extension with luer lock connectors. |
| Sodium Chloride | MilliporeSigma | S3014 | Solid sodium chloride salt. |
| Spike-in Control I (High Microbial Load) | ZymoBIOMICS | D6320 | Spike-in microbes (I. halotolerans and A. halotolerans) for absolute microbial load calculations |
| Stericup Quick Release Sterile Vacuum Filtration System | MilliporeSigma | S2GPU02RE | 250 mL 0.22 µm vacuum filtration chamber. |
| Super Sani-Cloth Germicidal Disposable Wipes | Professional Disposables International | H04082 | Disposable germicidal wipes for sterilization. |
| Trace Metals Mixture, 1000x | ThermoFisher Scientific | NC0112668 | Concentrated solution of physiological trace metals. |
| Unlined Aluminum Seal, 30 mm | DWK Life Sciences | 224187-01 | Aluminum seals crimped over top of rubber stoppers. |
| USP Medical Grade Air Tank | Airgas | AI USP200 | Compressed air tank for input to sparging system. |
| USP Medical Grade Oxygen Tank | Airgas | OX USP200 | Compressed oxygen tank for input to sparging system. |
Lab Prep
Airway microbial communities are thought to play an important role in the progression of cystic fibrosis (CF) and other chronic pulmonary diseases. Microbes have traditionally been classified based on their ability to use or tolerate oxygen. Supplemental oxygen is a common medical therapy administered to people with cystic fibrosis (pwCF); however, existing studies on oxygen and the airway microbiome have focused on how hypoxia (low oxygen) rather than hyperoxia (high oxygen) affects the predominantly aerobic and facultative anaerobic lung microbial communities. To address this critical knowledge gap, this protocol was developed using an artificial sputum medium that mimics the composition of sputum from pwCF. The use of filter sterilization, which yields a transparent medium, allows optical methods to follow the growth of single-celled microbes in suspension cultures. To create hyperoxic conditions, this model system takes advantage of established anaerobic culturing techniques to study hyperoxic conditions; instead of removing oxygen, oxygen is added to cultures by daily sparging of serum bottles with a mixture of compressed oxygen and air. Sputum from 50 pwCF underwent daily sparging for a 72-h period to verify the ability of this model to maintain differential oxygen conditions. Shotgun metagenomic sequencing was performed on cultured and uncultured sputum samples from 11 pwCF to verify the ability of this medium to support the growth of commensal and pathogenic microbes commonly found in cystic fibrosis sputum. Growth curves were obtained from 112 isolates obtained from pwCF to verify the ability of this artificial sputum medium to support the growth of common cystic fibrosis pathogens. We find that this model can culture a wide variety of pathogens and commensals in CF sputum, recovers a community highly similar to uncultured sputum under normoxic conditions, and creates different culture phenotypes under varying oxygen conditions. This new approach might lead to a better understanding of unanticipated effects induced by the use of oxygen in pwCF on airway microbial communities and common respiratory pathogens.
Airway microbial communities are thought to play an important role in the progression of cystic fibrosis (CF) and other chronic pulmonary diseases. Microbes have traditionally been classified based on their ability to use or tolerate oxygen. Supplemental oxygen is a common medical therapy administered to people with cystic fibrosis (pwCF); however, existing studies on oxygen and the airway microbiome have focused on how hypoxia (low oxygen) rather than hyperoxia (high oxygen) affects the predominantly aerobic and facultative anaerobic lung microbial communities. To address this critical knowledge gap, this protocol was developed using an artificial sputum medium that mimics the composition of sputum from pwCF. The use of filter sterilization, which yields a transparent medium, allows optical methods to follow the growth of single-celled microbes in suspension cultures. To create hyperoxic conditions, this model system takes advantage of established anaerobic culturing techniques to study hyperoxic conditions; instead of removing oxygen, oxygen is added to cultures by daily sparging of serum bottles with a mixture of compressed oxygen and air. Sputum from 50 pwCF underwent daily sparging for a 72-h period to verify the ability of this model to maintain differential oxygen conditions. Shotgun metagenomic sequencing was performed on cultured and uncultured sputum samples from 11 pwCF to verify the ability of this medium to support the growth of commensal and pathogenic microbes commonly found in cystic fibrosis sputum. Growth curves were obtained from 112 isolates obtained from pwCF to verify the ability of this artificial sputum medium to support the growth of common cystic fibrosis pathogens. We find that this model can culture a wide variety of pathogens and commensals in CF sputum, recovers a community highly similar to uncultured sputum under normoxic conditions, and creates different culture phenotypes under varying oxygen conditions. This new approach might lead to a better understanding of unanticipated effects induced by the use of oxygen in pwCF on airway microbial communities and common respiratory pathogens.
Procedura
Airway microbial communities are thought to play an important role in the progression of cystic fibrosis (CF) and other chronic pulmonary diseases. Microbes have traditionally been classified based on their ability to use or tolerate oxygen. Supplemental oxygen is a common medical therapy administered to people with cystic fibrosis (pwCF); however, existing studies on oxygen and the airway microbiome have focused on how hypoxia (low oxygen) rather than hyperoxia (high oxygen) affects the predominantly aerobic and facultative anaerobic lung microbial communities. To address this critical knowledge gap, this protocol was developed using an artificial sputum medium that mimics the composition of sputum from pwCF. The use of filter sterilization, which yields a transparent medium, allows optical methods to follow the growth of single-celled microbes in suspension cultures. To create hyperoxic conditions, this model system takes advantage of established anaerobic culturing techniques to study hyperoxic conditions; instead of removing oxygen, oxygen is added to cultures by daily sparging of serum bottles with a mixture of compressed oxygen and air. Sputum from 50 pwCF underwent daily sparging for a 72-h period to verify the ability of this model to maintain differential oxygen conditions. Shotgun metagenomic sequencing was performed on cultured and uncultured sputum samples from 11 pwCF to verify the ability of this medium to support the growth of commensal and pathogenic microbes commonly found in cystic fibrosis sputum. Growth curves were obtained from 112 isolates obtained from pwCF to verify the ability of this artificial sputum medium to support the growth of common cystic fibrosis pathogens. We find that this model can culture a wide variety of pathogens and commensals in CF sputum, recovers a community highly similar to uncultured sputum under normoxic conditions, and creates different culture phenotypes under varying oxygen conditions. This new approach might lead to a better understanding of unanticipated effects induced by the use of oxygen in pwCF on airway microbial communities and common respiratory pathogens.
