Pipeline for Multi-Scale Three-Dimensional Anatomic Study of the Human Heart
Summary
This protocol presents a comprehensive pipeline to analyze samples obtained from human hearts that span the microscopic and macroscopic scales.
Abstract
Detailed study of non-failing human hearts rejected for transplantation provides a unique opportunity to perform structural analyses across microscopic and macroscopic scales. These techniques include tissue clearing (modified immunolabeling-enabled three-dimensional (3D) imaging of solvent-cleared organs) and immunohistochemical staining. Mesoscopic examination procedures include stereoscopic dissection and micro-computed tomographic (CT) scanning. Macroscopic examination procedures include gross dissection, photography (including anaglyphs and photogrammetry), CT, and 3D printing of the physically or virtually dissected or whole heart. Before macroscopic examination, pressure-perfusion fixation may be performed to maintain the 3D architecture and physiologically relevant morphology of the heart. The application of these techniques in combination to study the human heart is unique and crucial in understanding the relationship between distinct anatomic features such as coronary vasculature and myocardial innervation in the context of the 3D architecture of the heart. This protocol describes the methodologies in detail and includes representative results to illustrate progress in the research of human cardiac anatomy.
Introduction
As function follows form, understanding the architecture of the heart is fundamental for appreciation of its physiology. Although numerous investigations have revealed cardiac anatomy from micro- to macroscales1,2,3, multiple questions remain unresolved, especially those related to human cardiac anatomy. This is in part because basic studies focusing on functional anatomy generally utilized animal hearts4,5,6, which are often distinct from human hearts1,7,8. Furthermore, each individual study, even those using human heart samples, tends to focus on very specific structures, which renders it difficult to apply the findings in the context of the whole heart. This is even more so if the focused structures are at micro- or mesoscales, such as the perinexus9 and ganglionated plexuses10.
In this context, systemic structural study of the human heart rejected for transplantation provides a unique and rare opportunity to obtain a comprehensive atlas of cardiac structures in focus across microscopic and macroscopic scales11. Microscopic examination protocols include tissue clearing (modified immunolabeling-enabled three-dimensional (3D) imaging of solvent-cleared organs, iDISCO+)12,13, and immunohistochemical staining. Mesoscopic examination protocols include stereoscopic dissection, macro photography, and micro-computed tomographic (CT) scanning. Macroscopic examination protocols include gross dissection14, photography (including anaglyphs and photogrammetry)15,16,17, CT, virtual dissection18, and 3D printing of the physically or virtually dissected or whole heart17. In preparation for macroscopic examination, pressure-perfusion fixation is performed to maintain the 3D architecture and physiologically relevant morphology of the heart14,19,20,21. The combined application of these techniques is unique and crucial to correlate distinct anatomic features in the context of the 3D architecture of the human heart.
As the opportunity to obtain a non-pathological human heart sample is extremely limited, a multi-scale approach described herein maximizes the use of the sample. By applying various procedures described below, representative results will illustrate to the reader how the findings can be utilized for multiple purposes, including discovery in scientific research11 (comprehensive analyses of cardiac innervation, distribution of ganglionated plexuses), improvement of clinical procedures (simulation for surgical and interventional approaches), and anatomical education (real 3D demonstration of cardiac anatomy).
Protocol
This study used de-identified tissue samples collected from non-failing donor human hearts and was approved by the Institutional Review Board of the University of California, Los Angeles (UCLA). Samples were obtained from non-failing hearts that were rejected for transplantation. The hearts were pressure-perfused, fixed in 4% paraformaldehyde (PFA), and imaged before tissue processing per the following methods. Figure 1 summarizes the flow chart of the order of the study. The details of the reagents and the equipment used in the study are listed in the Table of Materials.
1. Micro-scale examination
- Tissue clearing using modified immunolabeling-enabled 3D imaging of solvent-cleared organs (iDISCO+) protocol.
- Dissect 4% PFA-fixed tissue with a scalpel to fit within the 3 mm × 16 mm × 25 mm chamber for confocal microscopy. To image thicker tissues, additional chambers and/or spacers may be stacked on the slide.
- Dehydrate specimens using graded methanol (MeOH) series (20%, 40%, 60%, 80%, and 100% MeOH in deionized H2O [vol/vol]) for 1 h each at room temperature (RT) with agitation.
- Wash with 100% MeOH for 1 h at RT and immerse in 66% dichloromethane/33% MeOH at RT with agitation overnight.
- The next day, wash twice in MeOH (100%) for 1 h at RT, chill at 4 °C, and treat with 5% H2O2 in MeOH (vol/vol) overnight at 4 °C.
- Rehydrate with graded MeOH series (80%, 60%, 40%, and 20% MeOH) and wash in 0.01 mol/L PBS for 1 h each at RT with agitation.
- Wash the tissues twice in 0.01 mol/L PBS with 0.2% Triton X-100 for 1 h at RT.
- Prepare for immunolabeling by permeabilizing in 0.01 mol/L PBS, 20% dimethyl sulfoxide (DMSO), 0.2% Triton X-100, and 0.3 mol/L glycine for 2 days at 37 °C with agitation.
- Block in 0.01 mol/L PBS with 10% DMSO, 0.2% Triton X-100, and 5% normal donkey serum for another 2 days at 37 °C with agitation.
- Label with a primary antibody that is compatible with MeOH conjugated to fluorophores diluted in 0.01 mol/L PBS with 10 mg/mL heparin (PTwH), 0.2% Tween-20, 5% DMSO, and 3% normal donkey serum for 3-4 days at 37 °C with agitation.
- Replenish the antibody solution and incubate for another 3-4 days at 37 °C with agitation.
- After 1-week incubation in primary antibody solution, wash 4 to 5 times in PTwH overnight at RT.
- Incubate with secondary antibody conjugated to fluorophores diluted in PTwH, 3% normal donkey serum for 3 days at 37 °C with agitation.
- Replenish the secondary antibody solution incubate for another 3 days at 37 °C with agitation.
- After 6-day incubation in secondary antibody solution, wash in PTwH 4-5 times overnight at RT.
- Dehydrate with a graded MeOH series (20%, 40%, 60%, 80%, 100%, and 100% MeOH). The sample may be stored overnight at RT.
- Incubate in 66% dichloromethane/33% MeOH for 3 h at RT with agitation.
- Wash twice in 100% dichloromethane for 15 min at RT with agitation.
- Incubate and store specimens in benzyl ether. Fill the tube to minimize air from oxidizing the sample.
- Imaging tissue-cleared specimen
- Affix a chamber containing adhesive to a slide and apply nail polish around the perimeter of the chamber. For thicker tissues, additional chambers and/or spacers may be stacked on the slide.
- Place cleared tissue in the chamber, fill with benzyl ether, and apply a coverslip.
- Apply nail polish around the coverslip to create a seal.
- Obtain tilescan and Z stack images using an upright laser scanning confocal microscope with a 5x or 10x lens to image at a depth up to the working distance of the lens.
- Image at a resolution of 1024 x 1024 using laser lines appropriate for emission spectra of fluorophores used. Muscle autofluorescence is visible using the 488 nm laser line.
- Ensure that the z-axis step size is commensurate with Nyquist sampling based on the numerical aperture of the specified objective11.
- Stitch images and use software for 3D visualization.
- Create figures using maximum intensity projection (MIP) images of Z stacks for individual and merged channels (Figure 2).
- Immunohistochemistry
NOTE: After the tissue is paraffin-embedded22, the following procedure is used to create slides for immunohistochemical study.- Preparation of refractive index matching solution (RIMS).
- Prepare 0.1 mol/L phosphate buffer by adding 10.9 g of Na2HPO4 (anhydrous) and 3.1 g of NaH2PO4 (monohydrate) to deionized H2O to a total volume of 1 L (pH 7.4). Filter-sterilize the solution and store it at RT.
- Dilute the phosphate buffer to 0.02 mol/L.
- Dissolve Histodenz in 30 mL of 0.02 mol/L phosphate buffer by stirring the solution for 10 min with a magnetic stir bar in the final storage bottle that may be sealed to minimize evaporation and contamination.
- Add sodium azide to a total concentration of 0.01% (w/v) and adjust the pH to 7.5 with NaOH.
- Adjust the RI by varying the final concentration of Histodenz.
- Store the RIMS at RT for months. Discard if microbial contamination is noted.
NOTE: Do NOT autoclave any solutions containing sodium azide.
- Preparation of slides for immunohistochemical study
- Create sections of 5 µm thickness with microtome. Apply tissue section to charged slides.
- Remove paraffin by incubating the slides in >75% xylene for 10 min. Move slides to a second container with xylene for an additional 10 min.
- Remove xylene by immersing the slides in 100% EtOH for 10 min, then in 95% EtOH for 5 min and 70% EtOH for 5 min.
- Rinse the slides with deionized H2O for 5 min.
- Immerse slides in antigen retrieval buffer for 25 min at 90-95°C.
- Allow the container to cool to RT for 1 h with agitation.
- Immerse the slides in soaking buffer (0.01 mol/L PBS + 0.4% Triton X-100) for 30 min at 4°C.
- Encircle the tissue with a PAP pen. Add PBS to each slide and place it in a humidified chamber to prevent desiccation.
- Wash slides with PBS at RT with agitation for 5 min.
- Block with blocking buffer (0.01 mol/L PBS + 10% Donkey Serum + 0.1% TX-100) for 1 h with agitation.
- Incubate with a primary antibody diluted in blocking buffer overnight at 4°C.
- The next day, allow slides to warm to RT for 15 min.
- Wash the slides 3 times with 0.01 mol/L PBS + 0.2% TritonX-100 for 5 min.
- Incubate with a secondary antibody diluted in blocking buffer for 1 h at RT with agitation.
- Wash the slides 3 times with 0.01 mol/L PBS + 0.2% TritonX-100 for 5 min.
- Wash the slides 3 times with 0.01 mol/L PBS for 5 min.
- Place 1 drop of RIMS with a dropper and apply a coverslip.
- Apply nail polish around the coverslip to create the seal.
- As a negative control, run a sample without the primary antibody to demonstrate the absence of specific staining.
- Imaging immunostained slides
- Image the slides with a laser scanning confocal microscope with 10x, 20x and 40x objective lenses.
- Image at a resolution of 1024 x 1024 using laser lines appropriate for the emission spectra of the secondary antibodies used.
- Create figures using maximum intensity projection (MIP) images of Z stacks for individual and merged channels (Figure 3).
- Preparation of refractive index matching solution (RIMS).
2. Meso-scale examination
- Stereoscopic dissection
- Perform delicate dissections focusing on tiny or thin structures, such as atrioventricular node, atrioventricular node artery, and cardiac nerve plexus (submillimeter to millimeters in scale) with either a magnifying desk lamp with clamp, surgical telescopes, or stereomicroscope.
- Micro-CT scanning
NOTE: CT imaging is performed after pressure perfusion and fixation and at any stages of dissection using a micro-positron emission tomography (PET)/CT scanner (Figure 4).- Warm up the CT X-ray source for 25 min before sample imaging.
- Place the heart sample on the scanner bed.
- Move the scanner bed to a horizontal position of 544 mm and a vertical position of 14 mm to center the heart in the CT field of view (FOV).
- Acquire CT image at 80 kVp, 150 µA, with 720 projections during 1 min scan time at a spatial resolution of 200 µm.
- Reconstruct the CT data with a 12 cm x 12 cm x 10 cm field of view and a matrix of 600 x 600 x 500 voxels, and save as a DICOM file.
3. Macro-scale examination
- Pressure perfusion and fixation
NOTE: The authors modify previously described pressure perfusion and fixation techniques and apply them to the non-failing human hearts rejected for transplantation14,19,20,21.- Use high-flow pumps for perfusion fixation. Use either 100% ethanol14, 4% PFA, or 10% formalin for the fixative.
NOTE: The heart is recovered with the ascending aorta, pulmonary trunk, and both vena cavae, and pulmonary veins resected as distally as possible and delivered in University of Wisconsin solution23. - Use two 20-24 Fr surgical cannulas for right- and left-heart perfusion. For right-heart perfusion, cannulate the superior vena cava, and place a vent at the pulmonary trunk or pulmonary artery with half-cut 12-30 mL sized plastic syringes with Luer-Lock tips attached to three-way stopcocks.
- Occlude the inferior vena cava and the other pulmonary artery with twine after putting an appropriately sized locked half-cut plastic syringe or 1.5-5.0 mL centrifuge tube.
- For antegrade perfusion of the left heart, cannulate one of the pulmonary veins and place a vent at the distal cut end of the aorta with the half-cut 12-30 mL sized plastic syringes with Luer-Lock tips attached to three-way stopcocks.
- For retrograde perfusion of the left heart, cannulate one of the aortic arch branches and place a vent at another branch of the aortic arch. Place the tips of the cannulas in both ventricles.
- Occlude other vessel orifices with twine after inserting an appropriately sized locked half-cut plastic syringe or 1.5-5.0 mL microcentrifuge tube. Use a thin gauze to cover the inserting portion of the syringes/ tubes/ cannulas to prevent leakage and slipping. Fix large leaks using suture, banding, or clumping. Small leaks are permissible.
- Suspend the heart in a plastic container.
- Connect 22-24 Fr soft plastic tubing to each cannula and insert the other end of the tube into the container filled with fixative.
- Circulate fixative through the right- and left-heart circuits using a high-flow pump set at approximately 100-300 mL/min for the right heart and 200-400 mL/min for the left heart to achieve approximately 20 mmHg in the right ventricle and 80 mmHg in the left ventricle, respectively.
- Maintain perfusion at 4 °C for 24 h.
- Wash the heart with 0.01 mol/L PBS for 30 min with agitation four times.
- Store heart in 0.01 mol/L PBS/0.02% sodium azide at 4 °C.
NOTE: The pressure-perfusion fixation is only effective for a fresh heart, not for a heart recovered from an embalmed cadaver.
- Use high-flow pumps for perfusion fixation. Use either 100% ethanol14, 4% PFA, or 10% formalin for the fixative.
- Gross dissection
- Perform progressive dissection with photographic recordings at each stage of dissection.
- To maintain clinical relevance, pay special attention to avoid distorting/deforming any structures to maintain the physiological morphology of the heart.
- Image target structures using clinically relevant orientation, such as right anterior oblique orientation.
- Photography
- Place the pressure-perfused and fixed heart on a tripod with a platform mounted with multiple prongs and the ability to rotate 360o.
- Photograph the heart using a digital single-lens reflex camera (Figure 5)24 while using multiple light-emitting diode light panels set on the C-Stands and wide black duvetyn background cloth.
- Photograph using the lens with a long focal length (200 mm) for a working distance of 4-6 ft to minimize distortion of the subject14.
- Anaglyphs
- To display anaglyphic images, reconstruct a pair of photographs or volume-rendered images from CT datasets with a 10° difference in the rotation angle on the horizontal plane.
- Convert a set of these two-dimensional (2D) images, referred to as a stereogram, into anaglyphs using freeware16.
- To view an anaglyph, use red/cyan glasses.
- Photogrammetry
NOTE: Photogrammetry is the applied science of generating a three-dimensional surface-rendered reconstruction from multiple two-dimensional photographs taken at varying angles17.- Suspend the sample on C-Stands or place it on the rotation table to obtain hundreds of multi-directional photographs with a smartphone.
- Generate the 3D model in FBX format using commercially available software.
- CT scanning
NOTE: CT scanning may be performed after pressure perfusion and fixation and at any stage of dissection.- Suspend the heart sample from a bar placed across the top of the container. To prevent the heart from swinging during the scan, support the base of the heart with plastic prongs fixed at the bottom of the container. Thus, the air will serve as a negative contrast.
- Perform the CT scan using a commercially available multi-detector row CT scanner with the following parameters: tube voltage of 120 kV, tube current of 800-900 mA, and a gantry rotation of 280 ms. The dose length product is generally 500-1200 mGy.cm.
- Reconstruct axial image data using the following parameters: a section thickness, 0.6 mm; an incremental interval, 0.3 mm; a field of view, as small as possible (generally 100-200 mm); and a matrix, 512 × 512.
- Virtual dissection
- Analyze the CT scan images using commercially available software to generate virtual dissection images.
NOTE: Virtual dissection is a modification of the volume-rendering process wherein the focus is shifted to the walls of the cardiac chambers and vessels18. In this process, manual thresholding virtually removes the enhanced chamber from the original datasets. - Visualize the non-enhanced walls, septa, and valves with virtual dissection to produce images similar to gross dissection. Unlike gross dissection of the heart specimens, the cut planes during virtual dissection are practically unlimited. Almost any view can be recreated to visualize the structures of interest as needed.
- Analyze the CT scan images using commercially available software to generate virtual dissection images.
- 3D printing
- Open the compatible file of the heart sample in 3D printer software.
- Use the profile of 0.10 mm Fast DETAIL for the print Settings in the 3D printer and reduce the print speed to 20 mm/s. Enable Generate support material.
- Use the profile of TPU filament for "Filament Settings" in the 3D printer.
- Use the profile of the Original Prusa MK4 Input Shaper 0.4 nozzle for "Printer Settings" in the printer.
- After slicing is completed, save the BGCODE file in a USB flash drive for 3D printing.
- Use the 1.75 mm TPU filament for 3D printing the human heart sample. Before 3D printing, dry the TPU filament for 6 h using a filament dryer.
- To reduce filament tension during 3D printing, place the filament spool on a spool holder with a built-in bearing to facilitate filament spool rotation. Perform 3D printing using a commercially available 3D printer with a textured powder-coated steel sheet.
- Carefully remove support materials when 3D printing is complete.
Representative Results
Microscale examinations
Applying tissue clearing allows imaging of larger volumes of tissue in 3D using confocal microscopy. In the heart, ganglia containing cardiac neurons and the neural patterning of myocardial innervation can be visualized (Figure 2). Figure 3 shows a confocal image of the human left ventricle myocardium immunostained for nerves and smooth muscle cells. Blood vessels are noted to traverse the myocardium, and numerous nerve fibers are identified, both in association with and independent of blood vessels.
Meso- and macroscale examinations
When using absolute ethanol for 24 h pressure perfusion and fixation, the natural color of the sample is bleached, the tissue is dehydrated25, and elasticity is significantly reduced. On the other hand, upon fixation with PFA and formalin, natural color and elasticity are remarkably maintained. For these reasons, PFA or formalin is mainly used as the preferred fixative.
Figure 6 shows representative images of the gross dissection, virtual dissection, STL polygon model, and three-dimensional printing. Figure 7 shows representative images of the anaglyphs created from both gross and virtual dissection images. Depth perception can be obtained with anaglyphic glasses. The single photogrammetric model captured can be observed from almost all directions using commercially available software and demonstrates complex anatomical features relevant to routine transcatheter cardiac procedures. By applying these techniques to the heart prepared with pressure perfusion and fixation, three-dimensional information about the heart can be almost eternally retained either digitally or physically and shared without boundaries. Figure 8 shows the 50%-scale3D printings replicated from the dissected hearts rejected for transplantation.
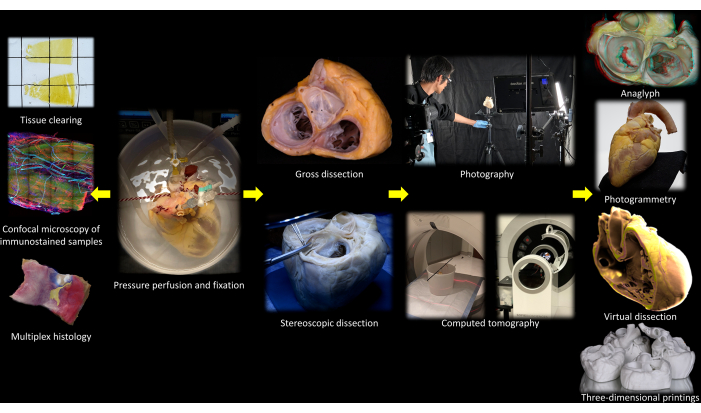
Figure 1: Flow chart of the protocol. Please click here to view a larger version of this figure.
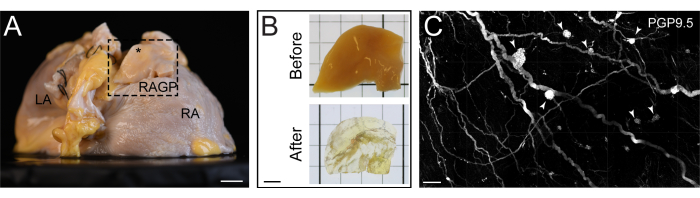
Figure 2: Tissue-cleared section of the human right atrium. (A) Right posterior oblique view of a human right atrium with the right atrial ganglionated plexus (RAGP) dissected for tissue clearing. (B) RAGP specimen before and after tissue clearing. (C) Maximum intensity projection of an iDISCO+-cleared portion of human RAGP demonstrating ganglia (arrowheads) immunostained with pan-neuronal marker protein gene product 9.5 (PGP9.5). Scale bars are 1 cm (A), 5 mm (B), and 500 µm (C). This figure is adapted from Hanna et al.11. Please click here to view a larger version of this figure.
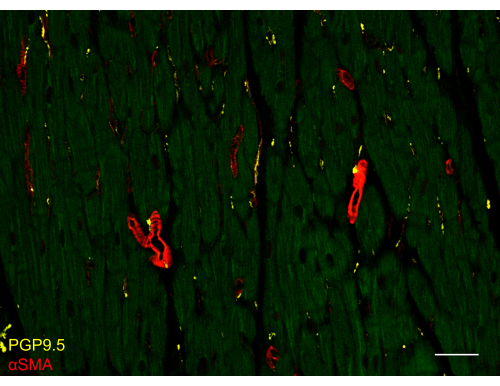
Figure 3: Immunostained slides of the human left ventricle. Confocal image from the human left ventricle myocardium slice immunostained with pan-neuronal marker protein gene product 9.5 (PGP9.5) and smooth muscle cell marker α-smooth muscle actin (αSMA). Muscle autofluorescence is visible using the 488 nm laser line (green). The scale bar is 50 µm. Please click here to view a larger version of this figure.
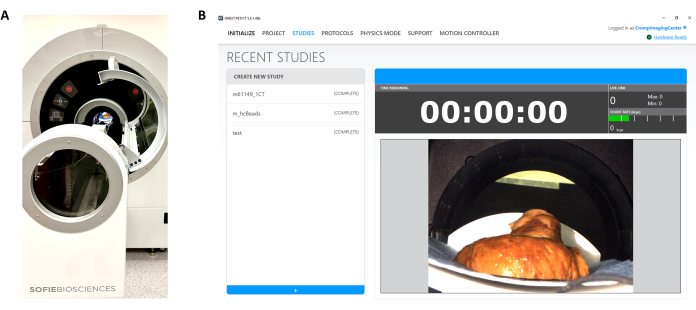
Figure 4: Micro-computed tomographic imaging of heart samples. (A) Micro-computed tomography setup for heart sample imaging. (B) User interface for micro-computed tomography imaging. Please click here to view a larger version of this figure.
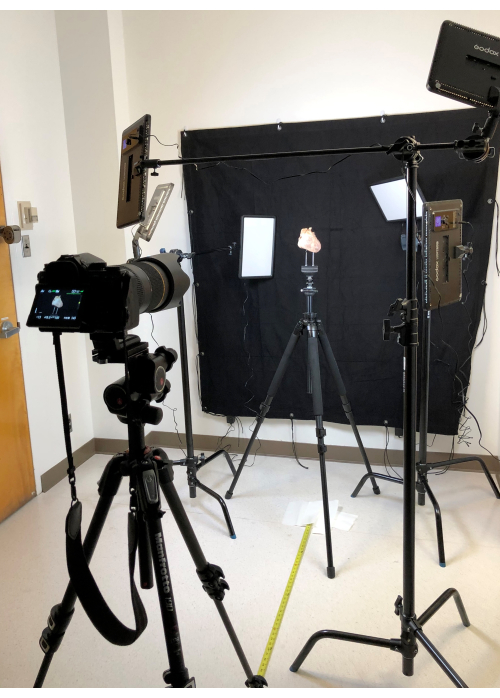

Figure 5: Photo-studio setting in UCLA Cardiac Arrhythmia Center. Please click here to view a larger version of this figure.
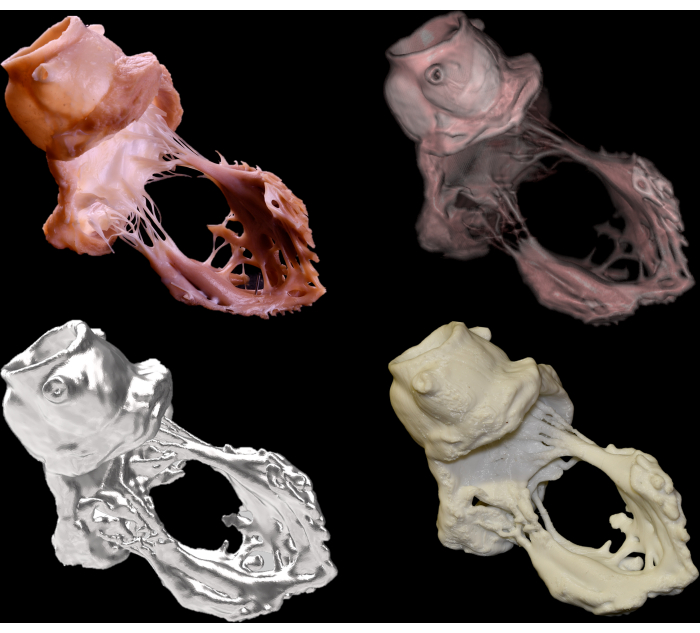
Figure 6: Gross dissection (upper left), virtual dissection (upper right), STL polygon model (bottom left), and three-dimensional printing (bottom right) images of the aortic and mitral valvular complex. Please click here to view a larger version of this figure.
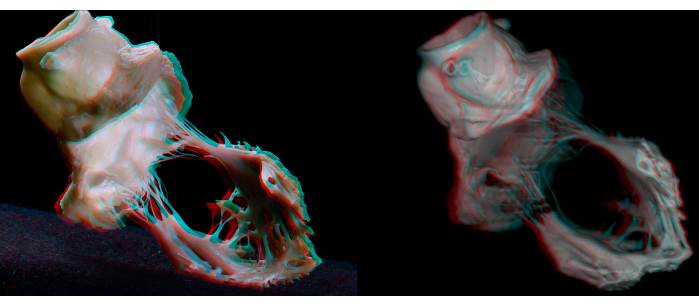
Figure 7: Anaglyphs of a gross dissection (left) and virtual dissection (right) of the aortic and mitral valvular complex. Anaglyphic glasses (red/cyan) are necessary to obtain depth perception. Please click here to view a larger version of this figure.
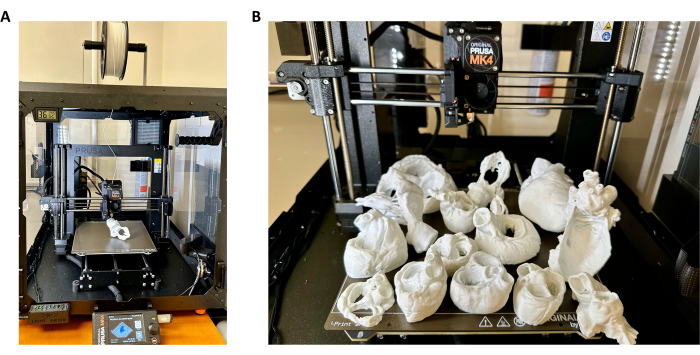

Figure 8: Three-dimensional printing of heart samples. (A) Three-dimensional (3D) printer setup for heart sample 3D printing with a TPU filament. (B) Representative heart 3D prints produced using methods described in this study. Please click here to view a larger version of this figure.
Discussion
The present study demonstrates the comprehensive pipeline to analyze samples obtained from whole human hearts. Representative results show micro- to macroscale anatomical examinations carried out routinely for a single heart. As a human heart sample is extremely precious, a multi-scale approach is ideal and effective so as not to waste any parts of the sample by applying multiple protocols for various purposes, including discovery in scientific research, improvement of clinical procedures, and anatomical education with maintaining anatomic correlation in the context of the whole heart.
Regarding microscale examination, immunostaining and microscopy may be applied to understand the cytoarchitecture of human cardiac specimens. Here, the application of tissue clearing and immunohistochemistry to study cardiac neuroanatomy at the cellular scale is demonstrated. The use of these techniques is helpful in the characterization of the cardiac nervous system and myocardial innervation patterns in relation to structures of interest, such as the cardiac conduction system and cardiac chambers. Although excellent spatial resolution is achieved, the use of confocal microscopy, particularly for tissue-cleared specimens, is time-consuming. Lightsheet microscopy may be used to reduce the time for image acquisition at the expense of spatial resolution.
Regarding meso- to macroscale examination, the spatial resolution of micro-CT scanners in the authors' institution ranges from 10-200 µm. The sample size is limited to 20 mm for a 10 µm scan, and 120 mm for a 100-200 µm scan. Micro-CT scanners in the authors' institution cannot accommodate the whole heart. Thus, at the authors' institution, a whole heart scan requires the use of a clinical CT scanner with 600 µm spatial resolution, although advances have allowed for the development of micro-CT scanners that can image the whole heart2. Technological development, such as photon counting CT, will surely expand further possibilities. Improvement of spatial resolution of the STL file should be the first step to further improve the quality of 3D printing. The higher cost of 3D printing limits the application of the technique to routine clinical practice. Photogrammetry images generated from any smartphone application are easy to develop and of acceptable quality but will require further sophisticated but expensive and time-consuming systems to improve resolution26. To visualize in 3D, extended reality with dedicated head gear27,28 and holographic monitors29 are additional tools but are also limited by higher cost.
In summary, through comprehensive structural analyses across microscopic and macroscopic scales, the microscale anatomy of each structure and its functional contribution can be understood in the context of the whole heart. Along with the development of high-resolution imaging, the distance between the micro- and macroscale anatomy is dramatically expanding. Experts in electron microscopic analysis of cardiomyocytes may not be familiar with the number of mitral leaflets and vice versa. To facilitate the comprehensive understanding of cardiac morphology, scientists must keep exploring further details of each tree, while maintaining the bird's eye view of the entire forest.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank the individuals who have donated their bodies for the advancement of education and research. We are grateful to the OneLegacy Foundation, which formed the basis for obtaining donor hearts for research. We are also grateful to Anthony A. Smithson and Arvin Roque-Verdeflor of the UCLA Translational Research Imaging Center (Department of Radiology) for their support in CT data acquisition. This project was supported by the UCLA Amara Yad Project. We are thankful to Drs. Kalyanam Shivkumar and Olujimi A. Ajijola for establishing and maintaining a human heart pipeline for research. We appreciate our Research Operations Manager, Amiksha S. Gandhi for her dedication to support our projects. This work was made possible by support from NIH grants OT2OD023848 & P01 HL164311 and Leducq grant 23CVD04 to Kalyanam Shivkumar, the American Heart Association Career Development Award 23CDA1039446 to PH, and the UCLA Amara-Yad Project (https://www.uclahealth.org/medical-services/heart/arrhythmia/about-us/amara-yad-project). The GNEXT microPET/CT scanner used in this study was funded by an NIH Shared Instrumentation for Animal Research Grant (1 S10 OD026917-01A1).
Materials
| 1x Phosphate buffered saline | Sigma-Aldrich | P3813 | |
| 3D Viewer | Microsoft | ||
| 647 AffiniPure Donkey Anti-Rabbit IgG | Jackson ImmunoResearch Laboratories | 711-605-152 | |
| 647 AffiniPure Donkey Anti-Sheep IgG | Jackson ImmunoResearch Laboratories | 713-605-147 | |
| AF Micro-NIKKOR 200 mm f/4D IF-ED lens | Nikon | ||
| Anti-Actin, α-Smooth Muscle – Cy3 antibody | Sigma-Aldrich | C6198 | |
| Antigen Retrieval Buffer (100x EDTA Buffer, pH 8.0) | Abcam | ab93680 | |
| Anti-PGP9.5 (protein gene product 9.5) | Abcam | ab108986 | |
| Anti-TH (tyrosine hydrox ylase) | Abcam | ab1542 | |
| Anti-VAChT (vesicular acetylcholine transporter) | Synaptic Systems | 139 103 | |
| Benzyl ether | Sigma-Aldrich | 108014 | |
| Bovine serum albumin | Sigma-Aldrich | A4503-10G | |
| Cheetah 3D printer filament (95A), 1.75 mm | NinjaTek | ||
| Coverslip, 22 mm x 30mm, No. 1.5 | VWR | 48393 151 | |
| Cy3 AffiniPure Donkey Anti-Rabbit IgG | Jackson ImmunoResearch Laboratories | 711-165-152 | |
| Dichloromethane | Sigma-Aldrich | 270997-100ML | |
| Dimethyl sulfoxide | Sigma-Aldrich | D8418-500ML | |
| Ethanol, 100% | Decon laboratories | 2701 | |
| Glycine | Sigma-Aldrich | G7126-500G | |
| GNEXT PET/CT | SOFIE Biosciences | ||
| Heparin sodium salt from porcine intestinal mucosa | Sigma-Aldrich | H3149-50KU | |
| Histodenz | Sigma-Aldrich | D2158-100G | |
| Hydrogen peroxide solution | Sigma-Aldrich | H1009-500ML | |
| Imaging software | Zeiss | ZEN (black edition) | |
| Imaging software | Oxford Instruments | Imaris 10 | |
| iSpacer | Sunjin Labs | iSpacer 3mm | |
| KIRI Engine | KIRI Innovation | ||
| Laser scanning confocal microscope | Zeiss | LSM 880 | |
| LEAD-2 – Vertical & Multi-channels Peristaltic Pump | LONGER | ||
| Lightview XL | Brightech | ||
| Methanol (Certified ACS) | Fischer Scientific | A412-4 | |
| Nikon D850 | Nikon | ||
| NinjaTek NinjaFlex TPU @MK4 | NinjaTek | ||
| Normal donkey serum | Jackson ImmunoResearch Laboratories | 017-000-121 | |
| Original Prusa MK4 3D printer | Prusa Research | ||
| PAP pen | Abcam | ab2601 | |
| Paraformaldehyde, 32% | Electron Microscopy Sciences | 15714-S | |
| Polycam | Polycam | ||
| Primary antibody | |||
| PrusaSlicer 2.7.1 | Prusa Research | ||
| SARA-Engine | pita4 mobile LLC | ||
| Scaniverse | Niantic | ||
| Secondary antibody | |||
| SlowFade Gold Antiface Mountant | Invitrogen | S36936 | |
| Sodium azide, 5% (w/v) | Ricca Chemical Company | 7144.8-32 | |
| SOMATOM Definition AS | Siemens Healthcare | ||
| Standard Field Surgi-Spec Telescopes, | Designs for Vision | ||
| Stereomicroscope System SZ61 | OLYMPUS | ||
| StereoPhoto Maker | Free ware developed by Masuji Suto | ||
| Superfrost Plus Microscope Slides, Precleaned | Fisher Scientific | 12-550-15 | |
| Triton X-100 | Sigma-Aldrich | T8787-50ML | |
| Tween-20 | Sigma-Aldrich | P9416-100ML | |
| Xylene | Sigma-Aldrich | 534056-4L | |
| Ziostation2 | Ziosoft, AMIN |
References
- Das reizleitungssystem des säugetierherzens. Eine anatomisch-histologische studie über das atrioventrikularbündel und die purkinjeschen fäden. Jena: Gustav fischer Available from: https://wellcomecollection.org/works/gnxjxg9e (1906)
- Stephenson, R. S., et al. High-resolution 3-dimensional imaging of the human cardiac conduction system from microanatomy to mathematical modeling. Sci Rep. 7 (1), 7188 (2017).
- Kawashima, T., Sato, F. First in situ 3D visualization of the human cardiac conduction system and its transformation associated with heart contour and inclination. Sci Rep. 11 (1), 8636 (2021).
- Bojsen-Moller, F., Tranum-Jensen, J. Whole-mount demonstration of cholinesterase-containing nerves in the right atrial wall, nodal tissue, and atrioventricular bundle of the pig heart. J Anat. 108 (Pt 3), 375-386 (1971).
- Zhao, Y., et al. Ganglionated plexi and ligament of marshall ablation reduces atrial vulnerability and causes stellate ganglion remodeling in ambulatory dogs. Heart Rhythm. 13 (10), 2083-2090 (2016).
- Chung, W. H., et al. Ischemia-induced ventricular proarrhythmia and cardiovascular autonomic dysreflexia after cardioneuroablation. Heart Rhythm. 20 (11), 1534-1545 (2023).
- Crick, S. J., Sheppard, M. N., Ho, S. Y., Gebstein, L., Anderson, R. H. Anatomy of the pig heart: Comparisons with normal human cardiac structure. J Anat. 193 (Pt 1), 105-119 (1998).
- Saburkina, I., Pauziene, N., Solomon, O. I., Rysevaite-Kyguoliene, K., Pauza, D. H. Comparative gross anatomy of epicardiac ganglionated nerve plexi on the human and sheep cardiac ventricles. Anat Rec (Hoboken). 306 (9), 2302-2312 (2023).
- Hoagland, D. T., Santos, W., Poelzing, S., Gourdie, R. G. The role of the gap junction perinexus in cardiac conduction: Potential as a novel anti-arrhythmic drug target. Prog Biophys Mol Biol. 144, 41-50 (2019).
- Aksu, T., Gopinathannair, R., Gupta, D., Pauza, D. H. Intrinsic cardiac autonomic nervous system: What do clinical electrophysiologists need to know about the "heart brain". J Cardiovasc Electrophysiol. 32 (6), 1737-1747 (2021).
- Hanna, P., et al. Innervation and neuronal control of the mammalian sinoatrial node a comprehensive atlas. Circ Res. 128 (9), 1279-1296 (2021).
- Rajendran, P. S., et al. Identification of peripheral neural circuits that regulate heart rate using optogenetic and viral vector strategies. Nature Communications. 10, 1944 (2019).
- Renier, N., et al. Mapping of brain activity by automated volume analysis of immediate early genes. Cell. 165 (7), 1789-1802 (2016).
- Mcalpine, W. Heart and coronary arteries: An anatomical atlas for clinical diagnosis, radiological investigation, and surgical treatment. Springer-verlag. , (1975).
- Mori, S., Shivkumar, K. Stereoscopic three-dimensional anatomy of the heart: Another legacy of dr. Wallace a. Mcalpine. Anat Sci Int. 96 (3), 485-488 (2021).
- Izawa, Y., Nishii, T., Mori, S. Stereogram of the living heart, lung, and adjacent structures. Tomography. 8 (2), 824-841 (2022).
- Sato, T., Hanna, P., Ajijola, O. A., Shivkumar, K., Mori, S. Photogrammetry of perfusion-fixed heart: Innovative approach to study 3-dimensional cardiac anatomy. JACC Case Rep. 21, 101937 (2023).
- Tretter, J. T., Gupta, S. K., Izawa, Y., Nishii, T., Mori, S. Virtual dissection: Emerging as the gold standard of analyzing living heart anatomy. J Cardiovasc Dev Dis. 7 (3), 30 (2020).
- Thomas, A. C., Davies, M. J. The demonstration of cardiac pathology using perfusion-fixation. Histopathology. 9 (1), 5-19 (1985).
- Glagov, S., Eckner, F. A., Lev, M. Controlled pressure fixation apparatus for hearts. Arch Pathol. 76, 640-646 (1963).
- Iaizzo, P. A. The visible heart(r) project and free-access website ‘atlas of human cardiac anatomy’. Europace. 18 (suppl 4), iv163-iv172 (2016).
- Yang, Y., Huang, H., Li, L., Yang, Y. Multiplex immunohistochemistry staining for paraffin-embedded lung cancer tissue. J Vis Exp. 201, e65850 (2023).
- Tripathy, S., Das, S. K. Strategies for organ preservation: Current prospective and challenges. Cell Biol Int. 47 (3), 520-538 (2023).
- Mori, S., Shivkumar, K. Atlas of cardiac anatomy (anatomical basis of cardiac interventions. Cardiotext. 1, (2022).
- Crosado, B., et al. Phenoxyethanol-based embalming for anatomy teaching: An 18 years’ experience with crosado embalming at the university of otago in new zealand. Anat Sci Educ. 13 (6), 778-793 (2020).
- Titmus, M., et al. A workflow for the creation of photorealistic 3d cadaveric models using photogrammetry. J Anat. 243 (2), 319-333 (2023).
- Silva, J. N. A., Southworth, M., Raptis, C., Silva, J. Emerging applications of virtual reality in cardiovascular medicine. JACC Basic Transl Sci. 3 (3), 420-430 (2018).
- Maresky, H. S., et al. Virtual reality and cardiac anatomy: Exploring immersive three-dimensional cardiac imaging, a pilot study in undergraduate medical anatomy education. Clin Anat. 32 (2), 238-243 (2019).
- Mori, S., Shivkumar, K. Real three-dimensional cardiac imaging using leading-edge holographic display. Clin Anat. 34 (6), 966-968 (2021).

.
