Enhanced Crosslinking Immunoprecipitation (eCLIP) Method for Efficient Identification of Protein-bound RNA in Mouse Testis
Summary
Here, we present an eCLIP protocol to determine major RNA targets of RBP candidates in testis.
Abstract
Spermatogenesis defines a highly ordered process of male germ cell differentiation in mammals. In testis, transcription and translation are uncoupled, underlining the importance of post-transcriptional regulation of gene expression orchestrated by RBPs. To elucidate mechanistic roles of an RBP, crosslinking immunoprecipitation (CLIP) methodology can be used to capture its endogenous direct RNA targets and define the actual interaction sites. The enhanced CLIP (eCLIP) is a newly-developed method that offers several advantages over the conventional CLIPs. However, the use of eCLIP has so far been limited to cell lines, calling for expanded applications. Here, we employed eCLIP to study MOV10 and MOV10L1, two known RNA-binding helicases, in mouse testis. As expected, we find that MOV10 predominantly binds to 3′ untranslated regions (UTRs) of mRNA and MOV10L1 selectively binds to Piwi-interacting RNA (piRNA) precursor transcripts. Our eCLIP method allows fast determination of major RNA species bound by various RBPs via small-scale sequencing of subclones and thus availability of qualified libraries, as a warrant for proceeding with deep sequencing. This study establishes an applicable basis for eCLIP in mammalian testis.
Introduction
Mammalian testis represents an excellent developmental model wherein an intricate cell differentiation program runs cyclically to yield numerous spermatozoa. An unique value of this model lies in the emergence of transcriptional inactivation at certain stages of spermatogenesis, typically when meiotic sex chromosome inactivation (MSCI) occurs1,2 and when round spermatids undergo drastic nuclear compaction during spermiogenesis3. These inconsecutive transcriptional events necessitate post-transcriptional gene regulation, in which RNA-binding proteins (RBPs) play a crucial role, shaping transcriptome and maintaining male fertility.
To identify the bona fide RNA targets of an individual RBP in vivo, the crosslinking immunoprecipitation (CLIP) method was developed4,5, based on but beyond the regular RNA immunoprecipitation (RIP)6,7, by incorporation of key steps including ultraviolet (UV) crosslinking, stringent wash and gel transfer to improve signal specificity. The advanced application of the CLIP combined with high-throughput sequencing has provoked large interest in profiling protein-RNA interaction at genome-wide levels8. In addition to genetic studies on RBP function, such biochemical methods that identify the direct interplay of endogenous protein and RNA have been indispensable to accurately elucidate the RNA regulatory roles of RBPs. For example, MOV10L1 is a testis-specific RNA helicase required for male fertility and the Piwi-interacting RNA (piRNA) biogenesis9. Its paralogue MOV10 is known as a ubiquitously expressed and multifunctional RNA helicase with roles in multiple aspects of RNA biology10,11,12,13,14,15,16,17,18. By employing the conventional CLIP-seq, we found that MOV10L1 binds and regulates primary piRNA precursors to initiate early piRNA processing19,20, and that MOV10 binds mRNA 3' UTRs and as well as noncoding RNA species in testicular germ cells (data not shown).
Nevertheless, CLIP is originally a laborious, radioactive procedure followed by sequencing library preparation with a remarkable loss of CLIP tags. In the conventional CLIP, a cDNA library is prepared using adapters ligated at both RNA extremities. After protein digestion, crosslinked short polypeptides remain attached to RNA fragments. This crosslinking mark partially blocks reverse transcriptase (RTase) progression during cDNA synthesis, resulting in truncated cDNAs which represent about 80% of the cDNA library21,22. Thus, only cDNA fragments resulting from RTase bypassing the crosslinking site (read-through) are sequenced. Recently, various CLIP approaches, such as PAR-CLIP, iCLIP, eCLIP and uvCLAP, have been employed to identify crosslink sites of RBPs in living cells. PAR-CLIP involves the application of 365 nm UV radiation and photoactivatable nucleotide analogs and is therefore exclusive to in-culturing living cells, and incorporation of nucleoside analogs into newly synthesized transcripts is prone to producing bias where RNA physically interacts with protein23,24. In iCLIP, only a single adapter is ligated to the 3' extremity of crosslinked RNA fragments. After reverse transcription (RT), both truncated and read-through cDNAs are obtained by intramolecularly circularization and re-linearization followed by polymerase chain reaction (PCR) amplification25,26. However, the efficiency of intramolecular circularization is relatively low. Although older CLIP protocols need labeling of crosslinked RNA with a radioisotope, ultraviolet crosslinking and affinity purification (uvCLAP), with a process of stringent tandem affinity purification, does not rely on radioactivity27. Nevertheless, uvCLAP is limited to cultured cells that must be transfected with the expression vector carrying the 3x FLAG-HBH tag for tandem affinity purification.
In eCLIP, adapters were ligated first at the 3' extremity of RNA followed by RT, and next at the 3' extremity of cDNAs in an intermolecular mode. Hence, eCLIP is able to capture all truncated and read-through cDNA28. Also, it is neither restricted to radioactive labeling, nor to using cell lines based on its principle, while maintaining single-nucleotide resolution.
Here, we provide a step-by-step description of an eCLIP protocol adapted for mouse testis. Briefly, this eCLIP protocol starts with UV crosslinking of testicular tubules, followed by partial RNase digestion and immunoprecipitation using a protein-specific antibody. Next, the protein-bound RNA is dephosphorylated, and adapter is ligated to its 3' end. After protein gel electrophoresis and gel-to-membrane transfer, RNA is isolated by cutting the membrane area of an expected size range. After RT, DNA adapter is ligated to the 3' end of cDNA followed by PCR amplification. Screening of subclones prior to high-throughput sequencing is taken as a library quality control. This protocol is efficient at identifying major species of protein-bound RNA of RBPs, exemplified by the two testis-expressing RNA helicases MOV10L1 and MOV10.
Protocol
All performed animal experiments have been approved by the Nanjing Medical University committee. Male C57BL/6 mice were kept under controlled photoperiod conditions and were supplied with food and water.
1. Tissue Harvesting and UV Crosslinking
- Euthanize 2 adult mice using carbon dioxide (CO2) for 1-2 min or until breathing stops. Next, perform cervical dislocation on each mouse.
- Harvest about 100 mg of testes from mice of appropriate age (one adult testis in this study) for each immunoprecipitation experiment, and place the tissues in ice-cold phosphate buffered saline (PBS).
- Remove the tunica albuginea gently with one pair of fine-tipped tweezers.
- Add 3 mL of ice-cold PBS in a tissue grinder and triturate the tissue by mild mechanical force using a loose glass pestle (type A glass pestle).
NOTE: The purpose of this step is not to lyse cells, but to pull apart the tissue. Preservation of cell viability and integrity is important. - Transfer the tissue suspension to a cell culture dish (10 cm in diameter) and add ice-cold PBS up to 6 mL.
- Shake the plate quickly so that liquid covers the bottom of the dish evenly. If the tissue is ground properly, evenly distributed seminiferous tubules will be visible, whereas the presence of tissue clumps indicates that tissue dispersion is suboptimal.
- Crosslink the suspension three times with 400 mJ/cm2 at 254 nm on ice. Mix suspension between each irradiation.
NOTE: For each new experiment, crosslinking should be optimized. - Collect the suspension in a 15 mL conical tube and pellet at 1,200 x g for 5 min at 4 °C. Remove the supernatant, resuspend the pellet in 1 mL of PBS and then transfer the suspension to a 1.5 mL centrifuge tube. Spin at 4 °C and 1,000 x g for 2 min, and remove supernatant.
- At this point, immediately proceed with the rest of the protocol, or snap freeze the pellets in liquid nitrogen and store at -80 °C until use.
2. Beads Preparation
- Add 125 µL of protein A magnetic beads per sample (pellet) to a fresh centrifuge tube.
NOTE: Use protein G magnetic beads for mouse antibodies. - Place the tube on the magnet to separate the beads from the solution. After 10 s, remove the supernatant. Wash beads twice with 1 mL of ice-cold lysis buffer.
NOTE: Subsequent separation of protein A magnetic beads follows this step. Lysis buffer composition is 50 mM Tris-HCl, pH 7.5; 100 mM NaCl; 1% NP-40; 0.1% SDS; 0.5% sodium deoxycholate; 1/50 ethylenediaminetetraacetic acid (EDTA)-free protease inhibitor cocktail (add fresh). - Resuspend beads in 100 µL of cold lysis buffer with 10 µg eCLIP antibody. Rotate tubes at room temperature for 45 min.
NOTE: The final antibody concentration for immunoprecipitation is 10 µg/mL. If the antibody concentration is unknown,the amount of antibody should be optimized. - Wash beads twice with 1 mL of ice-cold lysis buffer.
3. Tissue Lysis and Partial RNA Digestion
- Resuspend tissue pellets in 1 mL of cold lysis buffer (add 22 µL of 50x (EDTA)-free protein inhibitor cocktail and 11 µL of RNase inhibitor to 1 mL of lysis buffer).
- Resuspend two UV-crosslinked pellets and two non-crosslinked pellets per group of experiments: UV-crosslinked-1 pellets for eCLIP library (UV-1); UV-crosslinked-2 pellets for eCLIP library (UV-2); non-crosslinked-1 pellets for eCLIP library as a control (non-UV); non-crosslinked-2 pellets for IgG IP to demonstrate the specificity of antibodies.
NOTE: The ideal control for the eCLIP library of the UV-crosslinked wide-type testes is that of the UV-crosslinked knockout testes from mice of the same litter.
- Resuspend two UV-crosslinked pellets and two non-crosslinked pellets per group of experiments: UV-crosslinked-1 pellets for eCLIP library (UV-1); UV-crosslinked-2 pellets for eCLIP library (UV-2); non-crosslinked-1 pellets for eCLIP library as a control (non-UV); non-crosslinked-2 pellets for IgG IP to demonstrate the specificity of antibodies.
- Keep lysing the samples on ice for 15 min (to prevent degradation of protein and RNA).
- Sonicate in a digital sonicator at 10% amplitude for 5 min, at 30 s on/30 s off. Always place the sample on ice and clean the probe with nuclease-free water between each sample.
- Add 4 µL of DNase to each tube, and mix well. Incubate for 10 min at 37 °C, shaking at 1,200 rpm.
- Add 10 µL of diluted RNase I (4 U/µL RNase I in PBS), and mix well. Incubate for 5 min at 37 °C, shaking at 1,200 rpm.
- Clear the lysate by centrifugation at 15,000 x g for 20 min (at 4 °C).
- Carefully collect the supernatant. Leave 50 µL of lysate and discard the pellet with it.
- Save inputs samples as RWB (run for western blot) and RRI (run for RNA isolation). Save 20 µL (2%) of UV-1, UV-2, non-UV and IgG samples as inputs for RWB gel loading. Save 20 µL (2%) of UV-1 or UV-2 samples as inputs for RRI gel loading.
4. Immunoprecipitation
- Add 1 mL of the lysate (from step 3.7) to the beads (prepared in section 2) and rotate the samples at 4 °C for 2 h or overnight.
- Collect the beads with a magnetic stand and discard the supernatant.Wash the beads twice with 900 µL of high salt buffer (50 mM Tris-HCl, pH 7.5; 1 M NaCl; 1 mM EDTA; 1% NP-40; 0.1% SDS; 0.5% sodium deoxycholate), and then wash beads twice with 900 µL of wash buffer (20 mM Tris-HCl, pH 7.5; 10 mM MgCl2; 0.2% Tween-20).
NOTE: For the IgG sample, pause the procedure here and store on ice in wash buffer. - Wash beads once with 500 µL of 1x dephosphorylation buffer (10 mM Tris-HCl, pH 7.5; 5 mM MgCl2; 100 mM KCl; 0.02% Triton X-100).
5. Dephosphorylation of RNA 3' Ends
- Collect the beads with a magnetic stand and discard the supernatant. Remove residual liquid using fine pipette tips. Add 100 µL of dephosphorylation master mix (10 µL of 10x dephosphorylation buffer [100 mM Tris-HCl, pH 8.0; 50 mM MgCl2; 1 M KCl; 0.2% Triton X-100; 1 mg/mL bovine serum albumin (BSA)]; 78 µL of nuclease-free water; 2 µL of RNase Inhibitor; 2 µL of DNase; 8 µL of alkaline phosphatase) to each sample, and incubate for 15 min at 37 °C, shaking at 1,200 rpm.
- Add 300 µL of polynucleotide kinase (PNK) master mix (60 µL of 5x PNK pH 6.5 buffer [350 mM Tris-HCl, pH 6.5; 50 mM MgCl2]; 223 µL of nuclease-free water; 5 µL RNase inhibitor; 2 µL DNase; 7 µL of PNK enzyme; 3 µL of 0.1 M dithiothreitol) to each sample, incubate for 20 min at 37 °C, shaking at 1,200 rpm.
- Collect the beads with a magnetic stand and discard the supernatant. Wash beads once with 500 µL of cold wash buffer. Then wash beads once with 500 µL of cold high salt buffer. Repeat these washes once more in this order.
- Wash beads once with 500 µL of cold wash buffer and then twice with 300 µL of cold 1x ligation buffer (no dithiothreitol, 50 mM Tris-HCl, pH 7.5; 10 mM MgCl2).
6. RNA Adapter Ligation to RNA 3' Ends
- Discard the supernatant, and remove residual liquid with fine pipette tips. Add 25 µL of 3' ligation master mix to each sample. Mix carefully by pipetting. This step is prone to producing bubbles.
NOTE: Ligation master mix contains 3 µL of 10x ligation buffer [500 mM Tris-HCl, pH 7.5; 100 mM MgCl2]; 9 µL of nuclease-free water; 0.4 µL of RNase Inhibitor; 0.3 µL of 0.1 M ATP; 0.8 µL of dimethyl sulfoxide (DMSO); 9 µL of 50% polyethylene glycol (PEG) 8000; 2.5 µL of RNA ligase [30 U/µL]. - Add 2.5 µL of RNA adapter X1A (Table 1) and 2.5 µL of RNA adapter X1B (Table 1) to each sample. Mix carefully by pipetting or flicking, and incubate for 75 min at 25 °C, flicking to mix every 10 min.
- Wash beads once with 500 µL of cold wash buffer (resume IgG sample here).
- Wash beads once with 500 µL of cold high salt buffer and then with 500 µL cold wash buffer. Repeat these washes once more.
- Magnetically separate beads, and remove residual liquid with fine pipette tips.
- Resuspend the beads in 100 µL of cold wash buffer (pause the IgG sample here and store on ice in wash buffer). Move 20 µL to new tubes as RWB samples. Magnetically separate the remaining 80 µL as RRI samples. Remove the RRI samples' supernatant and resuspend the beads in 20 µL of wash buffer.
- Add 37.5 µL of 4x lithium dodecyl sulfate (LDS) sample buffer and 15 µL of 10x sample reducing agent to the IgG sample. Add 7.5 µL of 4x LDS sample buffer and 3 µL of 10x sample reducing agent to the (remaining) samples. (Do not mix by pipetting). Incubate for 10 min at 70 °C, shaking at 1,200 rpm. Cool on ice for 1 min, and centrifuge at 1,000 x g for 1 min at 4 °C.
7. SDS-PAGE and Membrane Transfer
- Load gels.
- For RRI gel (4-12% Bis-Tris protein gel, 10-well, 1.5 mm) place tubes on a magnet and separate protein eluate from the beads. Load 30 µL of sample per well. Samples are spaced by pre-stained protein size marker.
- For RWB gel (4-12% Bis-Tris protein gel, 10-well, 1.5 mm) place tubes on a magnet and separate protein eluate from the beads. Load 15 µL of sample per well. Save the remaining samples at -20 °C as backups.
- Add 500 µL of antioxidant to 500 mL of 1x SDS running buffer.Run at 200 V in 1x SDS running buffer for 50 min or until dye front is at the bottom.
NOTE: Antioxidant contains N, N-Dimethylformamide, sodium bisulfite, which migrates with reduced proteins to prevent reoxidation of sensitive amino acids such as methionine and tryptophan. - Transfer protein-RNA complexes from the gel to a nitrocellulose membrane at 10 V for 70 min in 1xtransfer buffer with 10% methanol (vol/vol). Rinse the RRI membrane in cold PBS, wrap it in plastic wrap, and store at -80 °C.
- Block the RWB membrane in 5% milk in TBST at room temperature for 1 h. Rinse the membrane in TBST. Incubate with primary antibody in TBST at 4 °C overnight. Wash twice with TBST for 5 min. Incubate with secondary antibody (1:5,000 HRP goat anti-rabbit IgG) in TBST at room temperature for 1 h. Wash three times with TBST for 5 min, 10 min and 15 min, respectively.
- Mix equal volumes of electrochemiluminescence (ECL) Buffer A and Buffer B, add to membrane and incubate for 1 min. Cover the membrane with plastic wrap, and expose it to an X-ray film at room temperature for 2-3 min. Then develop the film.
NOTE: The film with overexposure will clearly show the shape of the membrane's edges, by which the film can be aligned back to the RWB membrane. Then, align the RWB and RRI membranes based on the positions of markers therein. Layered in order from bottom to top are the film, the RWB membrane and the RRI membrane in sequence.
8. RNA Isolation
- Cut the region from the protein band to 75 kDa (about 220 nt of RNA) above it, using the RWB membrane and film as guides.
NOTE: As a protein-protected RNA molecule may have a maximum length of 225 bases, with about 340 Da per base, it is reasonable to cut the region about 75 kDa above the RBP band to retrieve all protein-RNA complexes. - Cut the excised piece of membrane into several small slices, and place them into a fresh 1.5 mL centrifuge tube. Add 200 µL of proteinase K (PK) buffer (100 mM Tris-HCl pH 7.5; 50 mM NaCl; 10 mM EDTA) with 40 µL of PK to the membrane pieces. Mix and incubate for 20 min at 37 °C, shaking at 1,200 rpm.
- Add 200 µL of PK urea buffer (100 mM Tris-HCl pH 7.4; 50 mM NaCl; 10 mM EDTA; 7 M Urea) to each sample. Incubate for 20 min at 37 °C, shaking at 1,200 rpm.
- Add 400 µL of acid phenol/chloroform/isoamyl alcohol (25:24:1), mix well and incubate for 5 min at 37 °C, shaking at 1,200 rpm.
- Place 2 mL phase lock gel (PLG) heavy tube in the centrifuge and spin at 15,000 x g for 25 s.
- Transfer all contents except membrane slices to PLG heavy tube. Incubate for 5 min at 37 °C, shaking at 1,200 rpm.
- Spin at room temperature and 15,000 x g for 15 min. Transfer the aqueous layer into a new 15 mL conical tube.
- Use RNA purification and concentration columns to extract RNA.
- Add 2 volumes (800 µL) of RNA binding buffer to each sample (from step 8.7) and mix. Add an equal volume (1200 µL) of 100% ethanol and mix.
- Transfer 750 µL of sample (from step 8.8.1) to the RNA purification and concentration columns in a collection tube and centrifuge at 15,000 x g for 30 s. Discard flow-through.
- Repeat step 8.8.2 until all samples are passed through the column. Add 400 µL of RNA prep buffer to the column and centrifuge at 15,000 x g for 30 s. Discard flow-through, apply 700 µL of RNA wash buffer and centrifuge the column at 15,000 x g for 30 s. Discard flow-through.
- Add 400 µL of RNA wash buffer to the column and centrifuge at 15,000 x g for 2 min. Transfer the column carefully into a new 1.5 mL tube. Add 10 µL of nuclease-free water to the column matrix, let sit for 2 min, and centrifuge at 15,000 x g for 30 s. Store eCLIP samples at -80 °C until RT (step 11.1).
9. Dephosphorylation of Input RNA 3' Ends
- Add 15 µL of dephosphorylation master mix (2.5 µL of 10x dephosphorylation buffer [100 mM Tris-HCl, pH 8.0; 50 mM MgCl2; 1 M KCl; 0.2% Triton X-100; 1 mg/mL BSA]; 9.5 µL of nuclease-free water; 0.5 µL of RNase inhibitor; 2.5 µL of alkaline phosphatase) to 10 µL of input samples (from step 8.8.4). Incubate for 15 min at 37 °C, shaking at 1,200 rpm.
- Add 75 µL of PNK master mix (20 µL of 5x PNK PH 6.5 buffer [350 mM Tris-HCl, pH 6.5; 50 mM MgCl2]; 44 µL of nuclease-free water; 1 µL of RNase inhibitor; 2 µL of DNase; 7 µL of PNK enzyme; 1 µL of 0.1 M dithiothreitol) to samples. Incubate for 20 min at 37 °C, shaking at 1,200 rpm.
- Cleanup of input RNA
- Resuspend thenucleic acids extraction magnetic beadsin the vial (vortex for more than 30 s). Add 20 µL of nucleic acids extraction magnetic beads for each sample to new tubes. Collect the beads with a magnetic stand and discard the supernatant.
- Wash beads once with 1 mL of RNA purification lysis buffer (RLT buffer). Place the tube on a magnet for 30 s and discard the supernatant.
NOTE: Subsequent separation of nucleic acids extraction magnetic beads followed this step. - Resuspend beads with 300 µL of RLT buffer and add to the sample. Add 10 µL of 5 M NaCl and 615 µL of 100% ethyl alcohol (EtOH) and mix by pipetting. Rotate at room temperature for 15 min. Magnetically separate the beads and remove supernatant.
- Resuspend beads in 1 mL of 75% EtOH and transfer to a new tube. After 30 s, collect the beads with a magnetic stand and discard the supernatant. Wash twice with 75% EtOH, magnetically separate the beads, and discard residual liquid with fine pipette tips. Air dry for 5 min (avoid excessive drying).
- Resuspend the beads with 10 µL of nuclease-free water and incubate it for 5 min. Magnetically separate beads, and move 5 µL of supernatant to a new tube (for 3' adapter ligation below). The remaining RNA can be stored at -80 °C as backups.
10. RNA Adapter Ligation to Input RNA 3' Ends
- Add 1.5 µL of DMSO and 0.5 µL of RiL19 adapter (Table 1) to 5 µL of input RNA (from step 9.3.5), incubate for 2 min at 65 °C, and place on ice for more than 1 min. Add 13.5 µL of 3' ligation master mix to each sample, mix by pipetting, and incubate for 75 min at 25 °C, with flicking to mix every 15 min.
NOTE: Ligation master mix contains 2 µL of 10x ligation buffer [500 mM Tris-HCl, pH 7.5; 100 mM MgCl2; 10 mM dithiothreitol]; 1.5 µL of nuclease-free water; 0.2 µL of RNase inhibitor; 0.2 µL of 0.1 M ATP; 0.3 µL of DMSO; 8 µL of 50% PEG8000; 1.3 µL of RNA ligase [30 U/µL]. - Cleanup of ligated input RNA
- Magnetically separate 20 µL of nucleic acids extraction magnetic beads for each sample, and remove the supernatant. Wash beads once with 1 mL of RLT buffer.
- Resuspend beads in 61.6 µL of RLT buffer and transfer suspension to each sample. Add 61.6 µL of 100% EtOH. Use pipette to mix for 15 min every 5 min. Magnetically separate beads, and remove supernatant.
- Repeat step 9.3.4.
- Resuspend beads with 10 µL of nuclease-free water, and let it sit for 5 min. Magnetically separate the beads and transfer 10 µL of the supernatant to a new tube.
NOTE: This is a possible stopping point (input samples can be stored at -80 °C until next day).
11. Reverse Transcription, DNA Adapter Ligation to cDNA 3' Ends
- Add 0.5 µL of RT primer (Table 1) to 10 µL of input RNA (from step 10.2.4) and 10 µL of CLIP RNA (from step 8.8.4) respectively, incubate for 2 min in pre-heated PCR block at 65 °C, place on ice for more than 1 min.
- Add 10 µL of RT master mix (2 µL of RT buffer [500 mM Tris-HCl, pH 8.3; 750 mM KCl; 30 mM MgCl2]; 4 µL of nuclease-free water; 0.3 µL of RNase Inhibitor; 2 µL of 0.1 M dithiothreitol; 0.2 µL of 0.1 M dATP; 0.2 µL of 0.1 M dCTP; 0.2 µL of 0.1 M dGTP; 0.2 µL of 0.1 M dTTP; 0.9 µL of reverse transcriptase) to each sample, mix, incubate at 55 °C for 45 min on a pre-heated PCR block.
- Mix 20 µL of RT reaction product with 3.5 µL of PCR product cleanup reagent. Incubate for 15 min at 37 °C.
- Add 1 µL of 0.5 M EDTA, mix by pipetting. Add 3 µL of 1 M NaOH, mix by pipetting, and incubate for 12 min at 70 °C on a PCR block to hydrolyze the template RNA.
- Add 3 µL of 1 M HCl, pipette-mix to neutralize the buffer.
- Cleanup of cDNA
- Magnetically separate 10 µL of nucleic acid extraction magnetic beads for each sample, remove the supernatant. Wash once with 500 µL of RLT buffer.
- Resuspend beads in 93 µL of RLT buffer and transfer suspension to the sample. Add 111.6 µL of 100% EtOH, incubate it for 5 min, and pipette mix every 2 min. Collect the beads with a magnetic stand and discard the supernatant.Resuspend beads with 1 mL of 80% EtOH and move to a new tube.
- After 30 s, collect the beads with a magnetic stand and discard the supernatant. Wash twice with 80% EtOH. Magnetically separate and discard residual liquid with fine tip. Air dry for 5 min (avoid excessive drying). Resuspend beads in 5 µL of 5 mM Tris-HCl and incubate it for 5 min.
- Add 0.8 µL of DNA adapter (Table 1) and 1 µL of DMSO to the beads, incubate for 2 min at 75 °C. Place on ice for more than 1 min.
- Prepare 12.8 µL of ligation master mix (2 µL of 10x ligation buffer [500 mM Tris-HCl; 100 mM MgCl2; 10 mM dithiothreitol]; 1.1 µL of nuclease-free water; 0.2 µL of 0.1 M ATP; 9 µL of 50% PEG8000; 0.5 µL of RNA ligase [30 U/µL]), flick to mix, spin briefly in a centrifuge, and add it to each sample, stir sample with pipette tip slowly.
- Add another 1 µL of RNA ligase [30 U/µL] to each sample and flick to mix.Incubate for 30 s at 25 °C, shaking at 1,200 rpm. Incubate at 25 °C overnight. Flick to mix lightly 5 to 6 times, once per hour.
- Cleanup of ligated cDNA
- Magnetically separate 5 µL of nucleic acids extraction magnetic beads for each sample, and remove supernatant.
- Wash once with 500 µL RLT buffer.
- Resuspend beads in 60 µL of RLT buffer to beads and transfer suspension to each sample. Add 60 µL of 100% EtOH, incubate it for 5 min and pipette mix every 2 min. Magnetically separate, discard supernatant.
- Repeat step 9.3.4.
- Resuspend beads with 27 µL of 10 mM Tris-HCl, incubate it for 5 min. Magnetically separate, and move 25 µL of sample to a new tube. Dilute the 1 µL of ligated cDNA with 9 µL of nuclease-free water in a new tube. Store the remaining samples at -20 °C until step 13.1.
12. Quantification of cDNA by Real-time Quantitative PCR (qPCR)
- Add 9 µL of qPCR master mix (5 µL of 2x master mix; 3.6 µL of nuclease-free water; 0.4 µL of primer mix [10 µM PCR-F-D50X and 10 µM PCR-R-D70X]) to a 96-well qPCR plate. Add 1 µL of 1:10 diluted (in H2O) cDNA (from step 11.10.5), seal and mix.
- Run the qPCR program in a thermocycler: 2 min at 50 °C; 2 min at 95 °C; 3 s at 95 °C followed by 30 s at 68 °C for 40 cycles; 15 s at 95 °C followed by 60 s at 68 °C followed by 15 s at 95 °C for 1 cycle. Note Ct (cycle threshold) value.
13. PCR Amplification of cDNA
- Dispense 35 µL of PCR master mix (25 µL of 2x PCR master mix; 5 µL of nuclease-free water; 5 µL of primer mix [20 µM PCR-F-D50X and 20 µM PCR-R-D70X]) into 8-well strips. For CLIP group, add 12.5 µL of CLIP sample + 2.5 µL of H2O; for inputs group, add 10 µL of inputs + 5 µL of H2O. Mix well and spin briefly in centrifuge.
- Run the PCR program: 30 s at 98 °C; 15 s at 98 °C followed by 30 s at 68 °C followed by 40 s at 72 °C for 6 cycles; 15 s at 98 °C followed by 60 s at 72 °C for N cycles = (qPCR Ct values-3)-6; 60 s at 72 °C; 4 °C hold.
NOTE: It is better to perform 1 to 2 extra PCR cycles for the first couple of CLIPs.
14. Gel Purification
- Load samples on a 3% high resolution agarose gel. Leaving 1 empty well between samples, and use a ladder on both sides of the gel. Run at 100 V for 75 min in 1x Tris-Borate-EDTA (TBE) buffer.
- Under blue light illumination, cut gel slices 175-350 bp using fresh razor blades for each sample. Place them into 15 mL of conical tubes.
- Weigh the gel slice, and elute the gel using a gel extraction kit.
- Add 6x volumes of gel dissolving buffer to melt the gel (100 mg gel = 600 µL of gel dissolving buffer). Dissolve the gel at room temperature. (Shake to mix every 15 min until the gel slice has completely dissolved). Add 1 gel volume of 100% isopropanol and mix well.
- Transfer 750 µL of sample (from step 14.2.2) to the column in a collection tube and centrifuge at 17,900 x g for 1 min. Discard flow-through.
- Repeat step 14.2.3 until all samples have passed through the column, wash once with 500 µL of gel dissolving buffer.
- Add 750 µL of wash buffer (from gel extraction kit) to the column and centrifuge at 17,900 x g for 1 min. Discard flow-through, spin at 17,900 x g for 2 min. Place the column to a new 1.5 mL tube.
- Remove all remaining wash buffer from the plastic purple rim of the column. Air dry for 2 min. Add 12.5 µL of nuclease-free water to the center of the membrane. Let the column stand for 2 min at room temperature, and spin at 17,900 x g for 1 min (For an improved yield, repeat the elution step).
15. TOPO Clone of PCR Product
- Prepare the TOPO cloning reaction mix (1 µL of PCR product from step 14.2.6; 1 µL of cloning mix; 3 µL of sterile water). Mix gently and incubate for 5 min at 20-37 °C. Cool on ice.
- Thaw chemically competent E. coli bacteria on ice. Add 5 µL of the TOPO cloning product to 100 µL of competent bacteria29. Gently mix well. Incubate on ice for 30 min.
- Heat-shock for 60 s at 42 °C. Then cool on ice for 2 min. Add 900 µL of lysogeny broth (LB) medium, incubate at 37 °C for 1 h, shaking at 225 rpm. Spin at 1,000 x g for 1 min, and then remove 900 µL of supernatant. Resuspend the rest of solution.
- Plate the transformed bacteria onto 1.5% LB agar plates containing ampicillin (100 µg/mL), and incubate overnight at 37 °C.
- Prepare at least 20 1.5 mL centrifuge tubes for each construct. Fill one set with 500 µL of LB medium containing 100 µg/mL ampicillin. Use a sterile pipette tip to scratch off one bacterial colony randomly and mix (by pipetting) with 500 µL of LB medium. Incubate at 37 °C for 5 h, shaking at 225 rpm.
- Sequence the insert fragment using the M13 reverse primer: CAGGAAACAGCTATGAC by Sanger Sequencing30.
Representative Results
The eCLIP procedure and results are illustrated in Figure 1, Figure 2, Figure 3, Figure 4. Mice were euthanized with carbon dioxide and a small incision was made in the lower abdomen using surgical scissors (Figure 2A,B). Mouse testes were removed, detunicated and then UV-crosslinked after grinding (Figure 2C–I). Representative eCLIP results of using two known RNA-binding helicases in testis tissues are depicted in Figure 3 and 4. We performed MOV10 eCLIP in testes from adult wild-type mice, with common concentration of 40 U/mL of RNase I treating the crosslinked lysate. The top panel of Figure 3A shows that the target protein sized about 114 kDa was successfully enriched. Western blot of the immunoprecipitated MOV10L1 proteins was performed with two concentrations (5 or 40 U/mL) of RNase I during the eCLIP process (Figure 3A). Figure 3B shows qPCR using 1:10 diluted cDNA (already ligated with DNA adapter) from MOV10 and MOV10L1 UV-crosslinked, non-crosslinked, and the paired size matched input (SMInput) sample. Non-crosslinked samples show decreased RNA recovery. We observed that, the Ct values of the non-crosslinked group was generally 5 times more than UV-crosslinked group. Figure 4A displays PCR amplification and size selection via agarose gel electrophoresis (cut 175-350 bp). Primer-dimer product appears at about 140 bp. Figure 4B shows the UCSC genome browser view of two representative subclone sequences. MOV10-bound eCLIP tags are found to be located within the 3' UTR of gene Fto; The approximate rate of 3' UTR targets accounts for 75% (Figure 4C), consistent with the majority of MOV10 targets in HEK293 cells10 and in testes (data not shown). In contrast, MOV10L1-bound eCLIP tags are found to be located within a piRNA cluster indicating MOV10L1 targets piRNA precursors. The approximate rate of piRNA precursor targets accounts for 42% (Figure 4E), which reflects a trend from our previous conventional CLIP experiment20. MOV10L1 eCLIP with 40 U/mL RNase I digestion yields relatively more sequences with less than 20 bp (Figure 4D).
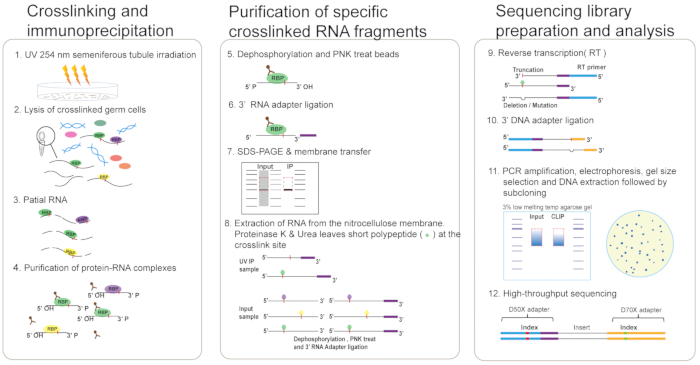
Figure 1: Schematic representation of eCLIP. UV-crosslinked mouse seminiferous tubules (step 1) are lysed in eCLIP lysis buffer and sonicated (step 2). Lysate is treated with RNase I to fragment RNA, after which protein-RNA complexes are immunoprecipitated using the anti-RBP antibody (step 3-4). Dephosphorylation of RNA fragments and ligation of 3′ RNA adapter are performed (step 5-6). Protein-RNA complexes are run on an SDS-PAGE gel and transferred to nitrocellulose membranes (step 7). RNA is recovered from the membrane by digesting the protein with proteinase K and Urea which leaves a short polypeptide remaining at the crosslink site. Dephosphorylation of RNA fragments of input samples and ligation of 3′ RNA adapter is performed (step 8). Perform RT of RNA and ligation of 3′ DNA adapter (step 9-10). Perform PCR amplification of cDNA library, gel extraction, and blunt-end PCR cloning for preliminary library quality control (step 11). Finally, perform high-throughput sequencing (step 12). Please click here to view a larger version of this figure.
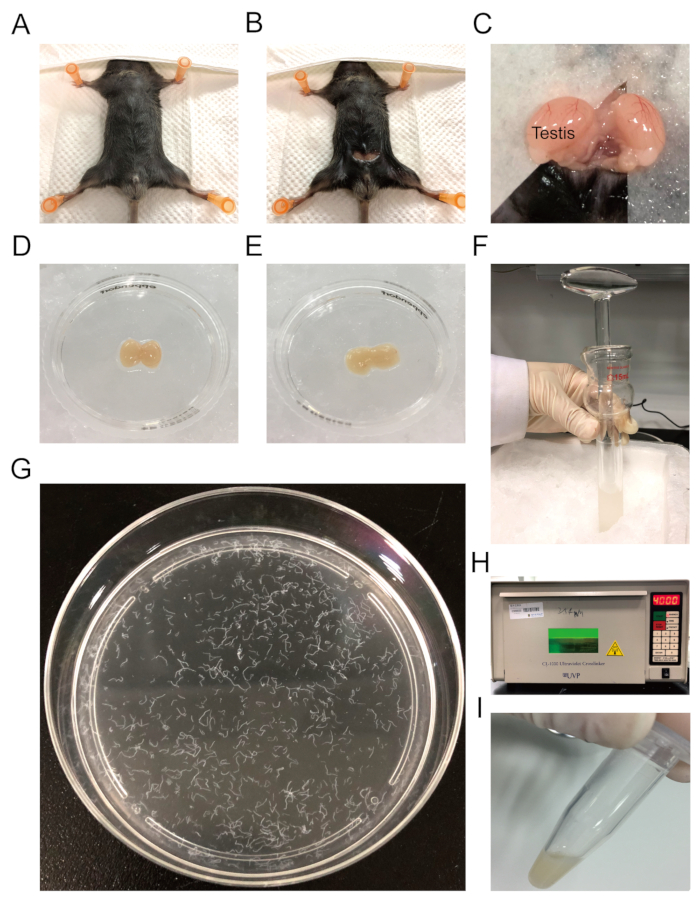
Figure 2: Testis tissue harvesting and UV crosslinking. (A) The exposure of the mouse abdomen. (B) A 0.5 cm incision in the abdominal wall exposing the peritoneum. (C) A pair of testes are taken out by pulling out the fat pads. (D) Testicular tissue is removed and placed in a small dish containing ice-cold PBS. (E) Gently remove the tunica albuginea. (F) Press the loose pestle to triturate the tissue in a tissue grinder dounce. (G) Distributed seminiferous tubules in a 10 cm plate. (H) UV crosslinking with 400 mJ/cm2 energy. (I) The crosslinked samples are collected in 1.5 mL centrifuge tubes. Please click here to view a larger version of this figure.
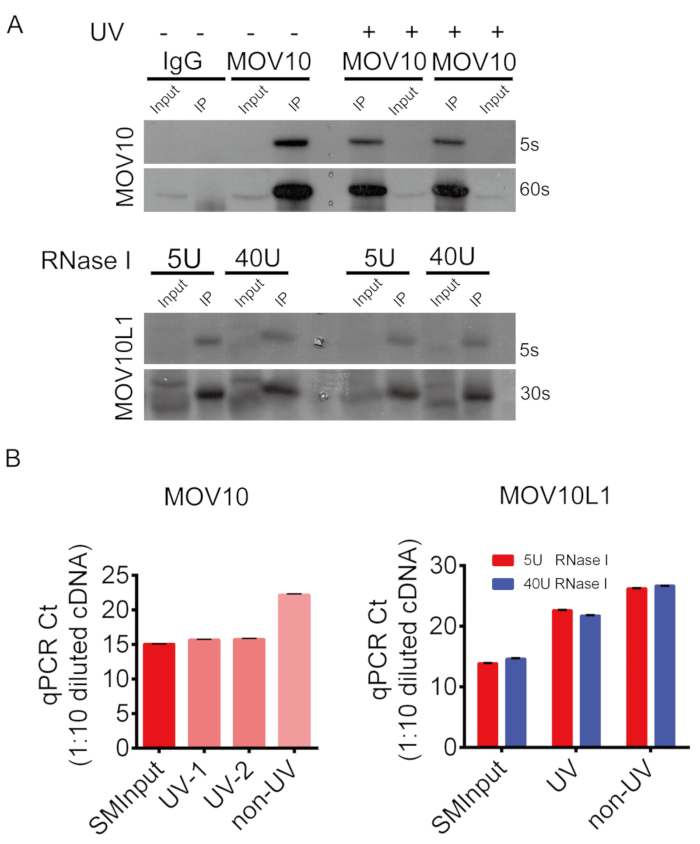
Figure 3: Representative results of MOV10 and MOV10L1 eCLIP. (A) Western blot validation of MOV10 and MOV10L1 immunoprecipitates. (B) qPCR on 1:10 diluted eCLIP libraries of MOV10 and MOV10L1, with replicates for UV, non-UV and the paired SMInput samples. Please click here to view a larger version of this figure.
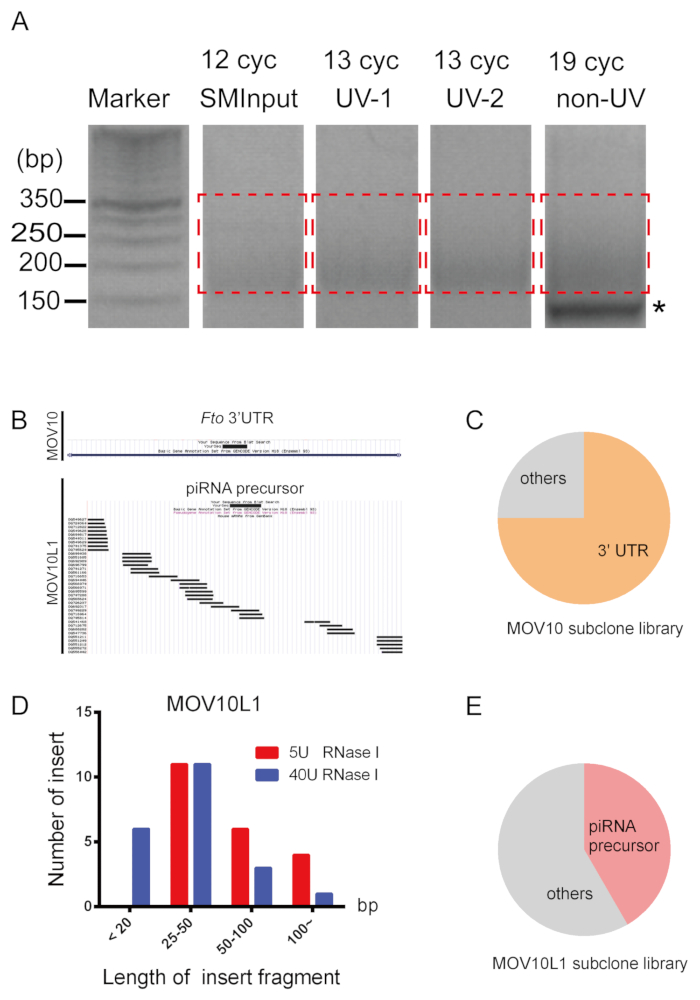
Figure 4: eCLIP library preparation and quality assessment. (A) The gel images of PCR amplification are shown. Asterisk indicates primer dimer. Red dotted line indicates regions excised for PCR product of cDNA, somewhere between 175 and 350 bp. (B) UCSC genome browser view of two representative subclone sequences31. (C) Small-scale subclone sequencing analysis of MOV10. (D) MOV10L1-bound tags display distinct patterns of length distribution when processed by two different RNase concentrations. (E) Small-scale subclone sequencing analysis of MOV10L1. Please click here to view a larger version of this figure.
| Sequence Name | Sequence Information | Description | |||||
| RNA adapters | |||||||
| RNA X1A | /5Phos/AUAUAGGNNNNNAGAUCGGAAGAGCGUCGUGUAG/3SpC3/ | stock at 200 µM; working at 20 µM | |||||
| RNA X1B | /5Phos/AAUAGCANNNNNAGAUCGGAAGAGCGUCGUGUAG/3SpC3/ | stock at 200 µM; working at 20 µM | |||||
| RiL19 | /5phos/AGAUCGGAAGAGCGUCGUG/3SpC3/ | stock at 200 µM; working at 40 µM | |||||
| DNA adapter | |||||||
| Rand103tr3 | /5Phos/NNNNNNNNNNAGATCGGAAGAGCACACGTCTG/3SpC3/ | stock at 200 µM; working at 80 µM | |||||
| RT primer | |||||||
| AR17 | ACACGACGCTCTTCCGA | stock at 200 µM; working at 20 µM | |||||
| PCR primers | |||||||
PCR-F-D 501 |
AATGATACGGCGACCACCGAGATCTACACTATAGCCTACACTCTTTCCCTACACGACGCTCTTCCGATCT | stock at 100 µM; working at 20 µM | |||||
| PCR-R-D 701 | CAAGCAGAAGACGGCATACGAGATCGAGTAATGTGACTGGAGTTCAGACGTGTGCTCTTCCGATC | stock at 100 µM; working at 20 µM | |||||
| (See Illumina customer service letter for D502-508, D702-712) | |||||||
Table 1: Adapter and primer sequences. The adapter contains an in-line random-mer (either N5 or N10) to determine whether two identical sequenced reads indicate two unique RNA fragments or PCR duplicates of the same RNA fragment. "5 Phos" stands for 5' Phosphorylation, which is needed if an oligo is used as a substrate for DNA/RNA ligase. "3SpC3" stands for 3' C3 Spacer, which can prevent ligation between adapters.
Discussion
With increasing understanding of the universal role of RBPs under both biological and pathological contexts, the CLIP methods have been widely utilized to reveal the molecular function of RBPs20,32,33,34,35. The protocol described here represents an adapted application of the eCLIP method to mouse testis.
One challenge in performing eCLIP in testis is maintaining viability and integrity of fresh testicular cells, which is also important for effective crosslinking. Shearing the testis with mild mechanical force using the loose pestle can prevent cell lysis32,36. Proper digestion of RNA is also critical for successful eCLIP assays. RNA fragments could be more convergent after digestion, but length less than 20 bp can be removed via pre-processing of the library reads. In order to adopt an ideal RNase dosage for an RBP candidate, we suggest a preliminary test based on the results of the subclone sequencing of eCLIP libraries that can be prepared by RNase treatment with concentrations ranging from 0 U to 40 U (per milliliter of lysate). The small-scale subclone sequencing analysis is a recommended step for a reliable examination of library quality in our eCLIP method. First, the percentage of inserts shorter than 20 bp should not be too high, or, the subsequent pre-processing of eCLIP library will cause a costly loss of reads. Secondly, the efficiency of correct ligation of both adapters should be checked. Substandard samples can be eliminated without deep sequencing to ensure successful deep sequencing, the results of which generally take much longer to analyze.
Although the feasibility of eCLIP in mouse testis is still limited by the specificity of the antibody for the step of immunoprecipitation, eCLIP is advantageous over conventional CLIP methods in several aspects. First, it is a non-radioactive method. By eCLIP, RNA targets of RBPs are directly captured in vivo without having to resort to labor-intensive techniques using radioactive materials. Secondly, the method is less time intensive. The whole procedure takes only 4 days through eCLIP library preparation. Third, sequence diversity. Compared with the conventional unified amplification cycle of 25-35 cycles, eCLIP refers to Ct values of qPCR to set the number of PCR cycles specifically. Lastly, it provides stronger signal-to-noise ratio. The size-matched input serves as an appropriate background for authentic targets.
In summary, our eCLIP results consolidate the conclusions that MOV10 and MOV10L1 have a binding preference to mRNA 3' UTR and piRNA precursors, respectively. The protocol we described herein represents the first employment of the eCLIP method in reproduction, an area in which RNA-RBP interaction knowledge is rather insufficient, although genetic studies have provided ample information about the biological roles of RBPs. Visualization of this eCLIP protocol may help guide its widespread applications in broader areas.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank Eric L Van Nostrand and Gene W Yeo for helpful guidance with the original protocol. K.Z. was supported by National Key R&D Program of China (2016YFA0500902, 2018YFC1003500), and National Natural Science Foundation of China (31771653). L.Y. was supported by National Natural Science Foundation of China (81471502, 31871503) and Innovative and Entrepreneurial Program of Jiangsu Province.
Materials
| Antibodies | |||
| Anti-mouse MOV10 antibody | Proteintech, China | 10370-1-AP | |
| Anti-mouse MOV10L1 antibody | Zheng et al.20109 | polyclonal antisera UP2175 | provided by P. Jeremy Wang lab(University of Pennsylvania) |
| HRP Goat Anti-Rabbit IgG | ABclonal | AS014 | |
| Rabbit IgG | Beyotime, China | A7016 | |
| Equipment | |||
| Centrifuge | Eppendorf, Hamburg, Germany | 5242R | |
| Digital sonifier | BRANSON,USA | BBV12081048A | 450 Watts; 50/60 HZ |
| DynaMag-2 Magnet | Invitrogen,USA | 12321D | |
| Mini Blot Module | Invitrogen,USA | B1000 | |
| Mini Gel Tank | Invitrogen,USA | A25977 | |
| Shaking incubator | Eppendorf, Hamburg, Germany | Thermomixer comfort | |
| Tissue Grinder, Dounce | PYREX, USA | 1234F35 | only the "loose" pestle is used in this protocol |
| TProfessional standard 96 Gradient | Biometra, Germany | serial no.: 2604323 | |
| Tube Revolver | Crystal, USA | serial no.: 3406051 | |
| UV-light cross-linker | UVP, USA | CL-1000 | |
| Materials | |||
| TC-treated Culture Dish | Corning, USA | 430167 | 100 mm |
| Tubes | Corning, USA | 430791 | 15 mL |
| Microtubes tubes | AXYGEN , USA | MCT-150-C | 1.5 mL |
| Reagents | |||
| Acid phenol/chloroform/isoamyl alcohol | Solarbio, China | P1011 | 25:24:01 |
| AffinityScript Enzyme | Agilent, USA | 600107 | |
| Antioxidant | Invitrogen,USA | NP0005 | |
| DH5α competent bacteria | Thermo Scientific, USA | 18265017 | these economical cells yield >1 x 106 transformants/µg control DNA per 50 µL reaction. |
| DMSO | Sigma-Aldrich, USA | D8418 | |
| DNA Ladder | Invitrogen, USA | 10416014 | |
| dNTP | Sigma-Aldrich, USA | DNTP100-1KT | |
| Dynabeads Protein A | Invitrogen, USA | 10002D | |
| ECL reagent | Vazyme, China | E411-04 | |
| EDTA | Invitrogen, USA | AM9260G | |
| EDTA free protease inhibitor cocktail | Roche, USA | 04693132001 | add fresh |
| Exo-SAP-IT | Affymetrix, USA | 78201 | PCR Product Cleanup Reagent |
| FastAP enzyme | Thermo Scientific, USA | EF0652 | |
| LDS Sample Buffer | Thermo Scientific, USA | NP0007 | |
| MetaPhor Agarose | lonza, Switzerland | 50180 | |
| MgCl2 | Invitrogen, USA | AM9530G | |
| MiniElute gel Extraction | QIAGEN, Germany | 28604 | column store at 4 ℃; buffer QG=gel dissolving buffer; buffer PE= wash buffer(for step 14) |
| MyONE Silane beads | Thermo Scientific, USA | 37002D | nucleic acids extraction magnetic beads |
| NaCl | Invitrogen,USA | AM9759 |
|
| NP-40 | Amresco, USA | M158-500ML | |
| NuPAGE Bis-Tris Protein Gels | Invitrogen, USA | NP0336BOX | 4%–12%,1.5 mm, 15-well |
| NuPAGE MOPS SDS Buffer Kit | Invitrogen, USA | NP0050 | |
| PBS | Gibco, USA | 10010023 | |
| Phase-Locked Gel (PLG) heavy tube | TIANGEN, China | WM5-2302831 | |
| PowerUp SYBR Green Master Mix | Applied Biosystems, USA | A25742 | |
| proteinase K | NEB, New England | P8107S | |
| Q5 PCR master mix | NEB, New England | M0492L | |
| RLT buffer | QIAGEN, Germany | 79216 | RNA purification lysis buffer |
| RNA Clean & Concentrator-5 columns | ZYMO RESEARCH, USA | R1016 | RNA purification and concentration columns |
| RNase I | Invitrogen, USA | AM2295 | |
| RNase Inhibitor | Promega, USA | N251B | |
| RQ1 DNase | Promega,USA | M610A | |
| Sample Reducing Agent | Invitrogen,USA | NP0009 | |
| SDS Solution | Invitrogen, USA | 15553027 | 10% |
| Sodium deoxycholate | Sigma-Aldrich, USA | 30970 | protect from light |
| T4 PNK enzyme | NEB, New England | M0201L | |
| T4 RNA ligase 1 high conc | NEB, New England | M0437M | |
| TA/Blunt-Zero Cloning Mix | Vazyme, China | C601-01 | |
| TBE | Invitrogen,USA | AM9863 | |
| Tris-HCI Buffer | Invitrogen, USA | 15567027 | |
| Triton X-100 | Sangon Biotech, China | A600198 | |
| Tween-20 | Sangon Biotech, China | A600560 | |
| Urea | Sigma-Aldrich, USA | U5378 | |
| X-ray Films | Caresteam, Canada | 6535876 |
References
- Turner, J. M., Mahadevaiah, S. K., Ellis, P. J., Mitchell, M. J., Burgoyne, P. S. Pachytene asynapsis drives meiotic sex chromosome inactivation and leads to substantial postmeiotic repression in spermatids. Developmental Cell. 10 (4), 521-529 (2006).
- Turner, J. M. A. Meiotic sex chromosome inactivation. Development. 134 (10), 1823-1831 (2007).
- Kimmins, S., Sassone-Corsi, P. Chromatin remodelling and epigenetic features of germ cells. Nature. 434 (7033), 583-589 (2005).
- Ule, J., et al. CLIP identifies Nova-regulated RNA networks in the brain. Science. 302 (5648), 1212-1215 (2003).
- Ule, J., Jensen, K., Mele, A., Darnell, R. B. CLIP: a method for identifying protein-RNA interaction sites in living cells. Methods. 37 (4), 376-386 (2005).
- Trifillis, P., Day, N., Kiledjian, M. Finding the right RNA: identification of cellular mRNA substrates for RNA-binding proteins. RNA. 5 (8), 1071-1082 (1999).
- Tenenbaum, S. A., Carson, C. C., Lager, P. J., Keene, J. D. Identifying mRNA subsets in messenger ribonucleoprotein complexes by using cDNA arrays. Proceedings of National Acadamy of Sciences U S A. 97 (26), 14085-14090 (2000).
- Chi, S. W., Zang, J. B., Mele, A., Darnell, R. B. Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps. Nature. 460 (7254), 479-486 (2009).
- Zheng, K., et al. Mouse MOV10L1 associates with Piwi proteins and is an essential component of the Piwi-interacting RNA (piRNA) pathway. Proceedings of National Acadamy of Sciences U S A. 107 (26), 11841-11846 (2010).
- Gregersen, L. H., et al. MOV10 Is a 5′ to 3′ RNA helicase contributing to UPF1 mRNA target degradation by translocation along 3′ UTRs. Molecular Cell. 54 (4), 573-585 (2014).
- Banerjee, S., Neveu, P., Kosik, K. S. A coordinated local translational control point at the synapse involving relief from silencing and MOV10 degradation. Neuron. 64 (6), 871-884 (2009).
- Choi, J., Hwang, S. Y., Ahn, K. Interplay between RNASEH2 and MOV10 controls LINE-1 retrotransposition. Nucleic Acids Research. 46 (4), 1912-1926 (2018).
- Goodier, J. L., Cheung, L. E., Kazazian, H. H. MOV10 RNA helicase is a potent inhibitor of retrotransposition in cells. PLoS Genetics. 8 (10), e1002941 (2012).
- Haussecker, D., et al. Capped small RNAs and MOV10 in human hepatitis delta virus replication. Nature Structural & Molecular Biology. 15 (7), 714-721 (2008).
- Kenny, P. J., et al. MOV10 and FMRP regulate AGO2 association with microRNA recognition elements. Cell Reports. 9 (5), 1729-1741 (2014).
- Messaoudi-Aubert, S. E., et al. Role for the MOV10 RNA helicase in Polycomb-mediated repression of the INK4a tumor suppressor. Nature Structural & Molecular Biology. 17 (7), 862-868 (2010).
- Sievers, C., Schlumpf, T., Sawarkar, R., Comoglio, F., Paro, R. Mixture models and wavelet transforms reveal high confidence RNA-protein interaction sites in MOV10 PAR-CLIP data. Nucleic Acids Research. 40 (20), e160 (2012).
- Skariah, G., et al. Mov10 suppresses retroelements and regulates neuronal development and function in the developing brain. BMC Biology. 15 (1), 54 (2017).
- Zheng, K., Wang, P. J. Blockade of pachytene piRNA biogenesis reveals a novel requirement for maintaining post-meiotic germline genome integrity. PLoS Genetics. 8 (11), e1003038 (2012).
- Vourekas, A., et al. The RNA helicase MOV10L1 binds piRNA precursors to initiate piRNA processing. Genes & Development. 29 (6), 617-629 (2015).
- Hocq, R., Paternina, J., Alasseur, Q., Genovesio, A., Le Hir, H. Monitored eCLIP: high accuracy mapping of RNA-protein interactions. Nucleic Acids Research. 46 (21), 11553-11565 (2018).
- Huppertz, I., et al. iCLIP: protein-RNA interactions at nucleotide resolution. Methods. 65 (3), 274-287 (2014).
- Hafner, M., et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell. 141 (1), 129-141 (2010).
- Hafner, M., et al. PAR-CliP–a method to identify transcriptome-wide the binding sites of RNA binding proteins. Journal of Visualized Experiments. (41), (2010).
- Konig, J., et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nature Structural & Molecular Biology. 17 (7), 909-915 (2010).
- Konig, J., et al. iCLIP–transcriptome-wide mapping of protein-RNA interactions with individual nucleotide resolution. Journal of Visualized Experiments. (50), (2011).
- Maticzka, D., Ilik, I. A., Aktas, T., Backofen, R., Akhtar, A. uvCLAP is a fast and non-radioactive method to identify in vivo targets of RNA-binding proteins. Nature Communications. 9 (1), 1142 (2018).
- Van Nostrand, E. L., et al. Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP). Nature Methods. 13 (6), 508-514 (2016).
- Radulovich, N., Leung, L., Tsao, M. S. Modified gateway system for double shRNA expression and Cre/lox based gene expression. BMC Biotechnology. 11, 24 (2011).
- Breunig, C. T., et al. A Customizable Protocol for String Assembly gRNA Cloning (STAgR). Journal of Visualized Experiments. (142), (2018).
- Kuhn, R. M., Haussler, D., Kent, W. J. The UCSC genome browser and associated tools. Briefings in Bioinformatics. 14 (2), 144-161 (2013).
- Vourekas, A., et al. Mili and Miwi target RNA repertoire reveals piRNA biogenesis and function of Miwi in spermiogenesis. Nature Structural & Molecular Biology. 19 (8), 773-781 (2012).
- Preitner, N., et al. APC is an RNA-binding protein, and its interactome provides a link to neural development and microtubule assembly. Cell. 158 (2), 368-382 (2014).
- Meyer, C., et al. The TIA1 RNA-Binding Protein Family Regulates EIF2AK2-Mediated Stress Response and Cell Cycle Progression. Molecular Cell. 69 (4), 622-635 (2018).
- Gerstberger, S., Hafner, M., Tuschl, T. A census of human RNA-binding proteins. Nature Reviews Genetics. 15 (12), 829-845 (2014).
- Vourekas, A., Mourelatos, Z. HITS-CLIP (CLIP-Seq) for mouse Piwi proteins. Methods in Molecular Biology. 1093, 73-95 (2014).

