Metabolic Pathway Confirmation and Discovery Through 13C-labeling of Proteinogenic Amino Acids
Summary
13C-isotope labeling is a useful technique for determining the cell central metabolism for various types of microorganisms. After cells have been cultured with a specific labeled substrate, GC-MS measurement can reveal functional metabolic pathways based on unique labeling patterns in proteinogenic amino acids.
Abstract
Microbes have complex metabolic pathways that can be investigated using biochemistry and functional genomics methods. One important technique to examine cell central metabolism and discover new enzymes is 13C-assisted metabolism analysis 1. This technique is based on isotopic labeling, whereby microbes are fed with a 13C labeled substrates. By tracing the atom transition paths between metabolites in the biochemical network, we can determine functional pathways and discover new enzymes.
As a complementary method to transcriptomics and proteomics, approaches for isotopomer-assisted analysis of metabolic pathways contain three major steps 2. First, we grow cells with 13C labeled substrates. In this step, the composition of the medium and the selection of labeled substrates are two key factors. To avoid measurement noises from non-labeled carbon in nutrient supplements, a minimal medium with a sole carbon source is required. Further, the choice of a labeled substrate is based on how effectively it will elucidate the pathway being analyzed. Because novel enzymes often involve different reaction stereochemistry or intermediate products, in general, singly labeled carbon substrates are more informative for detection of novel pathways than uniformly labeled ones for detection of novel pathways3, 4. Second, we analyze amino acid labeling patterns using GC-MS. Amino acids are abundant in protein and thus can be obtained from biomass hydrolysis. Amino acids can be derivatized by N-(tert-butyldimethylsilyl)-N-methyltrifluoroacetamide (TBDMS) before GC separation. TBDMS derivatized amino acids can be fragmented by MS and result in different arrays of fragments. Based on the mass to charge (m/z) ratio of fragmented and unfragmented amino acids, we can deduce the possible labeled patterns of the central metabolites that are precursors of the amino acids. Third, we trace 13C carbon transitions in the proposed pathways and, based on the isotopomer data, confirm whether these pathways are active 2. Measurement of amino acids provides isotopic labeling information about eight crucial precursor metabolites in the central metabolism. These metabolic key nodes can reflect the functions of associated central pathways.
13C-assisted metabolism analysis via proteinogenic amino acids can be widely used for functional characterization of poorly-characterized microbial metabolism1. In this protocol, we will use Cyanothece 51142 as the model strain to demonstrate the use of labeled carbon substrates for discovering new enzymatic functions.
Protocol
1. Cell culture (Figure 1)
- Grow cells in minimal medium with trace elements, salts, vitamins, and specifically labeled carbon substrates that are best for pathway investigation. Use either shaking flasks or bioreactors for cell culture. Organic nutrients, such as yeast extract, may interfere with the measurement of amino acid labeling and thus cannot be present in the culture medium.
- Monitor cell growth by the optical density of the culture at an optimal wavelength (e.g., OD730 for Cyanothece 51142) with a UV/Vis spectrophotometer.
- Cells can first be grown in a non-labeled medium. The middle-log growth phase cells are preferred to be used for inoculation (3% (v/v) by volume inoculation ratio) of the labeled medium. The labeled culture should be sub-cultured (3% v/v inoculation ratio) in the same labeled medium to avoid the introduction of non-labeled carbon from the initial inoculum.
2. Amino acid extraction
- Harvest sub-cultured cells (10mL) in the middle-log growth phase by centrifugation (10 min, 8000×g).
- Resuspend the pellet in 1.5mL of 6M HCl and transfer it to a clear glass, screw-top GC vial. Cap the vials and place them in a 100°C oven for 24 hours to hydrolyze the biomass proteins into amino acids. Hydrolysis of biomass pellets can yield 16 of the 20 common amino acids (Figure 2) 5. Cysteine and tryptophan are degraded, and glutamine and asparagine are converted to glutamate and aspartate, respectively.
- Centrifuge the amino acid solution at 20,000×g for 5 min using 2 ml Eppendorf tubes, and transfer the supernatants to new GC vials. This step removes solid particles in the hydrolysis solution.
- Remove the GC vial lids and dry the samples completely under a stream of air using a Thermo Scientific Reacti-Vap evaporator (note: a freeze dryer can also be used to dry samples). This step can be done overnight.
3. Amino acid derivatization and GC-MS conditions
Analysis of amino acids or charged/highly polar metabolites via GC requires that these metabolites be derivatized, so that the amino acids are volatile and can be separated by gas chromatography 2.
- Dissolve the dried samples with 150 μL of tetrahydrofuran (THF) and 150 μL of N-(tert-butyldimethylsilyl)-N-methyltrifluoroacetamide derivatization reagent.
- Incubate all samples in an oven or a water bath between 65 and 80°C for 1 hour. Vortex occasionally to make sure the metabolites in the vial are dissolved.
- Centrifuge the samples at 20,000×g for 10 min, and then transfer the supernatant to new GC vials. The supernatant should be a clear and yellowish solution. Due to saturation of the detectors, GC-MS measurement accuracy can be affected by the high concentration of injected TBDMS derivatized amino acids (these samples often shows dark brown color), therefore, we should dilute these samples using THF before GC-MS measurement 6.
- Analyze the samples by GC-MS (use a 1:5 or 1:10 split ratio, injection volume = 1 μL, carrier gas helium = 1.2 mL/min). Use the following GC temperature program: hold at 150°C for 2 minutes, increase at 3°C per min to 280°C, increase at 20°C per min to 300°C, and then hold for 5 minutes. Solvent delay can be set as ~5 min (for a 30 meter GC column). The range of the mass to charge ratio (m/z) in MS can be set between 60 and 500.
4. GC-MS data analysis
- TBDMS derivatized amino acid measurement can also be affected by isotope discrimination in GC separation. Light isotopes move slightly faster than heave isotopes in GC column. To reduce the potential measurement bias, we may average the mass spectrum of the whole amino acid peak range 6
- The GC and MS spectra of TBDMS derivatized metabolites have been reported before7. The GC retention time and the unique m/z peaks for each amino acid are illustrated in Figure 3.
- Derivatization of amino acids or central metabolites introduces significant amounts of naturally-labeled isotopes, including 13C (1.13%), 18O (0.20%), 29Si (4.70%), and 30Si (3.09%). The measurement noise from natural isotopes in the raw mass isotopomer spectrum can be corrected by using published software5, 8. The final isotopic labeling data are reported as mass fractions, e.g., M0, M1, M2, M3 and M4 (representing fragments containing zero to four 13C labeled carbons).
- Measurement of amino acids can provide isotopic labeling information about eight crucial precursor metabolites: 2-oxo-glutarate, 3-P-glycerate, acetyl-CoA, erythrose-4-P, oxaloacetate, phosphoenolpyruvate, pyruvate, and ribose-5-P. The labeling patterns in these metabolites can be used to identify several central metabolic pathways (Figure 2) 9. The outcome of the labeling experiments can be further confirmed using other biochemistry methods (e.g., RT-PCR).
5. Pathway analysis using labeled amino acid data
By investigating only a few key amino acids produced from well-designed 13C tracer experiments, we may reveal several unique pathways or enzyme activities without performing sophisticated 13C-metabolic flux analysis of entire central metabolism.
- Entner–Doudoroff pathway: [1-13C] glucose can be used as the carbon source. If the pathway is active, serine labeling will be significantly lower than labeling in alanine 10.
- Branched TCA cycle: [1-13C] pyruvate can be used as the carbon source. If the TCA cycle is broken, aspartate can be labeled by two carbons, while glutamate is labeled with only one carbon 11, 12.
- CO2 fixation by Calvin-Benson-Bassham cycle in a mixotrophic metabolism: Non-labeled CO2 and labeled carbon substrates are both used as the carbon sources. If Calvin cycle is functional, serine and histidine labeling will be significantly diluted, comparing to other amino acids. Such method can determine the relative CO2 fixation when organic carbon sources are present in the medium 13.
- Oxidative pentose phosphate pathway: [1-13C] glucose can be used as the carbon source. If the pathway is active, non-labeled alanine will be >50% 12.
- Anaplerotic pathway (e.g., PEP + CO2 à oxaloacetate): 13CO2 and non-labeled carbon substrates (e.g., glycerol or pyruvate) can be used as the carbon source. If the pathway is active, aspartate labeling will be significantly enriched, comparing to alanine and serine 13.
- Re-citrate synthase: [1-13C] pyruvate can be used as the carbon source. If the enzyme is active, glutamate is labeled in β-carboxyl group 3, 4.
- Citramalate pathway: [1-13C] pyruvate, [2-13C] glycerol, or [1-13C] acetate can be used as the carbon source. If the pathway is active, leucine and isoleucine labeling amounts are identical 14.
- Serine-isocitrate lyase cycle: [1-13C] pyruvate or [1-13C] lactate can be used as the carbon source. If the pathway is active, the third position carbon in serine will be labeled 15.
- Utilization of nutrients (i.e., exogenous amino acids): a culture medium with fully labeled carbon substrates and non-labeled amino acids can be used. If the cells selectively utilize these supplemented non-labeled nutrients, we will see significant labeling dilution of these amino acids in the biomass. This method can be used to investigate which nutrient supplements are preferred by the cell 16.
6. Representative Results
Recent bioenergy studies have revived interests in using novel phototrophic microorganisms for bioenergy production and CO2 capture. In the past years, quite a few 13C-metabolism analyses, including advanced 13C-Metabolic Flux Analyses (13C-MFA), have been applied to investigate central metabolisms in phototrophic bacteria, because biochemical knowledge of the central metabolic pathways is not well-founded in these non-model organisms10, 11, 17-20. Here, we present an example of the discovery of an alternate isoleucine pathway in Cyanothece 51142 21. Cyanothece 51142 does not contain the enzyme (EC 4.3.1.19, threonine ammonia-lyase), which catalyzes conversion of threonine to 2-ketobutyrate in the typical isoleucine synthesis pathway. To resolve the isoleucine pathway, we grow Cyanothece 51142 (20 mL) in ASP2 medium 22 with 54 mM glycerol (2-13C, >98%). Cyanothece 51142 utilizes 2nd position labeled glycerol as the main carbon source. We observe that threonine and alanine have one labeled carbon, while isoleucine is labeled with three carbons. Therefore, synthesis in Cyanothece 51142 cannot be derived from the threonine route employed by most organisms (Figure 4). On the other hand, leucine and isoleucine have identical labeling patterns based on fragment (M-15)+ and fragment (M-159)+. For example, the isotopomer data from [M-15]+ (containing unfragmented amino acids) show identical labeling for leucine (M0=0.01, M1=0.03, M2=0.21, M3=0.69) and isoleucine (M0=0.01, M1=0.03, M2=0.24, M3=0.67). Thus leucine and isoleucine must be synthesized from the same precursors (i.e., pyruvate and acetyl-CoA). This observation is consistent with the labeled carbon transition in the citramalate pathway for isoleucine synthesis. To confirm this pathway, we search the Joint Genome Institute database and find the presence of a citramalate synthase CimA (cce_0248) in Cyanothece.
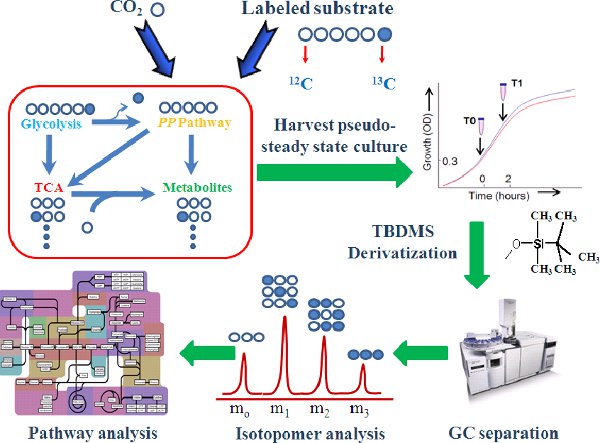
Figure 1.
The 13C-assisted pathway analysis steps.
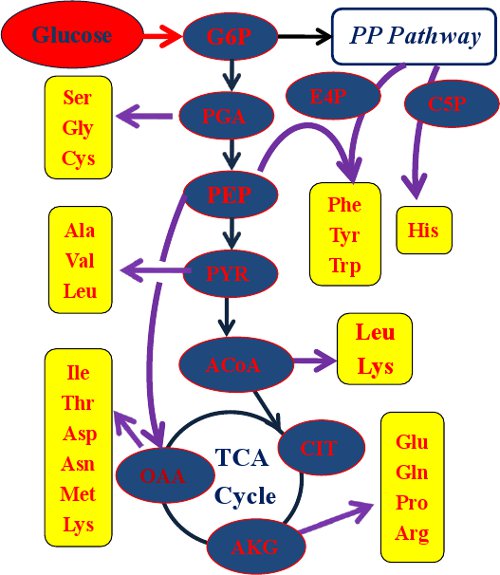
Figure 2.
Amino acids used for acquiring the labeling pattern of their metabolic precursors. ACoA, acetyl-CoA; AKG, α-Ketoglutarate; C5P, ribose 5-phosphate; CIT, citrate; E4P, erythrose 4-phosphate; G6P, glucose 6-phosphate; OAA, oxaloacetate; PEP, phosphoenolpyruvate; PGA, 3-phosphoglycerate; PYR, pyruvate.
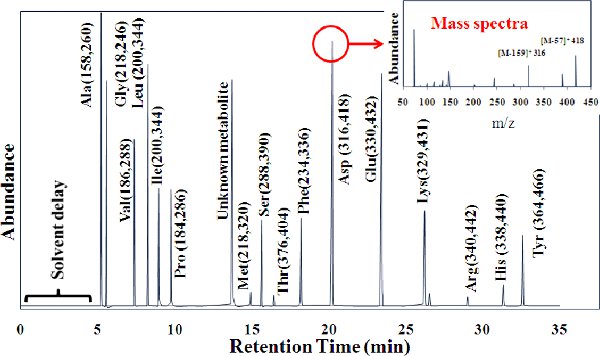
Figure 3. GC peaks for 16 amino acids. TBDMS derivatized amino acids are cracked by MS into two fragments: (M-57)+, containing the entire amino acid, and (M-159)+, which lacks the α carboxyl group of the amino acid. For leucine and isoleucine, the (M-57)+ was overlapped by other mass peaks. We suggest using fragment (M-15)+ to analyze the entire amino acid labeling. The (f302)+ group is detected in most amino acids, which contains only the first (α-carboxyl group) and second carbons in an amino acid backbone. Because this MS peak often has high noise-to-signal ratios, (f302)+ is not recommended for quantitatively analyzing the metabolic fluxes7.
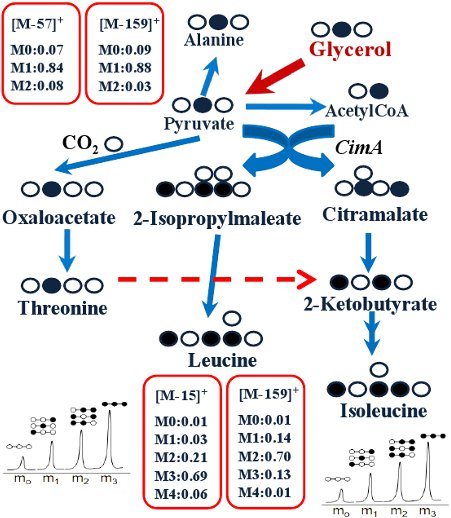
Figure 4.
Labeling transitions in isoleucine pathways in Cyanothece 51142 (modified from our previous paper)21.
Discussion
This protocol consists of feeding the cell with a labeled substrate and measuring the resulting isotopic labeling patterns in the amino acids via GC-MS. Since MS data (m/z ratios) give just the overall amount of labeling of MS ions, we have to assess the isotopomer distributions of amino acids by examining the m/z ratios of both unfragmented (M-57)+ and fragmented amino acids (i.e., (M-159)+ and (f302)+). Furthermore, we can perform several cell cultures with a chemically identical medium but substrates that have different labeling patterns (1st position labeled, 2nd position labeled, etc.). The labeling information about metabolites from these experiments can be integrated to decode the actual carbon transition routes through the central metabolic pathways.
For pathway analysis, the choice of a labeled substrate is important. In general, singly labeled carbon substrates are easier to use in tracing the fate of the labeled carbon when 13C percolates through central pathways, while multiple-carbon labeled substrates may confound carbon tracing. Also, singly labeled substrates are more informative to elucidate unique molecule structures in metabolites than uniformly labeled substrates.4 For example, the (Re)-type citrate synthase shows different reaction stereochemistry from normal citrate synthase, and thus causes citrate to have different molecular chirality. On the other hand, substrates are different in their suitability to detect their associated pathways. Glucose is best for detecting the split ratio between the glycolysis and pentose phosphate pathways, while pyruvate or acetate are best for analyzing the TCA cycle and some amino acid pathways. Therefore, it is necessary to use different substrates to investigate the overall picture of cell metabolisms.
13C-isotope labeling is a useful technique for determining functional pathways in microorganisms. However, this technique has several limitations. First, it is suited only for analysis of carbon metabolism using organic substrates, as it cannot directly resolve metabolism in autotrophic metabolisms if CO2 is used as the sole carbon source. Autotrophic cultures using CO2 label all amino acids to the same extent as the input 12CO2/13CO2 mixture 23. This makes pathway analysis difficult, as metabolism analysis has to be inferred from a rearrangement of 13C carbons in metabolites by different metabolic pathways. Second, this paper presents solely qualitative results discriminating between “active” and “non-active” pathways. Precise quantification of metabolism requires a sophisticated modeling approach (i.e., 13C-MFA) to decipher metabolic fluxes from isotopomer data. Third, the scope of 13C-metabolism analysis is limited by technical challenges in determining low abundance and unstable metabolites. Broader metabolic network can be probed by analyzing free metabolites besides amino acids. Measurement of free metabolites requires both highly-efficient metabolite extraction methods and highly-sensitive analytical platforms. LC-MS, FT-ICR MS, and CE-MS have been used for identifying the labeling patterns of free metabolites, and provide more insight into cell metabolisms2. Fourth, 13C-assisted pathway analysis is best done in minimal medium, because addition of non-labeled nutrient supplements leads to falsely lower labeling concentrations and make quantitative 13C-MFA studies not so straightforward. Also, cells may utilize exogenous amino acids extensively for protein synthesis, and thus give very weak labeling signals for these proteinogenic amino acids 24. If a rich medium is required to grow cells, measurement of intracellular metabolites, instead of amino acids, can effectively reduce the interference in labeling data that arises from exogenous non-labeled carbon nutrients.
Finally, an increasing number of genome sequences for non-model microbial species are being published each year. However, functional characterization of these species has lagged far behind the pace of genomic sequencing. 13C-labeling approaches can play important roles in the confirmation and discovery of metabolic pathways in many non-model organisms. Furthermore, the labeling information can be integrated with metabolic modeling (13C-MFA) to decipher absolute carbon fluxes in microorganisms 25. Therefore, this technique can be widely used in analyzing biological systems related to biofuel, ecological and medical applications.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This study was supported by an NSF Career Grant (MCB0954016) and a DOE Bioenergy Research Grant (DEFG0208ER64694).
Materials
| Name of the reagent | Company | Catalogue number | Comments (optional) |
|---|---|---|---|
| TBDMS | Sigma-Aldrich | 19915 | – |
| THF | Sigma-Aldrich | 34865 | – |
| Labeled carbon substrate | Cambridge Isotope Laboratories | Depend on the experimental requirement | Website: http://www.isotope.com |
| Gas chromatograph | Agilent Technologies | Hewlett-Packard, model 7890A | – |
| GC Columns | J&W Scientific, Folsom, CA | DB5 (30m) | – |
| Mass spectrometer | Agilent Technologies | 5975C | – |
| Reacti-Vap Evaporator | Thermo Scientific | TS-18825 | For drying amino acid samples |
References
- Zamboni, N., Sauer, U. Novel biological insights through metabolomics and 13C-flux analysis. Curr. Opin. Microbiol. 12, 553-558 (2009).
- Tang, Y. J. Advances in analysis of microbial metabolic fluxes via 13C isotopic labeling. Mass. Spectrom. Rev. 28, 362-375 (2009).
- Tang, Y. J. Investigation of carbon metabolism in Dehalococcoides ethenogenes strain 195 via isotopic and transcriptomic analysisa. J. Bacteriol. 191, 5224-5231 (2009).
- Tang, Y. J. Pathway confirmation and flux analysis of central metabolic pathways in Desulfovibrio vulgaris Hildenborough using GC-MS and FT-ICR mass spectrometry. Journal of Bacteriology. 189, 940-949 (2007).
- Dauner, M., Sauer, U. GC-MS analysis of amino acids rapidly provides rich information for isotopomer balancing. Biotechnology Progress. 16, 642-649 (2000).
- Wittmann, C. Fluxome analysis using GC-MS. Microbial Cell Factories. 6, 6-6 (2007).
- Antoniewicz, M. R., Kelleher, J. K., Stephanopoulos, G. Accurate assessment of amino acid mass isotopomer distributions for metabolic flux analysis. Anal. Chem. 79, 7554-7559 (2007).
- Wahl, S. A., Dauner, M., Wiechert, W. New tools for mass isotopomer data evaluation in 13C flux analysis: mass isotope correction, data consistency checking, and precursor relationships. Biotechnology and Bioengineering. 85, 259-268 (2004).
- Shaikh, A., Tang, Y. J., Mukhopadhyay, A., Keasling, J. D. Isotopomer distributions in amino acids from a highly expressed protein as a proxy for those from total protein. Analytical Chemistry. 80, 886-890 (2008).
- Tang, K. -. H., Feng, X., Tang, Y. J., Blankenship, R. E. Carbohydrate metabolism and carbon fixation in Roseobacter denitrificans OCh114. PLoS One. 4, e7233-e7233 (2009).
- Tang, K. -. H. Carbon flow of Heliobacterium modesticaldum is more related to Firmicutes than to the green sulfur bacteria. J. Biol. Chem. 285, 35104-35112 (2010).
- Feng, X. Characterization of the Central Metabolic Pathways in Thermoanaerobacter sp. X514 via Isotopomer-Assisted Metabolite Analysis. Appl. Environ. Microbiol. 75, 5001-5008 (2009).
- Feng, X. Mixotrophic and photoheterotrophic metabolisms in Cyanothece sp. ATCC 51142 under continuous light. Microbiology. 156, 2566-2574 (2010).
- Tang, Y. J. Flux analysis of central metabolic pathways in Geobacter metallireducens during reduction of soluble Fe(III)-NTA. Appl. Environ. Microbiol. 73, 3859-3864 (2007).
- Tang, Y. J., Meadows, A. L., Kirby, J., Keasling, J. D. Anaerobic central metabolic pathways in Shewanella oneidensis MR-1 reinterpreted in the light of isotopic metabolite labeling. Journal of Bacteriology. 189, 894-901 (2007).
- Zhuang, W. Q. Selective utilization of exogenous amino acids by Dehalococcoides ethenogenes strain 195 and the enhancement resulted to dechloronation activity. Appl. Environ. Microbiol. , (2011).
- Feng, X., Tang, K. -. H., Blankenship, R. E., Tang, Y. J. Metabolic flux analysis of the mixotrophic metabolisms in the green sulfur bacterium Chlorobaculum tepidum. J. Biol. Chem. 285, 35104-35112 (2010).
- McKinlay, J. B., Harwood, C. S. Carbon dioxide fixation as a central redox cofactor recycling mechanism in bacteria. Proc. Natl. Acad. Sci. U.S.A. 107, (2010).
- McKinlay, J. B., Harwood, C. S. Calvin cycle flux, pathway constraints, and substrate oxidation state together determine the H2 biofuel yield in photoheterotrophic bacteria. MBio. 2, (2011).
- Erb, T. J. Synthesis of C5-dicarboxylic acids from C2-units involving crotonyl-CoA carboxylase/reductase: The ethylmalonyl-CoA pathway. PNAS. 104, 10631-10636 (2007).
- Wu, B. Alternative isoleucine synthesis pathway in cyanobacterial species. Microbiology. 156, 596-602 (2010).
- Reddy, K. J., Haskell, J. B., Sherman, D. M., Sherman, L. A. Unicellular, aerobic nitrogen-ï¬xing cyanobacteria of the genus Cyanothece. J. Bacteriol. 175, 1284-1292 (1993).
- Shastri, A. A., Morgan, J. A. A transient isotopic labeling methodology for 13C metabolic flux analysis of photoautotrophic microorganisms. Phytochemistry. 68, 2302-2312 (2007).
- Tang, Y. J. Invariability of central metabolic flux distribution in Shewanella oneidensis MR-1 under environmental or genetic perturbations. Biotechnol Prog. 25, 1254-1259 (2009).
- Zamboni, N., Fendt, S. M., Ruhl, M., Sauer, U. 13C-based metabolic flux analysis. Nature Protocols. 4, 878-892 (2009).

