Low Molecular Weight Protein Enrichment on Mesoporous Silica Thin Films for Biomarker Discovery
Summary
We developed a technology based on mesoporous silica thin film for the selective recovery of low molecular weight proteins and peptides from human serum. The physico-chemical properties of our mesoporous chips were finely tuned to provide substantial control in peptide enrichment and consequently profile the serum proteome for diagnostic purposes.
Abstract
The identification of circulating biomarkers holds great potential for non invasive approaches in early diagnosis and prognosis, as well as for the monitoring of therapeutic efficiency.1-3 The circulating low molecular weight proteome (LMWP) composed of small proteins shed from tissues and cells or peptide fragments derived from the proteolytic degradation of larger proteins, has been associated with the pathological condition in patients and likely reflects the state of disease.4,5 Despite these potential clinical applications, the use of Mass Spectrometry (MS) to profile the LMWP from biological fluids has proven to be very challenging due to the large dynamic range of protein and peptide concentrations in serum.6 Without sample pre-treatment, some of the more highly abundant proteins obscure the detection of low-abundance species in serum/plasma. Current proteomic-based approaches, such as two-dimensional polyacrylamide gel-electrophoresis (2D-PAGE) and shotgun proteomics methods are labor-intensive, low throughput and offer limited suitability for clinical applications.7-9 Therefore, a more effective strategy is needed to isolate LMWP from blood and allow the high throughput screening of clinical samples.
Here, we present a fast, efficient and reliable multi-fractionation system based on mesoporous silica chips to specifically target and enrich LMWP.10,11 Mesoporous silica (MPS) thin films with tunable features at the nanoscale were fabricated using the triblock copolymer template pathway. Using different polymer templates and polymer concentrations in the precursor solution, various pore size distributions, pore structures, connectivity and surface properties were determined and applied for selective recovery of low mass proteins. The selective parsing of the enriched peptides into different subclasses according to their physicochemical properties will enhance the efficiency of recovery and detection of low abundance species. In combination with mass spectrometry and statistic analysis, we demonstrated the correlation between the nanophase characteristics of the mesoporous silica thin films and the specificity and efficacy of low mass proteome harvesting. The results presented herein reveal the potential of the nanotechnology-based technology to provide a powerful alternative to conventional methods for LMWP harvesting from complex biological fluids. Because of the ability to tune the material properties, the capability for low-cost production, the simplicity and rapidity of sample collection, and the greatly reduced sample requirements for analysis, this novel nanotechnology will substantially impact the field of proteomic biomarker research and clinical proteomic assessment.
Protocol
1. Chip Fabrication
- Create the coating solution for the chip by starting with the hydrolyzed silicate precursor solution. Mix 14 mL of tetraethylorthosilicate (TEOS) with 17 mL of ethanol, 6.5 mL of deionized water and 0.5 mL of 6M HCl under strong stirring (1200 rpm) using a stir hot plate. Heat this solution at 80 °C for 2 hours, keeping the stirring constant.
- Prepare polymer solutions by adding the desired tri-block coploymer (pluronic F127, L121 and P123) to 10 mL of ethanol at room temperature with strong stirring. Complete the mixture by adding 7.5 ml of the silicate solution (from step 1.1) into the tri-block copolymer solution followed by 2 hours of strong stirring at room temperature. This represents the final coating solution.
- Apply 1 mL of coating solution to a 4 inch silicon wafer by spin-coating at a rate of 1500 rpm for 20 seconds. Then heat at 80 °C for 12 hours.
- Heat the films to remove organic surfactant by raising temperature 1 °C per minute to 425 °C, and then bake for 5 hours.
- Pretreat the mesoporous silica (MPS) chip surface with oxygen plasma ashing (Plasma Asher – March Plasma System). (O2 Flow rate: 80 sccm, power: 300 W, time: 10 minutes).
- Optional surface chemical modification: silanate chips in 3% organosilane in a Methanol:DI water (19:1) solution for 72 hours at room temperature in an N2 glove box. Rinse sequentially with Methanol and DI water. Cure chips at 110 °C for 15 minutes in a fan-operated oven.
2. Sample Pre-treatment
- Add TFA and ACN to each serum sample such that the final concentrations are 0.01% TFA and 5% ACN. Vortex to mix.
- Shake these samples on a table vortex shaker at room temperature for 30 minutes.
3. Serum Fractionation
- Pre-bake chips over night in an oven at 160 °C. Alternatively, store the chips in a desiccator until ready to use to prevent hydration of the surface by ambient water in air.
- Use compressed air to dust off any particles that may be on the surface of the chip.
- Cut prefabricated, purchased CultureWell chambered coverglass to include the desired number of wells. Clean coverglass with 100% ethanol and then place on MPS chip surface. Press coverglass down with forceps to ensure a complete seal with chip.
- Pipette 10 μL of serum sample into each 3 mm well. Incubate for 30 minutes in a humidified chamber at room temperature.
- Pipette up serum from wells and discard. Pipette 10 μL of deionized water to each well to wash away larger proteins. Repeat 4 times.
- Pipette 5 μL elution buffer (0.1%TFA + 50% ACN) to each sample well. Pipette up and down 30 times while moving the pipette tip around in the well to mix the elution buffer. (Only apply elution buffer to 1-2 samples at a time to prevent elution buffer from evaporating so quickly).
- After mixing, pipette all the elution buffer and place in a microcentrifuge tube until ready to perform MALDI-TOF analysis.
- In order to emulate the complexity of a biological sample and to evaluate the enrichment effect in harvesting low molecular weight species with our on-chip fractionation system, we selected and assembled a standard mixture of proteins and peptides counting twenty-six different species with a broad range of molecular weights (900-66 500 Da) and pIs (4.0-10.2) and concentrations (0.5-8 pmol/μl ) (see protein list in Table 1).
4. MALDI-TOF Analysis of Peptides
- Spot 0.5 μL of sample to MALDI target plate and allow to dry.
- Spot 0.5 μL matrix (α-cyano-4-hydroxycinnamic acid (CHCA), 5 g/L) or saturated solution of trans-3,5-dimethoxy-4-hydroxycinnamic acid (SA) in 50% acetonitrile containing 0.1% TFA and allow to co-crystallization.
- Spot 0.5 μL of calibration solution to each calibration spot and allow to dry.
- Insert target plate into MALDI-TOF mass spectrometer. The machine should be set to positive reflector mode with a laser intensity of 4200 and 3000 shots per sample. The selected mass range should be 800 to 5000 Da with a target mass of 2000 Da.
- Perform same MALDI-TOF analysis in linear mode but change the mass range to 900 to 10,000 Da or 3000 to 70,000 Da and a target mass of 5000 Da.
5. Data Analysis
- The raw spectra were processed with the ConvertPeakList software and the data was exported to SpecAlign software for preprocessing. All spectra were aligned using the PAFFT correlation method and intensities were normalized to total ion current (TIC) in each corresponding spectrum. All spectra were smoothed and de-noised with factor of 4 and 0.5 respectively. Peaks were detected with a baseline of 0.5, mass window of 21 and height ratio 1.5, negative values were removed before analysis.
- Hierarchical clustering was performed using Cluster 3.0 and visualized with MapleTree software. MALDI MS Data (m/z peak intensities) was log-transformed, normalized, and median centered. Pearson correlation was used to calculate the distance between the samples, and complete linkage clustering was performed.
- An independent Student t-test was used for comparison between groups (n = 2 groups) for each detected MS peak prior to unsupervised hierarchical clustering analysis. A P-value of 0.02 or lower was considered significant to select differentially harvested peptides and proteins among the different mesoporous proteomic chips (Large pores vs. Small pores).
6. Representative Results
As shown in Figure 1, in this study we fabricated a series of mesoporous silica thin films with a variety of nanotextures and comprehensively explored their use in selective capturing and enriching LMW peptides and proteins from human serum. Figure 2a and b show the MS spectra of the unprocessed serum sample for peptides in the range of 900 to 10,000 Da and for proteins in the range of 3,000 ~ 70,000 Da respectively. These spectra illustrate the signal suppression in the LMW region due to the presence of well ionized, highly abundant, high molecular weight (HMW) proteins such as Albumin. Figure 2c and d depict the MS spectra of the serum sample after fractionation by the MPS L121 (pore size, 6 nm). The majority of the large molecules have been depleted, resulting in a significant enrichment of the LMW components. As a control, the same serum sample was applied onto a nonporous pure silica surface to evaluate the specificity of MPS thin films for LMWP recovery. As can be seen in Figure 2e and f, there was no significant harvesting of peptides or proteins from the nonporous silica. Thus it can be concluded that it was the mesoporous architecture and not the silica surface affinity that constitutes the predominate factor in the enrichment of LMWP.
Precisely controlled variations in pore size can be achieved through the use of copolymers with differing hydrophobic block lengths. By using the designed proteins and peptides mixture, the effect of pore size on the LMW peptide and protein recovery efficacy was investigated using MPS thin films prepared from four Pluronic surfactants (F127, P123, L121, and L121 plus swelling agent) with different volume ratios of the hydrophilic and hydrophobic components to form pore sizes of 3.7 nm, 5.2 nm, 7.4 nm, and 9.0 nm respectively. This range of pore sizes led to the recovery of a different repertoire of peptides and proteins from the same serum sample via size and shape exclusion (Figure 3). The protein spectra at high molecular weight demonstrate the molecular cut-off of each type of chip. In addition to the size-dependant depletion of HMW proteins, the on-chip fractionation of the standards solution displays a differential and selective enrichment of LMW species associated with the pore sizes. The two-way hierarchical clustering presented in Figure 3b shows the LMW standards enrichment pattern obtained with the different MSC. Even if all the peptides are below the molecular cut-off of the chips, there is a positive correlation between the pore sizes and the molecular weight of the trapped species. The MSC with large pores, up to 9 nm, preferentially harvest bigger peptides, while smaller peptides are recovered more efficiently by the chips with smaller pores. The structural transformation of the mesoporous arrangement was carried out by tuning the concentration of the template polymer. Increasing the concentration of the template polymer resulted in a reduced interfacial curvature between the phases of the water, the copolymer, and the silicate, consequently initiating the interrelated progression from a spherical to a cylindrical structure. Pluronic F127, with its high molecular weight, possesses this high degree of structural periodicity. By increasing the concentration of F127 in starting solution, different MPS thin film periodic nanostructures can be obtained from 3D nanostructure to 2D nanostructure. The 3D cubic and honeycomb hexagonal nanostructures, possessing more desirable nanopore interconnectivity and more accessible nanopore morphology, exhibit superior performance in selectively enriching LMW peptides than the 2D hexagonal structure, even though they share similar pore size distributions and the same molecular cut-off for serum fractionation (Figure 4). We also streamlined the conjugation of organo-silane on MPS chips by introducing oxygen plasma ashing to pretreat the chip surface. In order to qualitatively study the electrostatic effect on selective on-chip enrichment, we use the proteins and peptides mixture. MS analysis of the proteomic standards solution fractionated on MPS chips prepared with L121 and conjugated with the chemical functional groups is presented in Figure 5. The positively charged and negatively charged the peptides and LMW proteins are captured on the anionic and the cationic chips respectively. The quantitative comparison of multiple MPS chips in recovering the peptides with positive net charge is displayed in Figure 5a. The chips with negative charge and the chips without any modification (with a minor negative charge originally) exhibit significantly higher enrichment for those peptides than the chips modified with APTES (-NH2). Conversely, the positively charged MPS chips possess exceptional capability to recover those peptides with negative net charge, as demonstrated in Figure 5b. While α-Endorphin does not show a significant change due to its PI around 6.

Figure 1. Principle of MPS chips fractionation and LMW enrichment. After sample spotting on the surface, LMW proteins and peptides are trapped into the pores while the larger species remained outside the pores and are removed during the washing steps. The enriched fractions are then eluted and analyzed by MALDI.
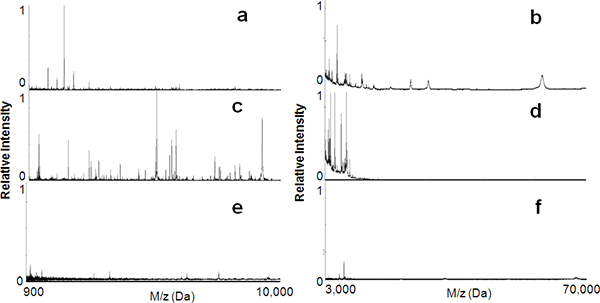
Figure 2. Peptide enrichment using the mesoporous silica thin film chips. MALDI MS profiles in both the low mass range (900 to 10 000 Da) and the high mass range (3000 to 70 000 Da) before (a,b) and after (c,d) serum processing on the mesoporous silica thin films (L121, 6 nm). The molecular recovery is significantly reduced when using blank nonporous silica surfaces (e,f).

Figure 3. Molecular cut-off and size-dependent enrichment of the MPS chips. (a) Magnified view of the MALDI spectra demonstrating the characteristic molecular cut-off of each MPS chips correlating to the pore size. (b) Two-way hierarchical clustering of the peptide mix features among the different chips. The intensity of the red or yellow color indicates the relative peptide concentration. Larger pores enhanced the harvesting of larger peptides (from 3600 to 8500 Da), while the small peptides (from 900 to 3500 Da) were preferentially recovered from the chips with smaller pores.
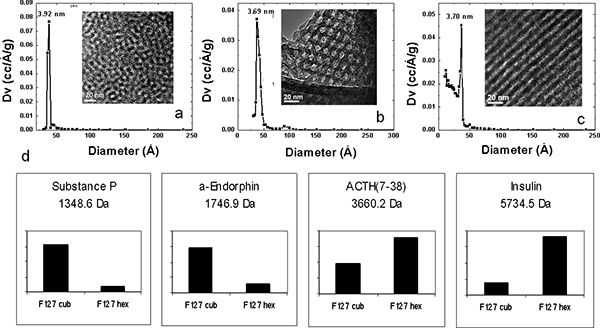
Figure 4. Physical characterizations of MPS thin films and selective recovery with different nanostructures. XRD patterns (a, b, c), TEM (inset a, b, c), Pluronic F127 at different concentrations in the precursor solution: 4.0×10-3 M (a), 6.0×10-3 M (b), and 8.0×10-2 M (c). (d) Bar graph of the intensity of detection illustrating the selective peptides recovery on 3D cubic and 3D hexagonal F127 proteomic chips (Cub and Hex, respectively). The different structural modifications present a selective enrichment.
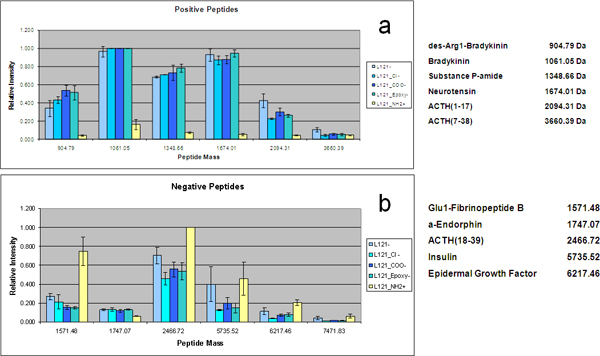
Figure 5. Charge-specific recovery for the chips with different surface functions. Bar graph of the MS intensity of detection of selectively captured peptides on the functionalized chips. According to their iso-electric point, the peptides are positively or negatively charged at pH 7.0. (a) Positive peptides ((1) des-Arg1-Bradykinin, (2) Bradykinin, (3) substance P-amide, (4) Neurotensin, (5) ACTH(1-17), (6) ACTH(7-38)) are specifically enriched on the negatively charged surfaces. (b) Negative peptides ((7) Glu1-fibrinopeptide B, (8) α-endorphin, (9) ACTH(18-39), (10) insulin, (11) EGF, (12) insulin-like GFII) are specifically enriched on the positively charged surfaces. Please click here to see a larger version of this figure.
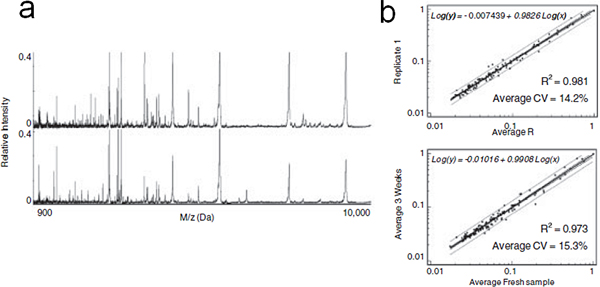
Figure 6. On-chip stabilization of fractionated serum. (a) Representative MALDI profiles of LMW peptides and proteins eluted immediately after serum fractionation (top) or after 3 wk of on-chip storage at room temperature (bottom). (b) From top to bottom: Linear regression analysis of average intensities of detected MS peaks in each replicate compared to replicate 1 for freshly fractionated serum and fractionated serum after 3wk MPS chips storage at room temperature. Equation, CV and coefficient of determination (R2) are indicated. Please click here to see a larger version of this figure.
Discussion
Evidence is mounting that the low molecular-weight region of the circulatory proteome is a rich source of diagnostic biomarkers for the early detection of disease. In this technology, we presented a series of mesoporous silica chips with different pore sizes, pore structures and modifications to selectively enrich peptides and low molecular weight proteins. To evaluate protein stability, the MPS chips were incubated with human serum, dried after washing, and stored for 3wk at room temperature. The protein/peptide patterns obtained were comparable with those of freshly fractionated serum (Figure 6a), as confirmed by the results of the statistical analysis showed in Figure 6b. The variability of the peak signals measured by the average CV was estimated at 12.7% for crude serum and at 14.2% for the fractionated samples. The marginal variations could be due to the internal variability of the MALDI instrument and suggested that the on-chip pretreatment and storage did not induce any significant alteration of the MS protein profiles. The same experiment has been performed on non porous silicon. The dried serum recovered after storage on silicon surface was fractionated on the MPS chips before MALDI analysis. The poor MS profile obtained demonstrated the stabilization advantage of the mesoporous surface. In analogy with previously postulated mechanisms, we hypothesize that the LMW species trapped inside the nanopores were preserved from degradation through the size exclusion of proteases, or by steric inhibition of their proteolytic activity in the confined space of the nanopores. The MPS chip based method could be a powerful tool in the peptide and LMW protein profiling of complex biological fluids study. The MPS chips are inexpensive to manufacture and allow for scaled up production to attain the simultaneous processing of a large number of samples, providing advantageous features for exploratory screening and biomarker discovery.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was funded by Alliance of NanoHealth Pre-center Award (W81XWH-11-2-0168) and Texas Center for Cancer Nanomedicine (1U54CA151668-01).
Materials
| Name of the reagent | Company | Catalogue number | Comments |
| Tetraethoxysilane | Sigma-Aldrich | 131903 | 98% |
| Spin coater | Brewer Science | Cee 200X | |
| Plasma Asher | Nordson March | AP-600 | |
| Spectroscopic Ellipsometer | J. A. Woollam Co. | M-2000DI | |
| MALDI-TOF | Applied Biosystems | Voyager-DE-STR | |
| α-cyano-4-hydroxycinnamic acid | Sigma-Aldrich Co. | C8982 | Matrix for MALDI-TOF |
| trans-3,5-dimethoxy-4-hydroxycinnamic acid | Sigma-Aldrich Co. | 85429 | Matrix for MALDI-TOF |
| Stir hot plate | Thermo Scientific | 11-475-30Q | |
| CultureWell chambered coverglass | Sigma-Aldrich | GBL103350 | 3 mm diam. ×1 mm depth, 3-10 μL, sterile |
| Pluronic F 127 | BASF | PEO106-PPO70-PEO106 | |
| Pluronic L121 | BASF | PEO5-PPO70-PEO5 | |
| Pluronic P123 | BASF | PEO20-PPO70-PEO20 |
Table 1. Physico-chemical properties and designed concentrations (Molecular weight and Iso-electric point) of the selected peptide and protein standards.
References
- Wulfkuhle, J. D., Liotta, L. A., Petricoin, E. F. Proteomic applications for the early detection of cancer. Nat. Rev. Cancer. 3, 267-275 (2003).
- Etzioni, R. The case for early detection. Nat. Rev. Cancer. 3, 243-252 (2003).
- Hanash, S. M., Pitteri, S. J., Faca, V. M. Mining the plasma proteome for cancer biomarkers. Nature. 452, 571-579 (2008).
- Liotta, L. A., Ferrari, M., Petricoin, E. Clinical proteomics: written in blood. Nature. 425, 905-905 (2003).
- Petricoin, E. F., Belluco, C., Araujo, R. P., Liotta, L. A. The blood peptidome: a higher dimension of information content for cancer biomarker discovery. Nat. Rev. Cancer. 6, 961-967 (2006).
- Anderson, N. L., Anderson, N. G. The human plasma proteome: history, character, and diagnostic prospects. Mol. Cell Proteomics. 1, 845-867 (2002).
- Gorg, A., Weiss, W., Dunn, M. J. Current two-dimensional electrophoresis technology for proteomics. Proteomics. 4, 3665-3685 (2004).
- Liu, H., Sadygov, R. G., Yates, J. R., 3rd, . A model for random sampling and estimation of relative protein abundance in shotgun proteomics. Anal. Chem. 76, 4193-4201 (2004).
- Rabilloud, T. Two-dimensional gel electrophoresis in proteomics: old, old fashioned, but it still climbs up the mountains. Proteomics. 2, 3-10 (2002).
- Hu, Y. Tailoring of the nanotexture of mesoporous silica films and their functionalized derivatives for selectively harvesting low molecular weight protein. ACS Nano. 4, 439-451 (2010).
- Bouamrani, A. Mesoporous silica chips for selective enrichment and stabilization of low molecular weight proteome. Proteomics. 10, 496-505 (2010).

