Analyzing and Building Nucleic Acid Structures with 3DNA
Summary
The 3DNA software package is a popular and versatile bioinformatics tool with capabilities to analyze, construct, and visualize three-dimensional nucleic acid structures. This article presents detailed protocols for a subset of new and popular features available in 3DNA, applicable to both individual structures and ensembles of related structures.
Abstract
The 3DNA software package is a popular and versatile bioinformatics tool with capabilities to analyze, construct, and visualize three-dimensional nucleic acid structures. This article presents detailed protocols for a subset of new and popular features available in 3DNA, applicable to both individual structures and ensembles of related structures. Protocol 1 lists the set of instructions needed to download and install the software. This is followed, in Protocol 2, by the analysis of a nucleic acid structure, including the assignment of base pairs and the determination of rigid-body parameters that describe the structure and, in Protocol 3, by a description of the reconstruction of an atomic model of a structure from its rigid-body parameters. The most recent version of 3DNA, version 2.1, has new features for the analysis and manipulation of ensembles of structures, such as those deduced from nuclear magnetic resonance (NMR) measurements and molecular dynamic (MD) simulations; these features are presented in Protocols 4 and 5. In addition to the 3DNA stand-alone software package, the w3DNA web server, located at http://w3dna.rutgers.edu, provides a user-friendly interface to selected features of the software. Protocol 6 demonstrates a novel feature of the site for building models of long DNA molecules decorated with bound proteins at user-specified locations.
Introduction
Understanding the three-dimensional structures of DNA, RNA, and their complexes with proteins, drugs, and other ligands, is crucial for deciphering their diverse biological functions, and for allowing the rational design of therapeutics. Exploration of such structures entails three separate, yet closely related components: analysis (to extract patterns in shapes and interactions), modeling (to assess energetics and molecular dynamics), and visualization. Structural analysis and model building are essentially two sides of the same coin, and visualization complements both of them.
The 3DNA suite of computer programs is an increasingly popular structural bioinformatics toolkit with capabilities to analyze, construct, and visualize three-dimensional nucleic acid structures. Earlier publications outlined the capabilities of the software1, provided recipes to perform selected tasks2, introduced the web-based interface to popular features of the software3, presented databases of structural features collected using 3DNA4, 5 and illustrated the utility of the software in the analysis of both DNA and RNA structures6, 7.
The goal of this article is to bring the 3DNA software kit to laboratory scientists and others with interests and/or needs to investigate DNA and RNA spatial organization with state-of-the-art computational tools. The protocols presented here include step-by-step instructions (i) to download and install the software on a Mac OS X system, (ii-iii) to analyze and modify DNA structures at the level of the constituent base-pair steps, (iv-v) to analyze and align sets of related DNA structures, and (vi) to construct models of protein-decorated DNA chains with the user-friendly w3DNA web interface. The software has the capability to analyze individual structures solved using X-ray crystallographic methods as well as large ensembles of structures determined with nuclear magnetic resonance (NMR) methods or generated by computer-simulation techniques.
The structures examined here include (i) the high-resolution crystal structure of DNA bound to the Hbb protein from Borrelia burgdorferi8 (the tick-borne bacterium that causes Lyme disease in humans9, 10), (ii) two large sets of sequentially related DNA molecules produced with molecular simulations11 – 4,500 snapshots of d(GGCAAAATTTTGCC)2 and d(CCGTTTTAAAACGG)2 collected at 100-psec increments during the calculations, and (iii) a small ensemble of NMR-based structures of the O3 DNA operator bound to the headpieces of the Escherichia coli Lac repressor protein12. The instructions below include information on how to access the files of atomic coordinates associated with each of these structures as well as how to use 3DNA (a copy of this file is found on the 3DNA forum at http://forum.x3dna.org/jove) to examine and modify these structures.
Protocol
1. Installation of the Software Package
- Connect to the 3DNA website at http://x3dna.org and click the link to the 3DNA Forum. Within the Forum select the ‘register’ link and follow the instructions to create a new account.
- The following instructions detail installation of the software on an OS X-based Macintosh computer with a default ‘bash’ shell. The procedure for Linux or Windows (Cygwin, MinGW/MSYS) systems with commonly used shells (including ‘tcsh’) is found within the 3DNA Forum.
- Once you have established a user name and password, log into the Forum. Select the ‘Downloads’ link in the Welcome section and on the new page choose ‘3DNA download’. This will bring you to the 3DNA download page where various versions of the software can be downloaded.
- Select the Mac OS X Intel link to download a compressed tarball file (tar.gz) containing the software.
- Double click on the (tar.gz) file to create a folder named x3dna-v2.1. This folder, which contains the 3DNA software suite, can be moved to any location. For the purpose of this recipe, we drag and drop it into the ‘Applications’ directory. At this stage you are finished with the tar.gz file and can delete it.
- Open the Terminal application, typically found under ‘Utilities’, and change to the x3dna-v2.1 directory created in Step 5 using the following command (ended here and in all of the instructions below by a carriage return):
cd /Applications/x3dna-v2.1 - Run the setup script needed for 3DNA to function properly by typing:
./bin/x3dna_setup - Copy the two-line export settings printed in Step 7. If the user has placed the software in the ‘Applications’ directory, the two lines should appear as follows:
export X3DNA=/Applications/x3dna-v2.1
export PATH=/Applications/x3dna-v2.1/bin:$PATH
A different word besides ‘Applications’ will appear in the export command if the user has chosen to install the software in a different directory. This alternate directory name will replace ‘Applications’ in the above two commands. Note that if the default shell is ‘tcsh’ instead of ‘bash’, the content of the two lines would be: setenv X3DNA /Applications/x3dna-v2.1 and setenv PATH /Applications/x3dna-v2.1/bin:$PATH. - Check in ‘Finder’ to see if a file named ‘.bashrc’ already exists on your computer.
- Open the TextEdit application found in the ‘Applications’ directory and select ‘Make Plain Text’ under the ‘Format’ menu.
- If the ‘.bashrc’ file from Step 9 exists, open the file in TextEdit and paste the export settings from Step 8 to the end of the file. If no ‘.bashrc’ file exits, paste the export settings from Step 8 into a blank file and save with the file name ‘.bashrc’ in your home directory. (The home directory is denoted by a ‘house’ icon on the Macintosh.)
- In order for the new settings to be available, you must run the newly saved ‘.bashrc’ file using the following command within the Terminal program:
source ~/.bashrc - The 3DNA suite is now installed on your system and you can perform Protocols 2-5 within the Terminal program. Protocol 6 is performed using w3DNA, a web interface to selected features of the 3DNA software.
2. Analysis of a Crystal Structure
- We illustrate the analysis of a nucleic acid structure using the DNA bound to the Hbb protein from Borrelia burgdorferi8. The file of atomic coordinates associated with this structure can be found at the Protein Data Bank13 (PDB; http://www.rcsb.org), where it is assigned the structural identifier 2np2. The accompanying Supplementary Data include a copy of this file, which can also be found on the 3DNA Forum at http://forum.x3dna.org/jove.
- The first step in the 3DNA analysis of the structure is the creation of a file that lists all of the paired bases. This is done with the ‘find_pair’ program by typing the following:
find_pair 2np2.pdb 2np2.bps
Here 2np2.pdb is the input file of atomic coordinates described above and 2np2.bps is the output text file that contains the base-pairing information. The latter base-pair file can be examined and edited by the user if necessary. - The next step in the analysis is to determine the geometric parameters that characterize the structure. These include standard chemical torsion angles14, pseudorotation angles that describe the puckering of the sugar ring15, rigid-body parameters that specify the arrangements of bases and successive base pairs16, and other measures of three-dimensional chemical structure. The command ‘analyze’ takes as input the base-pair file generated in Step 2 and creates several output files, which appear in the present working directory. These values are calculated by typing the following command:
analyze 2np2.bps
The output files used here include 2np2.out, which contains an overview of the calculated parameters, and bp_step.par, which contains a list of the rigid-body parameters described above. The latter file can be easily read into a spreadsheet program for further analysis and plotting. See Representative Results (Figure 1) for an example.
3. Construction of a DNA Structure from Rigid-body Parameters
- In addition to the analysis of a nucleic-acid structure, the 3DNA software provides the ability to build a structural model from rigid-body parameters using the ‘rebuild’ command. In this Protocol we show how to construct two DNA helical models, first using the rigid-body parameters from the Hbb-DNA complex as calculated in Protocol 2 and then using a modified set of parameters in which selected roll angles are changed. The roll angle measures the degree of bending of successive base pairs into the grooves of the DNA16. Positive values of roll are associated with the narrowing of the major groove and negative values with the narrowing of the minor groove. The standard reference frame17 adopted by 3DNA and other popular nucleic acid analysis software, e.g. Curves+18, assures numerical consistency in the computed parameters of Watson-Crick base pairs.
- By default, the 3DNA ‘rebuild’ function will create exact atomic models that contain only the coordinates of atoms in the nitrogenous bases. In order to create an approximate atomic model, which includes coordinates from both backbone and base atoms, type the following command:
x3dna_utils cp_std bdna
This command instructs 3DNA to introduce a standard B-DNA backbone conformation in the construction of models from rigid-body parameters. - To build a double-helical model with the rigid-body parameters found in the Hbb-DNA structure, type the following:
rebuild -atomic bp_step.par 2np2_rbld_org.pdb
Here ‘-atomic’ specifies the construction of an all-atom model with atomic coordinates for all heavy (non-hydrogen) atoms, ‘bp_step.par’ is the file generated in Step 3 of Protocol 2 which contains the inputted rigid-body parameters, and ‘2np2_rbld_org.pdb’ is the output file with the atomic coordinates of the created structure. The latter file is formatted in accordance with the specifications of the PDB and thus terminates with ‘.pdb’. - The ‘bp_step.par’ file can be edited to generate a modified structure. On a Macintosh, the changes can be performed using the TextEdit application as described in Protocol 1 Step 10.
- Here we modify the extreme roll angles formed at the two sites of greatest bending. The values, which are expressed in degrees, are changed to zero from values of 64.95 in line 17 and 60.93 in line 26. We save the modified file with a new name, ‘bp_step_roll0.par’, and close the TextEdit program.
- We build the structure, as in Step 3, with the modified set of step parameters as the input file by typing:
rebuild -atomic bp_step_roll0.par 2np2_rbld_roll0.pdb - The two models can be viewed in a standard molecular viewer, such as PyMOL (www.pymol.org), using the generated coordinate files (.pdb) as input. See Representative Results (Figure 2) for images of the rebuilt structures.
4. Analysis of Multi-model Structure Files
- In contrast to Protocol 2, in which we analyze a single nucleic-acid-containing structure, here we show how to analyze a large ensemble of structures. We examine the variation over time of the rigid-body parameters along a series of 14 base-pair DNA chains obtained from molecular-dynamics (MD) simulations11. The analysis considers 4,500 structures, which are spaced at 100-psec increments in the MD calculation, and saved in 4,500 separate files. The accompanying Supplementary Data include copies of these files as well as a set of 4,500 other files for the simulation of a DNA with the reverse sequence (see the introductory text for the precise order of bases). The two sets of files and a smaller subset of data can also be found on the 3DNA Forum at http://forum.x3dna.org/jove.
- The first step in the analysis is the same as that performed in Protocol 2, identification of the paired bases by running the ‘find_pair’ program. Since the entire set of structures shares the same base-pairing scheme one needs to run ‘find_pair’ only once on a representative file. Here we choose the 1001st structure in the simulation of DNA chains containing a central AT step, with the file name md_AT_set1.pdb.1001. The following instruction generates the base-pairing information and stores it in the file md_AT_set1.bps:
find_pair md_AT_set1.pdb.1001 md_AT_set1.bps
The user can examine the identified base paring and make manual changes in the file based on knowledge of the structure. - The next step is to determine the geometric parameters of the entire set of structures using the ‘x3dna_ensemble analyze’ command. The program uses as input the base-pair file generated in Step 2 (md_AT_set1.bps) as well as a file containing a listing of the structures to be analyzed. The latter file, here called md_AT_set1.list, contains the 4,500 files to be analyzed, with the file names entered on individual lines in the order generated by the MD simulation. A copy of this file is included in the Supplementary Material and also offered at the 3DNA Forum. The user should be warned, because of the large number of files, that the computational time can be long (~20 min per 1,000 files on an AMD Phenom II X4 965 processor). The command is run by typing the following:
x3dna_ensemble analyze -b md_AT_set1.bps -l md_AT_set1.list -o md_AT_set1.out
In the above example ‘-b’ pertains to the base-pair file created in Step 2, ‘-l’ specifies that a list of files is given, and ‘-o’ allows the user to assign a name to the output file. There are many options that can be used in conjunction with the ‘x3dna_ensemble analyze’ command, such as selection of specific models. The user can type ‘x3dna_ensemble analyze -h’ to obtain more information on these options. - The next step is to extract the desired rigid-body parameters from the overview output file, md_AT_set1.out, generated in Step 3. This is performed using the ‘x3dna_ensemble extract’ command. A listing of available geometric parameters can be found by typing ‘x3dna_ensemble extract -l’. Here we extract the roll angles and create a comma-separated text file (CSV) of the roll angles with the following command:
x3dna_ensemble extract -p roll -s , -f md_AT_set1.out -o md_AT_set1_roll.csv
The ‘-p roll’ in this command specifies the parameter to be extracted, the ‘-s ,’ signifies a comma-separated file, the ‘-f’ denotes the output file obtained from the ensemble analyze command, and the ‘-o’ allows the user to name the CSV output file of roll angles along the DNA in all analyzed structures. This file can be read into other programs for further analysis and plotting. See Representative Results (Figure 3) for analysis of the variation of roll in the two MD-generated datasets described above and the 3DNA Forum for the MATLAB script used to generate these images.
5. Superposition of Multi-model Structures onto a Common Reference Frame
- The most recent version of 3DNA (version 2.1) has a new feature that allows a user to look at multiple structures from a common perspective. The ‘x3dna_ensemble reorient’ command superimposes a collection of related structures on a common base pair or base-pair step. The following protocol applies this capability to the NMR-derived structures of the O3 DNA operator bound to the headpieces of the Lac repressor protein12. These structures are stored at the PDB under the identifier 2kek in a multi-model structural file, which contains all the structure models in a single file, as opposed to separate files like those used for each of the 4,500 models in Protocol 4. The accompanying Supplementary Data include a copy of this file, which can also be found on the 3DNA Forum at http://forum.x3dna.org/jove.
- As in previous protocols, the first step is to calculate the base pairing of the DNA with the ‘find_pair’ program. In this case we use as input the file ‘2kek.pdb’ and type the following:
find_pair 2kek.pdb 2kek.bps
The base pairing is generated using the first model in the pdb file. Manual corrections to the outputted base-pair file, ‘2kek.bps’, can be made based on knowledge of the structure. - The program ‘x3dna_ensemble reorient’ will align the individual models in a multi-model structure on a common reference frame. This program requires both the pdb file and the base-pairing file from the previous step as input and is run with the following command:
x3dna_ensemble reorient -b 2kek.bps -f 1 -e 2kek.pdb -o 2kek_fr1.pdb
Here the ‘-b’ option specifies the base-pairing file, the ‘-f 1’ denotes the base pair on which all structures are aligned, in this case base pair 1 at the 5´-end of the sequence-bearing strand, the ‘-e’ specifies the input ensemble file, and the ‘-o’ allows the user to provide an output file name, in this case ‘2kek_fr1.pdb’. More information on command options can be obtained by typing ‘x3dna_ensemble reorient -h’. See Representative Results (Figure 4) for discussion of the aligned structures.
6. Construction of a Protein-decorated DNA Molecule
- The w3DNA web interface includes some popular features of the 3DNA software package. Here we draw attention to the capability to construct three-dimensional models of protein-decorated DNA with the web server. The bound DNA fragments adopt the rigid-body parameters of the selected protein-DNA complexes, here the values of the DNA bound to the Hbb protein analyzed in Protocol 2. The unbound regions of DNA can adopt one of three user-selected helical forms (A, B, or C). The A, B, and C refer to three canonical models of DNA, obtained from early fiber diffraction experiments, with respectively 11, 10, and 9 base pairs per turn of the double helix19-21.
- The first step in building a protein-decorated DNA model is to visit the w3DNA website located at http://w3dna.rutgers.edu. Selection of ‘Reconstruction’ from the menu at the top left-hand side of the page activates the online model-building capabilities.
- The next step is to select the ‘Bound Protein-DNA template’ link found in the lightly shaded box in the middle of the new page. This selection will activate a pull-down menu, which allows the user to specify the number of bound proteins. Here we select two bound proteins and click the ‘Continue’ button. Note that models are limited in size to 1,000 base pairs, or 2,000 nucleotides, and thirty bound proteins.
- Selection of the number of bound proteins leads to a specification page similar to that shown in Figure 5. This page allows the user to specify the DNA sequence, the helical form of the unbound DNA, and the identities and locations of the bound proteins.
- The user types or pastes in the desired sequence in the text box in the middle of the specification page (Figure 5, Label 1). Note only standard residues – A, T, C, G, and U – are accepted. Here we enter an 81 base-pair homopolymer, composed entirely of A·T pairs with the A in the sequence-bearing strand.
- There is a pull-down menu (Figure 5, Label 2), below the text box, to select the helical conformation of the unbound regions of DNA. In this example we choose the B-form conformation.
- There is a text box near the bottom of the specification page to enter the binding position(s) and file name(s) of the bound protein(s) (Figure 5, Label 3). The binding position specifies the location of the center of the protein-bound fragment on the DNA sequence. If the bound DNA contains an odd number of base pairs, as in the Hbb-DNA structure, the binding position denotes the location of the middle base pair of the fragment. In the case of Hbb-bound DNA, this middle base pair is a T·T mispair. In this example, where we bind two copies of the Hbb protein to the DNA template, the T·T pair is placed at positions 20 and 63 of the 81 base-pair DNA sequence. Note that the sequence of the bound fragment does not have to coincide with that entered in Step 5. If the protein-bound region contains an even number of base pairs, the binding position denotes the location of the middle base-pair step on the DNA. That is, the central base-pair step of the bound fragment is placed between base pairs n and n +1, where n is the specified number along the segment. Also note that user can decorate the DNA with any protein so long as the structure of its complex with DNA is known and stored in standard PDB format13.
- At the bottom of the specification page there is a box that allows the user to generate a preview of the protein-bound DNA (Figure 5, Label 4). Here we check that box and click the ‘Continue’ button. This action generates a review page listing the selected parameters as well as any errors that may have arisen from the overlap of the selected binding sites. The user can select the ‘Back’ button if any changes need to be made or the ‘Build’ button to proceed.
- The next page displays a static image of the DNA-protein complex in its most extended arrangement and allows for on-line interactive visualization via Jmol and Webmol. The user can also download a coordinate (.pdb) file that is formatted in accordance with the specifications of the PDB. This is done by selecting the ‘Download the rebuilt pdb file’ link below the generated image. The latter file can be easily read into a molecular graphics program for further visualization. See Representative Results (Figure 6) for examples of the Hbb-decorated DNA described above and a slightly longer (86 base-pair) chain with the Hbb centered at positions 20 and 67.
Representative Results
The 3DNA software tools are routinely used to analyze nucleic acid structures. For example, the identities of base pairs and the rigid-body parameters that characterize the arrangements of bases in double-helical fragments of DNA and RNA structures are automatically computed and stored for each new entry in the Nucleic Acid Database22, a worldwide repository of nucleic acid structural information. The values of the rigid-body parameters determined with Protocol 2 readily reveal distortions in three-dimensional structure, such as the two sites of extreme DNA bending into the major groove, with large positive roll angles (64.95° and 60.93°), found at AT·AT steps 13 and 22 in the crystal complex with the Borrelia burgdorferi Hbb protein8 (Figure 1).
The capability of the software to rebuild structures from these quantities with Protocol 3 makes it possible to determine how individual base and base-pair steps contribute to the overall molecular fold. As illustrated in Figure 2, the global bending of DNA induced by Hbb reflects more than the two extreme roll distortions noted above. That is, the DNA remains highly curved when reconstructed with these base-pair steps straightened, i.e. with null roll angles at the two sites. The same technique has previously revealed the contributions of specific base-pair steps and deformations to the superhelical pitch of the DNA wrapped on the surface of the nucleosome core particle6 and to the width of the minor groove of the DNA bound to the bacterial nucleoid-associated protein Fis23.
The new capability in 3DNA, described in Protocol 4, to examine large numbers of related structures makes it possible to extract both sequence- and time-dependent patterns in the spatial arrangements of simulated DNA and RNA molecules. For example, the (yellow) color-coding of the roll angles between successive base pairs in two large sets of simulated DNA structures11 reveals the preferential bending of these molecules at pyrimidine-purine base-pair steps (Figure 3). The higher values of roll, depicted in red, that persist for short periods at the ends of the DNA are suggestive of localized melting and reannealing of the double-helical structure. The variational patterns of other rigid-body parameters, such as the angles and distances between complementary bases, can help to decipher the precise structural distortions.
The capability of the 3DNA software, presented in Protocol 5, to reorient related molecules in a common reference frame, reveals features of overall structure hidden in many of the files stored in the PDB. For example, the conventional alignment of related structures on the basis of a root-mean-square fit of corresponding atoms produces a series of similar spatial pathways that roughly superimpose upon one another, here the ten NMR-based models of the O3 DNA operator bound to the headpieces of the Lac repressor protein12 (Figure 4 left). The superposition of the same structures on a common coordinate frame on the 5´-terminal base pair of each duplex reveals sizable distortions of global structure, in which the molecules flex in appreciably different directions (Figure 4 right). The structural variability may influence the ease with which the Escherichia coli Lac repressor protein binds O3 and induces a loop between O3 and sequentially distant operators in the lac operon24.
The steps outlined in Protocol 6, for building models of long DNA fragments decorated at arbitrary sites with proteins and other ligands, adds a new perspective on the organization of large macromolecular assemblies. Such models help to understand how the multi-molecular complexes interact during biological processing. As illustrated in Figure 6, the precise placement of an architectural protein like Hbb can have a dramatic effect on the overall folding of DNA. If two copies of the known high-resolution structure8 are separated by 43 base pairs, an 81-base-pair DNA fragment closes into a tight, nearly closed configuration. If the two proteins are separated by an additional five base pairs, the DNA follows an open, meandering pathway. The very different arrangements of the protein-decorated duplex show how the spacing of architectural proteins can affect the cyclization or looping of DNA25, 26.
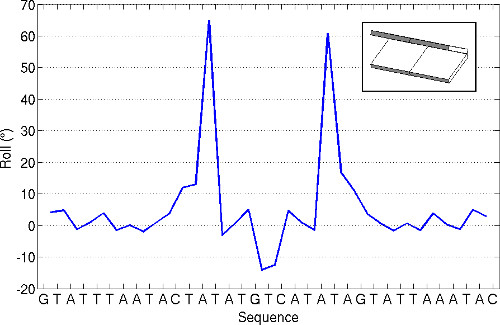
Figure 1. Variation of the roll angle between successive base pairs (see insert for visual depiction) along the DNA chain bound to the Hbb protein from Borrelia burgdorferi8. Values obtained using the ‘analyze’ command of 3DNA and the structural data described in Protocol 2. Note the extreme values of roll at the AT steps that bracket the central third of the structure.
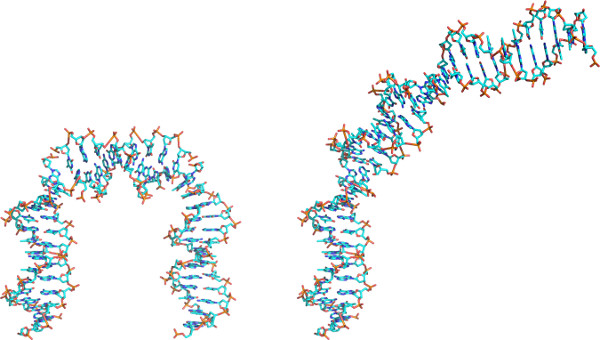
Figure 2. Approximate atomic models of DNA constructed using the ‘rebuild’ function of 3DNA and rendered in PyMOL with color-coded atoms (C-cyan; N-blue; O-red; P-gold). Models based on (left) the rigid-body step parameters of the Hbb protein and (right) a modified set of step parameters, where the two largest values of roll have been set to zero. See Protocol 3 for step-by-step instructions. Note the unfolding of DNA induced by the imposed changes in roll.
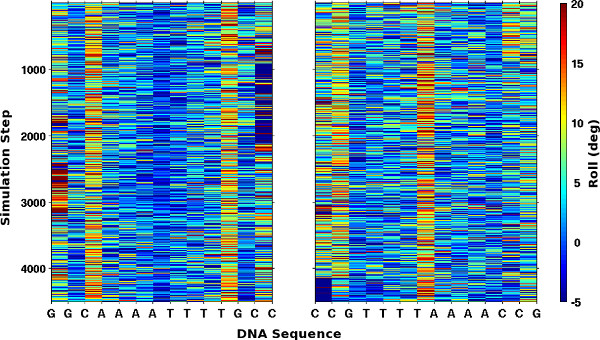
Figure 3. Mosaic images of the roll angles along the DNA in two sets of simulated structures11. Values of roll, extracted using Protocol 4, are color-coded from blue to red over the range [-5°, 20°]. Note the reverse order of bases and large values of roll, highlighted by yellow/red columns, which occur at pyrimidine-purine steps in the two 14 base pair self-complementary sequences.
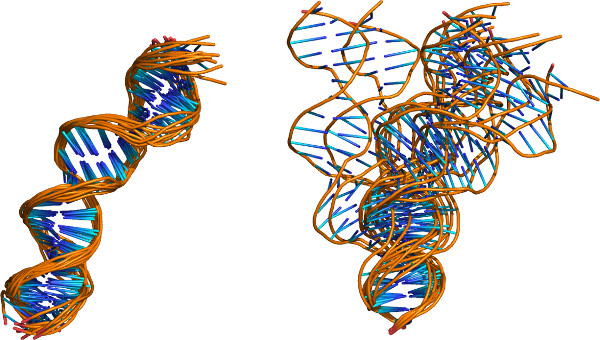
Figure 4. Cartoon images of the DNA models found in the NMR-derived structures of the O3 DNA operator with the Lac repressor protein headpieces12 illustrative of the capabilities of the ‘x3dna_ensemble reorient’ command. Images rendered in PyMOL (backbones show as gold tubes and bases as blue sticks) and aligned using (left) the coordinates in the PDB entry (2kek) and (right) the structural superposition presented in Protocol 5. Note the large differences among the structures when placed in a common reference frame on the 5´-terminal base pair.
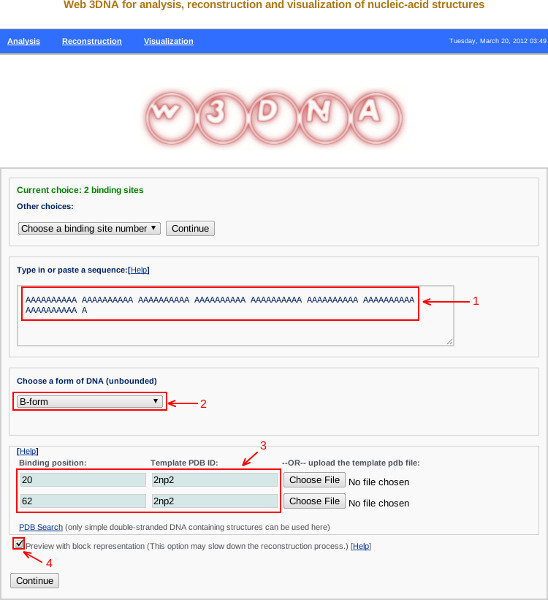
Figure 5. Screen shot from the w3DNA web server illustrating specifications of the DNA sequence (Label 1), the helical form of unbound DNA (Label 2), the positions and identities of proteins (Label 3), and the preview image check box (Label 4) described in Protocol 6. Click here to view larger figure.
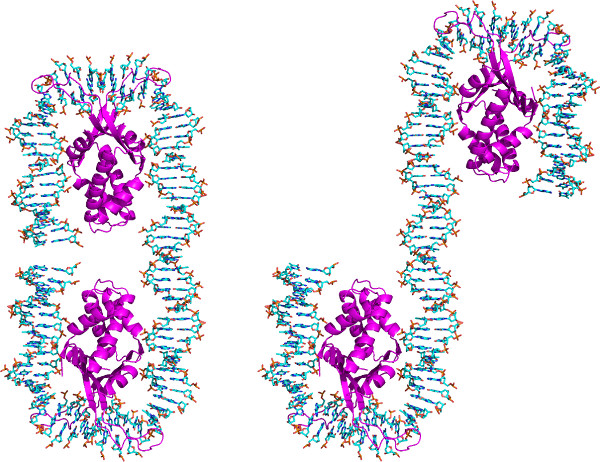
Figure 6. Approximate atomic models of two Hbb proteins8 bound to a long DNA fragment. Structures created with the w3DNA web server as described in Protocol 6 and rendered in PyMOL. The protein chains are show as violet ribbons while the DNA is colored-coded by atom type (C-cyan; N-blue; O-red; P-gold). The central base pair of each protein-binding sites is set at positions (left) 20 and 62 along the 81 base-pair DNA chain and (right) 20 and 67 along the 86 base-pair DNA chain. Note the major change in the folding of the structures associated with the increased (five base-pair) displacement of the two proteins.
Discussion
The set of protocols presented in this article only touch upon the capabilities of the 3DNA suite of programs. The tools can be applied to RNA structures to identify non-canonical base pairs, to determine the secondary structural contexts in which such pairing occurs, to quantify the spatial disposition of helical fragments, to measure the overlap of bases along the chain backbone, etc. The rebuild command allows the user to construct simple and informative block representations of the bases and base pairs like that shown in the inset to Figure 1. The building tools also include features to ‘thread’ different sequences on a given structural template, to generate models of numerous double-, triple-, and four-stranded DNA structures, to orient models in a specific direction, etc. Finally, 3DNA has played a significant role in a number of other projects, such as: the SwS solvation web service for nucleic acids27; the ARTS web server for aligning RNA tertiary structures28; the MDDNA web-based tool for the analysis of molecular dynamics results and structure prediction29; the HADDOCK information-driven protein-DNA docking method30; the SARA server for function annotation of RNA structures31; the 3D-DART DNA structure modeling server32; the 3D-footprint database for the structural analysis of protein-DNA complexes33; the RNA FRABASE 2.0 database for the identification of three-dimensional fragments within RNA structures34; and the SETTER web server for the pairwise comparison of RNA structures35. To the best of our knowledge, there is currently no other nucleic-acid structure software package with a comparable broad combination of features and the robust performance record of 3DNA.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We are grateful to Jiří Šponer for sharing the coordinates of DNA double helices generated in molecular dynamics simulations. We also acknowledge Nada Spackova for assistance in downloading these structures. Support of this work through USPHS Research Grants GM34809 and GM096889 is gratefully acknowledged.
References
- Lu, X. -. J., Olson, W. K. 3DNA: a software package for the analysis, rebuilding, and visualization of three-dimensional nucleic acid structures. Nucleic Acids Res. 31, 5108-5121 (2003).
- Lu, X. -. J., Olson, W. K. 3DNA: a versatile, integrated software system for the analysis, rebuilding, and visualization of three-dimensional nucleic-acid structures. Nature Protocols. 3, 1213-1227 (2008).
- Zheng, G., Lu, X. -. J., Olson, W. K. Web 3DNA-a web server for the analysis, reconstruction, and visualization of three-dimensional nucleic-acid structures. Nucleic Acids. Res. 37, W240-W246 (2009).
- Xin, Y., Olson, W. K. BPS: a database of RNA base-pair structures. Nucleic Acids Res. 37, D83-D88 (2009).
- Zheng, G., Colasanti, A. V., Lu, X. -. J., Olson, W. K. 3DNALandscapes: a database for exploring the conformational features of DNA. Nucleic Acids Res. 38, 267-274 (2010).
- Tolstorukov, M. Y., Colasanti, A. V., McCandlish, D., Olson, W. K., Zhurkin, V. B. A novel ‘roll-and-slide’ mechanism of DNA folding in chromatin. Implications for nucleosome positioning. J. Mol. Biol. 371, 725-738 (2007).
- Lu, X. -. J., Olson, W. K., Bussemaker, H. J. The RNA backbone plays a crucial role in mediating the intrinsic stability of the GpU dinucleotide platform and the GpUpA/GpA miniduplex. Nucleic Acids Res. 38, 4868-4876 (2010).
- Mouw, K. W., Rice, P. A. Shaping the Borrelia burgdorferi genome: crystal structure and binding properties of the DNA-bending protein Hbb. Mol. Microbiol. 63, 1319-1339 (2007).
- Burgdorfer, W., Barbour, A. G., Hayes, S. F., Benach, J. L., Grunwaldt, E., Davis, J. P. Lyme disease-a tick-borne spirochetosis?. Science. 216, 1317-1319 (1982).
- Benach, J. L., Bosler, E. M., Hanrahan, J. P., Coleman, J. L., Habicht, G. S., Bast, T. F., Cameron, D. J., Ziegler, J. L., Barbour, A. G. Spirochetes isolated from the blood of two patients with Lyme disease. N. Engl. J. Med. 308, 740-742 (1983).
- Lankaš, F., Špačková, N., Moakher, M., Enkhbayar, P., Šponer, J. A measure of bending in nucleic acids structures applied to A-tract DNA. Nucleic Acids Res. 38, 3414-3422 (2010).
- Romanuka, J., Folkers, G. E., Biris, N., Tishchenko, E., Wienk, H., Bonvin, A. M. J. J., Kaptein, R., Boelens, R. Specificity and affinity of Lac repressor for the auxiliary operators O2 and O3 are explained by the structures of their protein-DNA complexes. J. Mol. Biol. 390, 478-489 (2009).
- Berman, H. M., Westbrook, J., Feng, Z., Gilliland, G., Weissig, H., Shindyalov, I. N., Bourne, P. E. The Protein Data Bank. Nucleic Acids. Res. 28, 235-242 (2000).
- Joint, I. U. P. A. C. -. I. U. B. Commission on Biochemical Nomenclature (JCBN) Abbreviations and symbols for the description of conformations of polynucleotide chains. Eur. J. Biochem. 131, 9-15 (1983).
- Altona, C., Sundaralingam, M. Conformational analysis of the sugar ring in nucleosides and nucleotides. A new description using the concept of pseudorotation. J. Am. Chem. Soc. 94, 8205-8212 (1972).
- Dickerson, R. E., Bansal, M., Calladine, C. R., Diekmann, S., Hunter, W. N., Kennard, O., von Kitzing, E., Lavery, R., Nelson, H. C. M., Olson, W. K., et al. Definitions and nomenclature of nucleic acid structure parameters. J. Mol. Biol. 205, 787-791 (1989).
- Olson, W. K., Bansal, M., Burley, S. K., Dickerson, R. E., Gerstein, M., Harvey, S. C., Heinemann, U., Lu, X. -. J., Neidle, S., Shakked, Z., et al. A standard reference frame for the description of nucleic acid base-pair geometry. J. Mol. Biol. 313, 229-237 (2001).
- Lavery, R., Moakher, M., Maddocks, J. H., Petkeviciute, D., Zakrzewska, K. Conformational analysis of nucleic acids revisited: Curves+. Nucleic Acids Res. 37, 5917-5929 (2009).
- Franklin, R. E., Gosling, R. G. Molecular configuration in sodium thymonucleate. Nature. 171, 740-741 (1953).
- Watson, J. D., Crick, F. H. C. Genetical implications of the structure of deoxyribonucleic acid. Nature. 171, 964-967 (1953).
- Marvin, D. A., Spencer, M., Wilkins, M. H. F., Hamilton, L. D. A new configuration of deoxyribonucleic acid. Nature. 182, 387-388 (1958).
- Berman, H. M., Olson, W. K., Beveridge, D. L., Westbrook, J., Gelbin, A., Demeny, T., Hsieh, S. -. H., Srinivasan, A. R., Schneider, B. The Nucleic Acid Database: a comprehensive relational database of three-dimensional structures of nucleic acids. Biophys. J. 63, 751-759 (1992).
- Stella, S., Cascio, D., Johnson, R. C. The shape of the DNA minor groove directs binding by the DNA-bending protein Fis. Genes Dev. 24, 814-826 (2010).
- Swigon, D., Coleman, B. D., Olson, W. K. Modeling the Lac repressor-operator assembly: the influence of DNA looping on Lac repressor conformation. Proc. Natl. Acad. Sci. U.S.A. 103, 9879-9884 (2006).
- Czapla, L., Swigon, D., Olson, W. K. Effects of the nucleoid protein HU on the structure, flexibility, and ring-closure properties of DNA deduced from Monte-Carlo simulations. J. Mol. Biol. 382, 353-370 (2008).
- Czapla, L., Peters, J. P., Rueter, E. M., Olson, W. K., Maher, L. J. Understanding apparent DNA flexibility enhancement by HU and HMGB proteins: experiment and simulation. J. Mol. Biol. 409, 278-289 (2011).
- Auffinger, P., Hashem, Y. SwS: a solvation web service for nucleic acids. Bioinformatics. 23, 1035-1037 (2007).
- Dror, O., Nussinov, R., Wolfson, H. J. The ARTS web server for aligning RNA tertiary structures. Nucleic Acids Res. 34, 412-415 (2006).
- Dixit, S. B., Beveridge, D. L. Structural bioinformatics of DNA: a web-based tool for the analysis of molecular dynamics results and structure prediction. Bioinformatics. 22, 1007-1009 (2006).
- de Vries, S. J., van Dijk, M., Bonvin, A. M. The HADDOCK web server for data-driven biomolecular docking. Nat. Protoc. 5, 883-897 (2010).
- Capriotti, E., Marti-Renom, M. A. SARA: a server for function annotation of RNA structures. Nucleic Acids Res. 37, 260-265 (2009).
- van Dijk, M., Bonvin, A. M. 3D-DART: a DNA structure modelling server. Nucleic Acids Res. 37, W235-W239 (2009).
- Contreras-Moreira, B. 3D-footprint: a database for the structural analysis of protein-DNA complexes. Nucleic Acids Res. 38, D91-D97 (2010).
- Popenda, M., Szachniuk, M., Blazewicz, M., Wasik, S., Burke, E. K., Blazewicz, J., Adamiak, R. W. RNA FRABASE 2.0: an advanced web-accessible database with the capacity to search the three-dimensional fragments within RNA structures. BMC Bioinformatics. 11, 231 (2010).
- Čech, P., Svozil, D., Hoksza, D. SETTER: web server for RNA structure comparison. Nucleic Acids Res. , (2012).

