Circulating MicroRNA Quantification Using DNA-binding Dye Chemistry and Droplet Digital PCR
Summary
A sensitive and accurate method for cell-free microRNAs quantification using a dye-based chemistry and droplet digital PCR technology is described.
Abstract
Circulating (of cell-free) microRNAs (miRNAs) are released from cells into the blood stream. The amount of specific microRNAs in the circulation has been linked to a disease state and has the potential to be used as disease biomarker. A sensitive and accurate method for circulating microRNA quantification using a dye-based chemistry and droplet digital PCR technology has been recently developed. Specifically, using Locked Nucleic Acid (LNA)-based miRNA-specific primers with a green fluorescent DNA-binding dye in a compatible droplet digital PCR system it is possible to obtain the absolute quantification of specific miRNAs. Here, we describe how performing this technique to assess miRNA amount in biological fluids, such as plasma and serum, is both feasible and effective.
Introduction
MicroRNAs (miRNAs) are released into blood circulation by potentially all the cells of the organism, as a consequence of active release or necrotic and apoptotic processes. Cell-free miRNAs have been detected in the bloodstream either as free stable molecules or linked to lipoproteins or enveloped inside exosomes and microvesicles 1-3. They are believed to function as cell-to-cell communicators 4, and their amount changes in the presence of cancer, cardiac disorders or autoimmune diseases 5-7. Their accurate and reproducible quantification is the basis for their evaluation as disease biomarkers. However, for several reasons already described elsewhere 8,9, miRNA quantification in serum or plasma, as well as other body fluids, could be very challenging 10,11. We recently developed a method for the absolute quantification of circulating miRNAs, based on miRNA-specific LNA primers and DNA-binding dye droplet digital PCR (ddPCR) technology 12. This methodology has been applied to the validation of miRNA breast cancer biomarkers 13,14.
After the partitioning of each reverse-transcribed miRNA molecule inside a nanoliter-sized droplet, it is possible to count the copy number of each miRNA in each sample, basically counting the number of green, and therefore positive, fluorescent droplets. As soon as a PCR reaction occurs, a positive count is achieved, without the need to establish a standard curve or taking PCR efficiency into account in target amount calculation, as it happens with quantitative RT-PCR (RT-qPCR). In addition, ddPCR proved to be more sensitive and accurate than RT-qPCR in circulating miRNA quantification 15. In this article we present the detailed protocol of this methodology, discussing the most relevant steps andspecifically considering serum and plasma clinical samples.
Protocol
MicroRNA Isolation from Plasma or Serum
Note: Plasma and serum preparation is a relevant step in circulating miRNA quantification. There is no preferred procedure for plasma and serum preparation. The only important thing to consider is that all the samples from the same experiment must be processed using exactly the same workflow. Start from 200 µl serum or plasma. Total RNA can be isolated from serum or plasma using commercially available kits.
1. Protocol for Total RNA (including miRNA) Isolation
- Thaw serum/plasma samples on ice.
- Add 1 ml of Lysis Reagent (e.g., QIAzol) to 200 µl serum/plasma.
- Mix by vortexing and place the tube containing the homogenate on the bench at RT for 5 min.
- Add 3 µl of a 4.16 nM solution of the synthetic miRNA cel-miR-39-3p from C. elegans.
- Add 200 µl of chloroform. Shake the tube vigorously for 15 sec. Place the tube on the bench at RT for 2 min.
- Centrifuge for 15 min at 12,000 x g at 4 °C to obtain phases separation: the upper aqueous phase contains RNA.
- Transfer the aqueous phase to a new tube, about 700 µl, avoid transfer of any white interphase material.
- Add 1 ml of 100% ethanol and mix by inversion.
- Pipet up 700 µl of the sample into a mini spin column placed on a 2 ml collection tube. Close the lid and centrifuge at 12,000 x g for 15 sec. Discard the flow-through.
- Repeat this step using the remaining sample.
- Add 700 µl Buffer RWT to the mini spin column, close the lid and centrifuge for 15 sec at 12,000 x g to wash the column. Discard the flow-through.
- Pipet 500 µl Buffer RPE into the mini spin column. Close the lid and centrifuge for 15 sec at 12,000 rpm. Discard the flow-through. Repeat this step again.
- Centrifuge the mini spin column at full speed for 2 min to dry the spin column membrane from residual ethanol.
- Transfer the mini spin column to a new 1.5 ml collection tube and pipet 35 µl RNase-free water on the membrane of the column.
- Centrifuge for 1 min at 12,000 x g to elute the RNA.
- Store the RNA at -80 °C.
Note: Since RNA concentration cannot be determined accurately, use fixed volumes as a measure for input amount.
2. MicroRNA Reverse Transcription
- Thaw the reverse transcription (RT) reagent mix components: 5x reaction buffer, nuclease-free water. Mix gently inverting the tubes and place on ice. Immediately before use, remove the enzyme mix from the freezer and place on ice. Spin down all reagents.
- Prepare the RT reagent mix on ice as described in Table 1. Prepare at least 10% exceeding mix.
- Mix the reaction by pipetting, and then spin down.
- Incubate for 60 min at 42 ˚C.
- Heat-inactivate the reverse transcriptase for 5 min at 95 ˚C.
- Immediately cool to 4 °C.
- Store the cDNA at -20 °C.
3. cDNA Dilution
- Dilute the amount of cDNA template needed for the planned ddPCR reactions in nuclease free water. Dilute the cDNA reaction between 1:50 and 1:500 in water, dependent on target miRNA abundance. Store the diluted cDNA at -20 °C. The diluted cDNA proved to be stable for at least 3 months at -20 °C.
4. Droplet Generation and PCR
Note: Droplet generation should be performed on 8 samples at a time. Technical replicates are not required, due to the high reproducibility of this technology 12,15.A No Template Control (NTC) sample should be run in every plate and for each different ddPCR condition.
- Thaw and equilibrate the master mix (e.g., EvaGreen Master Mix), microRNA primer set and cDNA at RT.
- Mix the Master Mix thoroughly by inverting the tube several times. Spin down all the reagents.
- Prepare the ddPCR mix as described in Table 2. When multiple ddPCR reactions are performed, prepare a ddPCR mix working-solution. Prepare at least 10% exceeding mix.
- Once assembled, thoroughly mix and spin down the ddPCR mix.
- Dispense 12 µl of ddPCR mix into a 96-well PCR plate or PCR tubes.
- Add 8 µl of diluted cDNA template to each tube/well.
- Insert the droplet generator cartridge into the holder.
- Transfer 20 µl of each prepared sample to the sample wells (middle row) of the droplet generator cartridge carefully while avoiding air bubbles at the bottom of the well.
- Fill each oil well (bottom row) with 70 µl of droplet generator oil.
- Hook the gasket over the cartridge holder and insert into the droplet generator.
- Close the lid to start droplet generation according to manufacturer's protocol. When droplet generation has finished, open the lid, remove the disposable gasket. The top wells of the cartridge contain the droplets.
- Pipet slowly and smoothly 40 µl of the contents of the eight top wells (the droplets) into a single column of a 96-well PCR plate.
- Seal the PCR plate with foil immediately after transferring droplets to avoid evaporation. Use pierceable foil plate seals that are compatible with the needles in the droplet reader.
- Begin thermal cycling (PCR) within 30 min of sealing the plate.
- Perform the cycling protocol according to Table 3.
5. Droplet Reading
- Power on the droplet reader.
- Move the plate from the thermal cycler to the droplet reader.
- Place the 96-well PCR plate containing the post-PCR droplets into the base of the plate holder.
- Place the top of the plate holder on the PCR plate. Firmly press both release tabs down to secure the PCR plate in the holder.
- Start the software from the system PC.
- Click Setup to define the experiment (Absolute quantification) then click Run to start the droplet reading.
- When droplet reading is complete, click "Analyze" button to open and analyze the data.
- Use the 2D amplitude plot in the analysis software to select the positive droplets (lasso tool) (Figure 1B).
- Use the Events tab to check the number of positive and total droplets. A total number of 18,000 – 21,000 droplets is usually achieved with probeless ddPCR.
- Once the positive droplets are selected, export the miRNA concentration values using the export .csv option from the Concentration tab.
Representative Results
The absolute amount of specific miRNAs per ml of plasma or serum can be determined using a green fluorescent DNA-binding dye and droplet digital PCR technology. Figure 1 presents the process of positive-droplets selection, which determines the final miRNA concentration (copies/µl) in the amplification reaction calculated by the analysis software. The amount of each miRNA in the blood is very different, being some miRNA species more abundant than others. Using a 1:50 diluted cDNA in ddPCR reaction it is possible to obtain an adequate number of positive and negative droplets. In case of positive droplet saturation (e.g., no remaining negative droplets), a further dilution of cDNA samples could be necessary.
Every LNA primer set should be optimized to work properly on the droplet digital PCR system. The optimization process for miR-181a-5p is represented in Figure 2. Specifically, changing the amount of primer (usually in the range of 0.25 – 1 µl per ddPCR reaction) and the annealing temperature (usually in the range of 56 – 60 °C), it is possible to select the combination that determines a better separation between positive and negative droplets. In the case of miR-181a-5p, 60 °C annealing temperature and both 0.5 and 1 µl primer amounts provide the better result in terms of droplet amplitude.
It could happen that a primer set gives some non-specific amplification, probably due to primer-dimers formation. When working with circulating miRNAs, the positive signal from samples could be very low and therefore similar to the non-specific signal. If the "cloud" of non-specific target amplification is not overlapping with the true signal (Figure 1C), the miRNA can still be quantified selecting only the true positive droplets. In this case, the NTC sample is essential for droplet selection. All the samples to be included in the same analysis need to be processed using the same methodology and identical ddPCR condition.
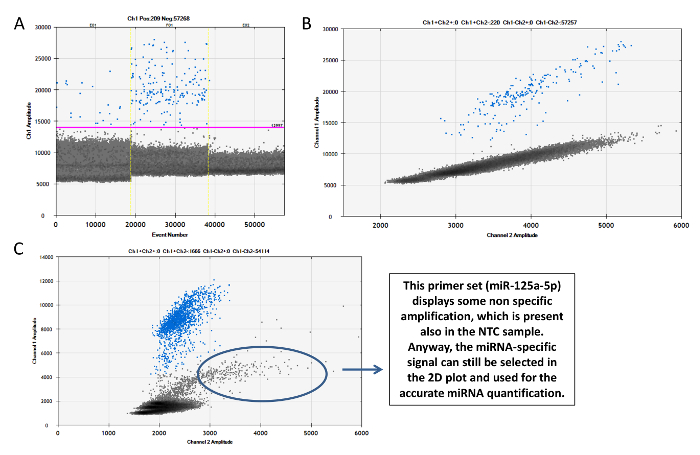
Figure 1. Positive Droplets Selection. Positive droplets are manually selected during the analysis using a threshold above the negative droplets in 1D plot (A) or performing a circle around the positive droplet cloud in 2D plot (B). If some primer set gives a nonspecific amplification, this is evidenced by a second cloud of positive droplets that appears in the negative control (No Template Control, NTC) sample (C). The "true" positives can still be selected excluding the nonspecific signal in 2D plot. Please click here to view a larger version of this figure.
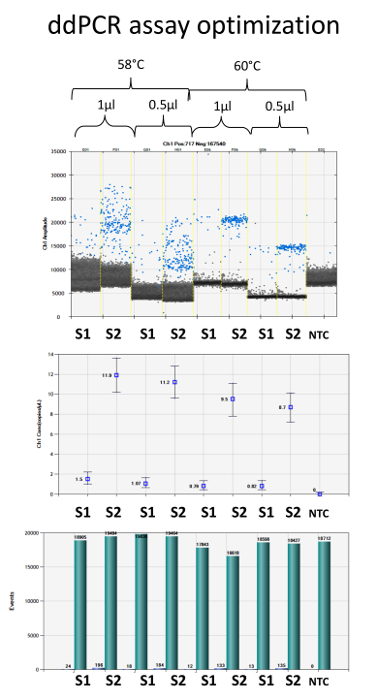
Figure 2. Primer Optimization for Cell-free miRNA Quantification with Probeless ddPCR. Upper panel: Quantification of miR-181a-5p in two plasma samples (S1 and S2) using 4 different ddPCR conditions. MiR-181a-5p LNA primer concentration (0.5 and 1 µl) and the annealing temperature (58 and 60 °C) affect the amplitude of positive (blue) and negative (black) droplets in DNA-intercalating dye ddPCR. A better separation for this miRNA can be obtained performing the PCR at 60 °C and using either 0.5 or 1 µl primer. The control sample (NTC, No Template Control) has no positive droplets, as expected. Central panel: Final concentration of miR-181-5p in the above-mentioned conditions. Results are presented as copies per microliter in the amplification reaction. Error bars represent the Poisson 95% confidence interval. Lower panel: Number of positive droplets (blue) on the total number of droplets (green). Please click here to view a larger version of this figure.
| Reagent | Volume (μl) |
| 5x Reaction buffer | 4 |
| Nuclease-free water | 11 |
| Enzyme mix | 2 |
| Template total RNA | 3 |
| Total volume | 20 |
Table 1 – Reverse Transcription Reagent Mix Components
| Reagent | Volume (μl) |
| 2x DNA-binding dye Supermix | 10 |
| LNA primer set | variable (0.5 – 1)* |
| Nuclease-free water | variable |
| Diluted cDNA template | 8 |
| Total volume | 20 |
| * a first ddPCR reaction should be perform with few samples to establish the best-working amount of primer | |
Table 2 – ddPCR Reagent Mix Components
| Cycling Step | Temperature | Time | Ramping Rate | Cycles |
| Enzyme activation | 95 °C | 5 min | ~2 °C/sec | 1 |
| Denaturation | 95 °C | 30 sec | 40 | |
| Annealing/extension | 60 °C | 1 min | ||
| Signal stabilization | 4 °C | 5 min | 1 | |
| 90 °C | 5 min | 1 | ||
| Hold (optional) | 4 °C | infinite | 1 | |
| Use a heated lid set to 105 °C and set the sample volume to 40 μl. | ||||
Table 3 – Thermal Cycling Conditions for Droplet Digital PCR
Discussion
Circulating miRNAs are present in blood at extremely low concentrations and the amount of RNA that can be extracted from plasma and serum samples is low. For this reason, they are difficult to quantify with other techniques such as microarray and RNA sequencing. Moreover, there is a generalized lack of agreement on data normalization and the presence of endogenous "reference" miRNAs in the blood. In this context, a sensitive technology like droplet digital PCR, capable of counting the number of miRNA copies per ml of serum or plasma even if very low, is very attractive. We paired this technology with a universal cDNA system that reverse-transcribes all the circulating miRNAs in one reaction therefore saving time and, most important, RNA material. In our approach, the specificity is inherent to the use of LNA primers, which performed very well in miRNA quantification when compared to other solutions 16.
Droplet digital PCR is an end-point assay that allows the calculation of target genes absolute concentration. Unlike quantitative PCR, inadequate amplification efficiencies do not affect the final ddPCR quantification. Therefore, considering the amount of PCR inhibitors present for instance in blood samples, ddPCR provides a robust tool for circulating nucleic acids assessment.
Due to its high sensitivity, the technology provides an adequate quantification also of the less abundant miRNAs, even with limited amount of starting material. In addition, assuming that each miRNA molecule is reverse transcribed in a cDNA molecule, we can calculate the absolute copies in 1 µl plasma/serum multiplying the obtained concentration in ddPCR reaction value for a dilution factor (145.83).
The presented workflow can be easily performed on 8-96 samples at a time, thus providing a reliable and time effective tool for microRNA quantification in clinical samples. Droplet digital PCR has the potential to become an essential tool for nucleic acids biomarkers assessment in a diagnostic setting, thereby bringing liquid biopsy from the bench to the bedside.
Disclosures
The authors have nothing to disclose.
Acknowledgements
Supported by funding from the Italian Association for Cancer Research (AIRC) to MF (MFAG 11676) and to MN (Special Program Molecular Clinical Oncology – 5 per mille n. 9980, 2010/15) and from the Italian Ministry of Instruction, University and Research FIRB 2011 to MN (Project RBAPIIBYNP).
Materials
| miRNeasy Mini Kit | Qiagen | 217004 | Columns for total RNA, including miRNA, extraction from serum/plasma |
| 100 nmole RNA oligo Cel-miR-39-3p | Integrated DNA Technologies | Custom | Sequence: UCACCGGGUGUAAAUCAGCUUG |
| Universal cDNA synthesis kit II, 8-64 rxns | Exiqon | 203301 | Kit for microRNA reverse transcription |
| MicroRNA LNA PCR primer set | Exiqon | 204000-206xxx and 2100000-21xxxxx | Primers for miRNA amplification inside droplets |
| QX200 droplet generator | BioRad | 186-4002 | Instrument used for droplet reading |
| QX200 droplet reader | BioRad | 186-4003 | Instrument used for droplet generation |
| QuantaSoft software | BioRad | 186-3007 | Software for data collection and analysis |
| PX1 PCR plate sealer | BioRad | 181-4000 | Plate sealer |
| DG8 droplet generator cartridges and gaskets | BioRad | 186-4008 | Cartridges used to mix sample and oil to generate droplets |
| QX200 ddPCR EvaGreen supermix | BioRad | 186-4033/36 | PCR supermix |
| QX200 droplet generator oil for EvaGreen dye | BioRad | 186-4005 | Oil for droplet generation |
References
- Arroyo, J. D., et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc Natl Acad Sci U S A. 108, 5003-5008 (2011).
- Skog, J., et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol. 10, 1470-1476 (2008).
- Vickers, K. C., Palmisano, B. T., Shoucri, B. M., Shamburek, R. D., Remaley, A. T. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat Cell Biol. 13, 423-433 (2011).
- Braicu, C., et al. Exosomes as divine messengers: are they the Hermes of modern molecular oncology. Cell Death Differ. 22, 34-45 (2015).
- Creemers, E. E., Tijsen, A. J., Pinto, Y. M. Circulating microRNAs: novel biomarkers and extracellular communicators in cardiovascular disease. Circ Res. 110, 483-495 (2012).
- Guay, C., Regazzi, R. Circulating microRNAs as novel biomarkers for diabetes mellitus. Nat Rev Endocrinol. 9, 513-521 (2013).
- Schwarzenbach, H., Nishida, N., Calin, G. A., Pantel, K. Clinical relevance of circulating cell-free microRNAs in cancer. Nat Rev Clin Oncol. 11, 145-156 (2014).
- Moldovan, L., et al. Methodological challenges in utilizing miRNAs as circulating biomarkers. J Cell Mol Med. 18, 371-390 (2014).
- Tiberio, P., Callari, M., Angeloni, V., Daidone, M. G., Appierto, V. Challenges in Using Circulating miRNAs as Cancer Biomarkers. Biomed Res Int. 2015, 731479 (2015).
- Jarry, J., Schadendorf, D., Greenwood, C., Spatz, A., van Kempen, L. C. The validity of circulating microRNAs in oncology: five years of challenges and contradictions. Mol Oncol. 8, 819-829 (2014).
- Witwer, K. W. Circulating MicroRNA Biomarker Studies: Pitfalls and Potential Solutions. Clin Chem. 61, 56-63 (2015).
- Miotto, E., et al. Quantification of Circulating miRNAs by Droplet Digital PCR: Comparison of EvaGreen- and TaqMan-Based Chemistries. Cancer Epidemiol Biomarkers Prev. 23, 2638-2642 (2014).
- Ferracin, M., et al. Absolute quantification of cell-free microRNAs in cancer patients. Oncotarget. , (2015).
- Mangolini, A., et al. Diagnostic and prognostic microRNAs in the serum of breast cancer patients measured by droplet digital PCR. Biomarker Research. , (2015).
- Hindson, C. M., et al. Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat Methods. 10, 1003-1005 (2013).
- Mestdagh, P., et al. Evaluation of quantitative miRNA expression platforms in the microRNA quality control (miRQC) study. Nat Methods. 11, 809-815 (2014).

