A Burrowing/Tunneling Assay for Detection of Hypoxia in Drosophila melanogaster Larvae
Summary
The protocol describes a simple assay to identify Drosophila melanogaster larvae that are experiencing hypoxia under normal atmospheric oxygen levels. This protocol allows hypoxic larvae to be distinguished from other mutants that show overlapping phenotypes such as sluggishness or slow growth.
Abstract
Oxygen deprivation in animals can result from exposure to low atmospheric oxygen levels or from internal tissue damage that interferes with oxygen distribution. It is also possible that aberrant behavior of oxygen-sensing neurons could induce hypoxia-like behavior in the presence of normal oxygen levels. In D. melanogaster, development at low oxygen levels results in inhibition of growth and sluggish behavior during the larval phases. However, these established manifestations of oxygen deficit overlap considerably with the phenotypes of many mutations that regulate growth, stress responses or locomotion. As result, there is currently no assay available to identify i) cellular hypoxia induced by a mutation or ii) hypoxia-like behavior when induced by abnormal neuronal behavior.
We have recently identified two distinctive behaviors in D. melanogaster larvae that occur at normal oxygen levels in response to internal detection of hypoxia. First, at all stages, such larvae avoid burrowing into food, often straying far away from a food source. Second, tunneling into a soft substratum, which normally occurs during the wandering third instar stage is completely abolished if larvae are hypoxic. The assay described here is designed to detect and quantitate these behaviors and thus to provide a way to detect hypoxia induced by internal damage rather than low external oxygen. Assay plates with an agar substratum and a central plug of yeast paste are used to support animals through larval life. The positions and state of the larvae are tracked daily as they proceed from first to third instar. The extent of tunneling into the agar substratum during wandering phase is quantitated after pupation using NIH ImageJ. The assay will be of value in determining when hypoxia is a component of a mutant phenotype and thus provide insight into possible sites of action of the gene in question.
Introduction
The sophisticated array of molecular genetic tools available in D. melanogaster make it a valuable organism for the study of evolutionarily conserved biological processes. Key molecular responses to oxygen availability have proved to be conserved across evolution and prior studies in D. melanogaster have generated insights into the universal components of these signaling pathways 1,2,3,4,5,6.
As part of a study aimed at dissecting sensory neuron function in D. melanogaster larvae, we identified two behavioral responses that proved to be activated by tissue hypoxia at normal oxygen levels 7. One of these, failure to burrow into food, is highly related to the response to low oxygen levels reported by Wingrove and O'Farrell 8. The second behavior, failure to tunnel into a soft substratum during the late third instar wandering phase, had not been previously identified as hypoxia-related. We determined that exposing wild type wandering larvae to low oxygen levels also inhibits substratum tunneling 7, thus establishing that both these behaviors originate from hypoxia – either induced by tissue damage or by low oxygen intake levels. Here we describe an assay we have developed to quantitate these two hypoxia-induced behaviors, which starts with observations immediately after larval hatching.
Hypoxic responses in the early larval stages have not been examined previously and therefore performing an analysis throughout larval life is a valuable component of our assay. Most of the obvious manifestations of hypoxia – slow development, poor growth, and locomotor sluggishness – overlap with larval phenotypes produced by many mutations. But we have found that only third instar larvae with hypoxia show a complete failure to tunnel 7. Thus, we determined that even larvae more compromised in terms of growth and locomotion than our hypoxic larvae, still performed some tunneling, whereas hypoxic larvae never tunneled 7. Another valuable element of this assay is thus that it provides a way to establish when hypoxia is the source of a particular set of pleiotropic phenotypes, as opposed to some other stress or metabolic malfunction. As a demonstration of the assay, here we describe its use in characterizing the responses of larvae with reduced tracheal expression of uninflatable, a gene that functions in the larval airways 9.
We envisage that this assay will be of value to researchers engaged in characterizing larval phenotypes that include poor growth and sluggish behavior. As a result, new genes that influence the distribution, usage, or responses to, oxygen throughout the body could be identified. Further, incorporating this assay into a mutant screening protocol would provide a direct route to identifying mutations that produce hypoxia. This assay will also be valuable in analyzing the circuitry that elicits the hypoxia-induced innate behaviors described here. Neural network analysis of this type is a focus of much current research and the simple nervous system of the D. melanogaster larva is a valuable system for dissecting out automated behaviors. Sensory neurons involved in larval oxygen perception have already been identified, providing a first step towards defining the complete circuitry for hypoxia-induced responses 10,11. Using our assay in combination with selective neural knockdown via the GAL4-UAS system12 is a clear route for delineating further components of the neural network.
Protocol
1. Preparation of larvae
- Two or more days before beginning the assay, set up crosses of the desired experimental and control combinations (minimum 20 females and 10 males each) in egg-lay dishes as described by Wieschaus and Nusslein-Volhard 13 but using 50 mL polypropylene beakers and 6.0 cm disposable Petri dishes containing grape agar and smeared with yeast paste just before use. Keep flies in egg-lay dishes in darkness at room temperature until needed.
- Grape agar recipe – combine 100 mL frozen 100% grape juice concentrate, 350 mL deioinized water, 5 mL glacial acetic and 13 g D. melanogaster agar in a 1L glass beaker. Microwave in 1-2 min bursts, stirring between bursts, until agar is all dissolved. Cool the solution for a few minutes then add 10 mL of 10% (w/v) of Nipagen (methyl-p-hydroxybenzoate) in 95% ethanol. Mix, then pipette 7 mL per plate into disposable 6.0 cm Petri dishes, allow to gel and dry at room temperature for 3-4 h. Store at 4 0C in plastic bags.
- Yeast paste recipe – 7 g dried yeast, 10 mL deionized water, mix with a spatula until uniform consistency.
- Timed egg collections to generate experimental and control larvae. Using new, yeast-smeared, grape plates, collect eggs at room temperature for 4 h in darkness during the morning hours. Remove these 4 h collection plates and replace with fresh plates. Incubate the 4 h collection plates overnight at 25 0C until the afternoon of the next day, when most of the larvae will have hatched. Collect these first instar larvae and use them to set up the experiment.
2. Setting up assay plates
- Preparation of assay plates. For a single run of the assay, prepare five assay plates each of 10 experimental larvae and five plates of 10 control larvae. Use a 1.5 cm cork borer to remove a central core of agar from each plate creating a hole for the food. Put yeast paste (0.8 – 0.9 g) into the hole and pat gently with a spatula to make a mound filling the hole.
- Agar plate preparation – combine 700 mL deionized water with 16 g D. melanogaster agar in a 2L glass beaker and microwave for 5 min. Remove from microwave and stir with a glass rod to bring undissolved agar into the solution. Add 17.5 mL of 10% (w/v) Nipagen in 95% ethanol, mix, then continue microwave heating in 30 s bursts followed by stirring, until all agar is completely dissolved. Cool for a minute or two, then pipette 15 mL aliquots into 10 cm plastic Petri dishes. Allow agar to gel, cover plates with lids and let them dry at room temperature for a few hours or overnight. Store plates in sealed plastic bags at 18 °C.
Note 1: To avoid formation of Nipagen crystals on the surface of the plates due to excess dessiccation, use plates within a few days of preparation. If necessary, crystals can be re-dissolved by dropping a few microliters of 95% alcohol onto them.
Note 2: batches of agar can have different gelling properties. The dry gel plates should be firm to the touch and sufficiently resilient that a clean, intact circle of agar can be removed when using the cork borer to create the food hole (see above). - Setting up larvae in assay plates. Use mechanical pressure to curve the tip of a plastic microspatula and dip the tip in yeast paste to provide “adhesive” for picking up larvae. Pick up first instar larvae individually and place them on the agar plate close to the food mound. Prepare at least five replicate plates of 10 larvae for both experimental and control genotypes. Once completed, cap and label each plate and set the plates (lid side up) in a dark space at room temperature (in our lab this is 22 °C).
3. Monitoring assay plates
- Examine plates daily under a dissecting microscope and record the number of larvae on top of the food or out on the agar surface. Note any visible dead or dying larvae. Note when, and if, tunneling into the agar substratum is seen as larvae move into third instar. Note dates on which pupae begin to form and note any pupal abnormalities. Continue daily observations until all larvae have died or pupated.
- Remove pupae carefully from the plate with a bent teasing needle and transfer to a grape plate. If desired, continue monitoring pupae to determine how many eclose as adults.
4. Preparing assay plates for imaging
- Carefully remove and discard all yeast food from the central well of each plate.
- Gently flood each plate with tap water and use a soft artist’s paint brush to carefully rub off debris that has been tracked onto the agar surface. Replace the water several times and continue gentle brushing until each plate is completely clean. Give plates a final rinse with deionized water, cap them and leave at room temperature overnight to dry.
Note: Tunneling into the agar is most intense around the food mound (see Figure 2). As a result, the rim of the food hole is usually expanded beyond the initial 1.5 cm diameter. In addition, the fragility of the worked agar in this region may cause small pieces of tunneled agar to be lost from the circumference of the hole during cleaning. Increasing the gel agar concentration could help with these problems, but corrections are possible during the Image J quantitation steps (see below).
5. Quantitation of tunneling
- Image the plates. Prepare an image of each plate taken against a black background so as to show the tunnels in the agar as bright white lines.
NOTE: We use a system commonly used to image DNA gels for this step (see Table of Materials and Reagents). Alternative imaging systems capable of generating tiff files, such as digital cameras or cell phone cameras could be adjusted to produce suitable images. - Use the public domain program NIH Image J to quantitate larval tunneling. Download the program from http://rsb.info.nih.gov/ij/download.html. For more information about the program go to http://imagej.nih.gov/ij/docs/intro.html.
NOTE: Other imaging systems could be used to quantitate tunneling. The instructions below apply to NIH Image J. - After opening a tiff file image of a tunneling plate, use the Oval tool to define an area for quantitation that represents the entire agar surface but crops out the plastic edge of the plate.
- Apply the automatic threshold to produce a reverse image of the plate, with the tunnels showing as black lines. Use the Color picker and Paintbrush tools to white out any black areas that represent damage to the agar rather than tunnels.
- Under the Analyze pull-down menu, select Set scale | Click to remove scale and then on the Set measurements window (also under Analyze), check Area and Limit to threshold.
- Press Cntrl key + M key to show the black area (tunnels) in pixels.
- Copy and paste the pixel value for each plate is into a spreadsheet program (such as Excel) that will permit quantitative analysis.
- The central hole is usually expanded due to intense tunneling in this region. To quantitate the missing tunneling, uncheck “Limit to threshold” and use the Wand tool to define the central hole. Press Cntrl key + M key to obtain the pixel value of this area. Subtract from this value the pixel value of a 1.5 cm hole and add the difference to the tunneling value obtained in step 5.6.
- For some plates, the central hole may be missing a piece of tunneled agar that includes a region of untunneled agar adjacent to the hole. When quantitating the central hole for these plates use the Color picker and Paint brush tools to add a black bar across the missing region to define the tunneled region and exclude the non-tunneled area from quantitation.
Note 1: If the experimental larvae perform any tunneling, this quantitation permits calculation of average tunneling values for sets of plates and statistical comparison of control and experimental values.
Representative Results
As a demonstration of the value of this assay we have used it to investigate potential hypoxia in larvae with impaired function of the gene uninflatable (uif) in the tracheae. uif encodes a large transmembrane protein that is strongly expressed on the apical surface of the larval tracheal cells. Mutants of uif were previously noted to exhibit aberrant behavior that could indicate tissue hypoxia due to tracheal malfunction 9. We specifically suppressed uif expression in the larval trachea using the Gal4-UAS system. Two Gal4 lines were used: i) breathless (btl)-Gal4, which is expressed strongly in the tracheae from the beginning of their embryonic development and ii) cut(ue)-Gal4, which begins expression in a small posterior region of the main dorsal trunk tracheae at the end of embryogenesis and continues strong expression in this region throughout larval life 7. A UAS-uif RNAi line was obtained from the Vienna Drosophila Research Center (VDRC ID #1050).
As described in the protocol above, five replicate plates of newly hatched first instar larvae were set up for each of four crosses as follows.
Experiment 1
1) Control 1 – cut(ue)-Gal4 x Canton-S (+)
2) Experimental 1 – cut(ue)-Gal4 x UAS-uif RNAi
Experiment 2
3) Control 2 – btl-Gal4 x Canton-S (+)
4) Experimental 2 – btl-Gal4 x UAS-uif RNAi
The behavior of the cut(ue)-Gal4>UAS-uif RNAi larvae of Experiment 1 was comparable to that of the control cut(ue)-Gal4> + animals in terms of burrowing, tunneling and survival to pupation, (Figure 1 and Figure 2). In contrast, down-regulating uif expression throughout the entire tracheal system from early in development (Experiment 2) produced markedly different responses. The btl-Gal4>UAS-uif RNAi larvae displayed both behavioral manifestations of tissue hypoxia – reduced food burrowing and a total absence of substratum tunneling later in larval life (Figure 1 and Figure 2). The experimental larvae were also noticeably smaller, thinner and more sluggish than controls (Figure 3), and showed a high death rate during the course of the assay. As discussed above, reduced growth, sluggishness and organismal death are known symptoms of larval hypoxia. In addition, most of the experimental larvae failed to attempt pupation and remained as third instar larvae long after control larvae had pupated. Some larvae survived more than 15 days before eventually dying without pupating (Figure 1A). A few of the experimental larvae tried to pupate but in all cases abnormal pupae were formed that did not generate viable adults.
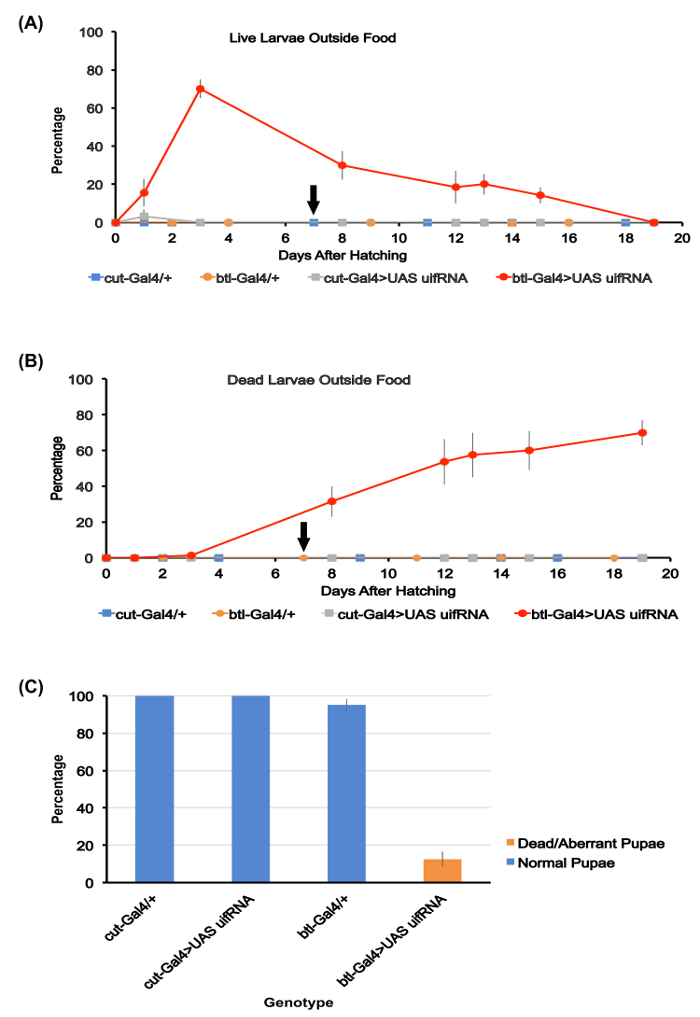
Figure 1. Quantitation of food burrowing and larval development.
(A) Percentage of live larvae outside the food mounds for the four genotypes studied here. Averages for the five assay plates used for each genotype are shown. By day 3 after hatching, ~70% of the btl-Gal4>UAS uif RNAi larvae were outside the food, whereas no larvae were detected outside the food or on the agar surface for the other three genotypes. The btl-Gal4>UAS uif RNAi larvae die slowly between days 4-20, with ~15% of them still alive 15 days after hatching. The black arrow along the x axis here and in (B) indicates the day by which all larvae of the three other genotypes had pupated.
(B) Percentage of dead larvae visible outside the food for the four genotypes. Averages for the five assay plates for each genotype are shown. Dead btl-Gal4>UAS uif RNAi larvae accumulate over time with most dying outside the food. The other genotypes all survive to pupation
(C) Percentage survival to pupation for the four genotypes studied. Averages for the five assay plates for each genotype are shown. No normal pupae of the btl-Gal4>UAS uif RNAi genotype were produced. Error bars throughout figure represent SEMs. Please click here to view a larger version of this figure.
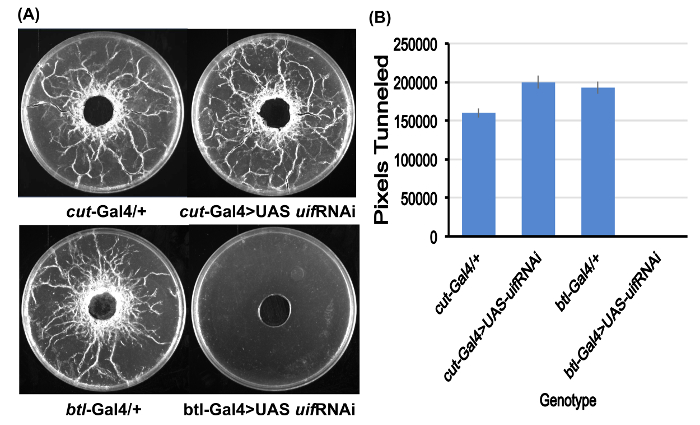
Figure 2. Quantitation of larval tunneling.
(A) Examples of the assay plates for all four genotypes studied after preparation for tunneling quantitation. Note complete absence of tunneling for the btl-Gal4>UAS uif RNAi larvae. Assay plates are 10 cm Petri plates.
(B) Tunneling quantitation derived from Image J for the four genotypes. Average pixel values for the five assay plates for each genotype. No tunneling was observed for the btl-Gal4>UAS uif RNAi larvae. Error bars = SEMs. Please click here to view a larger version of this figure.

Figure 3. Comparison of btl-Gal4>UAS uif RNAi and btl-Gal4> + larvae
Larvae of similar age (day 6 after hatching) for the btl-Gal4>UAS uif RNAi (A) and btl-Gal4> + (B) genotypes. Red arrows point to the dorsal trunk tracheae. Both larvae imaged at the same magnification. Scale bar = 0.5 mm. Please click here to view a larger version of this figure.
Discussion
We have presented here a simple assay designed to detect tissue hypoxia in D. melanogaster larvae. Diagnosis is based on decreased burrowing into food mounds in early larval life and the absence of substratum tunneling late in larval life. Larval crowding can cause premature migration away from a food source and thus one critical aspect of the assay is that a small number of larvae are assayed in the presence of a large excess food. Inclusion of the fungicide methyl-p hydroxyl benzoate (Nipagen) into the agar plates is also essential to prevent mold growth during the assay.
We find that the agar plates can be a source of variability in the assay. Typically, larvae of the same genotype, from different batches of parents, or from different collections of larvae, show relatively limited variation in their behavior in the assay. In contrast, agar plates made on different days or with different batches of agar can yield differences in tunneling. One stipulation therefore is that control and experimental larvae should all be tested using agar plates from the same batch preparation. Agar from different manufacturers or even shipments from the same manufacture can vary in their gelling strength, and so it may be necessary to the adjust the agar concentration upwards from the 2.2% used here to achieve an optimal gel. We have found that wild type larvae can readily burrow through 3% agar gels.
In order to demonstrate the value of this assay, we have used it to investigate potential tissue hypoxia in larvae with suppressed function of uninflatable in the tracheae. Our findings provide strong support for the hypothesis that loss of tracheal expression of this gene can produce hypoxia: btl-Gal4 >UAS-uif RNAi larvae showed failure to burrow into food and complete absence of substratum tunneling during the third instar. In our prior studies of other genotypes, we observed that hypoxia-induced loss of food burrowing is not as complete as loss of substratum tunneling and the btl-Gal4>UAS-uif RNAi larvae studied here behaved similarly. The failed tunneling component of this assay therefore provides the strongest indication of hypoxia.
Although the btl-Gal4>uif RNAi larvae showed behavioral traits diagnostic of hypoxia, the cut(ue)-Gal4>UAS-uif RNAi did not exhibit these abnormalities. The btl and cut(ue) Gal4 drivers are expressed at different stages, and in different patterns, within the larval tracheae. The btl-Gal4 driver is expressed throughout the tracheal system starting from its development in embryogenesis and continuing through larval life. In contrast, Gal4 expression from the cut(ue)-Gal4 driver only begins at the end of embryonic life, after morphogenesis of the tracheae, and is limited to the extreme posterior sections of the dorsal trunks, the major longitudinal vessels of the tracheal system. uif knockdown with this Gal4 line may not therefore reduce uif expression early enough or broadly enough to produce some threshold level of hypoxia required to trigger the behaviors measured in this assay.
A prior study found that third instar larvae exposed to low (10%) oxygen levels show decreased growth and delayed onset of pupariation 14. The btl-Gal4>UAS-uif RNAi larvae studied here progressed to the third instar but the effects on their growth and pupation rates were more pronounced: they were considerably smaller than controls with very little fatty tissue under the epidermis (Figure 3) and only a minor fraction (~10 %) attempted pupariation. These differences suggest the btl-Gal4>UAS-uif RNAi larvae experienced a greater degree of hypoxia, either because uif knockdown in the tracheae was present throughout their entire larval lives, or because it produced a more severe oxygen deprivation in third instar. How loss of uif function in the tracheae might prevent oxygen transport is unclear at this point. The tracheae of the btl-Gal4> UAS-uif RNAi larvae were readily visible through the cuticle (Figure 3), an indication that they contained air and were not damaged to the point that fluid entry compromised function. It is formally possible therefore that the tracheal damage created by loss of uif function does not elicit hypoxia but rather some other defect that inhibits tunneling. For the genotypes studied previously, we determined that failure to tunnel is associated with elevated levels of LDH mRNA7, the canonical indicator of glycolysis and hypoxia in late third instar larvae 15. Thus, final confirmation of hypoxia for the btl-Gal4>UAS-uif RNAi larvae (and in larvae examined in future use of this assay) would involve RT-PCR to assess LDH mRNA levels or use of a commercially available indicator to measure intracellular oxygen levels (for example see 16).
Disclosures
The authors have nothing to disclose.
Acknowledgements
Karen M. Qiang was the 2016 recipient of the George J. Schroepfer Research Award at Rice University. Fanli Zhou is the recipient of a Teaching Fellowship from Rice University. The services of the Bloomington Drosophila Stock Center, the Harvard TRiP facility, the Vienna Drosophila Resource Center are gratefully acknowledged.
Materials
| REAGENTS | |||
| Dehydrated yeast | |||
| Frozen grape juice concentrate | Welch's | Available at most large supermarkets | |
| Glacial acetic acid | Sigma-Aldrich | 320099 | |
| Drosophila agar | Apex Bioresearch Products | 66-103 | |
| Methyl-para-hydroxybenzoate | Apex Bioresearch Products | 20-658 | |
| EQUIPMENT | |||
| 50 ml polypropylene beakers | |||
| 6.0 cm disposable Petri dishes | Falcon | 08757100B | |
| 10 cm disposable plastic Petri dishes | E+K Scientific | EK-24104 | |
| Plastic microspatulas | Corning Incorporated | 3012 | |
| Bent teasing needle | Nasco | S08848MH | |
| Dissecting microscope | Any microscope with 10-30X magnification |
References
- O’Farrell, P. H. Conserved responses to oxygen deprivation. J. Clin. Invest. 107 (6), 671-673 (2001).
- Lavista-Llanos, S., et al. Control of the hypoxic response in D. melanogaster by the basic helix-loop-helix pas protein Similar. Mol. Cell. Biol. 22 (19), 6842-6853 (2002).
- Reiling, J. H., Hafen, E. The hypoxia-induced paralogs Scylla and Charybdis inhibit growth by down-regulating S6K activity upstream of Tsc in D. melanogaster. Genes Dev. 18 (23), 2879-2892 (2004).
- Gorr, T. A., Gassmann, M., Wappner, P. Sensing and responding to hypoxia via Hif in model invertebrates. J. Insect Physiol. 52 (4), 349-364 (2006).
- Romero, N. M., Dekanty, A., Wappner, P. Cellular and developmental adaptations to hypoxia: A D. melanogaster perspective. Meth. Enzymol. 435, 123-144 (2007).
- Dijkers, P. F., O’Farrell, P. H. Dissection of a hypoxia-induced, nitric oxide-mediated signaling cascade. Mol. Biol.Cell. 20 (18), 4083-4090 (2009).
- Zhou, F., Qiang, K. M., Beckingham, K. M. Failure to burrow and tunnel reveals roles for jim lovell in the growth and endoreplication of the D. melanogaster larval tracheae. PLOS ONE. 11, e0160233 (2016).
- Wingrove, J. A., O’Farrell, P. H. Nitric oxide contributes to behavioral, cellular, and developmental responses to low oxygen in D. melanogaster. Cell. 98 (1), 105-114 (1999).
- Zhang, L., Ward, R. E. Uninflatable encodes a novel ectodermal apical surface protein required for tracheal inflation in D. melanogaster. Dev. Biol. 336 (2), 201-212 (2009).
- Langlais, K. K., Stewart, J. A., Morton, D. B. Preliminary characterization of two atypical soluble guanylyl cyclases in the central and peripheral nervous system of D. melanogaster melanogaster. J. Expt. Biol. 207 (13), 2323-2338 (2004).
- Morton, D. B. Behavioral responses to hypoxia and hyperoxia in D. melanogaster larvae. Fly. 5 (2), 119-125 (2011).
- Brand, A., Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development. 118 (2), 401-415 (1993).
- Wieschaus, E., Nusslein-Volhard, C., Roberts, D. B. Looking at embryos. Drosophila: a practical approach. , 199-227 (1986).
- Callier, V., Shingleton, A. W., Brent, C. S., Ghosh, S. M., Kim, J., Harrison, J. F. The role of reduced oxygen in the developmental physiology of growth and metamorphosis initiation in D. melanogaster. J. Expt. Biol. 216 (23), 4334-4340 (2013).
- Li, Y., et al. Hif- and non-Hif-regulated hypoxic responses require the estrogen-related receptor in D. melanogaster. PLoS genetics. 9 (1), e1003230 (2013).
- Grifoni, D., Sallazzo, M., Fontana, E., Froldi, F., Pession, A. Multiple strategies of oxygen supply in Drosophila malignancies identify tracheogenesis as a novel cancer hallmark. Scient. Rep. 5 (9061), (2015).

