Extraction and Quantification of Soluble, Radiolabeled Inositol Polyphosphates from Different Plant Species using SAX-HPLC
Summary
Here we describe strong anion exchange high-performance liquid chromatography of [3H]-myo-inositol-labeled seedlings which is a highly sensitive method to detect and quantify inositol polyphosphates in plants.
Abstract
The phosphate esters of myo-inositol, also termed inositol phosphates (InsPs), are a class of cellular regulators playing important roles in plant physiology. Due to their negative charge, low abundance and susceptibility to hydrolytic activities, the detection and quantification of these molecules is challenging. This is particularly the case for highly phosphorylated forms containing ‘high-energy’ diphospho bonds, also termed inositol pyrophosphates (PP-InsPs). Due to its high sensitivity, strong anion exchange high-performance liquid chromatography (SAX-HPLC) of plants labeled with [3H]-myo-inositol is currently the method of choice to analyze these molecules. By using [3H]-myo-inositol to radiolabel plant seedlings, various InsP species including several non-enantiomeric isomers can be detected and discriminated with high sensitivity. Here, the setup of a suitable SAX-HPLC system is described, as well as the complete workflow from plant cultivation, radiolabeling and InsP extraction to the SAX-HPLC run and subsequent data analysis. The protocol presented here allows the discrimination and quantification of various InsP species, including several non-enantiomeric isomers and of the PP-InsPs, InsP7 and InsP8, and can be easily adapted to other plant species. As examples, SAX-HPLC analyses of Arabidopsis thaliana and Lotus japonicus seedlings are performed and complete InsP profiles are presented and discussed. The method described here represents a promising tool to better understand the biological roles of InsPs in plants.
Introduction
Almost four decades ago, inositol phosphates (InsPs) emerged as signaling molecules, after Ins(1,4,5)P3 (InsP3) was identified as a second messenger that activates the receptor-mediated release of Ca2+ in animal cells1,2. To date, no InsP3 receptor (IP3-R) has been identified in plants, which questions a direct signaling role for InsP3 in plant cells3. Regardless, InsP3 serves as a precursor for other InsPs involved in several plant developmental processes, including the regulation of specific signaling pathways3,4,5,6,7,8. For instance, InsP3 can be further phosphorylated to InsP6, also known as “phytic acid”, which represents a major source of phosphate, myo-inositol and cations, and was shown to play key roles in plant defense against pathogens, mRNA export and phosphate homeostasis5,9,10,11,12.
Inositol pyrophosphates (PP-InsPs) are a class of InsPs that contain at least one high-energy di-phospho bond, initially identified in animal cells, amoeba and yeast, where they play critical roles in various cellular processes13,14,15. Despite seminal work on PP-InsPs in plants16,17,18,19,20,21,22,23,24,25,26, the biological functions and isomer identity of these molecules still remain largely enigmatic. In the model plant Arabidopsis thaliana, cellular InsP8 was proposed to regulate defenses against insect herbivores and necrotrophic fungi via coincidence-detection of InsP8 and active jasmonate by the ASK1-COI1-JAZ receptor complex17. Furthermore, roles of InsP8 and other PP-InsPs in energy homeostasis and nutrient sensing, as well as phosphate homeostasis have been proposed17,23,24,25,26.
Regardless of the biological system employed, one major methodological challenge when studying InsPs has been the reliable detection and precise quantification of these molecules. Mass spectrometry-based methods have been used to detect InsPs, including PP-InsPs, from cell extracts. However, those studies failed to differentiate distinct isomers26,27. Another approach to analyze InsPs employs pull-down of InsPs from cell lysates using TiO2 beads, followed by polyacrylamide gel electrophoresis (PAGE) of the eluted InsPs. The InsPs can then be stained by either toluidine blue or DAPI24,28,29. However, it is so far not possible to reliably detect InsPs lower than InsP5 from plant extracts using this method. Recently, a method using [13C]-myo-inositol for nuclear magnetic resonance (NMR) analysis of InsPs was published as an alternative to strong anion exchange high-performance liquid chromatography (SAX-HPLC)30. This technique has been reported to achieve a similar sensitivity compared to SAX-HPLC and to allow the detection of 5-InsP7, as well as the discrimination of different non-enantiomeric InsP5 isomers from cell extracts. However, the implementation of the NMR-based method requires chemically synthesized and commercially not available [13C]-myo-inositol. Therefore, the method employed in most cases is radiolabeling samples with [3H]-myo-inositol, followed by SAX-HPLC31,32,33. This technique is based on the uptake of radioactive myo-inositol into the plant and its conversion into different InsPs by the combined activity of dedicated cellular kinases and phosphatases.
The [3H]-labeled InsPs are then acid-extracted and fractionated using SAX-HPLC. Because of their negative charge, the InsPs strongly interact with the positively charged stationary phase of the SAX-HPLC column and can be eluted with a buffer gradient containing increasing phosphate concentrations to outcompete InsPs from the column. Elution times thus depend on charge and geometry of the InsP species to be separated. In the absence of chiral columns, only non-enantiomeric isomers can by separated by this protocol. However, radiolabeled standards can be used to assign the isomeric nature of a specific InsP peak. Multiple efforts in the past by various laboratories to generate labeled and unlabeled standards with (bio)chemical methods or to purify them from various cells and organisms have helped assigning peaks to certain InsP species, and also to narrow down the isomeric identity of individual InsP species5,7,21,34,35,36,37,38,39,40,41,42,43. Also, the recent elucidation of enzymatic pathways leading to the formation of PP-InsPs in plants, as well as the discovery of a bacterial type III effector with a specific 1-phytase activity, provide information on how to generate useful standards for these analyses10,17,18,22,23.
The resulting fractions can be measured in a liquid scintillation counter due to the β-decay of tritium (3H). With increasing labeling time, a steady-state isotopic equilibrium is reached, after which the obtained InsP profiles should represent the InsP status of the plant31. The major advantage of this protocol in comparison to other available techniques is the high sensitivity achieved by the use of the direct precursor for InsPs and the measurement of a radioactive signal.
SAX-HPLC of samples extracted from [3H]-myo-inositol-labeled plants or other organisms is commonly used for the detection and quantification of InsPs ranging from lower InsP species to PP-InsPs, representing a valuable tool to better understand the metabolism, function and modes of action of InsPs. So far, this method is also the most appropriate choice for researchers with special interest in lower InsP species. While the basics of this procedure, on which the protocol described here builds on, have been previously described7,21,31,34, a detailed protocol tailored to the analysis of plant-derived InsPs and especially of PP-InsPs is still missing. Previous publications reported difficulties to reliably detect the low abundant PP-InsPs, especially InsP8, due to one or more of the following factors: relatively low amounts of plant material, [3H]-myo-inositol with low specific activity (> 20 Ci/mmol), usage of extraction buffers that are either not based on perchloric acid or are less concentrated than 1 M, different neutralizing buffers, as well as sub-optimal gradients or detection of [3H] with an on-line detector. In comparison to those studies, the protocol presented here is designed for the reliable detection of PP-InsPs7,21,34.
Here we present a detailed workflow, starting from the setup of the equipment to plant cultivation and labeling, InsP extraction and the SAX-HPLC run itself. Although the method was optimized to the model plant A. thaliana, it can be easily modified to study other plant species, as shown here with the first reported InsP profile of the model legume Lotus japonicus. Although the use of a different plant species might require some optimization, we envisage that those will be minor, making this protocol a good starting point for further research in plant InsPs. In order to facilitate possible optimizations, we indicate every step within the protocol in which modifications are possible, as well as all critical steps that may be challenging when establishing the method for the first time. Additionally, we report how data obtained by this method can be used for the quantification of specific InsPs and how different samples can be analyzed and compared.
Protocol
1. Setting up the HPLC system
- Set up a system consisting of two independent HPLC pumps (binary pump), one for each buffer. Both pumps need to be controlled together via a computer with respective software or by having a master pump. Implement a piston seal wash for both pumps, either via gravitational force or through a third low pressure pump. Designate one pump for buffer A (termed pump A) and one for buffer B (termed pump B).
NOTE: Both have to be able to generate pressures up to 60 bar (6 MPa) and flow rates of at least 0.5 mL/min. - Connect both pumps to a dynamic mixer.
- Connect the mixer to an injection valve with a sample loop of at least 1 mL capacity.
- Connect the injection valve to the column with a capillary via the corresponding end fittings.
- Connect the column to the fraction collector by using a capillary with an appropriate length.
NOTE: This description is based on our HPLC system (see the Table of Materials), which requires more manual steps than newer and more sophisticated systems. Our system allows easy access and modification of all components. Quaternary pumps (with the binary gradient described here) can also be used and will lead to elution profiles and overall quality of the analyses similar to those achieved with binary pumps.
2. Preparation of buffers, column and HPLC system
- Prepare the buffers for the extraction of soluble InsPs: extraction buffer (1 M HClO4) and neutralization buffer (1 M K2CO3). Prepare both buffers with ultra-pure deionized water. They are stable at room temperature for several months. Immediately prior to extraction, add EDTA to both solutions to a final concentration of 3 mM (e.g., from a filtered 250 mM EDTA stock solution).
CAUTION: HClO4 (perchloric acid) is strongly corrosive. - Prepare the buffers for the SAX-HPLC run: buffer A (1 mM EDTA) and buffer B (1 mM EDTA, 1.3 M (NH4)2HPO4; pH 3.8 with H3PO4). Prepare both using ultra-pure deionized water followed by vacuum filtration with 0.2 µm pore-sized membrane filters. These are stable at room temperature for several months.
NOTE: EDTA should be included in all buffers to prevent interactions of cations with InsPs, which could result in altered InsP charge or even insoluble InsP salt complexes. - Program the gradient as follows: 0‒2 min, 0% buffer B; 2‒7 min, up to 10% buffer B; 7‒68 min, up to 84% buffer B; 68‒82 min, up to 100% buffer B; 82‒100 min, 100% buffer B, 100‒101 min, down to 0% buffer B; 101‒125 min, 0% buffer B. The optimal flow-rate for this gradient is 0.5 mL/min.
- During the run, collect fractions every minute, starting from minute 1 to minute 96. The remaining 30 min of the gradient serve to wash the column and the system, and do not have to be collected for scintillation counting.
- If possible, set the maximal reachable pressure before the emergency shutdown of the HPLC pumps to 80 bar (8 MPa). This prevents critical damage to the column’s resin.
- When using a new SAX HPLC column, wash it thoroughly (>50 mL) with filtered ultra-pure deionized water before the first use.
NOTE: This will ensure removal of the contained methanol, thus preventing salt precipitation in later steps. If possible, use a separate HPLC pump. If this is not available, make sure that the HPLC has flushed with water before washing the column. The flow-rate should not exceed 2 mL/min. After washing, the column is ready for the analysis and, when properly handled, can be used for 20‒40 runs. After that, the resolution will successively decrease. Prolonged washing with buffer A (>1 h) and performing step 2.6 can help increase the lifetime of the column. If the decrease in resolution persists, the column needs to be exchanged. The gradient can be adjusted to increase the separation between specific inositol polyphosphate species or to decrease the overall runtime. Using different HPLC systems (with different void volume or different volume of the capillaries) will strongly affect the retention times. Also, column changes have minor effects on the retention times. - Perform a "mock run". Instead of an extracted sample, inject filtered ultra-pure deionized water in the HPLC system and run the standard gradient. The fractions do not have to be collected.
NOTE: Step 2.6 is optional. However, it should be performed if one of the following situations apply: A new column is installed; The HPLC system has been used for a different method beforehand; The HPLC system has not been used for longer than 3 days; There was a problem with the preceding run.
3. Plant cultivation and labeling with [3H]-myo-inositol
NOTE: The following steps should be performed with sterile components and under sterile conditions, while wearing gloves to protect hands from contamination with the radiolabel. Plant media, especially when containing sucrose, are prone to microbial contamination.
- Sterilize A. thaliana seeds with 1 mL of 1.2% sodium hypochlorite for 3 min followed by 1 mL of 70% ethanol for 3 min. Then add 1 mL of 100% ethanol, pipette the seeds with the ethanol onto a circular filter paper and allow them to air-dry under a laminar-flow on a clean bench.
- When using L. japonicus seeds, place them in a mortar and scrub seeds with sandpaper before sterilization to ensure a sufficient germination rate.
- Sow out Arabidopsis seeds in 1–2 rows on square Petri dishes filled with solid growth media consisting of half-strength Murashige and Skoog (MS) salt solution, 1% sucrose, 0.7% gellan gum in deionized water adjusted to pH 5.7 with KOH and allow them to stratify for at least 1 day at 4 °C in the dark.
- For Lotus seeds, sow them out in 1 row on square Petri dishes filled with solid growth media consisting of 0.8% bacteriological agar in deionized water and allow them to stratify for at least 3 days at 4 °C in the dark.
- Place the plates vertically in a growth incubator or climate chamber and allow them to grow for 10–12 days under short-day conditions (8 h light at 22 °C, 16 h dark at 20 °C).
- Transfer 10–20 seedlings into one well of a 12-well clear flat-bottomed cell culture plate filled with 2 mL of half-strength MS salt solution supplemented with 1% sucrose and adjusted to pH 5.7.
- Add 45 µCi of [3H]-myo-inositol (30–80 Ci/mmol, dissolved in 90% ethanol) and mix by gentle swirling. Cover the plate with the corresponding lid and seal it with microporous surgical tape (e.g., micropore or leucopore tape), placing it back into the growth incubator.
CAUTION: [3H] is a low-energy beta emitter that can be a harmful radiation hazard when inhaled, ingested or absorbed through bare skin. Always wear gloves when handling radioactive material or equipment that has direct or indirect contact to radioactive material. Also follow the local rules for safe handling of radiochemicals (e.g., wearing additional protective clothes, use of a dosimeter and surveys of surfaces for contaminations on a regular basis). - After 5 days of labeling, remove seedlings from the media and wash them briefly with deionized water. Dry them with paper towels and transfer them into a 1.5 mL microcentrifuge tube. Do not overfill the tube, and place no more than 100 mg FW/tube, which corresponds to approximately 10‒20 17-day-old seedlings.
NOTE: An excess of plant material will dilute the acid during the extraction process and will strongly decrease the extraction efficiency.- Snap-freeze the tube in liquid nitrogen and store it at -80 °C until extraction.
NOTE: Samples can be kept at -80 °C for several weeks without compromising sample quality. The growth conditions (media, light, temperature, time) can be modified according to the needs of a specific experiment or plant species. However, care should be taken when diluting the [3H]-myo-inositol, in order to ensure quantifiable SAX-HPLC runs of good quality. Therefore, it is recommended to start with the [3H]-myo-inositol concentrations stated here and reduce it stepwise if desired. During labeling time, plants can be submitted to different treatments (e.g., environmental stresses or chemical agents) to assess the impact of those conditions on global InsPs. To reach steady-state labeling, we recommend to label plants for at least 5 days.
- Snap-freeze the tube in liquid nitrogen and store it at -80 °C until extraction.
4. Extraction of soluble InsPs
NOTE: Keep samples and reagents on ice during the whole extraction process. Always wear gloves and protective glasses due to the high risk of contact with radioactive material, especially during grinding. Everything that gets in contact with samples is considered as radioactive waste and should be disposed of according to the local rules for safe disposal of radioactive material.
- Prepare the working solutions for the extraction and neutralization buffer as in step 2.1. Each sample will require 600 µL of extraction buffer and 400 µL of neutralization buffer. Store the buffers on ice.
- Take the samples from -80 °C freezer and keep them in liquid nitrogen until further processing. Grind the samples with a microcentrifuge tube pestle until they start thawing and add 500 µL of ice-cold extraction buffer. Continue grinding until sample is completely homogenized and the solution has a deep green color (if leaves are present in the sample).
- Centrifuge the samples for 10 min at 4 °C at ≥ 18000 x g. Transfer the supernatant into a fresh 1.5 mL tube. Keep in mind that the tubes used for extraction are considered solid radioactive waste and need to be disposed of accordingly.
- Carefully add 300 µL of neutralization buffer to the extract. Precipitation of proteins and bubbling will start immediately. Mix by swirling with a pipette tip after a minute and wait for a few seconds before pipetting a small amount (5 µL) on pH paper (ideally range of pH 6–9). The pH should be between pH 7 and 8 in the end.
- If necessary, add small amounts (typically 10–20 µL) of either neutralization buffer or extraction buffer until the desired pH is reached. Let the samples rest on ice for at least 1 h with an open lid.
- Centrifuge the samples for 10 min at 4 °C at ≥ 18,000 x g. Transfer the supernatant into a fresh 1.5 mL tube.
NOTE: The samples can be either directly used in a SAX-HPLC run or kept on ice (if used later on the same day) or frozen in liquid nitrogen and stored at -80 °C for 2‒4 weeks. To ensure a high reproducibility and comparability, it is recommended to always freeze the samples in liquid nitrogen for 5 min, even if they will be directly used afterwards. Longer term storage of extracted samples at -80 °C is possible as long as samples are only thawed once. If frozen samples are used for the analysis, make sure that no particles are visible after thawing. Otherwise, centrifuge again for 10 min at 4 °C at ≥ 18,000 x g and transfer the supernatant into a fresh 1.5 mL tube.
5. Performing the HPLC run
- Equip the fraction collector with 96 small scintillation vials (capacity of ~6 mL) and fill each vial with 2 mL of a suitable scintillation cocktail (e.g., Ultima-Flo AP liquid scintillation cocktail) compatible with buffers with low pH and high ammonium phosphate concentration (see Table of Materials).
NOTE: The number of vials and the size of the vials depend on the fraction collector and scintillation counter used. It is important to at least collect the first 90 fractions, if the gradient described here is used, to obtain a full inositol polyphosphate profile. Also make sure to properly label every vial and its respective lid, to prevent mix-up of fractions or samples. - Start the HPLC system/pumps and have it ready to run. Activate the piston seal wash and keep it activated during the whole run. Load the sample by manually injecting the complete supernatant from step 4.5 (approximately 750 µL) using a suitable syringe (see Table of Materials). If automatic injection is possible, transfer the sample to the corresponding sample vial. Turn the valve from “load” to “inject” position and start the gradient and the fraction collector.
NOTE: Depending on the HPLC system used, the starting procedure might differ, especially when comparing older systems (as described here) with a fully software-controlled newer model. It is very important to ensure that the gradient, the sample injection and fraction collection start simultaneously. - While the HPLC run is ongoing, check the pressure regularly. The starting pressure should be around 18–24 bar (1.8–2.4 MPa) and should slowly rise to 50–60 bar (5–6 MPa) once 100% buffer B is reached.
CAUTION: Decreased pressure might indicate a leak in the system while increased pressure indicates a blockage. Pressure fluctuations (≥ 3 bar in a few seconds) can indicate the presence of air in the system. Keep in mind that everything that leaves the column, as well as every leakage that occurs at the injector or afterwards is radioactive.
NOTE: The pressure also depends on the HPLC system and can be lower or higher than stated here. It will slowly increase after approximately 15–20 runs. However, this does not necessarily influence the quality of the obtained runs. - After the run, close the vials tightly and mix the fractions with the scintillation cocktail by vigorous shaking. Proceed directly with the measurement or keep the vials in an upright position, ideally in the dark.
NOTE: Fractions mixed with scintillation cocktail are stable for weeks and can be measured later. Since the half-life of tritium is 12.32 years, the signal loss is negligible. - Once the run of the last sample of the day is finished, stop both HPLC pumps.
- (Optional) To increase the longevity of the system, especially when it is not used regularly, wash pump B and capillaries by placing the capillary from buffer B into a bottle with buffer A and let the pump run for 10–15 min. Before the next use, remember to replace the capillary into buffer B and to uncouple pump B from the mixer to flush it with buffer B. Once the pump and capillaries are filled again with buffer B, reconnect it with the mixer and the system is ready to use.
6. Measuring the fractions
- Insert the vials into scintillation counter racks and measure each vial for 5 min in a liquid scintillation counter.
- Ideally, use racks that directly fit small vials and avoid hanging in the vials in bigger (e.g., 20 mL) vials to reduce counting errors. The software settings used in this protocol are shown in Supplemental Figure 1.
NOTE: Regularly perform an SNC (self-normalization and calibration) protocol using unquenched [3H] standards. Shorter counting times (1–5 min) are possible to reduce the waiting time. However, to ensure a high counting reproducibility and accuracy, 5 min are recommended.
7. Data analysis
- Export the measurements from the scintillation counter as a spreadsheet file or a compatible/convertible file format. Evaluate the data with a computer equipped with Excel or similar software, and a suitable analysis software like Origin.
- Prepare a 2-D line chart where the measured counts per minutes (cpm) are plotted against the retention time (see Figure 1, Figure 2).
- To compare samples with each other, normalize the data by summing up the cpm from each eluted fraction from minute 25 to 96 for each individual sample.
NOTE: Minute 25 is used as cut-off to exclude unincorporated [3H]-myo-inositol, InsP1 and InsP2 from the analysis, as those tend to fluctuate strongly and cannot be well separated (at least with the gradient proposed in this protocol) and thus strongly change the normalization factor due to their high activity. - Normalize all data to the sample with the lowest total cpm (in fractions 25‒96) by dividing the total cpm from the sample with the lowest cpm (in fractions 25–96) by the total cpm (in fractions 25–96) of the other samples. The resulting factor can then be used to normalize the cpm from each fraction by multiplying the cpm of each fraction with the factor.
NOTE: In the end, the sum of the cpm values from minute 25 to the end should be equal for all samples compared with each other. Only normalized runs should be presented in the same graph/figure (when presented as actual profiles). Supplemental Figure 2 shows an example of how these calculation steps are made (using only fractions 25–35 of two samples for simplification). However, in some cases it is not necessary to normalize data. For instance, when peaks are quantified according to step 7.4 and presented as percentages of total InsPs (as shown in Figure 3D). As stated before, when presenting multiple analyses side by side as profiles, or when the actual measured activity is used for conclusions (e.g., treatment a) increases InsP7 by x% compared to control, referring to the cpm values of InsP7 of both samples and not to their percentage of total InsPs) normalization is needed. To analyze the effect of genotype or treatment differences on labeling efficiency, it is important not 세스 normalize, as this would invalidate these differences. However, absolute quantification with this method is challenging because the extraction efficiency with this protocol can be variable for various reasons and are sometimes even observed when replica of same genotype and treatment are analyzed. Keep in mind that depending on the HPLC system, column and gradient used for the analyses, the cut-off might need to be changed. - To perform relative quantifications of certain inositol polyphosphate peaks and to subsequently create bar graphs that contain data of replications for statistical analyses, continue the analysis with a specialized software that can calculate peak areas of chromatograms (e.g., Origin). See Supplemental Figure 3.
NOTE: Most HPLC systems that are software-controlled are supplied with a respective software capable of this task. Peaks are determined as the fractions with cpm values above background (that varies to a certain degree between runs) and retention times that are similar to previously published data. The retention time of a specific peak is determined in spreadsheet software (e.g., Excel) and used to assign peaks for calculation of definite integrals (e.g., in Origin). Supplemental Figure 3 illustrates this process of peak determination, background subtraction and integration of peaks.
Representative Results
The results shown here aim to illustrate possible outcomes obtained according to variations at technical and biological levels. The first is exemplified by analyses using new versus aged columns (Figure 1) and fresh versus stored samples (Figure 3), and the second by evaluating extracts from two different plant systems, A. thaliana (Figure 1, Figure 3) and L. japonicus (Figure 2).
An optimal SAX-HPLC run is depicted on Figure 1A‒C, which shows a complete inositol polyphosphate spectrum obtained from A. thaliana extracts after scintillation counting. Note that peaks are nicely separated and can be assigned to different isomers (or enantiomer-pairs) based on chromatographic mobilities described earlier5,7.
Figure 2 shows the representative result of a SAX-HPLC analysis of L. japonicus seedlings that were grown and labeled under the same conditions as the Arabidopsis seedlings. While presumably all InsP species and peaks that are known from Arabidopsis can be seen, there are substantial differences regarding the relative (e.g., ratios between isomers) amount of specific InsP isomers, when comparing the profiles of both species. For instance, the Lotus extracts showed increased InsP3c, InsP4b, InsP5b and reduced InsP3a, InsP4a, InsP5a and InsP5c compared to Arabidopsis which leaves room for further investigations. Figure 2D illustrates the different ratios between InsP isomers between Arabidopsis and Lotus.
Figure 3 shows two InsP profiles of a sample that was split after the extraction. The first half was immediately analyzed and the second half one day later, after storage at -80 °C. Note that only minor differences are observed between the different samples (i.e., black and red lines on Figure 3A‒C, and Figure 3D). This illustrates that one freeze-thaw cycle does not harm the sample and that the method itself generates reproducible results.
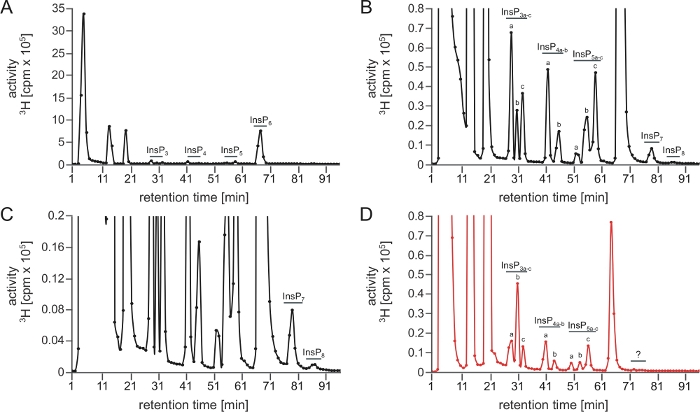
Figure 1: Typical InsP profile of a successful and of an unsuccessful SAX-HPLC analysis performed with this protocol. (A‒C) SAX-HPLC profile of 17-day-old wild-type (Col-0) Arabidopsis seedlings radiolabeled with [3H]-myo-inositol. Global InsP extraction and SAX-HPLC run were performed on the same day. (A) Full spectra; (B, C) Zoom-ins of the profile shown in A. All visible peaks are highlighted and assigned to the corresponding InsP species. Based on published chromatographic mobilities5,7, InsP4a likely represents Ins(1,4,5,6)P4 or Ins(3,4,5,6)P4, InsP5a represents InsP5 [2-OH], InsP5b represents InsP5 [4-OH] or its enantiomeric form InsP5 [6-OH], and InsP5c represents InsP5 [1-OH] or its enantiomeric form InsP5 [3-OH]. The isomeric natures of InsP3a-c, InsP4b, InsP7, and InsP8 are still unknown. Panel (D) shows a SAX-HPLC profile of identically grown plants but using an aged column (>40 runs). A clear reduction of InsP6 compared to other InsP species and the absence of PP-InsPs is visible. Please click here to view a larger version of this figure.
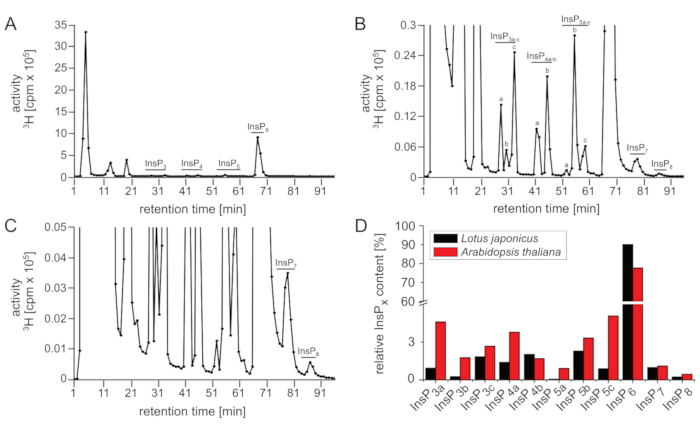
Figure 2: Representative InsP profile of L. japonicus plants. SAX-HPLC profile (A‒C) of 17-day-old wild-type (Gifu) L. japonicus seedlings radiolabeled with [3H]-myo-inositol. (A) Full spectra; (B, C) Zoom-ins of the profile shown in A. All visible peaks are highlighted and assigned to the corresponding InsP species. Based on published chromatographic mobilities5,7, InsP4a likely represents Ins(1,4,5,6)P4 or Ins(3,4,5,6)P4, InsP5b likely represents InsP5 [4-OH] or its enantiomeric form InsP5 [6-OH], and InsP5c likely represents InsP5 [1-OH] or its enantiomeric form InsP5 [3-OH]. The isomeric natures of InsP3a-c, InsP4b, InsP7, and InsP8 are unknown. (D) Comparison between the individual InsP species (in % of total activity from elution 25‒96) of A. thaliana (data from Figure 1A‒C) and L. japonicus (data from Figure 2A–C). Please click here to view a larger version of this figure.
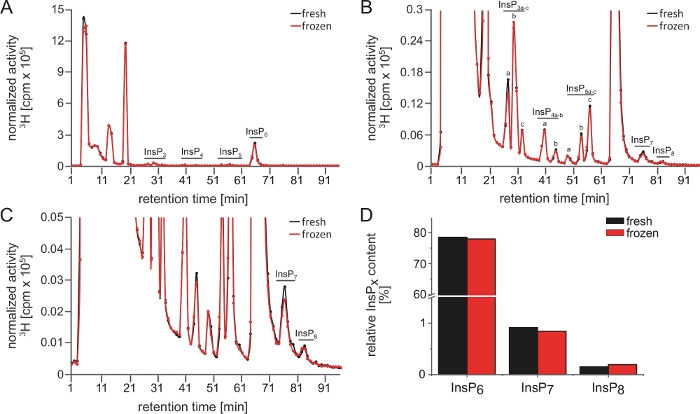
Figure 3: InsP profiles of a split sample illustrating the reproducibility of SAX-HPLC analyses. (A‒C) SAX-HPLC profiles of 17-day-old wild-type (Col-0) Arabidopsis seedlings radiolabeled with [3H]-myo-inositol. Prior the run, the sample was split and one half run immediately and the other half one day later after storage at -80 °C. (A) Full spectra; (B, C) Zoom-ins of the profile shown in A. All visible peaks are highlighted and assigned to the corresponding InsP species. Based on published chromatographic mobilities5,7, InsP4a likely represents Ins(1,4,5,6)P4 or Ins(3,4,5,6)P4, InsP5a represents InsP5 [2-OH], InsP5a represents InsP5 [2-OH], InsP5b represents InsP5 [4-OH] or its enantiomeric form InsP5 [6-OH], and InsP5c represents InsP5 [1-OH] or its enantiomeric form InsP5 [3-OH]. The isomeric natures of InsP3a-c, InsP4b, InsP7, and InsP8 are still unknown. Panel D shows the quantification of InsP6 and the PP-InsPs InsP7 and InsP8 of both runs. The values represent the amount (in %) of the respective InsP species relative to all InsP (total activity from elution 25–96). Please click here to view a larger version of this figure.
Supplemental Figure 1: Software settings for liquid scintillation counting using a light scintillation counter. Screenshots showing the software version, as well as settings used for scintillation counting of [3H] samples performed with this protocol are depicted. Please click here to download this figure.
Supplemental Figure 2: Representative example of data normalization. A screenshot of a worksheet shows all steps and formulas used to normalize SAX-HPLC runs to each other. For simplification, only fractions 25–35 of samples are shown. Please click here to download this figure.
Supplemental Figure 3: Peak determination, background subtraction and integration using analysis software. (A) Data from SAX-HPLC analysis is loaded into the software (minutes 28–96) and the peak analyzer tool is selected. (B‒E) The baseline is defined manually by setting points between individual peaks and the background is subtracted. (F) Peaks are determined manually based on appearance and published chromatographic mobilities5,7. (G) Peak ranges are defined manually by cpm values. (H) Peaks are integrated and calculated as % of all peaks. Please click here to download this figure.
Discussion
Here we present a versatile and sensitive method to quantify InsPs including PP-InsPs in plant extracts and provide practical tips on how to get this method established. Even though the protocol is generally robust, suboptimal runs and analyses can occur. In most cases, those runs can be identified by a strong reduction or even complete loss of highly phosphorylated InsPs, especially the PP-InsP species InsP7 and InsP8. Possible reasons can be microbial contaminations of the plant material and insufficient deactivation of endogenous plant PP-InsP hydrolases during extraction due to insufficient grinding and thawing of plant material that will not be in immediate contact with extraction buffer. Further reasons include inaccurate pH adjustment by insufficient or excess addition of neutralization buffer, or simply insufficient sample material. The latter can make it difficult to detect PP-InsPs, since those are often present in very low amounts in the cells. An excess of sample material or inefficient drying during step 3.5 may cause dilution of the perchloric acid, therefore also leading to insufficient enzyme deactivation and a specific loss of InsP6 and PP-InsPs. The amount of plant material, as well as radiolabel used in this protocol were optimized based on costs and performance, and is therefore close to the lowest amount that is still sufficient for providing optimal results. In addition, the column resin will gradually loose its resolution capacity. The first sign of this process is (for reasons not entirely clear to the authors) a specific loss of higher phosphorylated InsP species like the PP-InsPs in the HPLC spectrum. With further aging, even InsP6 will not be resolved properly by the column (Figure 1D). Therefore, the use of an adequate column, as well as meticulous handling of the sample and proper maintenance of the HPLC components is crucial for ensuring accurate results.
When comparing samples and runs, especially when generated with different equipment (e.g., HPLC systems and columns) or on different days, it is crucial to normalize the samples to each other (as described in step 7.3) and to analyze them in the same way. Only through normalization it is possible and accurate to show multiple samples in the same graph (Figure 3). For quantification of individual InsPs relative to total InsPs, or to another specific InsP species, it is not necessary to normalize, as long as only relative values and not absolute values are shown. Ideally, both the InsP profiles and the quantifications are shown. However, in some cases it is not possible to adequately show two or more runs in the same graph. Different retention times or different levels of background activity can make it difficult to compare unquantified SAX-HPLC profiles alone. The same is the true when many samples need to be compared. In such cases, a further evaluation using an additional software (e.g., Origin) for individual peak quantification is necessary.
The authors are aware that the protocol described here can be optimized and needs to be adapted to each individual research question. Although being optimized for Arabidopsis extracts7,17 in this protocol, this method is versatile and can help determining InsP profiles of other plant species as well. Here we exemplify this possibility by presenting for the first time a InsP profile for L. japonicus, which required no modifications of the labeling conditions, InsP extraction or SAX-HPLC run (Figure 2). Notably, while overall similar, differences are observed between L. japonicus and Arabidopsis InsP profiles. For instance, in L. japonicus InsP5 [4-OH] or its enantiomeric form InsP5 [6-OH] are more abundant than InsP5 [1-OH] or its enantiomeric form InsP5 [3-OH] in comparison to Arabidopsis, where InsP5 [1-OH] or its enantiomeric form InsP5 [3-OH] are the dominant InsP5 species. Likewise, we anticipate that alterations in the media composition, [3H]-myo-inositol concentration, plant age, environmental conditions (e.g., light and temperature), addition of chemical compounds or analyses of plant-microbial interactions among other factors, might need to be tested and adapted.
One important drawback of this method that needs to be considered is that the labeling is done in a (sterile) liquid culture, which does not represent a physiological environment for most land plants. In addition, due to the high costs of [³H]-myo-inositol, the volume of the labeling solution and the size of the culture vessel is generally limited, which also restricts the size of the plants that can be used. Cultivation in liquid culture can be avoided by directly infiltrating for instance leaves of soil-grown plants with [³H]-myo-inositol and subsequently following the protocol described here, as previously reported10.
There are several drawbacks of this protocol in comparison to alternative methods, such as TiO2 pull-down followed by PAGE or mass spectrometry based techniques. Due to the [3H]-myo-inositol labeling, only InsP species that directly originate from radiolabeled myo-inositol will be detected in the end. The method described here is blind to other Ins isomers such as scyllo-inositol and other isomers some of which have been identified in certain plants44. Furthermore, myo-InsPs derived from other pathways will be excluded, including those synthesized by de novo synthesis of myo-inositol and myo-inositol-3-phosphate via isomerization of glucose-6-phosphate, catalyzed by myo-inositol-3-phosphate synthase (MIPS) proteins45. Although [32P] or [33P]-ortho-phosphate can be used as alternative labels, their use poses a major disadvantage, since every phosphate-containing molecule, including the abundant nucleotides and its derivatives, will be labeled. Those molecules can also be extracted with this protocol and bind to the SAX column, which will result in a high level of background activity that will interfere with the identification of individual InsP peaks5. In addition, quantification of [32P]- or [33P] -labeled InsPs and PP-InsPs can be strongly influenced by phosphate and pyrophosphate moiety turnover and might not report a mass readout for inositol species.
On the other hand, [3H]-myo-inositol specifically labels myo-inositol-containing molecules. InsPs, inositol-containing lipids, such as phosphoinositides, and galactinol are in this case labeled. However, only InsPs will be analyzed with this protocol, since lipids are insoluble in the extraction buffer and galactinol does not bind to the SAX column.
So far, the differences from a plant InsP profile generated by [3H]-myo-inositol labeling compared with one determined by TiO2 pulldown/PAGE remains unknown, since such comparisons have not been performed in plants. A recent study in animal cells addressed this question46. In that work, a pool of InsP6 that is invisible by [3H]-myo-inositol labeling, which should thereby be directly derived from glucose-6-phosphate, was identified by comparing SAX-HPLC profiles with PAGE gels of mammalian cell lines. 24 h of phosphate starvation resulted in a 150% increase of InsP6 when quantifying PAGE gels of InsPs purified using TiO2 pulldown. SAX-HPLC analyses of [3H]-myo-inositol-labeled cells that were treated identically only showed an increase by 15% of [3H]-InsP6. As previously mentioned, InsPs lower than InsP5 are undetectable with PAGE analysis in most cases. Radiolabeling followed by SAX-HPLC appears to be the method of choice, as long as mass spectrometric protocols are not optimized to detect this group of highly negatively charged molecules.
Another remaining challenge is to distinguish enantiomers in SAX-HPLC analyses (or in any other method for InsP analysis)10,17. This challenge can be tackled by the addition of chiral selectors, i.e., enantiopure compounds like L-arginine amide that interact with the respective enantiomeric molecules to form diastereomeric complexes that can be separated10. To our knowledge, this approach has only been implemented to discriminate the enantiomeric InsP5 isomers InsP5 [1-OH] and InsP5 [3-OH] by NMR analyses10. Discrimination of other enantiomeric pairs or successful discrimination of enantiomers by chiral SAX-HPLC analysis or chiral PAGE-based methods have not yet been reported and should be further developed. Considering the conserved synthesis and the conserved regulation of PP-InsPs by phosphorous availability, we envision that especially non-radioactive methods such as PAGE or MS-based methods, together with nutrient analyses, will help ground truthing efforts to calibrate remote sensing data designed to diagnose nutrient deficiencies in crops17,18,24,25. However, the method presented here can currently still be considered the gold standard for InsP analyses and will be instrumental to discover new functions of these intriguing messengers in plants.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany’s Excellence Strategy – EXC-2070 – 390732324 (PhenoRob), the Research Training Group GRK2064 and individual research grants SCHA1274/4-1 and SCHA1274/5-1 to G.S.. We also thank Li Schlüter and Brigitte Ueberbach for technical assistance.
Materials
| Centrifuge | Eppendorf AG | model: 5430 R | |
| Diammonium hydrogen phosphate, ≥97 % | Carl Roth GmbH + Co. KG | 0268.1 | |
| Ethylenediamine tetraacetic acid disodium salt dihydrate, ≥99 %, p.a., ACS | Carl Roth GmbH + Co. KG | 8043.2 | |
| Falcon 12-well clear flat bottom TC-treated multiwell cell culture plate, with lid, individually wrapped, sterile | Corning Inc. | 353043 | |
| Fraction collector | LAMBDA Instruments GmbH | model: OMNICOLL single channel collector | |
| Growth incubator | poly klima GmbH | model: PK 520-LED | |
| HPLC pumps | Kontron Instruments | model: 420 | |
| HPLC syringe for Rheodyne valves, 1 mL | Hamilton Company | 81365 | |
| Injector for HPLC | Supelco | model: Rheodyne 9725 | |
| Inositol, myo-[1,2-3H(N)] | American Radiolabeled Chemicals Inc. | ART 0261 | |
| Liquid nitrogen | University, Chemistry Department | ||
| Liquid scintillation counter | PerkinElmer Inc | model: TRI-CARB 2900TR | |
| Micro pestle | Carl Roth GmbH + Co. KG | CXH7.1 | |
| Mixed cellulose Eester filter, ME range (ME 24), plain, 0.2 µm pore size, 47 mm circle | GE Healthcare Life Sciences | 10401770 | |
| Mixer for HPLC | Kontron Instruments | model: M 800 | |
| Murashige & Skoog medium, salt mixture | Duchefa Biochemie | M0221 | |
| OriginPro software | OriginLab Corp. | ||
| Orthophosphoric acid, ≥85 %, p.a., ISO | Carl Roth GmbH + Co. KG | 6366.1 | |
| Partisphere 5 µm SAX cartridge column, 125 x 4.6 mm | Hichrom Limited | 4621-0505 | |
| Perchloric acid, 70 %, 99.999 % trace metals basis | Sigma-Aldrich | 311421 | |
| Petri dish, square, PS, clear, 120/120/17 mm, sterile | Greiner Bio One International GmbH | 688161 | |
| pH-indicator paper pH 5.5 – 9.0, Neutralit | Merck KGaA | 109564 | |
| Phytagel | Sigma-Aldrich | P8169 | |
| Potassium carbonate | Carl Roth GmbH + Co. KG | P743.2 | |
| Safe-Lock tubes, 1.5 mL | Eppendorf AG | 30120086 | |
| Sample loop for 9725 injectors, volume 2 mL, PEEK | Supelco | 57648 | |
| SNAPTWIST scintillation vial, 6.5 mL | Simport Scientific Inc. | S207-5 | |
| Sterile bench | LaboGene | model: ScanLaf MARS 900 | |
| Sucrose, ≥99,5 %, p.a. | Carl Roth GmbH + Co. KG | 4621.1 | |
| Ultima-Flo AP liquid scintillation cocktail | PerkinElmer Inc | 6013599 | |
| Ultra-pure deionized water | Milli-Q | ||
| Wrenchless WVS End Fitting Kit | Hichrom Limited | 4631-1001 |
References
- Berridge, M. J., Irvine, R. F. Inositol Phosphates and Cell Signaling. Nature. 341 (6239), 197-205 (1989).
- Streb, H., Irvine, R. F., Berridge, M. J., Schulz, I. Release of Ca-2+ from a Nonmitochondrial Intracellular Store in Pancreatic Acinar-Cells by Inositol-1,4,5-Trisphosphate. Nature. 306 (5938), 67-69 (1983).
- Krinke, O., Novotna, Z., Valentova, O., Martinec, J. Inositol trisphosphate receptor in higher plants: is it real. Journal of Experimental Botany. 58 (3), 361-376 (2007).
- Kuo, H. F., et al. Arabidopsis inositol pentakisphosphate 2-kinase, AtIPK1, is required for growth and modulates phosphate homeostasis at the transcriptional level. Plant Journal. 80 (3), 503-515 (2014).
- Kuo, H. F., et al. Arabidopsis inositol phosphate kinases IPK1 and ITPK1 constitute a metabolic pathway in maintaining phosphate homeostasis. Plant Journal. 95 (4), 613-630 (2018).
- Gillaspy, G. E., Capelluto, D. G. S. . Lipid-mediated Protein Signaling. , 141-157 (2013).
- Stevenson-Paulik, J., Bastidas, R. J., Chiou, S. T., Frye, R. A., York, J. D. Generation of phytate-free seeds in Arabidopsis through disruption of inositol polyphosphate kinases. Proceedings of the National Academy of Sciences of the United States of America. 102 (35), 12612-12617 (2005).
- Lemtiri-Chlieh, F., MacRobbie, E. A. C., Brearley, C. A. Inositol hexakisphosphate is a physiological signal regulating the K+-inward rectifying conductance in guard cells. Proceedings of the National Academy of Sciences. 97 (15), 8687-8692 (2000).
- Lee, H. S., et al. InsP6-sensitive variants of the Gle1 mRNA export factor rescue growth and fertility defects of the ipk1 low-phytic-acid mutation in Arabidopsis. Plant Cell. 27 (2), 417-431 (2015).
- Blüher, D., et al. A 1-phytase type III effector interferes with plant hormone signaling. Nature Communications. 8, (2017).
- Murphy, A. M., Otto, B., Brearley, C. A., Carr, J. P., Hanke, D. E. A role for inositol hexakisphosphate in the maintenance of basal resistance to plant pathogens. Plant Journal. 56 (4), 638-652 (2008).
- Poon, J. S. Y., Le Fevre, R. E., Carr, J. P., Hanke, D. E., Murphy, A. M. Inositol hexakisphosphate biosynthesis underpins PAMP-triggered immunity to Pseudomonas syringae pv. tomato in Arabidopsis thaliana but is dispensable for establishment of systemic acquired resistance. Molecular Plant Pathology. 21 (3), 376-387 (2020).
- Wild, R., et al. Control of eukaryotic phosphate homeostasis by inositol polyphosphate sensor domains. Science. 352 (6288), 986-990 (2016).
- Shears, S. B. Inositol pyrophosphates: Why so many phosphates. Advances in Biological Regulation. 57, 203-216 (2015).
- Shears, S. B. Intimate connections: Inositol pyrophosphates at the interface of metabolic regulation and cell signaling. Journal of Cellular Physiology. 233 (3), 1897-1912 (2018).
- Desai, M., et al. Two inositol hexakisphosphate kinases drive inositol pyrophosphate synthesis in plants. Plant Journal. 80 (4), 642-653 (2014).
- Laha, D., et al. VIH2 Regulates the Synthesis of Inositol Pyrophosphate InsP8 and Jasmonate-Dependent Defenses in Arabidopsis. Plant Cell. 27 (4), 1082-1097 (2015).
- Laha, D., et al. Arabidopsis ITPK1 and ITPK2 Have an Evolutionarily Conserved Phytic Acid Kinase Activity. Acs Chemical Biology. 14 (10), 2127-2133 (2019).
- Dorsch, J. A., et al. Seed phosphorus and inositol phosphate phenotype of barley low phytic acid genotypes. Phytochemistry. 62 (5), 691-706 (2003).
- Flores, S., Smart, C. C. Abscisic acid-induced changes in inositol metabolism in Spirodela polyrrhiza. Planta. 211 (6), 823-832 (2000).
- Brearley, C. A., Hanke, D. E. Inositol phosphates in barley (Hordeum vulgare L) aleurone tissue are stereochemically similar to the products of breakdown of InsP(6) in vitro by wheat-bran phytase. Biochemical Journal. 318, 279-286 (1996).
- Laha, N. P., et al. ITPK1-Dependent Inositol Polyphosphates Regulate Auxin Responses in Arabidopsis thaliana. bioRxiv. , (2020).
- Laha, D., et al. Inositol Polyphosphate Binding Specificity of the Jasmonate Receptor Complex. Plant Physiology. 171 (4), 2364-2370 (2016).
- Dong, J. S., et al. Inositol Pyrophosphate InsP(8) Acts as an Intracellular Phosphate Signal in Arabidopsis. Molecular Plant. 12 (11), 1463-1473 (2019).
- Zhu, J., et al. Two bifunctional inositol pyrophosphate kinases/phosphatases control plant phosphate homeostasis. Elife. 8, (2019).
- Couso, I., et al. Synergism between Inositol Polyphosphates and TOR Kinase Signaling in Nutrient Sensing, Growth Control, and Lipid Metabolism in Chlamydomonas. Plant Cell. 28 (9), 2026-2042 (2016).
- Ito, M., et al. Hydrophilic interaction liquid chromatography-tandem mass spectrometry for the quantitative analysis of mammalian-derived inositol poly/pyrophosphates. Journal of Chromatography A. 1573, 87-97 (2018).
- Wilson, M. S. C., Saiardi, A. Inositol Phosphates Purification Using Titanium Dioxide Beads. Bio-Protocol. 8 (15), (2018).
- Loss, O., Azevedo, C., Szijgyarto, Z., Bosch, D., Saiardi, A. Preparation of Quality Inositol Pyrophosphates. Jove-Journal of Visualized Experiments. (55), e3027 (2011).
- Harmel, R. K., et al. Harnessing C-13-labeled myo-inositol to interrogate inositol phosphate messengers by NMR Electronic supplementary information (ESI) available. Chemical Science. 10 (20), 5267-5274 (2019).
- Azevedo, C., Saiardi, A. Extraction and analysis of soluble inositol polyphosphates from yeast. Nature Protocols. 1 (5), 2416-2422 (2006).
- Shears, S. B., Miller, G. J. . Inositol Phosphates: Methods and Protocols. , 1-28 (2020).
- Wilson, M. S. C., Saiardi, A. Importance of Radioactive Labelling to Elucidate Inositol Polyphosphate Signalling. Topics in Current Chemistry. 375 (1), (2017).
- Stevenson-Paulik, J., et al. Inositol phosphate metabolomics: Merging genetic perturbation with modernized radiolabeling methods. Methods. 39 (2), 112-121 (2006).
- Liu, C., Riley, A. M., Yang, X., Shears, S. B., Potter, B. V. L. Synthesis and Biological Activity of d- and l-chiro-Inositol 2,3,4,5-Tetrakisphosphate: Design of a Novel and Potent Inhibitor of Ins(3,4,5,6)P4 1-Kinase/Ins(1,3,4)P3 5/6-Kinase. Journal of Medicinal Chemistry. 44 (18), 2984-2989 (2001).
- Hughes, P. J., Hughes, A. R., Putney, J. W., Shears, S. B. The regulation of the phosphorylation of inositol 1,3,4-trisphosphate in cell-free preparations and its relevance to the formation of inositol 1,3,4,6-tetrakisphosphate in agonist-stimulated rat parotid acinar cells. Journal of Biological Chemistry. 264 (33), 19871-19878 (1989).
- Shears, S. B., Kirk, C. J., Michell, R. H. The pathway of myo-inositol 1,3,4-trisphosphate dephosphorylation in liver. The Biochemical journal. 248 (3), 977-980 (1987).
- Stevenson-Paulik, J., Odom, A. R., York, J. D. Molecular and Biochemical Characterization of Two Plant Inositol Polyphosphate 6-/3-/5-Kinases. Journal of Biological Chemistry. 277 (45), 42711-42718 (2002).
- Stephens, L. R., Hawkins, P. T., Downes, C. P. An analysis of myo-[3H]inositol trisphosphates found in myo-[3H]inositol prelabelled avian erythrocytes. The Biochemical journal. 262 (3), 727-737 (1989).
- Saiardi, A., et al. Mammalian inositol polyphosphate multikinase synthesizes inositol 1,4,5-trisphosphate and an inositol pyrophosphate. Proceedings of the National Academy of Sciences. 98 (5), 2306 (2001).
- Azevedo, C., Burton, A., Bennett, M., Onnebo, S. M. N., Saiardi, A., Barker, C. J. . Inositol Phosphates and Lipids: Methods and Protocols. , 73-85 (2010).
- Saiardi, A., Caffrey, J. J., Snyder, S. H., Shears, S. B. The Inositol Hexakisphosphate Kinase Family: CATALYTIC FLEXIBILITY AND FUNCTION IN YEAST VACUOLE BIOGENESIS. Journal of Biological Chemistry. 275 (32), 24686-24692 (2000).
- Brearley, C. A., Hanke, D. E. Inositol phosphates in the duckweed Spirodela polyrhiza L. Biochemical Journal. 314, 215-225 (1996).
- Pollard, J. K., Steward, F. C., Shantz, E. M. Hexitols in Coconut Milk – Their Role in Nurture of Dividing Cells. Plant Physiology. 36 (4), 492 (1961).
- Donahue, J. L., et al. The Arabidopsis thaliana Myo-inositol 1-phosphate synthase1 gene is required for Myo-inositol synthesis and suppression of cell death. Plant Cell. 22 (3), 888-903 (2010).
- Desfougeres, Y., Wilson, M. S. C., Laha, D., Miller, G. J., Saiardi, A. ITPK1 mediates the lipid-independent synthesis of inositol phosphates controlled by metabolism. Proceedings of the National Academy of Sciences of the United States of America. 116 (49), 24551-24561 (2019).

