Estimation of Crystalline Cellulose Content of Plant Biomass using the Updegraff Method
Summary
The Updegraff method is the most widely used method for the cellulose estimation. The main purpose of this demonstration is to provide a detailed Updegraff protocol for estimation of cellulose content in plant biomass samples.
Abstract
Cellulose is the most abundant polymer on Earth generated by photosynthesis and the main load-bearing component of cell walls. The cell wall plays a significant role in plant growth and development by providing strength, rigidity, rate and direction of cell growth, cell shape maintenance, and protection from biotic and abiotic stressors. The cell wall is primarily composed of cellulose, lignin, hemicellulose and pectin. Recently plant cell walls have been targeted for the second-generation biofuel and bioenergy production. Specifically, the cellulose component of the plant cell wall is used for the production of cellulosic ethanol. Estimation of cellulose content of biomass is critical for fundamental and applied cell wall research. The Updegraff method is simple, robust, and the most widely used method for the estimation of crystalline cellulose content of plant biomass. The alcohol insoluble crude cell wall fraction upon treatment with Updegraff reagent eliminates the hemicellulose and lignin fractions. Later, the Updegraff reagent resistant cellulose fraction is subjected to sulfuric acid treatment to hydrolyze the cellulose homopolymer into monomeric glucose units. A regression line is developed using various concentrations of glucose and used to estimate the amount of the glucose released upon cellulose hydrolysis in the experimental samples. Finally, the cellulose content is estimated based on the amount of glucose monomers by colorimetric anthrone assay.
Introduction
Cellulose is the primary load-bearing component of cell walls, which is present in both primary and secondary cell walls. The cell wall is an extracellular matrix that surrounds plant cells and is primarily composed of cellulose, lignin, hemicellulose, pectin, and matrix proteins. Approximately one third of plants biomass is cellulose1 and it plays significant roles in plant growth and development by providing strength, rigidity, rate and direction of cell growth, cell shape maintenance, and protection from biotic and abiotic stressors. Cotton fiber contains 95% cellulose2 content, while trees contain 40% to 50% of cellulose depending on the plant species and organ types3. The cellulose is composed of repeating units of cellobiose, a disaccharide of glucose residues connected by β-1,4 glycosidic bonds4. Cellulosic ethanol is produced from the glucose derived from the cellulose present in the plant cell walls5. Cellulosic fiber is made up of several micro fibrils in which each micro fibril acts as core unit with 500-15000 glucose monomers1,6. The cellulose homopolymer is synthesized by plasma membrane embedded cellulose synthase complexes (CSC's)1,7. Individual cellulose synthase A (CESA) proteins synthesize glucan chains and the adjacent glucan chains are connected by hydrogen bonds to form crystalline cellulose1,8. Cellulose exists in several crystalline forms with two predominant forms, cellulose Iα and cellulose Iβ as native forms9. In higher plants, cellulose exists in cellulose Iβ form while lower plant cellulose exists in Iα form10,11. Overall, the cellulose plays a significant role in imparting strength and rigidity to the plant cell walls.
First generation biofuels are primarily produced from corn starch, cane sugars, and beet sugars, which are food sources, while second-generation biofuels are focusing on the biofuel production from non-food plant biomass cell wall material12. Accurate estimation of crystalline cellulose content is not only important for fundamental research on cellulose biosynthesis and cell wall dynamics but also for applied biofuel and bio products research. Various methods have been developed and optimized for estimation of cellulose in the plant biomass, and the Updegraff method is the most widely used method for cellulose estimation. The first reported method for cellulose estimation was by Cross and Bevan in 190813. The method was based on the principle of alternate chlorination and extraction by sodium sulphate. However, the cellulose obtained by the original as well as modified protocols of Cross and Bevan method showed contamination of small fractions of lignin in addition to a substantial amount of xylans and mannans14. Despite several modifications to remove lignin and hemicelluloses from the cellulose fraction, the Cross-Bevan method retained a considerable amount of mannans along with cellulose. Later, Kurschner's method was developed by employing nitric acid and ethanol to extract cellulose15. This method stated that total lignin and 75% of pentosans were removed but the true cellulose results were the same as those estimated by chlorination method of Cross and Bevan. Another method (Norman and Jenkins) was developed by employing methanol-benzene, sodium sulphate, and sodium hypochlorite to extract cellulose16. This method also retained some fraction of lignin (3%) and significant amounts of pentosans leading to in accurate estimation of cellulose. Later, Kiesel and Semiganowsky used a different approach to hydrolyze cellulose using 80% concentrated sulfuric acid, and the hydrolyzed reduced sugars were estimated by Bertrand's method17. The two methods, Waksman's and Stevens18 and Salo14,19 which were developed based on Kiesel and Semiganowsky's method, also yielded 4-5% less cellulose content compared to earlier methods20.
The Updegraff method is the most widely used method for the estimation of crystalline cellulose content. This method was first described by Updegraff for the measurement of cellulose in 196921. The Updegraff method integrates the Kurschner method (use of nitric acid), Kiesel and Seminowsky methods (hydrolysis of cellulose into glucose monomers using sulfuric acid) with some modifications, and the anthrone assay of Viles and Silverman for simple colorimetric estimation of glucose and crystalline cellulose content22. The principle of this method is the use of acetic acid and nitric acid (Updegraff reagent) to eliminate hemicellulose and lignin from the homogenized plant tissues, which leaves acetic/nitric acid resistant cellulose for further processing and estimation15. The acetic/nitric acid resistant cellulose is treated with 67% sulfuric acid to break the cellulose into glucose monomers and the released glucose monomers are estimated by anthrone assay21,23. Several modifications of the original Updegraff method were used to simplify the procedure and cellulose estimation by anthrone assay24. Broadly, this method can be divided into five phases. In the first phase, the plant material is prepared. In the second phase, the crude cell wall is separated from the total biomass, as cellulose is the key component of plant cell walls. Later, in the third phase, the cellulose is separated from the non-cellulosic cell wall components by treating with Updegraff reagent. In the fourth phase, the acetic/nitric acid resistant cellulose is broken into glucose monomers by sulfuric acid treatment. Sulfuric acid treatment of cellulose results in formation of 5-hydroxymethylfurfural compounds from the reaction of glucose monomers with sulfuric acid. Finally, in the last phase, the anthrone generates a greenish blue complex by boiling with the furfural compound generated in the previous phase25. This anthrone based colorimetric method was first used in 1942 by Dreywood. Anthrone is a dye that identifies furfural compounds of pentose and hexose dehydrated products such as 5-hydroxymethylfurfural, under acidic conditions. Reaction with hexose produces an intense color and better response compared to pentoses25. The amount of bound glucose is measured by spectrophotometer absorbance at 620 nm and the intensity of the greenish blue complex is directly proportional to the amount of sugar in the sample. The measured absorbance values were compared with a glucose standard curve regression line to calculate the glucose concentration of the sample. The measured glucose content was used to estimate the cellulose content of the plant biomass.
Protocol
1. Experimental preparation
- Grind dried plant material into a fine powder.
- Protein Solubilization Buffer (PSB): Prepare stock solutions of 1 M Tris (pH 8.8), 0.5 M ethylenediaminetetraacetic acid (EDTA) (pH 8.0) and autoclave them. Make fresh PSB buffer from these stock solutions with final concentrations of 50 mM Tris, 0.5 mM EDTA and 10% sodium dodecyl sulfate (SDS) in sterile water.
- Prepare 100 mL of 70% ethanol (v/v): 70 mL of 100% ethanol and 30 mL of sterile water.
- Prepare 100 mL of methanol: chloroform in a 1:1 ratio (50 mL methanol and 50 mL chloroform).
- Prepare 82.5 mL of Updegraff reagent. Add 75 mL of 80% acetic acid to 7.5 mL of nitric acid so that the ultimate ratio of water: acetic acid: nitric acid is in 2:8:1 (v/v). To prepare 80% acetic acid, dissolve 80 mL of glacial acetic acid in 20 mL of sterile water (v/v).
- Prepare a fresh stock of 1 mg/mL glucose solution. Dissolve 10 mg of glucose in 10 mL of water (w/v).
- To prepare 100 mL of 67% sulfuric acid (v/v), add 67 mL of concentrated sulfuric acid to 33 mL of water. Always use a glass bottle and add acid slowly to the water. This step is exothermic (releases heat). Hence, prepare this solution on ice and cool in the refrigerator for at least 2 hours before use.
- Prepare fresh 0.2% anthrone (w/v) for each batch of experimental samples. Weigh 0.2 g of anthrone and dissolve in 100 mL of pre-chilled concentrated sulfuric acid in a glass bottle wrapped with aluminum foil. Keep in refrigerator for 1-2 hours before use.
NOTE: Pre-chilling concentrated sulfuric acid in the refrigerator on the day of the experiment, and fresh preparation of anthrone is highly recommended for accurate estimation of glucose content.
2. Preparation of plant biomass material
- Collect plant biomass samples from 2-month-old cotton experimental lines grown in the greenhouse with the same growth conditions, the same development stage, the same position of the plants and the same type of tissue (leaf/stem/root).
NOTE: Collect a minimum of three biological replicates for each sample. Wash them thoroughly with water to remove all the dirt from the root tissue. - Air-dry root tissue for 2 days on paper towels at room temperature to remove the moisture content (Figure 1).
NOTE: Air-dry desired tissue for cellulose estimation to prevent any fungal contamination. - Place the root samples into individual containers. Then label and dry them in the incubator at 49 °C for 10 days (Table of Materials).
NOTE: Alternatively, a freeze dryer can be used to dry the plant tissue in less time (1 or 2 days) without causing any chemical changes to the plant biomass material. - Cut the dried samples into small pieces, freeze them in liquid nitrogen, and grind into uniform fine powder by using a mortar and pestle, a freezer mill, or a biomass grinder (Figure 1).
NOTE: The freezer mill was used at a rate of 10 cps for 3 cycles. - Collect the grounded tissue (Figure 2) and proceed with the cell wall extraction.
NOTE: The process can be paused at this point by storing the samples in airtight containers at room temperature.
3. Extraction of crude cell walls from plant biomass
NOTE: The plant cell walls contain cellulose, lignin, non-cellulose components, pectin, matrix proteins, phenolic compounds, and water26. Since cellulose is present in cell walls, the first step is to separate cell wall component from non-cell wall components of the plant biomass26.
- Label and weigh individual blank 2 mL tubes before starting the cell wall extraction process.
- Make a note of empty tube weights in a lab notebook before proceeding further.
- Weigh 20 mg of powdered tissue from step 2.5 and transfer it to pre-weighed 2 mL tubes and label them.
- Add 1 mL of protein solubilization buffer (PSB) (50 mM Tris hydrochloride (HCl) buffer pH 8.8, 0.5 mM EDTA, and 10% sodium dodecyl sulfate (SDS) to solubilize proteins. Vortex and centrifuge at 25,200 x g for 5 min at room temperature (RT). After centrifugation, discard the supernatant and save the pellet.
NOTE: The supernatant can be saved if protein component needs to be analyzed. - Repeat step 3.4 two more times.
- To the retained pellet, add 1 mL of distilled water and vortex. Centrifuge at 25,200 x g for 5 min at room temperature (RT) and remove the supernatant.
- Repeat step 3.6 two more times.
- Add 1 mL of 70% ethanol to the saved pellet, vortex and heat it at 70 °C for 1 h in a water bath/heat block to remove soluble components and starch from the samples. Vortex and centrifuge at 25,200 x g for 5 min at RT. Discard the supernatant and save the pellet.
- Repeat step 3.8 one more time.
- To the pellet add 1 mL of 100% methanol and vortex. Centrifuge at 25,200 x g for 5 min at room temperature (RT) and remove the supernatant.
- Add 1 mL of chloroform/methanol (chloroform and methanol in 1:1 ratio) to the pellet and vortex. Centrifuge at 25,200 x g for 5 min at RT and remove the supernatant. The addition of methanol and chloroform solvent solubilizes and removes the lipid fraction from the biomass27.
- To the pellet, add 1 mL of 100% acetone and vortex. Incubate at room temperature for 5 min. Centrifuge at 25,200 x g for 5 min at RT and remove the supernatant. Acetone removes pigments such as chlorophyll and free fatty acids from the biomass28,29.
- Dry the pellet at 37 °C overnight or proceed further by vacuum drying.
NOTE: The dried pellet is the crude cell wall used for crystalline cellulose estimation. The process can be paused at this point or proceed further using vacuum drier for drying samples.
4. Treatment with Updegraff reagent (acetic and nitric acid) to remove non-cellulosic components
NOTE: The protocol involves use of acids and other chemicals. Wear personal protective equipment (PPE) throughout the process.
- Measure 5 mg of the dried crude cell wall pellet into a fresh 2 mL screw capped tube. Note the exact weight (Figure 3)30.
- Include a positive control at this point. Use 2 mg of filter paper (Table of Materials) as a positive control that yields 80% cellulose content.
- Add 1.5 mL of the Updegraff reagent to the weighed 5 mg of cell wall extract and positive control. Mix by vortexing.
NOTE: The positive control should be processed in the same way as the experimental samples from this step onwards. This step should be carried out in the fume hood with proper personal protective equipment (PPE).
CAUTION: This step should be carried out in screw cap tubes to prevent splashing of sample and popping of the tubes. Three biological replicates of each sample along with three replicates for positive control should be included. - Heat the suspension at 100 °C for 30 min in a boiling water bath and cool on a bench for 10 min. Centrifuge at 25,200 x g for 10 min at RT. Remove the supernatant by centrifugation and save the pellet.
NOTE: The waste generated should be collected separately for organic solvents, sulfuric acid, and Updegraff reagent (acetic acid and nitric acid). Waste with acetic acid and nitric acid should be kept at cool temperatures with ventilated caps and never mixed with any other organic solvents and other acids to prevent any explosion. - Add 1 mL of water to the pellet. Centrifuge at 25,200 x g for 10 min at RT. Remove 500 µL of the supernatant and add 1 mL of acetone to the tube.
- Centrifuge at 25,200 x g for 5 min at RT. Remove 1 mL of supernatant, and add 1 mL of acetone to the tube and centrifuge at 25,200 x g for 5 min at RT.
- After centrifugation, remove all the supernatant and suspend the pellet in 1 mL of acetone. Incubate the tubes at room temperature for 5 min and spin at 25,200 x g for 5 min at RT.
- Discard the supernatant and save the pellet. Dry the pellet at 37 °C overnight (Figure 3).
NOTE: The process can be paused at this point or continue further using vacuum drier for drying and proceed to next step.
5. Hydrolysis of cellulose by acid to produce glucose monomer units
- Add 1 mL of 67% sulfuric acid to the dried pellet. Vortex to mix the pellet completely in the acid.
- Shake tubes at room temperature for 1 h to dissolve the cellulose pellet in 67% sulfuric acid.
NOTE: As an alternative, solubilization of the cellulose pellet in 67% sulfuric acid can be improved by sonication of each sample for 10 min after 30 min of incubation in the shaker. After sonication, samples can be re-incubated in the shaker at RT. However, we observed complete solubility of cellulose pellet without sonication when we start with 5 mg of crude cell wall extract (Figure 3). This procedure worked well for various plant biomass samples3.
6. Measuring glucose content by the anthrone assay and estimation of cellulose content
- At this stage, the cellulose is in the form of free glucose monomers. Determine the amount of cellulose by measuring the amount of glucose present in the sample by spectrophotometry.
- Take 10 µL of the sample from the step 5 and add it to 490 µL of sterile distilled water to make a dilution of each sample to 500 µL. Vortex the diluted mixture of sample and water for 10 seconds.
- Add 1 mL of 0.2% freshly prepared anthrone reagent to each tube and mix immediately by vortexing.
- Boil samples at 100 °C for 10 min and cool the tubes on ice for 5 min. Transfer 200 µL of each sample in three wells of a 96 well plate. Load the 96 well plate into a spectrophotometer to measure the absorbance at 620 nm.
NOTE: For boiling at high temperature, use screw-capped tubes to prevent splashing of harmful chemicals and to avoid loss of samples.
7. Preparation of glucose standard curve
- To prepare glucose standard, make a fresh stock of 1 mg/mL glucose by dissolving 10 mg of glucose in 10 mL of sterile distilled water. Add this stock solution of 1 mg/mL in 20 µL increments (0, 20, 40, 60, 80,100,120,140, 160, 180 µL) to sterile distilled water (total volume 1 mL) to prepare glucose standards ranging from 0 µg to 180 µg/mL concentrations (Figure 3). Vortex each glucose standard after adding sterile water.
- From these 10 different concentrations, aliquot 500 µL of each glucose concentration into a fresh 2 mL screw capped tube.
- Add 1 mL of freshly prepared 0.2% anthrone to each tube and mix immediately by vortexing. Incubate on ice and boil at 100 °C for 10 min followed by incubation on ice for 5 min23.
- Transfer 200 µL of each standard to three wells in a 96 well micro titer plate for three technical replicates and measure the absorbance at 620 nm (Figure 3).
- Develop a standard glucose curve by plotting different glucose concentrations (0 to 180 µg/mL) against normalized absorbance values at 620 nm (Figure 7). Use the regression line Y= mx+c generated from the absorbance values to calculate the cellulose content in the prepared samples.
NOTE: Glucose standards must be freshly prepared for each set of experiment. The concentration of the glucose should be increased if the OD values are too high/ low so that these values fall within the range of the standard curve absorbance values. The regression line generates different m and c values based on the glucose standards used for each batch of samples. Mix tubes immediately after the addition of anthrone23. Anthrone, diluted samples and prepared standards were always kept cold until the anthrone assay was performed because of the exothermic nature of this reaction.
Representative Results
Cotton plants grown in the green house were selected for this study. Two different experimental lines of cotton were selected for comparative analysis of cellulose content. For each experimental line, the root tissue was collected from three biological replicates. A total of 500 mg of tissue was homogenized and 20 mg of it was used for crude cell wall extraction. Later, 5 mg of crude cell wall extract was used for Updegraff reagent treatment to remove hemicellulose and lignin from cellulose. The purified cellulose was hydrolyzed by sulfuric acid treatment to break down the cellulose into glucose units. The results of absorbance readings at 620 nm (Supplementary Table 1) were used to measure glucose concentration and estimate cellulose content by the anthrone assay. The average absorbance values of three technical replicates were normalized by subtracting the blank absorbance value of the 0 µg glucose concentration. The glucose standard curve was generated using a scattered bar graph (Figure 4).
The resultant regression line, y = mx + c from this standard curve, is used to calculate an unknown glucose concentration in the extracted test samples using the normalized absorbance readings. To calculate x (the unknown glucose concentration in µg/µL), the formula (x = normalized absorbance-c/m) was used. The c and m values from regression line of standard glucose curve (R2 = 0.99) were used to measure the glucose concentration in µg/µL. Then this value is divided by a dilution factor of 2 as the final volume is 500 µL and further divided by a 10 µL sample volume to measure the glucose concentration in µg/µL. Since the calculated value is for 5 mg of cell wall extract, it is further divided by 5 or with the respective crude cell wall weight used for each experimental sample to get glucose concentration per mg. The value is further divided by 1.1 to compensate for the water gained in the process of hydrolysis (glucose has a molecular weight of 180 and a water molecule has a molecular weight of 18 and hence the water generated in the process of releasing glucose monomers was removed by dividing the moisture factor, 1.1). The resultant value will be multiplied by 100 to calculate the crystalline cellulose content in percentage (Supplementary Table 1).
The results of the cellulose content show that they are consistent among the biological replicates, suggesting the technical accuracy of the Updegraff method. Further, the differences were plotted using 2D graphs. These results showed the cellulose content differences in the compared experimental sample numbers 1 and 2 (Figure 4). Sample 1 shows 43.4% while sample 2 shows 28.12%, suggesting that this method can distinguish differences between the experimental samples. A Student t-test was applied at 95% significance level to test their significant differences.
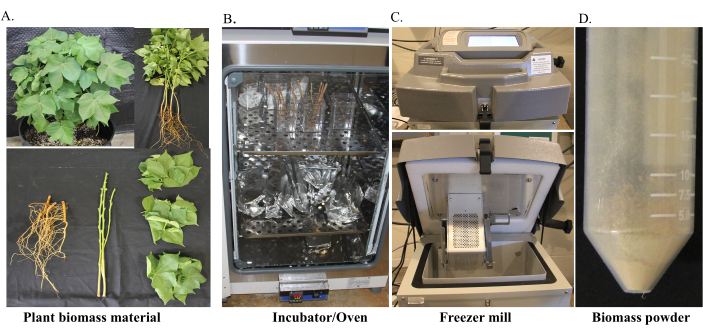

Figure 1: Preparation of cotton plant biomass for cellulose content estimation in roots. (A) Greenhouse grown plants were gently pulled from the soil and thoroughly washed with water. The plant root tissue was separated, kept on the paper towels and allowed to air dry for 2 days. (B) Then the root tissue was transferred to individually labeled containers and allowed to dry in the incubator at 49 °C for -10 days. (C) The dried tissue was cut into small pieces, frozen in liquid nitrogen, loaded in the grinding vials, and grounded into fine powder using freezer mill. (D) Finely grounded tissue powder was collected in the 50 mL tube. Please click here to view a larger version of this figure.
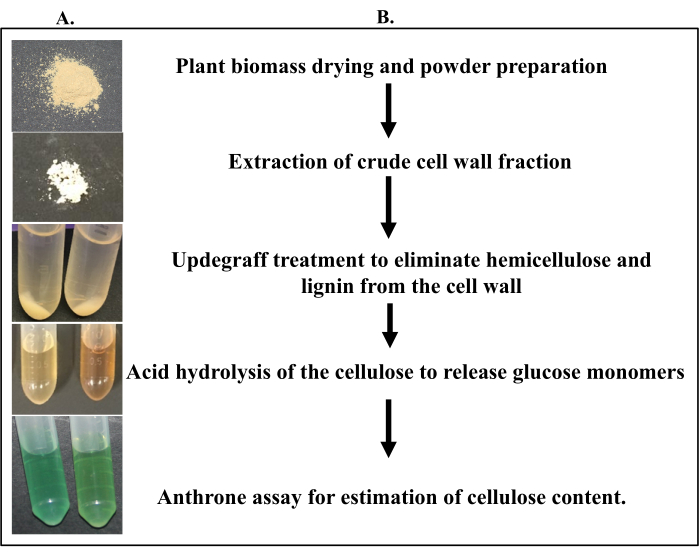
Figure 2: Flow chart of the main steps involved in the Updegraff method. (A) Pictorial representation of key steps in the protocol. (B) Explains critical steps that was shown in panel (A). Please click here to view a larger version of this figure.
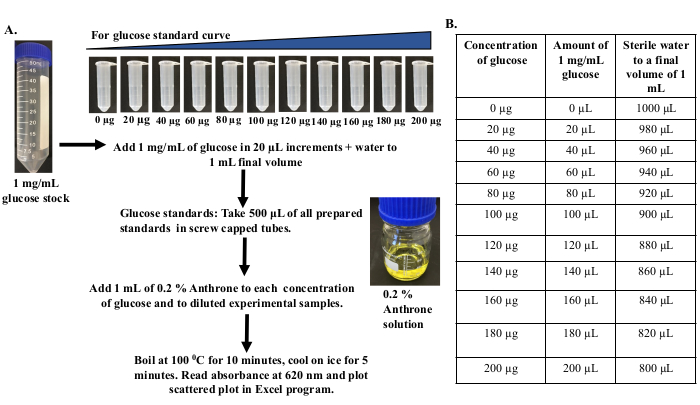
Figure 3: Preparation of different concentrations of glucose for the standard curve. (A) Prepare the stock solution of glucose by dissolving 10 mg of glucose in 10 mL of sterile distilled water so that the final concentration of the stock is 1 mg/mL. Add stock solution of 1 mg/mL glucose in 20 µL increments to a final volume of 1 mL with sterile distilled water to prepare different concentrations of glucose from 0 µg to 180 µg/mL. (B) Table showing amount of 1 mg/mL glucose and sterile distilled water to be added to make different concentrations of glucose. Please click here to view a larger version of this figure.
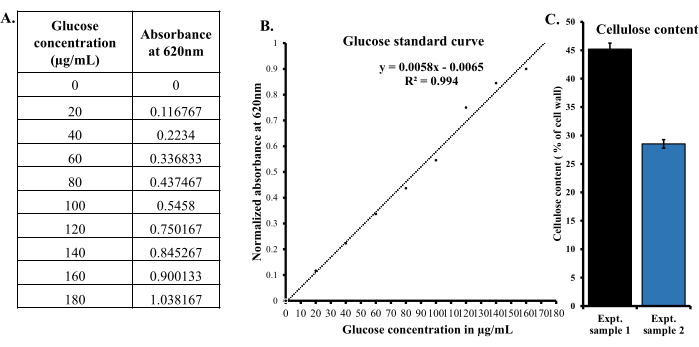
Figure 4: Glucose standard curve generated using absorbance values of different glucose concentrations at 620 nm. (A) Table showing different concentrations of glucose ranging from 0 to 180 µg/mL with the respective normalized absorbance values at 620 nm. (B) Glucose standard curve generated using a scatter chart from the absorbance values (A). (C) Bar graph showing crystalline cellulose content differences in the experimental samples 1 and 2. Please click here to view a larger version of this figure.
Supplementary Table 1: Estimation of cellulose content in the experimental samples. Please click here to download this Table.
Discussion
Cotton fibers are natural fibers produced from the cottonseed. Cotton fiber is a single cell with ~95% cellulose content2 with high crystalline cellulose content with extensive applications in textile industry31. As, cotton fiber contains ~95% cellulose, we have used cotton root tissues for demonstration of the estimation of crystalline cellulose content. Cotton root tissues are moderately rich in crystalline cellulose content and represents a commonly available plant biomass. The amount of total cell wall required for the crystalline cellulose content is only 5 mg and hence this method can be employed for estimating crystalline cellulose content in different developmental stages/tissue specific cellulose content estimation. The percentage of sulfuric acid (67%) also plays a critical role in breaking cellulose into glucose monomers and dissolving cellulose completely23 without a need for sonication. Sample loss is common after the Updegraff treatment as the crystalline cellulose pellet is not solid at the bottom of the tube at different steps in the estimation process. This can be minimized by removing small volumes of supernatant in each step of water and acetone washes after Updegraff treatment. Further, immediate vortexing of the sample and the standards with anthrone plays significant roles in color development and increased accuracy of the standards, leading to 99.5% coefficient of determination (R2) values (Figure 3). The glucose standard curve is key in determining the glucose concentration, which in turn is used to estimate the crystalline cellulose content. Hence, the concentrations of the glucose can be adjusted according to the required range of concentrations. Depending on the range of values to fit the experimental values, the standard curve can range from 0 to 100 µg/mL or 0 to 200 µg/mL up to 1000 µg/mL concentrations of glucose; this further changes the regression values c and m, improving the accuracy of crystalline cellulose estimation. It is important to always make fresh 0.2% anthrone for both samples and as well as for the standard. R-square, a statistical measurement, represents the variability of all data points on the regression line. It is important to boil both standards and samples after adding anthrone for the same amount of time. It is always important to include a positive control with a known cellulose content. For effective data reproducibility, the entire procedure must be followed accurately including solution preparation (standards, standard curve and Updegraff reagent).
Recently, cellulose is explored for non-textile applications such as cellulosic aerosols, computer parts and cellulosic ethanol32,33; hence, this method will have wide applications in the future. Scientific advancement is made through incremental progress in understanding concepts, proposing alternative hypotheses and providing/disproving the existing knowledge. Similar progress can be seen in the development of a reliable method for the estimation of cellulose content in the plant biomass. The cell wall is a complex matrix and the Updegraff method eliminates non-cellulose fractions from pure cellulose in a sequential manner, leading to accurate estimation of the cellulose content in the plant biomass. Further, the simple colorimetric method developed for the estimation of cellulose content by measuring the glucose concentration made this method highly adaptable and widely used. In the current study, the cellulose data was consistent among different biological replicates of the samples collected from a single line. Thus, it suggests that the data produced by this method is consistent, reliable, reproducible, and easy to measure the cellulose content by simple colorimetric assay.
The Updegraff method is the most widely used method for the cellulose estimation. With recent expansion of the industrial application of cellulose such as food, wood, paper, fibers, biofuels, clothes, cosmetic and pharmaceuticals, accurate estimation of cellulose content of the plant biomass is crucial for various applications. Though this is the most widely used method for cellulose estimation, it is important to take several precautions to obtain reproducible data. The biomass samples should be ground to the same size to obtain precise and reproducible estimation of the cellulose content. Care should be taken while generating the glucose standard curve. Glucose standards used for the glucose standard curve preparation should be increased or decreased based on the sample absorbance readings so that the unknown concentrations fall within the range of glucose concentrations that are used to prepare the standard curve. If not, m and c values might vary, resulting in the inaccurate estimation of crystalline cellulose content. Fresh preparation of 0.2 % anthrone for each batch of samples is another critical step of the Updegraff method. It is essential to use freshly prepared 0.2 % anthrone for the accurate estimation of the cellulose content.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank the Department of Plant & Soil Science and Cotton Inc. for their partial support of this study.
Materials
| Acetone | Fisher Chemical | A18-500 | Used in the protocol |
| Anthrone | Sigma Aldrich | 90-44-8 | For colorimetric assay |
| Centrifuge | Eppendorf | 5424 | For centrifugation |
| Chloroform | Mallinckrodt | 67-66-3 | Used in the protocol |
| Ethylenediaminetetraacetic acid (EDTA) | Sigma Aldrich | 6381-92-6 | Used in the protocol |
| Ethanol | Millipore Sigma | EM-EX0276-4S | Used in the protocol |
| Filter paper | Whatman | 1004-090 | Positive control |
| Glacial acetic acid | Sigma | SKU A6283 | Used in the protocol |
| Heat block/ ThermoMixer F1.5 | Eppendorf | 13527550 | For controlled temperatures |
| Incubator | Fisherbrand | 150152633 | Used for drying plant sample |
| Measuring Scale | Mettler Toledo | 30243386 | For specific quantities |
| Methanol 100 % | Fisher Chemical | A412-500 | Used in the protocol |
| Microplate (96 well) | Evergreen Scientific | 222-8030-01F | For anthrone assay |
| Nitric acid | Sigma Aldrich | 695041 | Used in the protocol |
| Polypropylene Microvials (2 mL) / screw capped tubes | BioSpec Products | 10831 | For high temperatures |
| Spectrophotometer(Multimode Detector) | Beckmancoulter DTX880 | 1000814 | For measuring absorbances |
| Spex SamplePrep 6870 Freezer / Mill | Spex Sample Prep | 68-701-15 | For grinding plant tissues into fine powder |
| Sulphuric acid | J.T.Baker | 02-004-382 | Used in the protocol |
| Sodium dodecyl sulfate (SDS) | Sigma Aldrich | 151-21-3 | Used in the PSB buffer |
| Tubes (2 mL) | Fisher Scientific | 05-408-138 | Used in the protocol |
| Tris Hydrochloride | Sigma Aldrich | 1185-53-1 | Used in the PSB buffer |
| Ultrapure distilled water | Invitrogen | 10977 | Used in the protocol |
| Vacuum dryer (vacufuge plus) | Eppendorf | 22820001 | For drying samples |
| Vortex mixer | Fisherbrand | 14-955-151 | For mixing |
| Waterbath | Thermoscientific | TSGP02PM05 | For temperature controlled conditions at specific steps |
| Weighing Paper | Fisher Scientific | 09-898-12A | Used in the protocol |
References
- Somerville, C. Cellulose synthesis in higher plants. Annual Review of Cell and Developmental Biology. 22, 53-78 (2006).
- Balasubramanian, V. K., Rai, K. M., Thu, S. W., Hii, M. M., Mendu, V. Genome-wide identification of multifunctional laccase gene family in cotton (Gossypium spp.); expression and biochemical analysis during fiber development. Scientific Reports. 6, 34309 (2016).
- Mendu, V., et al. Identification and thermochemical analysis of high-lignin feedstocks for biofuel and biochemical production. Biotechnology for Biofuels. 4, 43 (2011).
- Kraszkiewicz, A., Kachel-Jakubowska, M., Lorencowicz, E., Przywara, A. Influence of cellulose content in plant biomass on selected qualitative traits of pellets. Agriculture and Agricultural Science Procedia. 7, 125-130 (2015).
- Jordan, D. B., et al. Plant cell walls to ethanol. Biochemical Journal. 442, 241-252 (2012).
- Brett, C. T. Cellulose microfibrils in plants: biosynthesis, deposition, and integration into the cell wall. International Review of Cytology. 199, 161-199 (2000).
- Li, S., et al. Cellulose synthase complexes act in a concerted fashion to synthesize highly aggregated cellulose in secondary cell walls of plants. Proceedings of the National Academy of Sciences. 113, 11348-11353 (2016).
- Polko, J. K., Kieber, J. J. The regulation of cellulose biosynthesis in plants. The Plant Cell. 31, 282-296 (2019).
- Brown, R. M. The biosynthesis of cellulose. Journal of Macromolecular Science, Part A. 33, 1345-1373 (1996).
- Gautam, S. P., Bundela, P. S., Pandey, A. K., Jamaluddin, M. K., Sarsaiya, A., Sarsaiya, S. A review on systematic study of cellulose. Journal of Applied and Natural Science. 2, (2010).
- Coughlan, M. P. Enzymic hydrolysis of cellulose: An overview. Bioresource Technology. 39, 107-115 (1992).
- Robak, K., Balcerek, M. Review of second generation bioethanol production from residual biomass. Food Technology and Biotechnology. 56, 174-187 (2018).
- Cross, C. F., Bevan, E. J. Cellulose and chemical industry. Journal of the Society of Chemical Industry. 27, 1187-1193 (1908).
- Paloheimo, L., Eine, H., Kero, M. L. A method for cellulose determination. Agricultural and Food Science. 34, (1962).
- Kurschner, K., Hanak, A., Diese, Z. . Zeitschrift für Lebensmittel-Untersuchung und-Forschung. 59, 448-485 (1930).
- Norman, A. G., Jenkins, S. A new method for the determination of cellulose, based upon observations on the removal of lignin and other encrusting materials. Biochemical Journalournal. 27, (1933).
- Kiesel, A., Semiganowsky, N. Cellulose-Bestimmung durch quantitative verzuckerung. Berichte der deutschen chemischen Gesellschaft (A and B Series). 60, 333-338 (1927).
- Waksman, S. A. S., et al. A system of proximate chemical analysis of plant materials. Industrial Engineering Chemistry and Analytical Edition. 2, 167-173 (1930).
- Salo, M. -. L. Determination of carbohydrates in animal foods as seven fractions. Agricultural and Food Science. , 32-38 (1961).
- Giger-Reverdin, S. Review of the main methods of cell wall estimation: interest and limits for ruminants. Animal Feed Science and Technology. 55, 295-334 (1995).
- Updegraff, D. M. Semimicro determination of cellulose inbiological materials. Analytical Biochemistry. 32, 420-424 (1969).
- Viles, F. J., Silverman, L. Determination of starch and cellulose with anthrone. Analytical Chemistry. 21, 950-953 (1949).
- Scott, T. A., Melvin, E. H. Determination of dextran with anthrone. Analytical Chemistry. 25, 1656-1661 (1953).
- Kumar, M., Turner, S. Protocol: a medium-throughput method for determination of cellulose content from single stem pieces of Arabidopsis thaliana. Plant Methods. 11, 46 (2015).
- Yemm, E. W., Willis, A. J. The estimation of carbohydrates in plant extractsby anthrone. Biochemical Journal. 57, 508-514 (1954).
- Houston, K., Tucker, M. R., Chowdhury, J., Shirley, N., Little, A. The plant cell wall: A complex and dynamic structure as revealed by the responses of genes under Stress conditions. Frontiers in Plant Science. 7, (2016).
- Jiang, G., et al. Biomass extraction using non-chlorinated solvents for biocompatibility improvement of polyhydroxyalkanoates. Polymers. 10, 731 (2018).
- Li, T., et al. A saponification method for chlorophyll removal from microalgae biomass as oil feedstock. Marine Drugs. 14, 162 (2016).
- Wiltshire, K. H., Boersma, M., Möller, A., Buhtz, H. Extraction of pigments and fatty acids from the green alga Scenedesmus obliquus (Chlorophyceae). Aquatic Ecology. 34, 119-126 (2000).
- Foster, C. E., Martin, T. M., Pauly, M. Comprehensive compositional analysis of plant cell walls (lignocellulosic biomass) part II: carbohydrates. Journal of Visualized Experiments. (1837), (2010).
- Haigler, C., Betancur, L., Stiff, M., Tuttle, J. Cotton fiber: a powerful single-cell model for cell wall and cellulose research. Frontiers in Plant Science. 3, (2012).
- Spirk, S., Nypelö, T., Kontturi, E. Editorial: Biopolymer thin films and coatings. Frontiers in Chemistry. 7, (2019).
- Long, L. -. Y., Weng, Y. -. X., Wang, Y. -. Z. Cellulose aerogels: Synthesis, applications, and prospects. Polymers. 10, 623 (2018).

