Preparation of a Non-Cardiomyocyte Cell Suspension for Single-Cell RNA Sequencing from a Post-Myocardial Infarction Adult Mouse Heart
Summary
Here, we describe a protocol to isolate a sufficient amount of single non-cardiomyocytes with high viability from a post-myocardial infarction (MI) mouse heart. This can be used for subsequent single-cell sequencing, flow cytometry analysis, and primary cell culture.
Abstract
Myocardial infarction (MI) is one of the most common cardiovascular diseases, with increasing mortality worldwide. Non-cardiomyocytes account for more than half of the total cardiac cell population, and they contribute to adaptive compensations upon myocardial injury, including inflammatory responses, tissue repair, and scar formation. To study the post-MI cardiac microenvironment, single-cell RNA sequencing (scRNA-seq) is widely used to identify different cardiac cell types and intercellular communications. Among the procedures of scRNA-seq sample preparation, preparing the cell suspension is one of the most critical steps, because the cell viability can affect the quality of the scRNA-seq results. Therefore, we designed an experimental protocol for preparing a non-cardiomyocyte cell suspension from post-MI mouse hearts with an extra focus on improving the cell viability by choosing mild digestive enzymes, controlling the digestion time, and applying fluorescence-activated cell sorting (FACS). Finally, we isolated CD45+ cells from the non-cardiomyocyte cell suspension obtained through this protocol, and then we performed scRNA-seq.
Introduction
Myocardial infarction (MI) is one of the most common cardiovascular diseases, and its mortality increasing throughout the world1. MI is caused by an insufficient blood supply to the surrounding myocardium, which can be a result of a coronary artery blockage that occurs with atherosclerotic plaque rupture. Although percutaneous coronary intervention (PCI) has reduced the mortality rates of acute MI patients, the high prevalence of heart failure post MI remains a problem2. The key pathophysiology underlying post-MI heart failure is the body's compensatory response to cardiac injuries, which involves replacing the dead cardiac muscle with non-contractile fibrotic scars. These adaptive responses significantly rely on local inflammation mediated by the interactions between multiple cell types and cardiac tissue cells, and this inflammation is now considered as a potential therapeutic target to reduce fibrotic scar formation and, thus, protect against post-MI heart failure3,4. Interestingly, the microenvironment at the infarction site experiences a time-dependent transition in the infiltrating cell types and functions at different stages of MI1,5. Many studies have shown that non-cardiomyocytes (e.g., immune cells, fibroblasts, and endothelial cells) play central roles in post-MI inflammation and tissue repair5,6. In recent years, single-cell sequencing has been widely used as a powerful tool for elucidating the involvements and functions of non-cardiomyocytes in the post-MI microenviroment7,8. This provides insights into the pathophysiology of post-MI injury and repair and the development of potential therapies against post-MI heart failure.
High-throughput RNA sequencing (RNA-Seq) is a technique used to study entire transcriptomes in great detail using next-generation sequencing (NGS)7,8,9. Recently, the development of scRNA-Seq has revolutionized the biomedical research field. Compared with conventional bulk sequencing, scRNA-Seq analyses gene expression profiles and transcriptional heterogeneity at the single-cell level7,8. This technique significantly promotes the research of the cellular pathophysiology of MI9,10 by identifying different circulating cell types in the post-MI microenvironment and uncovering the interaction between cardiomyocytes and non-cardiomyocytes. These findings further contribute to uncovering novel therapeutic targets for post-MI heart failure. In general, the scRNA-seq-based post-MI experiment includes three main sections: (1) the establishment of post-MI animal model; (2) the preparation of the cell suspension; and (3) sample sequencing and data analysis. Noticeably, preparing the cell suspension is the most critical step in the preparation of the scRNA-Seq experiment because the quality of the cell suspension determines the accuracy of the results.
This protocol is designed to extract a non-cardiomyocyte cell suspension from the post-MI cardiac tissue; significantly, the specific details for maintaining cell viability and resolution are included. Meanwhile, the equipment used in this protocol, such as surgical kits for mice, rodent ventilators, and centrifuges, can be found in most animal experiment centers and biomedical laboratories, and, thus, the experiment cost of this protocol is relatively low. Furthermore, if considering time points and sites of infarction as variables, this protocol can be applied to simulate a wide range of clinical scenarios, especially for the post-MI complications.
Protocol
All the experiments were conducted in accordance with the guidelines for the care and use of laboratory animals at Zhejiang University and were approved by the Animal Advisory Committee at Zhejiang University.
1. Left anterior descending coronary artery ligation (LAD ligation) surgery
NOTE: Eight week old male C57BL/6J mice were used as models. The hearts were harvested 2 weeks after MI. Left anterior descending coronary artery ligation surgery was carried out as previously described and demonstrated11.
- Disinfect the surgical instruments with 70% ethanol before surgery.
- Weigh the mouse, and then anesthetize the mouse with intraperitoneal injections of 1% pentobarbital sodium (50 mg/kg). Observe the toe-pinch reflex, and measure the respiratory rate to evaluate the depth of anesthesia during the procedure.
- Place the mouse on the operating table with a 37 °C heating pad. Use ophthalmic ointment to prevent eye dehydration.
- Depilate its left precordial chest and neck with depilatory cream, and use cotton swabs to remove hairs. Disinfect its skin with iodophor, followed by 70% alcohol to clean the area three times.
- Perform a midline cervical incision (0.8-1.0 cm) under a microscope. Separate the skin, muscle, and tissue surrounding the trachea, and then insert a 20 G cannula into the trachea.
NOTE: Observe while inserting the cannula into the trachea under microscopic view to ensure that the tube is not inserted into the esophagus. - Connect the mouse to a rodent ventilator set at 0.2 mL tidal volume with 150 strokes per minute.
- Incise the mouse chest obliquely along the midclavicular line with a sterile scissor. Dissect the subcutaneous tissue to expose the ribs.
- Use sterile scissors to cut the skin obliquely along the left mid-clavicular line (0.8-1.0 cm). Perform left thoracotomy to separate the third and fourth ribs to expose the heart. Use a rib retractor for a clear view.
- Open the pericardium with curved forceps, and then ligate the left anterior descending artery (below the left atrium appendage) using a 7-0 silk suture. A pale anterior wall of the left ventricle indicates that the myocardial perfusion is successfully interrupted.
- Release the rib retractor, and stitch the thoracic incision in layers with a 5-0 silk suture while expelling air from the chest. Stitch the neck incision with a 5-0 suture silk suture.
- Remove the tracheal tube from the mouse once its spontaneous breathing is restored. Intraperitoneally inject 0.5 mL of pre-warmed sterile saline. Inject buprenorphine (0.1 mg/kg) every 12 h subcutaneously to minimize pain.
- Finally, place the mouse on the heating pad to wait for resuscitation. Monitor the mouse to ensure it does not suffer from excessive dyspnea or hemorrhaging via visual observation.
- Return the mouse to its cage after it wakes up. For the first week, monitor the mouse daily to verify that its mobility, grooming, and eating habits are adequate. According to the experiment's needs, harvest the heart at different time points after MI.
NOTE: In this protocol, the single-cell suspension was prepared with the heart 2 weeks after MI.
2. Preparation of the cardiac single-cell suspension
- Preparation of the solutions and media
- Cardiac tissue dissociation buffer: Mix 10 mg of collagenase IV (600 U/mL), 5 mg of dispase II (0.5 U/mL), and 50 µL of DNase I (100 µg/mL) in 5 mL of RPMI 1640. Pre-warm this medium to 37 °C in a water bath before use.
- Cell wash buffer: Prepare 1% BSA or 10% FBS in phosphate-buffered saline (PBS, PH 7.4), and keep at 4 °C or on ice.
- Perfusion solution: Pre-cool PBS (PH 7.4) as a perfusion solution.
- Red blood cell (RBC) lysis buffer: Use the RBC lysis buffer as mentioned by the manufacturer.
NOTE: Refer to the Table of Materials for the list of reagents and equipment used in this study.
- Euthanize the mouse by an intraperitoneal pentobarbital sodium (150 mg/kg) injection.
- Heart dissection
- Place the mouse in a supine position by fixing its hindlimbs in place with adhesive tape.
- Spray the body with 70% ethanol, and then cut vertically through the abdominal skin and muscles while avoiding piercing the liver. Carefully open the thorax while avoiding puncturing the heart.
- Use the pre-cooled PBS (PH 7.4) to perfuse the left ventricle until the colorless perfusion fluid is observed (approximately 15 mL of PBS [pH 7.4] is needed).
- Gently lift the heart from the thorax with forceps, and cut away the excess tissue attached to the outside of the heart with ophthalmic scissors. Place the heart in 1 mL of pre-cooled PBS (PH 7.4) in a 1.5 mL microcentrifuge tube.
- Enzymatic dissociation of the heart tissue
- Transfer the mouse heart into one well of a 24-well plate with 150 µL of the cardiac tissue dissociation buffer placed on ice, and cut it into small pieces (~1 mm) with a scissor.
- Transfer the minced heart tissue into a 15 mL centrifuge tube with 5 mL of the tissue dissociation buffer.
NOTE: The recommended cardiac tissue dissociation buffer volume is 5 mL/heart. - Place the centrifuge tube in the electrically thermostatic reciprocating shaker at 125 rpm for 1 h at 37 °C. Pipet the tissue suspension up and down 10 times every 20 min.
- Filter the cell suspension through a 70 µm cell strainer to remove undigested tissue and clumps. Add 25 mL of the cell wash buffer to rinse the original tube, and then transfer it to rinse the cell strainer. Next, filter the cell suspension through a 40 µm cell strainer to further remove cell clumps.
- Centrifuge the filtered cell suspension at 300 x g for 5 min at 4 °C. Discard the supernatant, and then resuspend the cell sample in 1x RBC lysis buffer for 5 min at room temperature (RT).
- Add four times more volume of the cell wash buffer, and centrifuge at 300 x g for 5 min at 4 °C.
- Remove the supernatant, and resuspend the cells in 10 mL of the cell wash buffer. Centrifuge at 300 x g for 5 min at 4 °C.
- Remove the supernatant, and resuspend the cells in 1 mL of the cell wash buffer.
- Mix 18 µL of the cell suspension with 2 µL of acridine orange (AO)/propidium iodide (PI) staining solution (9:1), and add 10 µL of the mixture to a counting chamber slide. Evaluate the cell numbers and viability using an automatic cell counter.
Representative Results
A single-cell suspension with high vitality was obtained by implementing section 2 of this protocol. However, cell fragments could still be observed (Figure 1A); hence, fluorescence-activated cell sorting (FACS) was performed to further improve the quality16.After FACS, the average cell size reduces from 9.6 µm to 9.1 µm (Table 1), which suggests that the proportion of cell fragments can be effectively reduced in the cell suspension by FACS (Figure 1B). The gating strategy used for FACS is shown in Figure 1C.
In addition to the images in Figure 1, the cell concentration, average cell size, and cell viability were measured by a cell counter (Table 1). The differences in cell concentrations were due to the volume of the cell suspension. Noticeably, the average cell size decreased after FACS (Table 1), which shows that FACS can effectively reduce cell clumping and remove cell fragments.
ScRNA-seq was performed to study the cardiac immunological microenvironment while verifying the cell suspension quality obtained through this protocol. After preparing the single-cell suspension, we firstly sorted the cells that expressed surface protein CD45, which is a common marker for a wide range of immune cells. Next, we performed scRNA-seq on CD45+ cells using the 10X Genomics platform17. In this way, we pooled 14,217 individual cells from three healthy heart samples into a merged data set after quality control (Table 2). The unsupervised clustering and reduction in t-distributed stochastic neighbor embedding (t-SNE) dimensionality showed an obvious separation of seven types of immune cells (Figure 2). Moreover, each immune cell type showed a high expression of classic markers. Conclusively, these results show that the single-cell suspension prepared by this protocol has a high potency for single-cell sequencing.
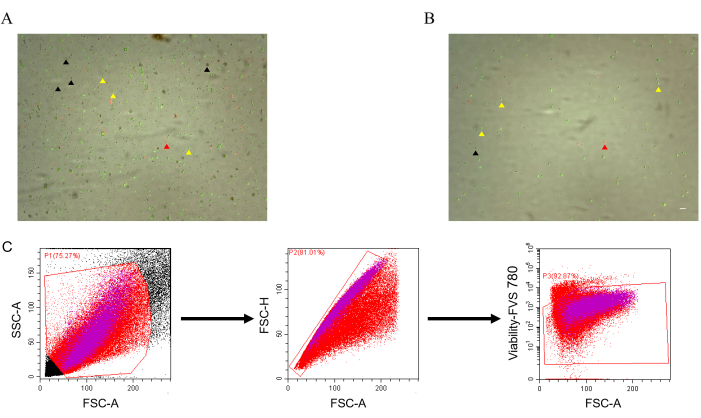
Figure 1: Cell viability in the cardiac single-cell suspension. (A) Cell viability without FACS. The black arrowheads indicate cell fragments (no fluorescence). (B) Cell viability with FACS. The yellow arrow indicates live cells (green fluorescence), and the red arrow indicates dead cells (red fluorescence).(C) Gating strategy. Scale bar: 50 µm. Please click here to view a larger version of this figure.
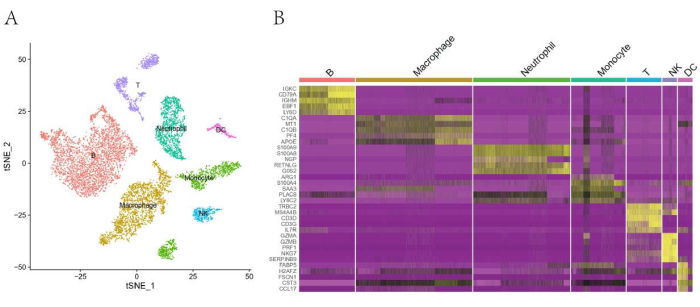
Figure 2: Single-cell RNA sequencing of cardiac CD45+ cells from three mouse heart samples. (A) tSNE plot of CD45+ cells showing the immune cell populations identified based on canonical marker genes. (B) Heatmap of the top five differentially expressed genes in each subpopulation. Please click here to view a larger version of this figure.
| Before FACS | After FACS | |
| Total cell concentration (cells/mL) | 1.51 x 106 | 3.40 x 105 |
| Live cell concentration (cells/mL) | 1.40 x 106 | 3.15 x 105 |
| Dead cell concentration (cells/mL) | 1.09 x 105 | 2.48 x 104 |
| Viability (%) | 92.8 | 92.7 |
| Average cell size (μm) | 9.6 | 9.1 |
Table 1: Cardiac single-cell suspension before and after FACS. The cell densities of the cardiac single-cell suspension obtained before and after FACS are compared using an automated cell counter.
| Estimates | |
| Estimated Number of Cells | 14,217 |
| Mean Reads per Cell | 81,017 |
| Median Genes per Cell | 1,374 |
| Total Genes Detected | 18,424 |
| Median UMI Counts per Cell | 4,021 |
| Valid Barcodes | 98.4% |
| Reads Mapped to Genome | 95.1% |
Table 2: Quality control statistics. The table lists the different estimates of the single-cell data of healthy mouse hearts.
Discussion
This article aimed to describe a protocol to isolate single non-cardiomyocytes from mouse hearts post MI. The protocol can be applied to isolate different types of cells in the post-MI microenvironment with high quality, including immune cells, endothelial cells, and fibroblasts. Three essential factors are crucial for obtaining a high-quality cell suspension for single-cell sequencing. The first one is the setting of the enzymatic digestion. It is important to control the time of digestion and the volume and concentration of the digestive enzymes to assure both cell viability and isolation efficiency. In this case, we appropriately increased the concentration of the enzyme and prolonged the digestion time. Here, we recommend a digestion duration of 45 min to 1 h. A shorter duration may increase the cell viability but may lead to insufficient cardiac tissue digestion and may increase the amount of cell clumps. In addition, we used DNase I to prevent cell aggregation because lytic cells can release free DNA to wrap the surrounding cells to form cell clumps13. The second essential factor is using a cell washing solution containing BSA or FBS. BSA or FBS can not only protect the activity of the enzymes but also improve the viability of the cells. The final essential factor is facilitating FACS to eliminate cell fragments and dead cells. Cell fragments can result in an inaccurate cell count and viability. These factors together enhance the viability and purity of the cell suspension, making it more compatible with single-cell sequencing.
This protocol firstly describes the cell suspension preparation for single-cell sequencing, and it also includes additional procedures to enhance the quality of the cell suspension at a relatively low cost. In comparison with other protocols used by previous studies, we avoid using digestive enzymes that can cause the aggregation of cells induced by free DNA, such as trypsin14. The removal of the cell debris is one of the major challenges for single-cell studies, and some studies use dead cell removal kits to improve cell viability before running the samples14,15. However, here, we suggest using FACS to enhance the sample purity due to its high effectiveness in reducing dead cells and debris.
The single-cell suspension obtained by this protocol can be further analyzed by flow analysis or primary cell culture (standard aseptic procedures)18. As the digestive enzymes are relatively mild, it is suitable to digest cardiac tissue taken in different stages of MI, as well as that from healthy hearts. However, this protocol has certain limitations. For example, prolonged digestion at 37 °C inevitably promotes the expression of stress genes19,20. This limitation can be solved by using low-temperature active protease or reducing the digestion time followed by FACS. Also, this protocol does not include isolating cardiomyocytes, and, thus, it is not suitable for studies on local tissue responses.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was supported by the Natural Science Funds of Zhejiang Province (LQ22H020010).
Materials
| Acridine Orange / Propidium Iodide Stain | Logos biosystems | F23001 | |
| Automated Cell Counter | Logos biosystems | L20001 | |
| Bovine Serum Albumin | Servicebio | G5001 | Cytoprotective effect |
| Cell Counting Slides | Logos biosystems | L12001 | |
| Collagenase Type IV | Gibco | 17104019 | Digestive enzymes |
| Dispase II | Sigma | D4693 | Digestive enzymes |
| Dnase I | Roche | 11284932001 | Prevent cell clumping |
| Falcon 40μm Cell Strainer | Falcon | 352340 | Remove cell clumps |
| Falcon 70μm Cell Strainer | Falcon | 352350 | Remove undigested tissue and clumps |
| Flow Cell Sorter | Beckman Coulter | B25982 | |
| Iodophor | OU QING SI | 10054963976859 | |
| Needle Holder | FST | 12061-01 | |
| Ophthalmic Forceps | RWD | F14012-10 | |
| Ophthalmic Scissors | RWD | S11036-08 | |
| Phosphate Buffered Saline | Servicebio | G4202-500ML | |
| RBC Lysis Buffer | Beyotime | C3702-120ml | Remove red blood cells |
| Rib Retractor | FST | 17005-04 | |
| Rodent Ventilator | Harvard | 730043 | |
| RPMI 1640 Medium | Gibco | 11875093 | Solvent solution of enzyme |
| Sterile Scissor | RWD | S14014-10 |
References
- Reed, G. W., Rossi, J. E., Cannon, C. P. Acute myocardial infarction. Lancet. 389 (10065), 197-210 (2017).
- Gu, J., et al. Incident heart failure in patients with coronary artery disease undergoing percutaneous coronary intervention. Frontiers in Cardiovascular Medicine. 8, 727727 (2021).
- Frangogiannis, N. G. The inflammatory response in myocardial injury, repair, and remodelling. Nature Reviews. Cardiology. 11 (5), 255-265 (2014).
- Kain, V., Prabhu, S. D., Halade, G. V. Inflammation revisited: Inflammation versus resolution of inflammation following myocardial infarction. Basic Research in Cardiology. 109 (6), 444 (2014).
- Tzahor, E., Dimmeler, S. A coalition to heal-the impact of the cardiac microenvironment. Science. 377 (6610), 4443 (2022).
- Song, K., et al. Heart repair by reprogramming non-myocytes with cardiac transcription factors. Nature. 485 (7400), 599-604 (2012).
- Jaitin, D. A., et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science. 343 (6172), 776-779 (2014).
- Massaia, A., et al. Single cell gene expression to understand the dynamic architecture of the heart. Frontiers in Cardiovascular Medicine. 5, 167 (2018).
- Papalexi, E., Satija, R. Single-cell RNA sequencing to explore immune cell heterogeneity. Nature Reviews. Immunology. 18 (1), 35-45 (2018).
- Jin, K., et al. Single-cell RNA sequencing reveals the temporal diversity and dynamics of cardiac immunity after myocardial infarction. Small Methods. 6 (3), 2100752 (2022).
- Tombor, L. S., et al. Single cell sequencing reveals endothelial plasticity with transient mesenchymal activation after myocardial infarction. Nature Communications. 12 (1), 681 (2021).
- Virag, J. A., Lust, R. M. Coronary artery ligation and intramyocardial injection in a murine model of infarction. Journal of Visualized Experiments. (52), e2581 (2011).
- Reichard, A., Asosingh, K. Best practices for preparing a single cell suspension from solid tissues for flow cytometry. Cytometry Part A. 95 (2), 219-226 (2018).
- Gladka, M. M., et al. Single-cell sequencing of the healthy and diseased heart reveals cytoskeleton-associated protein 4 as a new modulator of fibroblasts activation. Circulation. 138 (2), 166-180 (2018).
- Garcia-Flores, V., et al. Preparation of single-cell suspensions from the human placenta. Nature Protocols. , (2022).
- Guez-Barber, D., et al. FACS purification of immunolabeled cell types from adult rat brain. Journal of Neuroscience Methods. 203 (1), 10-18 (2012).
- Rheinländera, A., Schravenab, B., Bommhardt, U. CD45 in human physiology and clinical medicine. Immunology Letters. 196, 22-32 (2018).
- Hua, X., et al. Single-cell RNA sequencing to dissect the immunological network of autoimmune myocarditis. Circulation. 142 (4), 384-400 (2020).
- Grindberg, R. V., et al. RNA-sequencing from single nuclei. Proceedings of the National Academy of Sciences of the United States of America. 110 (49), 19802-19807 (2013).
- Vanden Brink, S. C., et al. Single-cell sequencing reveals dissociation-induced gene expression in tissue subpopulations. Nature Methods. 14, 935-936 (2017).

