Identification of the Genes Involved in Stomatal Development via Epidermal Phenotype Scoring
Summary
This paper describes two phenotyping methods without the use of epidermal peels to characterize the genes controlling stomatal development. The first method demonstrates how to analyze a stomatal phenotype using a toluidine blue O-stained plant epidermis. The second method describes how to identify stomatal ligands and monitor their biological activities.
Abstract
Stomata are small pores on the surface of land plants that are involved in gas exchange and water vapor release, and their function is critical for plant productivity and survival. As such, understanding the mechanisms by which stomata develop and pattern has tremendous agronomic value. This paper describes two phenotypic methods using Arabidopsis cotyledons that can be used to characterize the genes controlling stomatal development and patterning. Presented first are procedures for analyzing the stomatal phenotypes using toluidine blue O-stained cotyledons. This method is fast and reliable and does not require the use of epidermal peels, which are widely used for phenotypic analyses but require specialized training. Due to the presence of multiple cysteine residues, the identification and generation of bioactive EPF peptides that have a role in stomatal development have been challenging. Thus, presented second is a procedure used to identify stomatal ligands and monitor their biological activity by bioassays. The main advantage of this method is that it produces reproducible data relatively easily while reducing the amount of peptide solution and the time required to characterize the role of the peptides in controlling stomatal patterning and development. Overall, these well-designed protocols enhance the efficiency of studying the potential stomatal regulators, including cysteine-rich secretory peptides, which require highly complex structures for their activity.
Introduction
Proper patterning and differentiation of the plant stomata are critical for their function in two fundamental biological processes, photosynthesis and transpiration, and are enforced by EPF peptide signaling pathways. In Arabidopsis, three secreted cysteine-rich peptides, EPF1, EPF2, and STOMAGEN/EPFL9, control different aspects of stomatal development and are perceived by cell-surface receptor components, including ERECTA-family receptor kinases (ER, ERL1, and ERL2), SERKs, and TMM1,2,3,4,5,6,7,8,9,10. This recognition then leads to the downregulation of the transcription factors that promote stomatal differentiation by a MAPK-dependent process11. The discovery of these core stomatal genes is primarily achieved by the phenotypic screening of mutants exhibiting epidermal defects. This paper presents relatively simple and efficient phenotyping methods for visualizing the stomata and other epidermal cells, which are required to identify and characterize the potential genes controlling stomatal patterning and differentiation.
The observation of the details of the plant epidermis has typically been achieved by using epidermal peels with or without staining with a dye such as toluidine blue O (TBO) or safranin12,13,14. However, the main challenge of these methods is that they require specialized training to peel the leaf epidermis without tearing the tissues and to carefully observe and analyze the patterning data while avoiding the images taken from different parts of the leaf. Chemical treatments to clear the tissue samples with reagents such as chloral hydrate-based clearing solutions have also been widely used for a various range of biological materials8,15; these treatments do generate a great deal of phenotypic information by providing high-quality images but also require the use of dangerous chemicals (e.g., formaldehyde, chloral hydrate). This paper first presents a relatively easy and convenient phenotyping method that produces images sufficient for quantitative analysis but does not require the use of dangerous chemicals and epidermal leaf peels for the sample preparation. A TBO-stained cotyledon epidermis is also ideal for the study of stomatal development because the lack of trichomes and the smaller developmental gradient in cotyledons allow for the simple and tractable interpretation of the epidermal phenotypes.
Stomatal EPF peptides belong to the group of plant-specific, cysteine-rich peptides that have relatively large mature sizes and intramolecular disulfide bonds between conserved cysteine residues. Correct conformational folding is critical for their biological function, but cysteine-rich peptides, which are produced by either chemical synthesis or a heterologous recombination system, can be inactive and are a mixture of both properly folded and unfolded peptides3,7,16. Thus, the screening of bioactive peptides that have a role in controlling stomatal development has been a very challenging task. This manuscript additionally describes a bioassay for the better identification and characterization of bioactive stomatal peptides. In this method, Arabidopsis seedlings are grown in a multi-well plate containing media with and without potential peptides for 6-7 days. Then, the cotyledon epidermis is visualized using a confocal microscope. In general, to clearly visualize the biological activity of potential peptides in stomatal development, the genotypes that produce more and/or less stomatal lineage cells, such as the epf2 mutant, which produces more epidermal cells, and the STOMAGEN-ami line, which confers reduced epidermal cell density2,4,5, are used in addition to the wild-type Arabidopsis control (Col-0) for the bioassays.
Overall, the two protocols presented here can be used for the quick and efficient assessment of various epidermal phenotypes and for screening small peptides and hormones that have a role in controlling stomatal patterning and development.
Protocol
1. Staining Arabidopsis cotyledons with TBO
- Seed sterilization and growth conditions
- Sterilize ~30 Arabidopsis seeds per genotype in a microcentrifuge tube by adding 1 mL of a seed sterilization solution (33% commercial bleach, 0.1% Triton X-100), and gently rock for 10-12 min at room temperature (RT).
NOTE: Sterilize ~30 seeds of the wild-type Arabidopsis accession Columbia (Col-0) and/or ~60 seeds of transgenic plants carrying a chemically-inducible gene (e.g., Est::EPF27) to use transgenic seedlings grown on 1/2 Murashige and Skoog (MS) plates (2.16 g/L containing 0.8% agar [w/v]) without inducer (e.g., β-estradiol) as controls (Figure 1 and Figure 2). - Remove the sterilization solution, wash the seeds four times with 1 mL of sterile water in a laminar flow hood, and resuspend them in ~200 μL of sterile 0.1% agar.
- Sow ~30 seeds per genotype on 1/2 MS agar plates or 1/2 MS agar plates containing an inducer (e.g., 10 µM β-estradiol) for the chemical induction of the transgene (e.g., Est::EPF27) using a pipette, and seal with micropore tape.
NOTE: The seeds need to be evenly distributed on the plate to avoid placing them in close proximity, as this will prevent the seedlings from growing uniformly. - Stratify the seeds by placing the plates at 4 °C without light for 3-5 days to synchronize the germination.
- After stratification, incubate the plates in a growth chamber at 22 °C under 120 µmol·m−2·s−1 light with a photoperiod of 16 h of light and 8 h of dark for 10 days.
- Sterilize ~30 Arabidopsis seeds per genotype in a microcentrifuge tube by adding 1 mL of a seed sterilization solution (33% commercial bleach, 0.1% Triton X-100), and gently rock for 10-12 min at room temperature (RT).
- Sampling and TBO staining of Arabidopsis cotyledons
- At 10 days post-germination, carefully select and cut out one of the cotyledons from the individual seedlings that are growing uniformly with other seedlings on the plate to limit variability.
NOTE: Cotyledons are sampled from 10 day old seedlings because they do not have trichomes and immature epidermis cells, which simplifies the interpretation of the epidermal phenotypes. - Place each cotyledon into a microcentrifuge tube containing 1 mL of a fixing solution (9:1 ethanol to acetic acid) using forceps, and leave the sample in the fixing solution overnight, at minimum, and at room temperature.
NOTE: The samples can then be stored for up to a couple of years in this condition. The image Figure 2E was taken from a cotyledon sample that was stored in fixing solution for over 3 years. It is important to keep the cotyledon in a fixing solution immediately after cutting and to place no more than five cotyledons in each microcentrifuge tube for homogeneous sample preparation later. - Remove the fixing solution, and add 1 mL of 70% ethanol. After inverting the tube a couple of times, leave the tube containing the samples at RT for ~ 30 min.
- Repeat step 1.2.3 using 1 mL of 50% ethanol and then 20% ethanol.
- Replace the 1 mL of 20% ethanol with 1 mL of distilled water, and then leave the tube containing the cotyledon samples for ~30 min after inverting the tube a few times.
NOTE: The samples may stay in distilled water for more than 24 h, but this is not recommended for obtaining good-quality images (well-stained images for different epidermal cell types) for quantitative analysis. - Remove all the distilled water from the tube. Then, immediately add ~200 μL of TBO staining solution (0.5% TBO in H2O, filtered) for ~2 min.
NOTE: Make sure each cotyledon sample is evenly exposed to TBO solution by gently flicking the tubes during incubation. To prevent overstaining with TBO (the timing of the staining is essential and can be variable for each genotype), do not process more than six tubes containing samples at a time. - Remove the TBO staining solution as thoroughly as possible, and then immediately wash the samples a couple of times by adding 1 mL of fresh distilled water.
- At 10 days post-germination, carefully select and cut out one of the cotyledons from the individual seedlings that are growing uniformly with other seedlings on the plate to limit variability.
- Imaging and data analysis
- Take a microscope slide, and add a drop of 15% glycerol (~50 μL). Place the cotyledon with the abaxial side up into 15% glycerol on the slide using fine forceps. Then, gently cover it with a coverslip so that any air bubbles formed can be removed from the sample.
- Image the abaxial side of the cotyledon epidermis using a brightfield microscope (Figure 1 and Figure 2), and then examine the epidermal phenotype by counting the number of stomata and other epidermal cell types.
- For each genotype, calculate the epidermal phenotype using the following formulas described by Jangra et al.17:
Stomatal index (%) = (Number of stomata/Total number of epidermal cells) × 100
Stomatal density (mm−2) = Number of stomata/Area (mm2)
NOTE: Image at least eight cotyledons (N = 8) for each genotype, and document the epidermal phenotype of each genotype by comparing it to the phenotype of the wild-type plant and/or transgenic plants expressing a chemically-inducible transgene grown without an inducer.
2. Bioassays for stomatal peptides
NOTE: The procedure for bioassays is shown in Figure 3.
- Seed sterilization and growth conditions
- Sterilize ~25 seeds for each treatment as above, and sow approximately half of the seeds onto each of two 1/2 MS agar plates in a laminar flow hood.
NOTE: Use the genotype with high and/or low epidermal cells (e.g., epf22) instead of using a wild-type background (e.g., Col-0) to easily detect the biological activities of the peptides on the epidermal cell patterning and differentiation (Figure 4). - Stratify the seeds for 3 days at 4 °C in the dark.
- Take out each of the two plates at ~10 h intervals, and incubate the seeds on plates at 22 °C under 120 µmol·m−2·s−1 light with a photoperiod of 16 h of light and 8 h of dark for 1 day.
- Sterilize ~25 seeds for each treatment as above, and sow approximately half of the seeds onto each of two 1/2 MS agar plates in a laminar flow hood.
- Peptide treatment and imaging
- In a laminar flow hood, carefully transplant 10-12 one day old Arabidopsis seedlings from each of the two prepared plates into a 24-well plate containing 1.5 mL of 1/2 MS liquid medium in each well (~20 seedlings per well).
NOTE: The timing of the peptide treatment for the seedlings is critical to the phenotypic outcomes of the peptide treatment, so prepare the seeds on two separate plates to obtain two different seedling developmental stages for the treatment. - Add either a buffer alone (50 mM Tris-HCl [pH 8.0]) or two different concentrations of the peptide (typically, 1 µM and 2.5 µM peptide in 50 mM Tris-HCl [pH 8.0]) to each well containing ~20 one day old Arabidopsis seedlings germinated on 1/2 MS agar plates.
NOTE: Remove the peptide solution aliquots from a freezer, and keep them at RT for a few minutes before the treatment. Peptides stored in a freezer at −80 °C are stable for a couple of years, but freeze-thaw cycles should be avoided by storing the peptide solution in small aliquots. - After gently mixing the seedlings with a buffer alone or a peptide solution using a pipette, seal the plate with micropore tape. Incubate the assay plate under long-day conditions (16 h photoperiod, 120 µmol·m−2·s−1 light) for 5-7 days at 22 °C.
NOTE: Prevent the seedlings from growing too close together to allow each seedling to have even exposure to the peptide solution. Gently rotate the plate for 2-3 h before incubating the plate in a growth chamber to increase aeration. - Transfer each seedling from the well containing either buffer alone or peptide on a cover slide, and dissect the cotyledon of the seedling. Place the abaxial side of the cotyledon up using forceps, and cut it into small pieces.
- Take another clean microscope slide, and place on it a drop of propidium iodide solution (~25 μL, 2 mg/mL in H2O).
- Place one of the small pieces of cotyledon into a drop of propidium iodide solution using forceps, and gently place the coverslip. Apply additional propidium iodide solution on the edge of the coverslip to remove any air bubbles that have formed.
- Image the abaxial side of the cotyledon using a confocal microscope, and compare the images with the images from seedlings grown in 1/2 MS medium containing buffer only.
NOTE: The images presented in Figure 4 were taken from wild-type Col-0 or epf22,5 seedlings that were treated with buffer only (Figure 4A, B) or two batches of EPF2 peptide solution (Figure 4C-F) and were stored at −80 °C over 1 year.
- In a laminar flow hood, carefully transplant 10-12 one day old Arabidopsis seedlings from each of the two prepared plates into a 24-well plate containing 1.5 mL of 1/2 MS liquid medium in each well (~20 seedlings per well).
Representative Results
Various stomatal transgenic plants and mutants known to have less or more stomatal density and clustering (epf22,5, epf1 epf22,5, tmm12, a STOMAGEN-silenced line4, and transgenic lines carrying the estradiol-inducible Est::EPF1 or Est::EPF2 overexpression construct7) were used to demonstrate the effectiveness of the two phenotypic analyses presented here, which aimed to identify and characterize the genes that have a role in stomatal development and patterning. In order to produce high-quality epidermis images without the need for epidermal peels to quantify the different types of epidermal cells for analysis, it is critical to pre-adjust the sample staining time with TBO for each genotype, as each one may have different epidermal phenotypes compared to that of the wild-type control (Col-0) (Figure 1A and Figure 2A). Based on experience, genotypes with fewer stomata require longer staining times with TBO (Figure 1B and Figure 2D,E), while genotypes with more stomata and clustering require shorter staining times (Figure 1C and Figure 2B,C). As varying amounts of properly folded peptides in the total peptide solution can mask the biological activity of the peptides in stomatal development, it is good practice to use additional, higher total concentrations of the peptide solution (e.g., 2.5 μM EPF2), as well as genotypes that have less or more stomatal phenotypes (e.g., epf2), for the bioassays. The activity of EPF2 peptides, which have a role in inhibiting stomatal initiation, was easily detected by using epf2 mutants with more epidermal cells than Col-0, even if certain batches of the prepared EPF2 peptides had smaller amounts of the bioactive forms of the peptide (e.g., EPF2 peptide solution batch 1 in Figure 4).
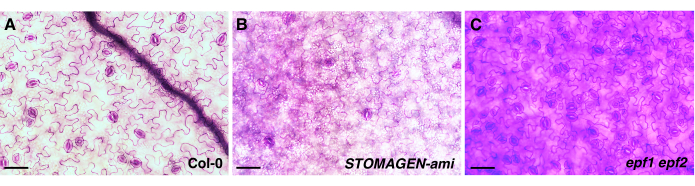
Figure 1: The effect of the incubation time with TBO on the genotypes exhibiting different epidermal phenotypes. Abaxial cotyledon images from 10 day old seedlings of three Arabidopsis genotypes: (A) wild-type (Col-0), (B) a STOMAGEN-silenced line (STOMAGEN-ami), and (C) epf1 epf2 mutants. Col-0 represents the wild-type Arabidopsis control, and the STOMAGEN-silenced line and the epf1 epf2 mutants represent the genotypes with low and high numbers of stomata, respectively. The images were taken using an inverted microscope with a 20x objective lens (0.35 mm2 field of view). The quality of the images was variable, although all the cotyledon samples from the different genotypes were stained with TBO (0.5% TBO in H2O) for the same amount of time (2 min) before imaging. Therefore, to obtain well-stained images for analysis, it is important to predetermine the adequate TBO staining time for each genotype. Scale bar = 50 µm. Please click here to view a larger version of this figure.
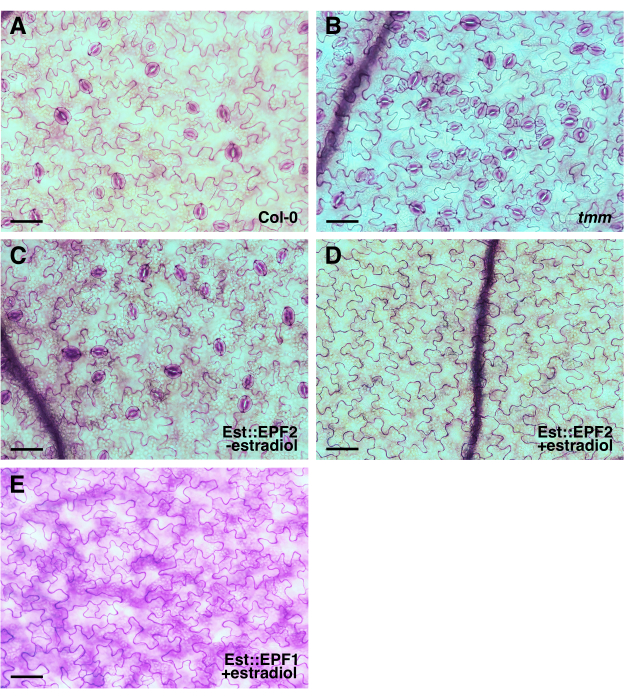
Figure 2: TBO-stained epidermis images without the use of epidermal peels for quantitative analysis. Representative epidermis images of cotyledons from 10 day old seedlings of (A) wild-type (Col-0), (B) tmm, and (C,D) transgenic lines carrying an estradiol-inducible EPF2 (Est::EPF2) or (E) EPF1 overexpression construct (Est::EPF1) can be used for the quantitative analysis of the epidermal phenotype.A cotyledon in a fixing solution for over 3 years was used to take the image presented in (E). The images were taken using an inverted microscope with a 20x objective lens (0.35 mm2 field of view). The cells were outlined by TBO staining, and half the width of the full-size images is presented for display. Scale bar = 50 µm. Please click here to view a larger version of this figure.
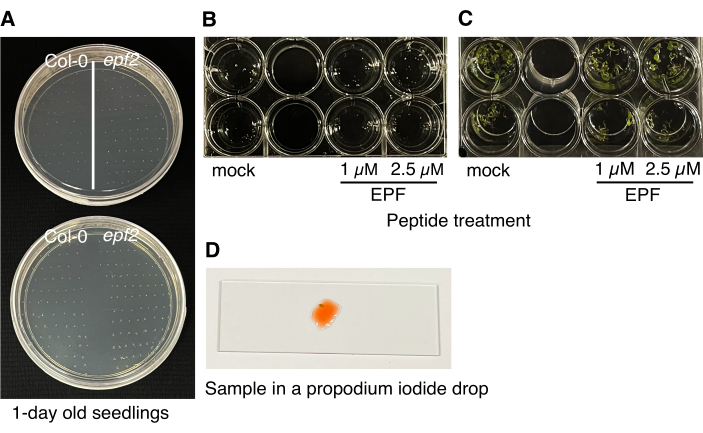
Figure 3: Procedure for the bioassays. (A) The seeds are sown on two 1/2 MS agar plates and taken out at 10 h intervals. (B) The 1 day old Arabidopsis seedlings are transplanted to 24-well plates containing 1/2 MS medium with either mock or two different peptide concentrations. (C) After 5-7 days, the seedlings are ready for imaging to determine the biological activity of the peptides in stomatal development and patterning. (D) A cotyledon slice is placed in a drop of propidium iodide solution on a microscope slide for imaging. Please click here to view a larger version of this figure.
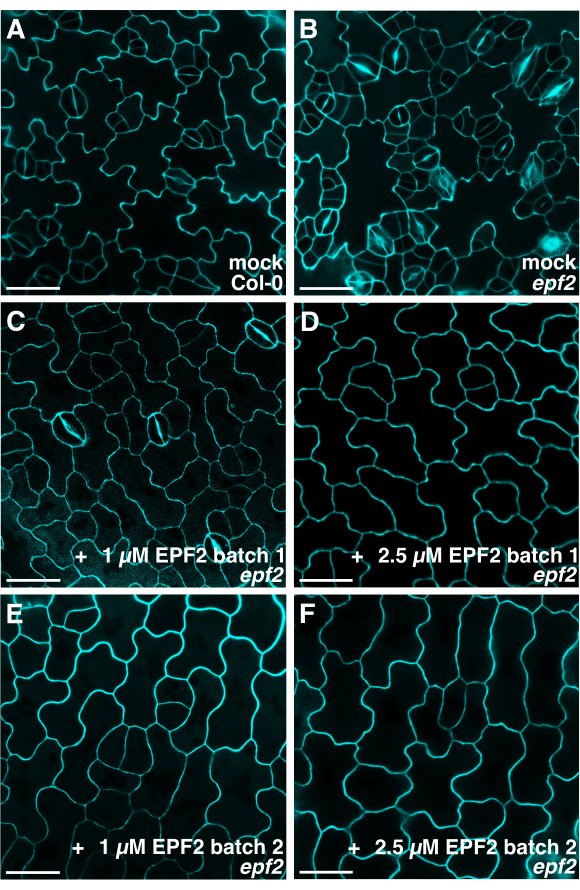

Figure 4: Confocal microscopy of the abaxial epidermis of Arabidopsis cotyledons following peptide treatment. Representative confocal images of (A) wild-type and (B) epf2 cotyledon epidermis grown for 6-7 days in a buffer solution and an (C–F) epf2 cotyledon epidermis grown with two different batches of EPF2 peptides. The images were taken using a confocal microscope using a 40x objective lens (excitation of 561 nm and a 561 nm long-pass emission filter) to capture the propidium iodide staining and visualize the cell outlines. Each EPF2 peptide solution prepared has different amounts of properly folded (bioactive) and misfolded (inactive) forms of the peptides. Therefore, for the initial screening of peptides with a potential role in stomatal development, it is recommended to use two different concentrations of the total peptide solutions as indicated for the bioassays. Scale bar = 30 µm. Please click here to view a larger version of this figure.
Discussion
The two phenotypic analysis methods for identifying and characterizing the genes controlling stomatal patterning and differentiation presented here are convenient and reliable assays since the protocols do not require the use of epidermal peels and specialized equipment (which are time-consuming and require special training for sample preparation) but do produce high-quality images for the quantitative analysis of epidermal phenotypes.
A limitation of this technique for phenotypic analysis using TBO-stained Arabidopsis cotyledons is that obtaining high-quality images with visual contrast for different epidermal cell types depends on the staining time and the unique characteristics of the tissue, which may be both species-dependent and genotype-dependent (Figure 1)18. Other phenotyping methods to observe epidermal cells using epidermal peels have the same challenges, and they also require more time and special training. Difficulties in obtaining properly stained epidermis images sufficient for data analysis can be avoided by staining cotyledons from genotypes with fewer stomata for longer periods of time and samples from genotypes with more stomata for shorter periods of time. The staining time for each genotype can also be adjusted by first checking only a few samples to optimize the staining time for genotypes with similar epidermal phenotypes.
Obtaining a sufficient amount of well-folded bioactive peptides, especially cysteine-rich peptides (which are ligands that have diverse functions in controlling plant development, including stomatal patterning), has been a challenge1,2,3,4,5,6,19,20,21,22,23. Furthermore, even if bioactive peptides are generated, the screening of peptides that have potential functions in specific biological processes is another challenging task because the bioactive peptides produced are often a mixture of well-folded, active, and inactive peptides. This paper presents an efficient bioassay method to discover and characterize potential stomatal peptides. This method has been successful in identifying various important peptides that control different aspects of stomatal development in Arabidopsis, as well as the grass Brachypodium3,4,7,17. However, the major limitation of this technique is the variability in the phenotypic responses, which most likely occurs due to the use of seedlings with slightly different developmental stages and peptide solutions containing different amounts of bioactive peptides. Early attempts at this method involved the placement of wild-type seeds directly in the wells of a plate containing a peptide solution after stratification. Without the use of genotypes that have more and/or less epidermal cells (e.g., epf22, Figure 4) and 1 day old Arabidopsis seedlings germinated on 1/2 MS agar plates, a lower number of seedlings are able to grow and show clear phenotypic responses after the peptide application. In this protocol, the variability issue was addressed by using ~20 one-day-old Arabidopsis seedlings with two different developmental stages and by using two different concentrations of peptide solution (1 μM and 2.5 μM).
The epidermal phenotypes of the peptide-treated samples can also be analyzed by the first phenotypic analysis method presented using TBO-stained samples. This method allows for relatively easy quantitative phenotypic analysis because a relatively large area of each sample can be analyzed. Additionally, this method offers flexibility in sample preparation after harvesting samples prepared in the same conditions for imaging. However, the images taken by this method are generally not of the highest quality. On the other hand, confocal imaging after propidium iodide staining gives higher-quality images with epidermal details. However, this method only allows for a small area of tissue to be focused on for analysis. In addition, only a certain number of the live tissues prepared can be analyzed at once.
In conclusion, the phenotypic analysis presented here can be used for the quick and effective examination of the potential genes controlling stomatal patterning and differentiation, thus improving the understanding of the mechanisms of stomatal development and peptide function. In addition, this bioassay protocol can be used to identify other small molecules that have a role in epidermal tissue patterning and as an initial screening method to determine the structure of well-folded bioactive peptides.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This research was funded through the Natural Resources and Engineering Research Council of Canada (NSERC) Discovery program and Concordia University. K.B. is supported by the National Overseas Scholarship from India.
Materials
| 18 mm x 18 mm cover slip | VWR | 16004-326 | |
| 24-well sterile plates with lid | VWR | CA62406-183 | |
| 3M Micropore surgical tape | Fisher Scientific | 19-027-761 | Microporous surgical paper tape used to seal MS plates |
| 76 x 26 mm Microscope slide | TLG | GEW90-2575-03 | |
| Acetic acid, ≥99.8% | Fisher Scientific | A38-212 | |
| Agar | BioShop | AGR001.1 | |
| Bleech | Household bleach (e.g., Clorox) | ||
| Confocal microscope | Nikon | Nikon C2 operated by NIS-Elements | |
| Ethanol | Greenfield | P210EAAN | |
| FIJI | Open-srouce | (Fiji Is Just) ImageJ v2.1/1.5.3j | Downloaded from https://imagej.net/software/fiji/ |
| Forceps | Sigma-Aldrich | F6521 | |
| Gamborg's vitamin mixture | Cassson Labs | GBL01-100ML | Store at 4 °C |
| Glycerol | Fisher Scientific | G33-4 | |
| Growth chambers | Conviron, model E15 | 16h light cycle, set at 21°C with a light intensity of 120 µmol·m-2·s-1. | |
| Lights | HD Supply | 25272 | Fluorescent lights in growth chambers, Sylvania F72T12/CW/VHO 72"T12 VHO 4200K |
| Microcentrifuge tube | Fisher Scientific | 14-222-155 | Tubes in which Arabidopsis thaliana seeds are placed to perform sterilization |
| Microscope | Nikon | Nikon Eclipse TiE equipped with a DsRi2 digital camera | |
| Murashige and Skoog basal salts | Cassson Labs | MSP01-1LT | Store at 4 °C |
| Petri Dish 100 mm x 20 mm | Fisher Scientific | 08-757-11Z | Petri dishes in which MS media is poured for the purpose of growing Arabidopsis thaliana |
| Propidium Iodide | VWR | 39139-064 | |
| Scalpel | Fisher Scientific | 08-916-5A | |
| Sucrose | BioShop | SUC700.5 | |
| Toluidine blue O | Sigma-Aldrich | T3260-5G | |
| Tris base | Sigma-Aldrich | T1503 | |
| Triton X-100 | Sigma-Aldrich | T8787-100ML | |
| β-Estradiol | Sigma-Aldrich | E2758 |
References
- Hara, K., Kajita, R., Torii, K. U., Bergmann, D. C., Kakimoto, T. The secretory peptide gene EPF1 enforces the stomatal one-cell-spacing rule. Genes & Development. 21 (14), 1720-1725 (2007).
- Hara, K., et al. Epidermal cell density is autoregulated via a secretory peptide, EPIDERMAL PATTERNING FACTOR 2 in Arabidopsis leaves. Plant & Cell Physiology. 50 (6), 1019-1031 (2009).
- Kondo, T., et al. Stomatal density is controlled by a mesophyll-derived signaling molecule. Plant & Cell Physiology. 51 (1), 1-8 (2010).
- Sugano, S. S., et al. Stomagen positively regulates stomatal density in Arabidopsis. Nature. 463 (7278), 241-244 (2010).
- Hunt, L., Gray, J. E. The signaling peptide EPF2 controls asymmetric cell divisions during stomatal development. Current Biology. 19 (10), 864-869 (2009).
- Hunt, L., Bailey, K. J., Gray, J. E. The signalling peptide EPFL9 is a positive regulator of stomatal development. New Phytologist. 186 (3), 609-614 (2010).
- Lee, J. S., et al. Direct interaction of ligand-receptor pairs specifying stomatal patterning. Genes & Development. 26 (2), 126-136 (2012).
- Shpak, E. D., McAbee, J. M., Pillitteri, L. J., Torii, K. U. Stomatal patterning and differentiation by synergistic interactions of receptor kinases. Science. 309 (5732), 290-293 (2005).
- Nadeau, J. A., Sack, F. D. Control of stomatal distribution on the Arabidopsis leaf surface. Science. 296 (5573), 1697-1700 (2002).
- Meng, X., et al. Differential function of Arabidopsis SERK family receptor-like kinases in stomatal patterning. Current Biology. 25 (18), 2361-2372 (2015).
- Lampard, G. R., Macalister, C. A., Bergmann, D. C. Arabidopsis stomatal initiation is controlled by MAPK-mediated regulation of the bHLH SPEECHLESS. Science. 322 (5904), 1113-1116 (2008).
- Yang, M., Sack, F. D. The too many mouths and four lips mutations affect stomatal production in Arabidopsis. The Plant Cell. 7 (12), 2227-2239 (1995).
- Theunissen, J. D. An improved method for studying grass leaf epidermis. Stain Technology. 64 (5), 239-242 (1989).
- Venkata, B. P., et al. crw1–A novel maize mutant highly susceptible to foliar damage by the western corn rootworm beetle. PLoS One. 8 (8), 71296 (2013).
- Tucker, M. R., et al. Somatic small RNA pathways promote the mitotic events of megagametogenesis during female reproductive development in Arabidopsis. Development. 139 (8), 1399-1404 (2012).
- Ohki, S., Takeuchi, M., Mori, M. The NMR structure of stomagen reveals the basis of stomatal density regulation by plant peptide hormones. Nature Communications. 2, 512 (2011).
- Jangra, R., et al. Duplicated antagonistic EPF peptides optimize grass stomatal initiation. Development. 148 (16), (2021).
- Zhao, C., Craig, J. C., Petzold, H. E., Dickerman, A. W., Beers, E. P. The xylem and phloem transcriptomes from secondary tissues of the Arabidopsis root-hypocotyl. Plant Physiology. 138 (2), 803-818 (2005).
- Uchida, N., et al. Regulation of inflorescence architecture by intertissue layer ligand-receptor communication between endodermis and phloem. Proceedings of the National Academy of Sciences of the United States of America. 109 (16), 6337-6342 (2012).
- Tameshige, T., et al. A secreted peptide and its receptors shape the auxin response pattern and leaf margin morphogenesis. Current Biology. 26 (18), 2478-2485 (2016).
- Abrash, E. B., Davies, K. A., Bergmann, D. C. Generation of signaling specificity in Arabidopsis by spatially restricted buffering of ligand-receptor interactions. The Plant Cell. 23 (8), 2864-2879 (2011).
- Uchida, N., Tasaka, M. Regulation of plant vascular stem cells by endodermis-derived EPFL-family peptide hormones and phloem-expressed ERECTA-family receptor kinases. Journal of Experimental Botany. 64 (17), 5335-5343 (2013).
- Kosentka, P. Z., Overholt, A., Maradiaga, R., Mitoubsi, O., Shpak, E. D. EPFL Signals in the boundary region of the SAM restrict its size and promote leaf initiation. Plant Physiology. 179 (1), 265-279 (2019).

