Cultivation of Human Neural Progenitor Cells in a 3-dimensional Self-assembling Peptide Hydrogel
Summary
Here we describe the use of a self-assembling 3-dimensional scaffold to culture human neural progenitor cells. We present a protocol to release the cells from the scaffolds to be analysed subsequently e.g. by flow cytometry. This protocol might be adapted to other cell types to perform detailed mechanistically studies.
Abstract
The influence of 3-dimensional (3D) scaffolds on growth, proliferation and finally neuronal differentiation is of great interest in order to find new methods for cell-based and standardised therapies in neurological disorders or neurodegenerative diseases. 3D structures are expected to provide an environment much closer to the in vivo situation than 2D cultures. In the context of regenerative medicine, the combination of biomaterial scaffolds with neural stem and progenitor cells holds great promise as a therapeutic tool.1-5 Culture systems emulating a three dimensional environment have been shown to influence proliferation and differentiation in different types of stem and progenitor cells. Herein, the formation and functionalisation of the 3D-microenviroment is important to determine the survival and fate of the embedded cells.6-8 Here we used PuraMatrix9,10 (RADA16, PM), a peptide based hydrogel scaffold, which is well described and used to study the influence of a 3D-environment on different cell types.7,11-14 PuraMatrix can be customised easily and the synthetic fabrication of the nano-fibers provides a 3D-culture system of high reliability, which is in addition xeno-free.
Recently we have studied the influence of the PM-concentration on the formation of the scaffold.13 In this study the used concentrations of PM had a direct impact on the formation of the 3D-structure, which was demonstrated by atomic force microscopy. A subsequent analysis of the survival and differentiation of the hNPCs revealed an influence of the used concentrations of PM on the fate of the embedded cells. However, the analysis of survival or neuronal differentiation by means of immunofluorescence techniques posses some hurdles. To gain reliable data, one has to determine the total number of cells within a matrix to obtain the relative number of e.g. neuronal cells marked by βIII-tubulin. This prerequisites a technique to analyse the scaffolds in all 3-dimensions by a confocal microscope or a comparable technique like fluorescence microscopes able to take z-stacks of the specimen. Furthermore this kind of analysis is extremely time consuming.
Here we demonstrate a method to release cells from the 3D-scaffolds for the later analysis e.g. by flow cytometry. In this protocol human neural progenitor cells (hNPCs) of the ReNcell VM cell line (Millipore USA) were cultured and differentiated in 3D-scaffolds consisting of PuraMatrix (PM) or PuraMatrix supplemented with laminin (PML). In our hands a PM-concentration of 0.25% was optimal for the cultivation of the cells13, however the concentration might be adapted to other cell types.12 The released cells can be used for e.g. immunocytochemical studies and subsequently analysed by flow cytometry. This speeds up the analysis and more over, the obtained data rest upon a wider base, improving the reliability of the data.
Protocol
1. Part 1: Culture of hNPCs in PuraMatrix
- In advance to the generation of a scaffold with a PuraMatrix concentration of 0.25% without laminin one needs to prepare the following solutions
- Prepare a solution containing 20% sucrose and a solution containing 10% sucrose dissolved in sterile distilled water.
- For solution 1 mix 120 μl of the 20% sucrose solution with 120 μl of distilled water in a 1.5 ml conical tube.
- For solution 2 mix 60 μl of the PuraMatrix solution with 60 μl of 20% sucrose solution in a 1.5 ml conical tube.
For a Preparation of a scaffold with a PuraMatrix concentration of 0.25% supplemented with laminin (8 μg per 100 μl Matrix) one needs to prepare the following solutions.
- For solution 1 mix 72 μl of distilled water with 120 μl of 20% sucrose solution and 48 μl laminin in a 1.5 ml conical tube.
- For solution 2 mix 60 μl PuraMatrix solution with 60 μl of the 20% sucrose solution in a 1.5 ml conical tube.
- For the embedment of cells in the PuraMatrix prepare a 4-well cell culture plate and place a sterilised glass cover slip (diameter 13 mm) in two wells. Store the plate in the clean bench for the later use. Glass cover slips are only used for the immunocytochemical studies. If no immunocytochemical stainings are intended, the step can be skipped.
To prepare the hNPCs for encapsulation in 3D scaffolds one has to prepare the following solutions. Dilute the Benzonase in the Trypsin/EDTA solution, using a dilution factor of 1:10.000. For a Trypsin-Inhibitor/Benzonase solution mix: DMEM/F12 + Benzonase 25U/μl + 1% HSA + Trypsin-inhibitor (0.5mg/ml).
- Detach the cells by adding 500 μl Trypsin/Benzonase solution and incubate for 5 min in an incubator (37°C, 5% CO2). Stop the reaction with Trypsin-Inhibitor/Benzonase solution. Subsequently centrifuge the detached cells for 5 min at 3000 x g. Wash cells with 5 ml 10% Sucrose solution and centrifuge again. Discard the supernatant.
- Resuspend the cells in 10% sucrose solution, the final cell solution should contain 1×106 cells/ml.
The next steps, 1.8 to 1.11, should be done as fast as possible, because of the low pH of the PuraMatrix solution, which might be harmful for the cells.
- Mix 60 μl of cell suspension with solution 1.
- Add 120 μl of solution 1, containing the cells, to the conical tube with solution 2 and mix carefully.
- Place 100 μl of the mixture immediately on each cover slips resided in the 4-well plate.
- Add slowly 200 μl cell culture medium per well and after 2-3 minutes another 200 μl. This will initiate the self-assembly of the matrix.
- Incubate cells at 37 °C, for 1h in the clean bench on a heating plate or the heated area of your clean bench. As far as possible do not move the plate as this might harm the assembling of the matrices.
- Remove most of the medium but not all, to avoid the matrices falling dry, with a 1000 μl pipette. Add 500 μl of medium to wash matrices for 10 min. Finally remove the medium and add 500 μl fresh medium and place the culture plate in an incubator (37°C, 5% CO2). As the matrices are sensitive to shock the plates should be moved as less as possible and stored, if possible, in a separate incubator.
- The here used hNPCs were proliferated for 7 days in the 3D-scaffolds and differentiation was initiated by withdrawal of the growth factors 13. Medium was changed every 2-3 days.
2. Part 2: Immunocytochemical staining of entire matrices
- In advance to the staining procedure one needs to prepare: a blocking buffer solution consisting of 5% normal goat serum and 0.3% of Triton X-100 dissolved in PBS, a 4% paraformaldehyde solution based on 0.1 M PBS and if needed a solution with 4′,6-Diamidin-2′-phenylindoldihydrochlorid (DAPI) for a nuclei staining containing 100 ng DAPI/ml dissolved in PBS. Samples were mounted with a mixture of Mowiol and DABCO.
- To wash the matrices remove the medium with a 1000 μl pipette and wash the scaffolds with PBS once for 5 minutes.
- To fix the scaffolds remove the PBS and add 400 μl of paraformaldehyde (4% in 0.1 M PBS) to each well and incubate the matrices for 30 minutes at room temperature. Fixed Scaffolds can be stored in PBS supplemented with 0.02% NaN3 at 4°C for several weeks up to several months.
- In advance of a staining wash the scaffolds with PBS for 5 minutes.
- Incubate the scaffolds in blocking buffer solution. Blocking solution was changed three times every 2-3 hours and subsequently scaffolds were incubated in blocking solution over night at 4°C.
- Remove the blocking buffer solution and add the primary antibody dissolved in PBS supplemented with 1% normal goat serum and incubate the matrices over night at 4°C.
- Remove the primary antibody solution and wash scaffolds 4 times for 2 h and subsequently over night at 4°C with PBS.
- Remove the PBS and add the solution with the secondary antibody, dissolved in PBS supplemented with 1% normal goat serum, for 4 h at room temperature in the dark.
- Wash the samples with PBS 4-6 times for 1 h and subsequently over night at 4°C in the dark.
- For a nuclei staining one can add 400 μl of the DAPI solution for 30 min.
- To mount the samples one needs microscope slides. Add 50 μl Mowiol/Dabco on the microscope slides. Remove the cover slips from the 4-well plate and put them carefully on the mounting medium.
- Here we used a Biozero 8000 microscope (Keyence, Germany, Karlsruhe) to obtain single micrographs and z-stacks. Each stack contains 30 single pictures with a distance of 1-2 μm. Using the corresponding analyser software the blur inherent to fluorescence was removed before full projections of the z-stacks were produced.
3. Part 3: Release of hNPCs from the scaffolds for flow cytometry analysis
- In advance to the release of the cells from scaffolds wash the scaffolds with 500 μl HBSS.
- Transfer the scaffolds by means of a 1000 μl pipette to a 15 ml conical tube containing cell culture medium.
- To disrupt the scaffolds mechanically resuspend the cell solution several times with a 1000 μl pipette and subsequently centrifuge the solution at 3000 x g for 5 min.
- Remove the supernatant with a pipette, add 500 μl of Trypsin/Benzonase solution and resuspend the cell pellet several times.
- Incubate the cells for 5 min at 37°C in the Trypsin/Benzonase solution. To stop the reaction add 1 ml Trypsin-inhibitor/Benzonase solution and pipette up and down several times.
- Centrifuge the cell suspension 5 min at 3000 x g, remove the supernatant and wash the cells with 2 ml HBSS buffer. Repeat this step two times. This procedure will remove matrix debris.
- Pass the cell suspension solution trough a cell strainer (pore size 70 μm) to remove aggregates and collect the cells in cell culture medium. The obtained cells can be seeded again e.g. on cover slips for functional studies as patch clamp or fluometric measurements. To process the cells for flow cytometry, fix the cells immediately (see 4.1).
4. Part 4: Quantification of βIII-tubulin positive cells by flow cytometry
- Fix the released cells obtained in step 3.7 with 1% PFA for 15 min at room temperature.
- Centrifuge the cell suspension for 5 min at 3000 x g and remove the supernatant. If needed, the cells can be prepared to be stored at 4°C for later analysis. Therefore resuspend cells in wash buffer (PBS supplemented with 0.5% BSA and 0.02% Na-azide).
- To proceed with the preparation for the flow cytometry, centrifuge the cell suspension for 10 min at 350 x g, discard the supernatant and resuspend the cells in 25 μl of the antibody solution. Therefore the first antibody is mixed with saponin buffer at a concentration 1:100 e.g. βIII-tubulin. The saponin buffer consists of PBS with a saponin concentration of 0.5%, a BSA concentration of 0.5% and a Na-azide concentration of 0.02%.
- Incubate the cell suspension with the first antibody for 2 h at room temperature on a shaker. As a negative control prepare a sample without the first antibody.
- For the first washing step add 300 μl saponin buffer directly to the cells. Centrifuge the cell suspension for 5 min at 350 x g. For the second washing step, discard the supernatant, add 300 μl buffer and centrifuge the cell suspension for 5 min at 350 x g.
- Remove the supernatant and add 25 μl solution containing the secondary antibody (e.g. Alexa Fluor 647, 1:1000, dissolved in saponin buffer). Incubate cells with the secondary antibody 1h at room temperature, in the dark.
- For the first washing step add 300 μl saponin buffer directly to the cells. Centrifuge the cell suspension for 5 min at 350 x g. For the second washing step, discard the supernatant, add 300 μl buffer and centrifuge the cell suspension for 5 min at 350 x g.
- Discard the supernatant and resuspend the cells in 500 μl wash buffer for the flow cytometry analysis.
5. Representative Results
An example of hNPCs cultured in the self-assembling hydrogel scaffold PuraMatrix is shown in figure 1. The hNPCs grow in spheric like structures in unmodified PM. In this culture conditions, cells can hardly be recognised in a transmission light, although bundles of processes between the spheres are visible (fig 1A). Depending on the culture conditions of the used cell type, the matrix can be modified e.g. by adding laminin. For the hNPC cell line ReNcell VM (Millipore) laminin is necessary to induce a growth pattern with less dense aggregates but a more homogeneous distribution of the cells (fig 1B). Independent of modifications, the matrix can be used to study e.g. neuronal differentiation. Figure 1C presents a staining of the hNPCs for the neuronal marker βIII-tubulin. In this example the cells were grown for 7 days in the matrix and subsequently differentiated for 4 days, where differentiation was induced by withdrawal of the growth factors EGF and bFGF.13 Cell bodies and a dense network of processes built up by the cells can be recognised easily. However, it is obvious that a quantification of the cell number is time consuming, as a large number of pictures of different regions of the matrix has to be analysed, to obtain a reliable data base for a statistical analysis. In fig 1D one can see an example of cells released from the matrix and subsequently platted on cover slips. These cells might be used for functional studies.
The release of the cells from the 3D-scaffolds offers the possibility to analyse different parameters like expression of marker proteins or survival rate of the cells by AnnexinV/PI staining or a TUNEL assay. Figure 2 shows an example of a flow cytometry analysis of the percentage of βIII-tubulin+ cells. The negative control (cells not marked for βIII-tubulin) is shown in figure 2A. These controls are used to determine the gate for the later detection of βIII-tubulin+ cells (fig 2B). A comparison of a manual counting of cells and an analysis by flow cytometry is shown in figure 2C. The percentage of βIII-tubulin+ cells was determined by counting the total cell number (by means of a nuclear DAPI staining) and the number of βIII-tubulin+ cells in the fluorescence pictures.
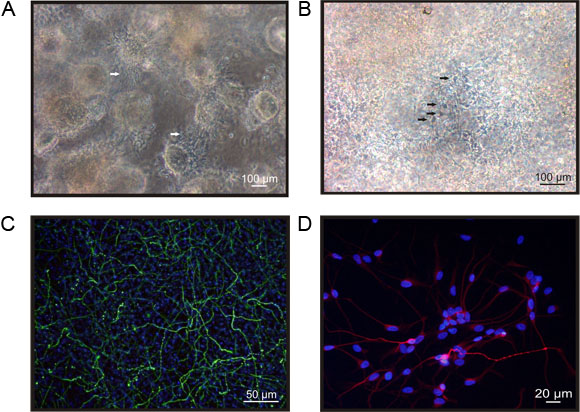
Figure 1. hNPCs cultured in the self-assembling peptide hydrogel PuraMatrix. A) hNPCs encapsulated in PuraMatrix (PM) grow in spheric, dense packed structures, where the diameter of the spheres can be up to several hundred μm. In between the spheres one can recognise bundles of processes built up by the cells (arrows). B) Cells encapsulated in PuraMatrix supplemented with laminin (PML) grow in less dense structures, more homogenously distributed. In the laminin supplemented matrix one can recognise single cells in less dense areas (arrows), however it is hardly possible to quantify the number of cells. C) hNPCs differentiated for 4 days in PML express neuronal marker like βIII-tubulin (green). To evaluate the percentage of positive cells one has to determine the total cell number by a nuclei staining like DAPI (blue). The microphotograph presents the full projection of a z-stack of pictures taken with Biozero-8000 microscope. D) Cells released from the matrix can be seeded on e.g. cover slips for further functional studies. The microphotograph shows cells cultured for 3d, subsequently to the release procedure. The staining for βIII-tubulin (red) reveals a comparable morphology to the cells hosted in the 3D scaffold.
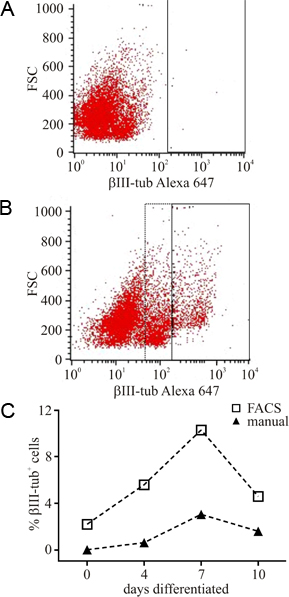
Figure 2. Flow cytometry analysis cells released from PuraMatrix. To overcome the time consuming quantification of microphotographs we used a protocol to release cells cultured in PuraMatrix giving access to faster analysis by flow cytometry. A) Unstained cells were used as negative control, to set the gate (black frame) for the subsequently analysis of e.g. βIII-tubulin cells. B) To quantify the percentage of βIII-tubulin+ cells the same gate, set in the negative control, was used. Positive cells appear in the right part of the x-axis, where also an intermediate population was observed (dotted frame), most likely representing cell debris. C) The comparison of manual counted cells and cells counted by flow cytometry revealed a much higher proportion of positive cells, where the time dependency of the number of βIII-tubulin+ was comparable, indicating the reliability of the flow cytometry protocol.
Discussion
The use of 3D-scaffolds offers the opportunity to study the development of different cell types in a cell culture situation closer to the in vivo situation. However, regarding the analysis of e.g. neuronal differentiation or functional studies one has to overcome some obstacles to gain reliable data for e.g. quantification of cell types.
Here we described the culture of hNPCs in the peptide hydrogel based scaffold PuraMatrix and the subsequently release of the cells to be used for the studies in a 2D situation providing an easy access to tools as FACS or functional assays. Recently we demonstrated the influence of the PuraMatrix concentration on the formation of the 3D-scaffold by atomic force microscopy and the influence on the survival and differentiation of the cells.13 However, one has to keep in mind that each cell type may need different culture conditions or matrices consisting of other materials e.g. matrigel or collagen.
The protocol to release the cells is based on a protocol provided by the manufacturer18 and is comparable with protocols used to prepare e.g. primary neuronal cultures where a mechanical isolation of the cells is followed by a digestion of surrounding material by enzymes. The cells tolerate this procedure and stay vital and built up processes and express neuronal marker like βIII-tubulin, once they are seeded again on a laminin coated surface. Here we performed flow cytometry studies with released cells to quantify the amount of βIII-tubulin+ cells. The comparison to a quantification done “manually” by counting cells in micrographs revealed a much higher proportion of βIII-tubulin+ cells. This is most likely due to the higher number of cells counted by the flow cytometer (50.000 per probe) in comparison to the manual analysis (several hundred per probe). However, the kinetics of the number of βIII-tubulin+ cells follows the same pattern in both samples, where we detect a decrease of βIII-tubulin+ cells over the time. This is in accordance with studies using the ReNcell VM cells.15-17 Other hurdles to overcome during a manual analysis are very dense areas of matrices which can hardly be analysed or high background of the matrix material resulting in an underestimation of the “real” cell number. We are convinced that this protocol can be adapted to other cell types providing a fast and reliable method to quantify several aspects of proliferation, differentiation or survival of the cells.
Disclosures
The authors have nothing to disclose.
Acknowledgements
The authors would like to thank Norman Krüger for his excellent technical support.
Materials
| Name of the reagent | Company | Catalogue number | Comments |
| PuraMatrix peptide hydrogel | BD Bioscience | 354250 | |
| Mouse laminin I | Cultrex | 400-2009090 | |
| Sucrose | Sigma | S9378-1KG | |
| Normal goat serum | Dako | X0907 | |
| Triton X 100 | Roth | 3051.3 | |
| PBS Dulbecco | Biochrom AG | L 1825 | |
| HBSS | Gibco | 14170-088 | Hanks’ Balanced Salt Solution 1X |
| βIII-tubulin antibody | Santa Cruz | Sc-51670 | Mouse, monoclonal, 1:500 |
| Alexa Fluor 488 | Invitrogen | A 11029 | Goat α mouse, 1:1000 |
| Alexa Fluor 568 | Invitrogen | A 11031 | Goat α mouse, 1:1000 |
| Alexa Fluor 647 | Invitrogen | A 21235 | Goat α mouse, 1:1000 |
| Mowiol 4-88 Reagent | Calbiochem | 475904 | |
| Dabco | Aldrich | D2,780-2 | 1,4-Diazabicyclo[2.2.2]octane 98% |
| Cell strainer | BD Biosciences | 352350 | 70 μm pore size |
| Saponin | Merck | 7695 | |
| Trypsin/ EDTA | GIBCO | 25300-054 | |
| Benzonase 250 U/μl | Merck | 1.01654.0001 | |
| Trypsin Inhibitor | Sigma | T6522 (500 mg) | |
| 20% HSA | Octapharma | Human-Albumin Kabi 20% |
References
- Zhang, S., Gelain, F., Zhao, X. Designer self-assembling peptide nanofiber scaffolds for 3D tissue cell cultures. Semin. Cancer. Biol. 15, 413-420 (2005).
- Gelain, F., Horii, A., Zhang, S. Designer self-assembling peptide scaffolds for 3-d tissue cell cultures and regenerative medicine. Macromol. Biosci. 7, 544-551 (2007).
- Blow, N. Cell Culture: building a better matrix. Nature. Methods. 6 (8), 619-622 (2009).
- Hauser, C. A., Zhang, S. Designer self-assembling peptide nanofiber biological materials. Chem. Soc. Rev. 39, 2780-2790 (2010).
- Teng, Y. D. Functional recovery following traumatic spinal cord injury mediated by a unique polymer scaffold seeded with neural stem cells. Proc. Natl. Acad. Sci. U. S. A. 99, 3024-3029 (2002).
- Silva, G. A. Selective differentiation of neural progenitor cells by high-epitope density nanofibers. Science. 303, 1352-1355 (2004).
- Gelain, F., Bottai, D., Vescovi, A., Zhang, S. Designer self-assembling peptide nanofiber scaffolds for adult mouse neural stem cell 3-dimensional cultures. PLoS. ONE. 1, e119-e119 (2006).
- Taraballi, F. Glycine-spacers influence functional motifs exposure and self-assembling propensity of functionalized substrates tailored for neural stem cell cultures. Front Neuroengineering. 3, 1-1 (2010).
- Zhang, S., Holmes, T., Lockshin, C., Rich, A. Spontaneous assembly of a self-complementary oligopeptide to form a stable macroscopic membrane. Proc. Natl. Acad. Sci. U. S. A. 90, 3334-3338 (1993).
- Zhang, S. Self-complementary oligopeptide matrices support mammalian cell attachment. Biomaterials. 16, 1385-1393 (1995).
- Horii, A., Wang, X., Gelain, F., Zhang, S. Biological designer self-assembling peptide nanofiber scaffolds significantly enhance osteoblast proliferation, differentiation and 3-D migration. PLoS. ONE. 2, e190-e190 (2007).
- Thonhoff, J. R., Lou, D. I., Jordan, P. M., Zhao, X., Wu, P. Compatibility of human fetal neural stem cells with hydrogel biomaterials in vitro. Brain. Res. 1187, 42-51 (2008).
- Ortinau, S. Effect of 3D-scaffold formation on differentiation and survival in human neural progenitor cells. Biomed. Eng. Online. 9, 70-70 (2010).
- Abu-Yousif, A. O., Rizvi, I., Evans, C. L., Celli, J. P., Hasan, T. PuraMatrix Encapsulation of Cancer Cells. J. Vis. Exp. (34), e1692-e1692 (2009).
- Morgan, P. J. Protection of neurons derived from human neural progenitor cells by veratridine. Neuroreport. 20, 1225-1229 (2009).
- Giese, A. K. Erythropoietin and the effect of oxygen during proliferation and differentiation of human neural progenitor cells. BMC. Cell Biol. 11, 94-94 (2010).
- Schmöle, A. C. Novel indolylmaleimide acts as GSK-3beta inhibitor in human neural progenitor cells. Bioorg. Med. Chem. 18, 6785-6795 (2010).
- PuraMatrix, B. D. Peptide Hydrogel. Guidelines for Use. Catalog No 354250, (2006).

