MicroRNA Detection in Prostate Tumors by Quantitative Real-time PCR (qPCR)
Summary
Quantitative Real Time polymerase chain reaction (qPCR) is a rapid and sensitive method to investigate the expression levels of various microRNA (miRNA) molecules in tumor samples. Using this method expression of hundreds of different miRNA molecules can be amplified, quantified, and analyzed from the same cDNA template.
Abstract
MicroRNAs (miRNAs) are single-stranded, 18–24 nucleotide long, non-coding RNA molecules. They are involved in virtually every cellular process including development1, apoptosis2, and cell cycle regulation3. MiRNAs are estimated to regulate the expression of 30% to 90% of human genes4 by binding to their target messenger RNAs (mRNAs)5. Widespread dysregulation of miRNAs has been reported in various diseases and cancer subtypes6. Due to their prevalence and unique structure, these small molecules are likely to be the next generation of biomarkers, therapeutic agents and/or targets.
Methods used to investigate miRNA expression include SYBR green I dye- based as well as Taqman-probe based qPCR. If miRNAs are to be effectively used in the clinical setting, it is imperative that their detection in fresh and/or archived clinical samples be accurate, reproducible, and specific. qPCR has been widely used for validating expression of miRNAs in whole genome analyses such as microarray studies7. The samples used in this protocol were from patients who underwent radical prostatectomy for clinically localized prostate cancer; however other tissues and cell lines can be substituted in. Prostate specimens were snap-frozen in liquid nitrogen after resection. Clinical variables and follow-up information for each patient were collected for subsequent analysis8.
Quantification of miRNA levels in prostate tumor samples. The main steps in qPCR analysis of tumors are: Total RNA extraction, cDNA synthesis, and detection of qPCR products using miRNA-specific primers. Total RNA, which includes mRNA, miRNA, and other small RNAs were extracted from specimens using TRIzol reagent. Qiagen’s miScript System was used to synthesize cDNA and perform qPCR (Figure 1). Endogenous miRNAs are not polyadenylated, therefore during the reverse transcription process, a poly(A) polymerase polyadenylates the miRNA. The miRNA is used as a template to synthesize cDNA using oligo-dT and Reverse Transcriptase. A universal tag sequence on the 5′ end of oligo-dT primers facilitates the amplification of cDNA in the PCR step. PCR product amplification is detected by the level of fluorescence emitted by SYBR Green, a dye which intercalates into double stranded DNA. Specific miRNA primers, along with a Universal Primer that binds to the universal tag sequence will amplify specific miRNA sequences.
The miScript Primer Assays are available for over a thousand human-specific miRNAs, and hundreds of murine-specific miRNAs. Relative quantification method was used here to quantify the expression of miRNAs. To correct for variability amongst different samples, expression levels of a target miRNA is normalized to the expression levels of a reference gene. The choice of a gene on which to normalize the expression of targets is critical in relative quantification method of analysis. Examples of reference genes typically used in this capacity are the small RNAs RNU6B, RNU44, and RNU48 as they are considered to be stably expressed across most samples. In this protocol, RNU6B is used as the reference gene.
Protocol
1. Prostate Sample Collection
- Collect the prostate samples at the time of prostatectomy. The specimen is oriented using anatomic landmarks. The prostate and seminal vesicles are painted as follows: right side green, left side blue.
- A random transverse midsection of the prostate is taken perpendicular to the rectal surface, frozen in liquid nitrogen, and stored at -80 °C9.
- Banked slices of specimens are photocopied, oriented (anterior, posterior, right and left), quadrisected. Sections are cut using the Cryostat.
- Sections are stained with H&E and reviewed by a pathologist to determine and delineate tumor versus normal areas on the stained slides and a corresponding image. The marked areas are used as a guide to indicate areas from which to extract the tumor tissue from which RNA will be extracted in the subsequent steps.
2. Isolating Total RNA, Including miRNA, from Samples
- Place frozen prostate samples on dry ice and referring to the delineated photocopy, cut out a small portion of the prostate tumor (between 50 to 100 mg).
- Homogenize the prostate tumor tissue in 1 mL of TRIzol Reagent. The quantities in the following steps are based on use of 1 mL of TRIzol Reagent.
Note: Here we have used TRIzol Reagent for extracting RNA, however other kits that isolate small RNA-containing total RNA can also be used.
- Incubate the homogenized samples for 5 minutes at room temperature.
- Add 0.2 mL of chloroform to the samples and shake vigorously for 15 seconds. Incubate samples for 3 minutes at room temperature, then centrifuge at 12,000 x g for 15 minutes at 4 °C.
- Transfer the colorless upper aqueous phase to fresh tubes, and add 0.5 mL of isopropyl alcohol. Incubate samples for 10 minutes at room temperature, then centrifuge at 12,000 x g for 10 minutes at 4 °C.
- Carefully aspirate the supernatant without disturbing the pellet containing the RNA. Wash the RNA pellet with 1 mL of 75% ethanol. Vortex the sample and re-sediment by centrifugation for 5 minutes at 7,500 x g at 4 °C.
- Carefully aspirate the supernatant and dry the RNA pellet for 5-10 minutes, making sure the RNA pellet is not completely dry. Re-dissolve in Nuclease-free water appropriate to pellet size. Measure the concentration of RNA using the NanoDrop 1000 spectrophotometer (measure absorbance at 260 nm and 280 nm).
- Check the quality and integrity of the RNA samples using Agilent Bioanalyzer.
3. Reverse Transcription of RNA
- Reverse transcription of RNA was performed using miScript Reverse Transcription Kit according to manufacturer’s instructions (Qiagen). This kit includes a reverse transcriptase and a poly(A) polymerase. The miScript RT Buffer includes Mg2+, dNTPs, oligo-dT primers, and random primers.
- Use between 10 pg and 1 μg of RNA to synthesize cDNA. If using more than 1 μg of RNA, scale up the reaction linearly to the appropriate volume.
- Prepare a master mix that contains 5X miScript RT Buffer (4 μl), miScript Reverse Transcription Mix (1 μl), and RNase-free water to bring reactions to final volume of 20 μl. Also include template RNA (up to 1 μg) in the master mix.
- Incubate the samples for 60 minutes at 37 °C followed immediately by an incubation for 5 minutes at 95 °C. This step can be performed in a PCR machine, heating block, or water bath. Thermocyclers are the most suitable and accurate method. Store the cDNA on ice for short term, and -20 °C for long term storage.
4. Generating a Standard Curve
- Prior to experiment with target miRNAs, a standard curve is generated by using cDNAs of known concentrations against their crossing points (CP) (Figure 2).
- Prepare a series of dilutions of 2-fold, 10-fold, 50-fold, 250-fold, and 1250-fold the original cDNA of a sample that is known to have a substantial expression of your gene of interest.
- Run the PCR as specified in Section 5 “Real-time PCR for detection of miRNA”, with the modification that cDNA is in serial dilutions not a static 40x dilution.
- Perform analysis using RelQuant software (Roche) to generate your standard curve.
Note: a new standard curve must be generated for each gene of interest.
5. Real-time PCR for Detection of miRNA
- Real time PCR for miRNAs was performed using miScript SYBR Green PCR Kit and miScript Primer Assay according to manufacturer’s instructions (Qiagen). Prepare a master mix containing 2x QuantiTect SYBR Green PCR Master Mix, 10x miScript Universal Primer, 10x miScript Primer Assay, and RNase-free water. Prepare a master mix for a 20 μl volume reaction.
- The Primer Assay is specific to the miRNA of interest. To reconstitute 10x miScript Primer Assay, centrifuge the vial briefly, and add 550 μl TE buffer, pH 8.0. Vortex the vial briefly to mix, aliquot primers to smaller volumes, and store at -20 °C. Two primers are required: primers for the target gene and the reference gene. RNU6B is usedas the reference gene.
- Dilute the cDNA 40x and store extra aliquots at -20 °C.
- cDNA serves as the template for the PCR. Use 2 μl of 40x diluted cDNA and dispense to the 20 μl light cycler capillaries (Roche).
- Add 18 μl of the master mix to each capillary, and centrifuge using a capillary adapter.
- Place the capillaries in a capillary based Real-Time cycler, such as LightCycler 3.5 Real-Time PCR System with a 32-capillary carousel format.
- Run the PCR cycling program as follows:
To activate HotStarTaq Polymerase that is in the 2x QuantiTect SYBR Green PCR Master Mix, pre-incubate at 95 °C for 15 minutes.
Followed by 50 cycles of:
Denaturation, 15 s, 94 °C;
Annealing, 30 s, 55 °C;
Extension, 30 s, 70 °C. - Select a sample to be the calibrator, and set its normalized target amount to 1. Compare the relative expression of the miRNA in all the other samples to the calibrator.
Note: Within a study, the same calibrating sample should be used to maintain consistency of results.
6. Analyzing Data
- Amplification curves for the PCR reactions are depicted graphically and numerically by Molecular Biochemicals LightCycler Software version 3.5 (Roche). Quantify reactions in the “Quantification” tab, and export the data to a text file.
- Import the data to the RelQuant analysis software (Roche) to generate quantification results. Import separate files for target gene, reference gene, and standard curve data.
- Specify the position of calibrator for both target and reference gene. Also specify positions of the samples. Data is expressed as the target to reference ratio of different samples divided by the target to reference ratio of the calibrator. The standard curve previously generated for a particular miRNA and housekeeping gene is used as a reference standard for extrapolating quantitative data for miRNA targets of unknown concentrations.
- Three replicates of samples are analyzed as a group and mean concentrations and standard deviations of the triplicate is calculated. If one of the triplicates is inconsistent with the rest of the set, it will be excluded by the program.
7. Representative Results
An example of qPCR analysis on prostate samples is shown in Figure 3. Results are depicted numerically, as well as graphically. The graphs showing the expression levels of the reference gene, U6, begin exponential amplification at about cycle 20, while expression of the target gene, miR-98, showed delayed amplification approximately at cycle 25. The data from this experiment was exported as text file and analyzed by RelQuant analysis software. Positions of the capillaries containing the calibrator and samples are specified. Figure 4 illustrates how the calibrator is set to be 1, and the expression of other samples relative to the calibrator.
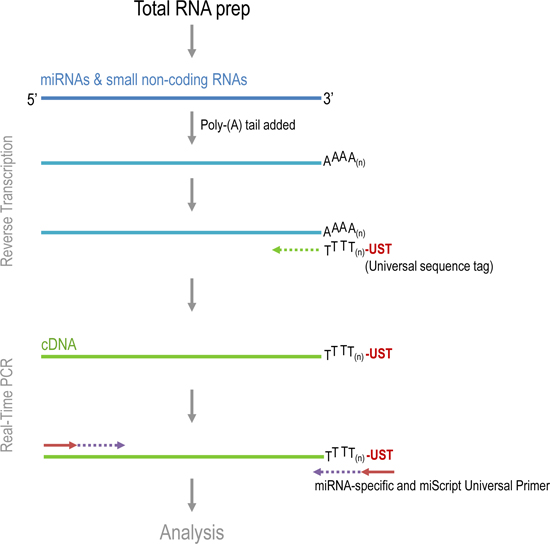
Figure 1. Various steps in miScript reverse-transcription and Real-time PCR.
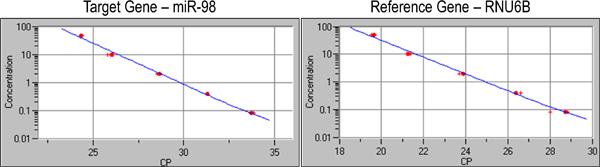
Figure 2. A standard curve is generated by using a series of dilutions of 2-fold, 10-fold, 50-fold, 250-fold, and 1250-fold the original cDNA sample.
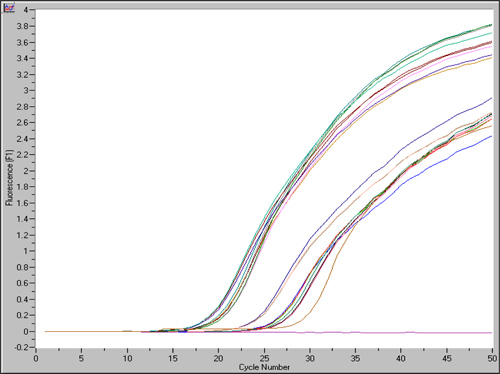
Figure 3. Roche Molecular Biochemicals LightCycler Software shows the entire information of the experiment graphically and by text. Quantitative Real-time PCR amplification plots show increased in fluorescence from different samples.
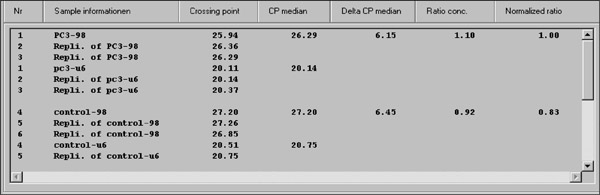
Figure 4. Data were quantified using RelQuant LightCycler analysis software. Usually, three replicates of samples are analyzed as a group and samples that produce clearly inconsistent results are excluded and mean concentrations and standard deviations of the triplicate is calculated.
Discussion
Aberrant expressions of some miRNAs have been consistently found in prostate tumors when compared to normal tissue10, and some of these miRNAs have been named as potential novel therapeutic agents against prostate cancer11. Hence the aberrant expression levels of miRNAs can be useful diagnostic and/or prognostic biomarkers. The Real-Time qPCR methodology presented here provides an assay for accurate quantification of miRNA levels in prostate tumor tissues. The miScript PCR system used can detect single nucleotide differences between mature miRNAs. The miScript miRNA qPCR Assays, however, are not intended for detection of stem loop precursor miRNAs, for which different miScript Precursor Assays are available.
The reliability of this technique depends on the quality of the input RNA, therefore concentration, integrity, and purity of RNA should be tested prior to Real-Time PCR. Moreover, ribonucleases are very stable and readily degrade RNA, thus extra caution should be taken in handling of the RNA. All reactions should be set up on ice to minimize RNA degradation. RNase inhibitors can also be added to the reaction prior to reverse transcription. Gloves should be frequently changed, while sterile and disposable plasticware must be used throughout the procedure.
If there is no PCR product or the amplification curve is detected late in Real-Time PCR, try increasing the number of PCR cycles, and make sure that the cycling program includes activation of HotStarTaq DNA polymerase for 15 minutes at 95 °C. Low amplification might also be due to inadequate starting cDNA template, therefore try to increase the amount of cDNA. Late amplification can also represent a false positive.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was funded by the Canadian Cancer Society Research Institute, grant no. 019038.
Materials
| Name of the reagent | Company | Catalogue number |
| TRIzol Reagent | Invitrogen | 15596 |
| miScript Reverse Transcription Kit | Qiagen | 218061 |
| miScript Primer Assays | Qiagen | Experiment specific |
| miScript SYBR Green PCR Kit | Qiagen | 218073 |
| LightCycler 3.5 Real-Time PCR System | Roche | |
| Light Cycler Capillaries | Roche | 04929292001 |
| NanoDrop 1000 spectrophotometer | Thermo Scientific | 2538 |
| Agilent 2100 Bioanalyzer | G2943CA |
References
- Reinhart, B. J. MicroRNAs in plants. Genes Dev. 16, 1616-1616 (2002).
- Xu, P. The Drosophila microRNA Mir-14 suppresses cell death and is required for normal fat metabolism. Curr. Biol. 13, 790 (2003).
- Bueno, M. J., Perez, d. e. C. a. s. t. r. o., I, ., Malumbres, M. Control of cell proliferation pathways by microRNAs. Cell Cycle. 7, 3143-3143 (2008).
- Friedman, R. C. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92 (2009).
- Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 116, 281-281 (2004).
- Croce, C. M. Causes and consequences of microRNA dysregulation in cancer. Nat. Rev. Genet. 10, 704 (2009).
- Coppola, V., De, M. R., Bonci, D. MicroRNAs and prostate cancer. Endocr. Relat. Cancer. 17, 1-1 (2010).
- Gordanpour, A. miR-221 Is down-regulated in TMPRSS2:ERG fusion-positive prostate cancer. Anticancer Res. 31, 403-403 (2011).
- Nam, R. K. Expression of the TMPRSS2:ERG fusion gene predicts cancer recurrence after surgery for localised prostate cancer. Br. J. Cancer. 97, 1690 (2007).
- Ambs, S. Genomic profiling of microRNA and messenger RNA reveals deregulated microRNA expression in prostate cancer. Cancer Res. 68, 6162 (2008).
- Liu, C. The microRNA miR-34a inhibits prostate cancer stem cells and metastasis by directly repressing CD44. Nat. Med. 17, 211 (2011).

