High-throughput CRISPR Vector Construction and Characterization of DNA Modifications by Generation of Tomato Hairy Roots
Summary
Using DNA assembly, multiple CRISPR vectors can be constructed in parallel in a single cloning reaction, making the construction of large numbers of CRISPR vectors a simple task. Tomato hairy roots are an excellent model system to validate CRISPR vectors and generate mutant materials.
Abstract
Targeted DNA mutations generated by vectors with clustered regularly interspaced short palindromic repeats (CRISPR)/Cas9 technology have proven useful for functional genomics studies. While most cloning strategies are simple to perform, they generally use multiple steps and can require several days to generate the ultimate constructs. The method presented here is based on DNA assembly and can produce fully functional CRISPR vectors in a single cloning reaction. Vector construction can also be pooled, further increasing the efficiency and utility of the process. A modification of the method is used to create CRISPR vectors with multiple gene targets. CRISPR vectors are then transformed into tomato hairy roots to generate transgenic materials with targeted DNA modifications. Hairy roots are a useful system for testing vector functionality as they are technically simple to generate and amenable to large-scale production. The methods presented here will have wide application as they can be used to generate a variety of CRISPR vectors and be used in a wide range of plant species.
Introduction
The ability to generate targeted DNA modifications with CRISPR/Cas9 has great potential for functional genomics studies. There are two components of the CRISPR/Cas9 system; the Cas9 nuclease, derived from Staphylococcus pyogenes and an approximately 100-nt guide RNA (gRNA) molecule that directs Cas9 to the targeted DNA site(s)1. Target recognition is conferred by the first ~20-nt of the gRNA, which allows for high-throughput production of targeting vectors2,3. Most organisms that can be engineered, already have been with CRISPR/Cas9 technology4,5.
In plants, constitutive promoters, such as the CaMV 35S promoter, are commonly used to drive the expression of the Cas9 nuclease6. The gRNAs are expressed using the RNA polymerase III U6 or U3 promoters which restricts the first base of the gRNA to either a G, for U6, or A for U3, for efficient transcription. However RNA polymerase II promoters, which are free of these restrictions, have also been used7,8.
Different gRNAs induce DNA mutations with different efficiencies, and so it can be important to first validate CRISPR vectors before investing in whole-plant transformations or setting up extensive phenotypic screens. Transient expression of CRISPR constructs in plants, by using agroinfiltration for example, generally results in a lower frequency of DNA modification as compared to stable plants6, making detection of mutations difficult and phenotypic assays impractical with such approaches. So-called hairy roots are a convenient, alternative system since a large number of independent, stably transformed materials can be generated within weeks, as opposed to months for stable plants. CRISPR vectors are very effective at inducing DNA mutations in hairy roots9,10.
DNA assembly methods efficiently ligate DNA fragments containing overlapping ends11. A major advantage of some DNA assembly methods is the ability to incorporate ssDNA (i.e., oligos) into the assembled products. Since gRNAs are only ~20-nt long and new targets can be made with synthesized oligos, these DNA assembly methods are well suited to CRISPR cloning. The protocols described here are based on the p201 series of CRISPR vectors that has been successfully used in soybean10, poplar12 and now tomato. The cloning procedure presented offers several advantages over the current cloning method10. Namely, fully functional vectors can be generated in a single cloning reaction in a single day. Vector construction can also be pooled to generate multiple CRISPR vectors in parallel, further reducing hands-on time and material costs. We also present a protocol for the generation of tomato hairy roots as an efficient method to produce transgenic materials with targeted gene deletions. Hairy roots are used to validate the CRISPR vectors and provide material for subsequent experiments.
Protocol
1. Guide RNA Design and Vector Construction
- Identify target sequences for the genes of interest. There are a variety of online CRISPR target-finding programs suitable for this step13,14.
NOTE: Here we use the GN20GG target motif, but other designs may be suitable depending on the application or vector system used. - Design 60-mer gRNA oligos to include the GN19 portion of the target motifs flanked by 5' and 3' 20-nt sequences that are required for DNA assembly. The final 60-mer motif is: 5'-TCAAGCGAACCAGTAGGCTT-GN19-GTTTTAGAGCTAGAAATAGC-3'.
NOTE: Standard, desalted primers work fine. - Prepare the four DNAs for assembly (Figure 1). See supplementary file for DNA sequences.
- Digest 1 – 5 µg of p201N:Cas910 plasmid with the restriction enzyme SpeI in 1x Buffer 4 at 37 °C for 2 hr. Column-purify the digest according to manufacturer's instructions. Resuspend in ~15 µl of 10 mM Tris-HCl, and quantify by UV spectrophotometry.
- Perform a second digest with the restriction enzyme SwaI in 1x Buffer 3.1 at 25 °C for 2 hr. Check 100 – 200 ng on a 0.8% agarose gel to confirm complete digestion.
NOTE: A correctly digested plasmid will have a single band at 14,313 bp. Note: Heat inactivation of the enzyme and dephosphorylation are not required. Digested plasmid may be stored at -20 °C. - PCR amplify the Medicago truncatula (Mt) U6 promoter and scaffold DNAs from the pUC gRNA Shuttle10 plasmid using the primers SwaI_MtU6F / MtU6R and ScaffoldF / SpeI_ScaffoldR, respectively. Use a high-fidelity polymerase with the PCR conditions: 95 °C for 3 min; 31 cycles (98 °C for 20 sec, 60 °C for 15 sec, 72 °C for 30 sec); and 72 °C for 5 min.
- Visualize ~3 µl aliquots of PCR products on a 1% agarose gel to confirm amplification. The MtU6 amplicon is 377 bp and the scaffold amplicon is 106 bp. Column-purify the remaining PCR products according to manufacturer's instructions, quantify by UV spectrophotometry and store at -20 °C.
- Resuspend the entire tube of gRNA oligos to 100 µM in laboratory-grade water. Add 1 µl of 100 µM oligo to 500 µl of 1x Buffer 2.1. Mix well. If pooling multiple oligos, add 1 µl of each 100 µM oligo to the 1x Buffer 2.1 tube. Diluted oligos can be stored at -20 °C.
- Program a thermal cycler to hold at 50 °C, using a heated lid. Combine 100 ng (0.011 pmol) of linearized vector, 50 ng (0.2 pmol) of MtU6 promoter, 12 ng (0.2 pmol) of scaffold, 1 µl (0.2 pmol) of diluted gRNA oligo(s), and water to a volume of 5 µl. Add 5 µl of 2x high-fidelity DNA assembly master mix, mix well and spin down. Incubate reaction for 60 min in the 50 °C thermal cycler. Then place the reaction on ice.
Note: Reaction volumes can be increased to accommodate low DNA yields. - Transform 2 µl of the reaction into competent E. coli cells using standard techniques and plate on LB supplemented with 50 mg L-1 kanamycin (Kan50). Grow plated-cells at 37 °C overnight. Save the remaining DNA assembly reaction at -20 °C.
- Colony screen by PCR using the primers StUbi3P218R and ISceIR. Dilute an aliquot of the remaining DNA assembly reaction with water 1:5. Use 1 µl as a positive control for the colony screen and 1 ng of circular p201N:Cas9 plasmid as a no-insert control.
- PCR amplify with the conditions; 95 °C for 3 min; 31 cycles (95 °C for 30 sec, 58 °C for 30 sec, 72 °C for 30 sec); and at 72 °C for 5 min. Visualize PCR products on a 1% agarose gel. Ensure that correct insertions have a band at 725 bp and vectors without inserts (e.g., uncut vector) will have a 310-bp band.
- Grow positive colonies in LB Kan50 liquid cultures overnight at 37 °C and purify plasmids according to manufacturer's instructions. Sanger sequence plasmids with the StUbi3P218R primer and align chromatograms to the MtU6 promoter, target, and scaffold sequences to ensure no errors were introduced during cloning.
- Perform a diagnostic digest by digesting ~1 µg of plasmid with EcoRV and StyI. Visualize on a 0.8% agarose gel. Fragments (in bp) are: 4192, 2001, 1743, 1544, 1381, 995, 831, 702, 460, 423, 318, and 174.
- Use DNA assembly to construct vectors with multiple gRNAs.
- To construct a vector with two gRNA cassettes, PCR amplify the first-position gRNA with the primers SwaI_MtU6F / UNS1_ScaffoldR and the second-position gRNA with the primers UNS1_MtU6F / SpeI_ScaffoldR from 1 ng of each of the respective plasmids constructed in steps 1.1 – 1.8. Use the PCR conditions in step 1.3.3. Visualize ~3 µl aliquots of PCR products on a 1% agarose gel to confirm amplification. Amplicons are ~500 bp.
- Combine and mix approximately equal amounts of un-purified PCR products in a 200 µl tube (typically 3 – 4 µl of each). For assembly, add 1 µl combined PCR products, 50 ng of SwaI and SpeI linearized p201N:Cas9, laboratory-grade water to 2.5 µl and 2.5 µl of 2x high-fidelity DNA assembly master mix. Incubate in a 50 °C thermal cycler for 15 min.
Note: The DNA assembly manuals recommends a 15 min incubation for 2 – 3 fragments. - Transform 1 – 2 µl into competent E. coli cells and plate on LB Kan50. Grow plated cells at 37 °C overnight. Save the remaining reaction at -20 °C.
- Colony screen by PCR with primers StUbi3P218R and ISceIR with the same conditions as step 1.6. Correct clones will have a 1,200 bp amplicon. Grow positive colonies in LB Kan50 liquid cultures overnight and purify plasmids.
- Sanger sequence plasmids with the primers StUbi3P218R (covers first-position gRNA) and p201R (covers second-position gRNA). Align chromatograms to the MtU6 promoter, target, and scaffold sequences. Diagnostic digest with EcoRV and StyI will result in the following fragments (in bp): 4192, 2476, 1743, 1544, 1381, 995, 831, 702, 460, 423, 318, 174. Bolded fragment contains the second gRNA.
- After meeting all the quality-control expectations, transform plasmids into Agrobacterium rhizogenes strain ARqua115. Add 1 µl of the plasmid prep to 50 µl of electrocompetent cells and electroporate in a 1 mm cuvette with the settings: 2.4 kV, 25 µF, and 200 Ω. Recover cells in ~500 µl of SOC and shake at 28 °C for 2 hr. Plate on LB Kan50 and grow at 28 °C for 2 days. Perform colony screen using the same primers and PCR conditions as 1.3.3. Make glycerol stocks from positive clones.
2. Hairy Root Transformation
- Sterilize tomato seeds in 20% household bleach for 15 min, with constant mixing. In a laminar flow hood, remove the bleach and wash in sterile laboratory-grade water 3 times.
- Plate 30 seeds in a GA-7 box containing ½ MS media (2.22 g L-1 MS salts + Gamborg Vitamins, 10 g L-1 sucrose, 3 g L-1 gellan gum, pH 5.8). Germinate for ~2 days in the dark, then move GA-7 boxes to light. Grow seedlings for ~4 more days in the light.
- The day before transformation, streak out A. rhizogenes cultures on solid LB Kan50 and grow at 28 °C overnight.
- Day of transformation, in the laminar flow hood assemble sterile materials: 12 ml culture tubes, 50 ml conical tube, ½ MS liquid (no gellan gum), 200 µM acetosyringone, filter paper, Petri dishes, forceps and scalpel, pipettes and pipette tips. Add 25 µl of acetosyringone to 50 ml of ½ MS and pour ~ 6 ml into each culture tube.
- Use a bent 200 µl tip to scrape A. rhizogenes cells from the plate and resuspend in the 6 ml of ½ MS liquid. Vortex the tube to completely resuspend the cells. After resuspending cells from each vector, measure the optical density (O.D.) of 1 ml of cells at a fixed wavelength of 600 nM. Ensure O.D. is between 0.2 and 0.4; dilute or add more cells as necessary. Repeat for each construct.
- Add ~ 2 ml of ½ MS liquid to a Petri dish with a sterile filter paper. Excise cotyledons from the seedlings and place on the moistened filter paper. Once all cotyledons have been collected, cut the distal ~1 cm off of the cotyledons, resulting in cotyledon pieces with two cut ends. Add explants to A. rhizogenes solutions, mix, and incubate for 20 min with occasional inverting.
Note: Approximately 12 explants per construct are routinely used, though as many as 80 explants have been inoculated in 6 ml of A. rhizogenes solution. - During the A. rhizogenes incubation, add filter paper to Petri dishes, one per construct transformed. Also set ½ MS solid media, no antibiotics, in the laminar flow hood to dry.
- Scoop cotyledons out of A. rhizogenes solution with sterile forceps and place on dry filter paper. Cover with Petri lid to ensure tissues do not dry out while proceeding to the next transformation. Blot cotyledons on the filter paper and transfer, abaxial side up, to ½ MS media. Wrap plates with surgical tape and co-cultivate in the dark at room temperature for 2 days.
- After co-cultivation, transfer cotyledons, abaxial side up, to ½ MS media supplemented with 300 mg L-1 ticarcillin/clavulanic acid (Tim300, to prevent the growth of residual A. rhizogenes) and Kan50 (for plant selection) and wrap with surgical tape. Maintain cultures under fluorescent lights at room temperature with a 16-hour photoperiod.
- After ~1.5 – 2 weeks roots at least 2 cm in length are excised from the cotyledons using sterile forceps and a scalpel and transferred to ½ MS Kan50 Tim300 media. Transfer 10 to 15 roots to a single plate. Mark the position of the root tips with a marker, wrap with surgical tape, and maintain in culture room.
- After one week, transformed roots are seen growing on the selective media. Harvest a subsample of transformed roots for DNA extraction using the preferred DNA-extraction method. Note: Plates with cotyledons can be kept for 2 – 3 weekly harvests, after which point the roots grow into a tangled mess and are difficult to isolate. Non harvested root tissues can be maintained indefinitely. A variety of assays can be performed on the isolated DNA samples to determine DNA mutation frequency and type. Results from a few such assays are described below.
Representative Results
CRISPR vector construction with DNA assembly typically generates tens to hundreds of independent clones. Colony screening by PCR easily identifies correct clones and can distinguish between plasmids with and without inserts (Figure 2A) which is useful for troubleshooting. Typically, all of the clones contain an insert and a user may opt to skip the colony screening steps altogether. Diagnostic digests (Figure 2B) and Sanger sequencing are used for quality control. When errors do occur, they are usually observed in the overlapping DNA regions, i.e., the 5' end of the MtU6 promoter, the gRNA, or the 3' end of the scaffold sequence. If multiple gRNA oligos are pooled into a single reaction, the gRNAs are determined by Sanger sequencing. The incorporation of individual gRNAs is evenly distributed and most of the gRNAs can be isolated from a single transformation (Figure 2C). The high-fidelity DNA assembly mix is preferred over the standard DNA assembly mix as the high-fidelity version is less error-prone (Figure 2C).
Tomato hairy roots are usually observed one to two weeks after transformation (Figure 3A). Roots ≥2 cm in length are excised and transferred to selective medium and kanamycin-resistant roots can be identified one week later (Figure 3B). Most roots will grow to some degree on selective media and root growth in transformed roots can be variable; if possible, select the most vigorously growing roots for analysis. One to two kanamycin-resistant roots are typically recovered per cotyledon inoculated. With regular transfers, transformed roots can be maintained indefinitely on selective media.
DNA is extracted from kanamycin-resistant roots and PCR is used to amplify the target region(s). PCR products can be used in a variety of assays to determine mutation efficiency, such as cloning and sequencing (Figure 3C), restriction fragment length polymorphism analysis, polyacrylamide gel electrophoresis16 (Figure 3D), T7E1 endonuclease16, or high-throughput amplicon sequencing10. As reported in other CRISPR-plant systems6, depending on the gRNA/target sequence selected, a range of mutations and mutation efficiencies may be observed.
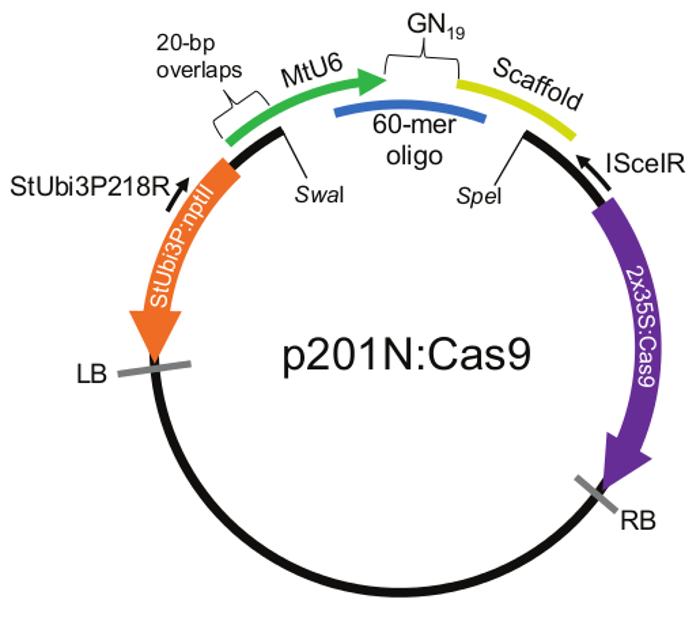
Figure 1. Principle of CRISPR Assembly. In the DNA assembly reaction, four DNAs are mixed together; p201N:Cas9 linearized with SwaI and SpeI (black circle), MtU6 promoter (green arrow), Scaffold (yellow bar) and the 60-mer gRNA oligo containing the GN19 target sequence (blue bar). The ends of each DNA molecule overlaps 20 bp with the adjacent piece. The DNA assembly reaction recombines these DNAs into a single, circular molecule. The StUbi3p218R and ISceIR primer binding sites are indicated with black arrows, the 2x35S:Cas9 cassette is purple, the StUbi3:nptII cassette is orange and left and right borders are indicated with grey bars. Note: Vector map is not to scale. Please click here to view a larger version of this figure.
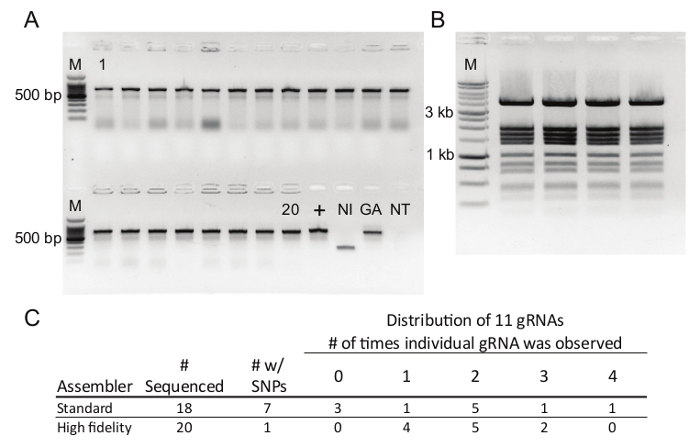
Figure 2. Colony Screening and Plasmid Quality Control. A) Colony screening of 20 clones from DNA assembly and transformation. M, 100-bp ladder; +, p201N:Cas9:GFPT2 positive control; NI, p201N:Cas9 no-insert control; GA, diluted DNA assembly reaction; NT, no-template control. B) Representative digest of four p201N:Cas9:gRNA vectors with the restriction enzymes EcoRV and StyI. M, 2-log ladder. C) Comparison of fidelity of the standard and high-fidelity DNA assembly mixes and distribution of recovered vectors when 11 gRNA oligos were pooled into a single reaction. Please click here to view a larger version of this figure.
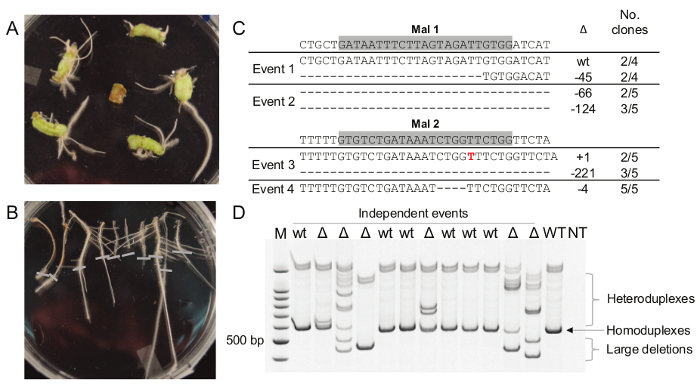
Figure 3. Hairy Root Cultures and Detection of DNA Modification. A) Hairy roots can be seen emerging from cotyledons 11 days after transformation. B) Roots are selected on ½ MS Kan50 media for approximately one week. Grey bars denote position of root tips at time of plating. C) Example of cloning and sequencing to determine DNA mutations with two different gene targets in four different hairy-root events. Grey box denotes GN20GG target sequence, Δ indicates type of mutation, and the number of clones with the indicated mutation. D) Example of polyacrylamide gel electrophoresis to determine DNA mutations. Large deletions and heteroduplex bands indicate the presence of DNA mutations at the target sequences. Six of 12 independent events were scored as mutants (Δ). WT, wild-type control; NT, no-template control. Please click here to view a larger version of this figure.
Discussion
Since DNA assembly is used to recombine any overlapping DNA sequences, this cloning method can be applied to any CRISPR vector construction. Most CRISPR cloning schemes use either gene synthesis of the gRNA, type IIS restriction enzymes17,18, or overlap-extension PCR19. Each of these techniques has inherent advantages and disadvantages, but they typically require multiple hands-on cloning steps. The primary advantage of the cloning method presented here is that the entire process occurs in a single, one-hour reaction and gRNA oligos can be pooled to further streamline the cloning procedure. Furthermore, the DNA assembly mechanism is independent of restriction enzymes, which makes this method compatible with virtually any DNA sequence. Conveniently, a master mix of the invariable DNA components (linearized vector, MtU6 promoter, and scaffold) can be made and stored at -20 °C for later assemblies.
While we use the GN20GG gRNA target motif, there are several alternative gRNA target motifs that can be used17. Shorter gRNAs have been demonstrated to increase specificity in animal cells20. By using a 5′ G with a U6 promoter in the gRNA sequence (or A in the case of a U3 promoter), regardless if it matches the intended target sequence, the more abundant target motif N21GG may be used. However, the GN20GG target motif does not present a major technical limitation as several suitable targets can be found in most plant genes. The cloning scheme presented here can be easily modified to accommodate any gRNA design; simply substitute bases in the gRNA oligo.
Another advantage of the DNA assembly cloning system is that the same reagents can be used to insert multiple gRNAs into a single vector. A protocol for two gRNAs is presented here, and with the same methodology, using unique nucleotide sequences21 as spacers between gRNAs, it should be possible to assemble six gRNAs or more in a single cloning reaction.
As presented in step 1.9, the sequential cloning of gRNA vectors into a final multi-gRNA vector will take several days to perform because of the multiple cloning and screening steps. A modification of this protocol permits the construction of multi-gRNA vectors in a single day by bypassing the single-gRNA vector cloning step. First assemble individual gRNAs with just the MtU6 amplicon, gRNA oligo, and scaffold amplicon. Individual reactions must be performed for each gRNA. Following step 1.4, omit the linearized vector and reduce the 50 °C incubation to 15 minutes. A 1:5 dilution of the reactions can be used as template in step 1.9.1.
A potential problem with step 1.9 is the carry-over of residual single-gRNA plasmids from the PCR amplification of the gRNAs since all plasmids are selected with kanamycin. While we have never observed single-gRNA vectors in any of our colony screens following the specified protocol, gel purification of the PCR products or DpnI treatment of the PCR reactions could be used should the problem occur.
The ability to generate CRISPR vectors in a high-throughput manner necessitates a high-throughput method of generating transgenic materials to use the CRISPR vectors. Hairy roots are well suited for this task as they are technically simple to generate, require relatively little laboratory space to maintain and can be produced in a wide range of plant species22,23. However one limitation is that hairy roots are generally not used to generate whole plants and root systems are not suitable models for investigating most above-ground phenotypes. The former limitation may be advantageous when targeting genes that are null-lethal in whole-plant systems.
An important consideration for any transformation experiment is that results may vary with the different genotypes of plants and bacterial strains used. We routinely induce hairy roots in the cultivar (cv.) Rio Grande but have had difficulty obtaining hairy roots with cv. Moneymaker. Here, we report the use of the A. rhizogenes strain ARqua1, but it should be noted that strain ATCC 15834 has also been reported to work in tomato9.
Another critical consideration for hairy root transformation is the potential for overgrowth of A. rhizogenes on the cotyledon explants. Bacterial overgrowth typically leads to cotyledon death and low transformation efficiencies. Cotyledons must be blotted dry after soaking with A. rhizogenes. Likewise, co-cultivation should be limited to two days to prevent excessive bacterial overgrowth. Some bacterial growth is typically observed on co-cultivation plates, but should be absent once cotyledons are transferred to media containing ticarcillin/clavulanic acid.
The ease of constructing CRISPR vectors and generating hairy roots enables large-scale reverse-genetic screens, similar to those that are used in animal cell culture systems2. Hairy roots have been used to study a variety of plant functions including root development9, symbiotic associations24,25, and nematode resistance26 and the use of hairy roots for screening immunity-related genes is an area of active research in our lab. In addition to overexpression, gene silencing, and other transgenic technologies, targeted gene deletion via CRISPR/Cas9 is a powerful tool for functional genomic studies.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This research was supported by National Science Foundation grant IOS-1025642 (GBM). We thank Maria Harrison for providing the ARqua1 strain.
Materials
| NEBuilder® (HiFi DNA assembly mix) | New England Biolabs | E5520 | |
| p201N:Cas9 | Addgene | 59175 | The p201H:Cas9 plasmid (59176) is also compatible with the reported overlaps and enzymes. |
| pUC gRNA Shuttle | Addgene | 47024 | |
| SwaI | New England Biolabs | R0604S | |
| SpeI | New England Biolabs | R0133S | |
| Zymo clean and concentrator-5 column purification | Zymo Research | D4003 | |
| NEB Buffer 2.1 | New England Biolabs | B7202S | |
| NEB CutSmart (Buffer 4) | New England Biolabs | B7204S | |
| NEB Buffer 3.1 | New England Biolabs | B7203S | |
| EconoSpin Mini Spin Column (plasmid prep) | Epoch Life Sciences | 1910-050/250 | |
| EcoRV-HF® | New England Biolabs | R3195S | |
| StyI-HF® | New England Biolabs | R3500S | |
| MS Salts + Gamborg Vitamins | Phytotechnology Laboratories | M404 | |
| Phytagel™ (gellan gum) | Sigma Aldrich | P8169 | |
| GA-7 Boxes | Sigma Aldrich | V8505 | |
| Micropore™ surgical tape | 3M | 1535-0 | |
| Timentin® (Ticarcillin/Clavulanic acid) | Various | 0029-6571-26 | |
| Primers 5' -> 3' | |||
| SwaI_MtU6F | GATATTAATCTCTTCGATGAAATTTATGCCTATCTTATATGATCAATGAGG | ||
| MtU6R | AAGCCTACTGGTTCGCTTGAAG | ||
| ScaffoldF | GTTTTAGAGCTAGAAATAGCAAGTT | ||
| SpeI_Scaffold R | GTCATGAATTGTAATACGACTCAAAAAAAAGCACCGACTCGGTG | ||
| StUbi3P218R | ACATGCACCTAATTTCACTAGATGT | ||
| ISceIR | GTGATCGATTACCCTGTTATCCCTAG | Cannot be used for Sanger sequencing since there is a second binding site on the plasmid | |
| UNS1_Scaffold R | GAGAATGGATGCGAGTAATGAAAAAAAGCACCGACTCGGTG | ||
| UNS1_MtU6 F | CATTACTCGCATCCATTCTCATGCCTATCTTATATGATCAATGAGG | ||
| p201R | CGCGCCGAATTCTAGTGATCG | ||
| Bolded sequences denote 20-nt overlaps with linearized p201N:Cas9. | |||
References
- Jinek, M., et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 337 (6096), 816-821 (2012).
- Shalem, O., et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science. 343 (6166), 84-87 (2014).
- Zhou, Y. X., et al. High-throughput screening of a CRISPR/Cas9 library for functional genomics in human cells. Nature. 509 (509), 487-491 (2014).
- Doudna, J. A., Charpentier, E. The new frontier of genome engineering with CRISPR-Cas9. Science. 346 (6213), (2014).
- Belhaj, K., Chaparro-Garcia, A., Kamoun, S., Patron, N. J., Nekrasov, V. Editing plant genomes with CRISPR/Cas9. Current Opinion in Biotechnology. 32, 76-84 (2015).
- Bortesi, L., Fischer, R. The CRISPR/Cas9 system for plant genome editing and beyond. Biotechnology Advances. 33 (1), 41-52 (2015).
- Jia, H. G., Wang, N. Targeted genome editing of sweet orange using Cas9/sgRNA. Plos One. 9, (2014).
- Upadhyay, S. K., Kumar, J., Alok, A., Tuli, R. RNA-Guided Genome Editing for Target Gene Mutations in Wheat. G3-Genes Genomes Genetics. 3 (12), 2233-2238 (2013).
- Ron, M., et al. Hairy root transformation using Agrobacterium rhizogenes. as a tool for exploring cell type-specific gene expression and function using tomato as a model. Plant Physiology. 166 (2), 455-U4442 (2014).
- Jacobs, T. B., LaFayette, P. R., Schmitz, R. J., Parrott, W. A. Targeted genome modifications in soybean with CRISPR/Cas9. BMC Biotechnology. 15, (2015).
- Gibson, D. G., Smith, H. O., Hutchison, C. A., Venter, J. C., Merryman, C. Chemical synthesis of the mouse mitochondrial genome. Nature Methods. 7 (11), 901-903 (2010).
- Zhou, X., Jacobs, T. B., Xue, L. J., Harding, S. A., Tsai, C. J. Exploiting SNPs for biallelic CRISPR mutations in the outcrossing woody perennial Populus reveals 4-coumarate:CoA ligase specificity and redundancy. New Phytologist. , (2015).
- Stemmer, M., Thumberger, T., Keyer, M. D., Wittbrodt, J., Mateo, J. L. CCTop: An Intuitive, Flexible and Reliable CRISPR/Cas9 Target Prediction Tool. Plos One. 10, (2015).
- Lei, Y., et al. CRISPR-P: A Web Tool for Synthetic Single-Guide RNA Design of CRISPR-System in Plants. Molecular Plant. 7 (9), 1494-1496 (2014).
- Quandt, H. J., Puhler, A., Broer, I. TRANSGENIC ROOT-NODULES OF VICIA-HIRSUTA – A FAST AND EFFICIENT SYSTEM FOR THE STUDY OF GENE-EXPRESSION IN INDETERMINATE-TYPE NODULES. Molecular Plant-Microbe Interactions. 6, 699-706 (1993).
- Zhu, X. X., et al. An efficient genotyping method for genome-modified animals and human cells generated with CRISPR/Cas9 system. Scientific Reports. 4 (8), (2014).
- Belhaj, K., Chaparro-Garcia, A., Kamoun, S., Nekrasov, V. Plant genome editing made easy: targeted mutagenesis in model and crop plants using the CRISPR/Cas system. Plant Methods. 9 (1), 39 (2013).
- Cong, L., et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 339 (6121), 819-823 (2013).
- Li, J. F., et al. Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotech. 31 (8), 688-691 (2013).
- Fu, Y. F., Sander, J. D., Reyon, D., Cascio, V. M., Joung, J. K. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nature Biotechnology. 32 (3), 279-284 (2014).
- Torella, J. P., et al. Unique nucleotide sequence-guided assembly of repetitive DNA parts for synthetic biology applications. Nat Protoc. 9 (9), 2075-2089 (2014).
- Ono, N. N., Tian, L. The multiplicity of hairy root cultures: Prolific possibilities. Plant Science. 180 (3), 439-446 (2011).
- Tepfer, D. Transformation of several species of higher plants by Agrobacterium rhizogenes.: sexual transmission of the transformed genotype and phenotype. Cell. 37 (3), 959-967 (1984).
- Li, H., Deng, Y., Wu, T. L., Subramanian, S., Yu, O. Misexpression of miR482, miR1512, and miR1515 Increases Soybean Nodulation. Plant Physiology. 153 (4), 1759-1770 (2010).
- Boisson-Dernier, A., et al. Agrobacterium rhizogenes-transformed roots of Medicago truncatula for the study of nitrogen-fixing and endomycorrhizal symbiotic associations. Molecular Plant-Microbe Interactions. 14 (6), 695-700 (2001).
- Collier, R., Fuchs, B., Walter, N., Kevin Lutke, W., Taylor, C. G. Ex vitro. composite plants: an inexpensive, rapid method for root biology. Plant Journal. 43 (3), 449-457 (2005).

