Design and Synthesis of a Reconfigurable DNA Accordion Rack
Summary
We describe the detailed protocol for design, simulation, wet-lab experiments, and analysis for a reconfigurable DNA accordion rack of 6 by 6 meshes.
Abstract
DNA nanostructure-based mechanical systems or DNA nanomachines, which produce complex nanoscale motion in 2D and 3D in the nanometer to ångström resolution, show great potential in various fields of nanotechnology such as the molecular reactors, drug delivery, and nanoplasmonic systems. The reconfigurable DNA accordion rack, which can collectively manipulate a 2D or 3D nanoscale network of elements, in multiple stages in response to the DNA inputs, is described. The platform has potential to increase the number of elements that DNA nanomachines can control from a few elements to a network scale with multiple stages of reconfiguration.
In this protocol, we describe the entire experimental process of the reconfigurable DNA accordion rack of 6 by 6 meshes. The protocol includes a design rule and simulation procedure of the structures and a wet-lab experiment for synthesis and reconfiguration. In addition, analysis of the structure using TEM (transmission electron microscopy) and FRET (fluorescence resonance energy transfer) is included in the protocol. The novel design and simulation methods covered in this protocol will assist researchers to use the DNA accordion rack for further applications.
Introduction
Mechanical systems based on DNA nanostructures or DNA nanomachines1,2,3,4,5 are unique because they produce complex nanoscale motion in 2D and 3D in the nanometer to ångström resolution, according to various biomolecular stimuli2,3,6. By attaching functional materials on these structures and controlling their positions, these structures can be applied to various areas. For example, DNA nanomachines have been proposed for a molecular reactor7, drug delivery8, and nanoplasmonic systems9,10.
Previously, we introduced the reconfigurable DNA accordion rack, which can manipulate a 2D or 3D nanoscale network of elements11 (Figure 1A). Unlike other DNA nanomachines that only control a few elements, the platform can collectively manipulate periodically arrayed 2D or 3D elements into various stages. We anticipate that a programmable chemical and biological reaction network or a molecular computing system can be built from our system, by increasing the number of controllable elements. The DNA accordion rack is a structure, in which the network of multiple DNA beams is connected to joints composed of single-stranded DNA (Figure 1B). The accordion rack generated by the DNA beams is reconfigured by the DNA locks, which hybridize to the sticky part of beams and change the angle between the beams according to the length of the bridging part of the locks (locked state). In addition, multi-step reconfiguration is demonstrated by adding new locks after formation of the free state by detaching DNA locks through toehold-based strand displacement12,13.
In this protocol, we describe the entire design and synthesis process of the reconfigurable DNA accordion rack. The protocol includes design, simulation, wet-lab experiments, and analysis for the synthesis of the DNA accordion rack of 6 by 6 meshes and a reconfiguration of these. The structure covered in the protocol is the basic model of the previous research11 and is 65 nm by 65 nm in size, consisting of 14 beams. In terms of the design and simulation, the structural design of the accordion rack is different from conventional DNA origami14,15 (i.e., tightly packed). Therefore, the design rule and molecular simulation have been modified from traditional methods. To demonstrate, we show the design technique using the modified approach of caDNAno14 and the simulation of the accordion rack using oxDNA16,17 with additional scripts. Finally, both protocols of TEM and FRET for analysis of configured accordion rack structures are described.
Protocol
1. Design of the 6 by 6 DNA Accordion Rack with caDNAno14
- Download and install caDNAno 2.0 software14 to design a DNA accordion rack (caDNAno 2.5 is also available on https://github.com/cadnano/cadnano2.5). Open caDNAno14 and click the Square Tool to add a new part with a square lattice.
- Number each beam of the accordion rack and draw on the left lattice panel of the caDNAno14 (Figure 2).
- Click the pencil tool and draw each beam on the right edit panel on the caDNAno14. Break beams every 32 bp, which is for joints between adjacent beams. Place staple crossovers in the same position as the joints. Use the Insert Tool and the Pencil Tool in caDNAno14 to let the joints have additional single-stranded crossovers.
- Click the Pencil Tool and connect the joints. Each beam has seven joints.
- Generate scaffold crossovers to merge the scaffolds into the single loop by using the previously reported scaffold routing algorithm11. Do not let the minimum binding domain between scaffold and staple strands be less than 8 bp (Figure 3).
- Place the scaffolds that are not used in the assembly at the vertices located on opposite sides of the accordion rack, as shown in Figure 3.
- Click the Break Tool. Break the strands where staple strands are circular or longer than 60 bp.
- Design the DNA lock strands.
- Click the Break Tool. Break 8 bp of a staple DNA region to make a sticky part and delete 8 bp of a staple DNA region. There are 18 sticky parts (Figure 1) in the 6 by 6 accordion rack.
- Place sequences that are reverse complementary to the sticky parts at both ends of lock strands and connect them by a bridging region, which consists of poly T strands of the desired length (Figure 1B).
- For the reconfiguration, add 8 bp of toehold sequences at the end of DNA locks for strand displacement. The toehold sequence used is in Table 2.
- Prepare poly A strands which are reverse complementary to the bridging region.
- Design strands that are reverse complementary to the DNA locks for the reconfiguration experiment.
- Click the Sequence Tool and click scaffold DNA. Choose the scaffold as standard M13mp18. Click the Export Tool and save sequence in csv format (Table 1).
2. Simulate the Structure with the oxDNA
- Download and install the oxDNA16,17. The latest source code is available on https://sourceforge.net/projects/oxdna/files/.
- Make starting configuration files from the caDNAno14 file using python script 'cadnano_interface.py', which is provided in the oxDNA16,17 package. The usage is as follows: ‘python cadnano_interface.py cadnano_file.json sq’. The topology file and configuration file are now generated.
Note: The topology file includes how many strands and nucleotides are in the structure and information regarding backbone-backbone bonds between nucleotides. The configuration file includes general information such as timestep, energy, and box size. Orientation information such as position vector, backbone-base vector, normal vector, velocity, and angular velocity of nucleotides is also included (Figure 4). - Change the information in the topology and configuration file from caDNAno14 to make them reflect the real structural information of the accordion rack. All beams are arranged in parallel when the topology and configuration files from caDNAno14 are visualized. However, the accordion rack is a lattice structure so the distance between bonded nucleotides are far for simulation (Figure 5).
- Rotate and move each beam to the desired lattice structure. The nine columns on the left in the configuration file are the position vector, backbone-base vector, and normal vector (Figure 4). To rotate a beam, rotate all of position, backbone-base, and normal vectors using rotational transformation. Then move a beam by changing the position vector to locate it as shown in Figure 5.
- Relax the structure using the script provided in the oxDNA package (see example in $oxDNA/EXAMPLES/RELAX_INITIAL_CONFIGURATION for further information).
- Run molecular dynamics simulation for ten million steps using the relaxed configuration file. The usage is as follows: ‘./oxDNA <input>’ Save data every 5000 or 10000 steps.
- Visualization
Note: The structures were visualized using cogli.- Download and install the latest version of the cogli (https://sourceforge.net/projects/cogli1/).
- Run the cogli with the topology and configuration files from the oxDNA simulation. The usage is as follows: ‘./cogli1 -t <topology file> <configuration file>’ .
- Hide the box by pressing b.
3. Synthesis of the Structure
Note: The synthesis method is adapted from the previous protocol15,18.
- Purchase the designed DNA staples from an oligonucleotide provider.
- Adjust the concentration of these DNA staples to 100 μM using nuclease-free water.
- Pool each DNA strand that constitutes a ‘free state’ structure into one tube and adjust the concentration to 2 μM for each strand.
- Pool DNA lock strands by length and number of lock sites into tubes and adjust the concentration to 2 μM for each strand. 18, 9, and 4 lock sites are used. Add poly A strands which are complementary to the bridging region at the same concentration.
- Pool strands that are reverse complementary to the DNA lock strands by length into tubes and adjust the concentration to 2 μM for each strand.
- Prepare MgCl2 solution of 300 nM by mixing 70 μL of nuclease-free water and 30 μL of the 1 μM MgCl2 solution. Prepare a 5x Tris EDTA solution by mixing 95 μL of nuclease-free water and 5 μL of the 100x Tris EDTA solution.
- Add 2 μL of staple DNA, 1.1 μL of MgCl2 solution, 2 μL of Tris EDTA solution, 7.6 μL of nuclease-free water and 7.3 μL of scaffold DNA of which the concentration is 110 nM to make 20 μL of mixed stock. Set the final concentration of the scaffold DNA to 40 nM, staple DNA to 200 nM, MgCl2 to 16 mM and Tris EDTA solution to 0.5x.
- Rapidly heat the mixed stock solution in a thermal cycler to 80 °C and cool to 60 °C at a rate of 4 min per °C and cool from 60 °C to 4 °C at a rate of 40 min per °C.
4. Purification of the Structure
Note: Samples of all structures were purified before analysis. In this section, we describe the protocol of PEG purification, which is adapted from previous literature19. The sample can also be purified by gel electrophoresis as described in previous literature15,18.
- Prepare 5 M of NaCl and 100x Tris-EDTA.
- Prepare precipitation-buffer by mixing 150 μL of PEG 8000, 500 μL of 100x Tris EDTA and 101 μL of 5 M NaCl and 249 μL of nuclease-free water.
- Prepare target-buffer by mixing 5.5 μL of 300 nM MgCl2 solution from Section 3.3, 10 μL of 5x Tris EDTA solution from Section 3.3 and 84.5 μL of nuclease-free water.
- Mix 20 μL of the synthesized structure from Section 3 and 20 μL of precipitation-buffer from Section 4.2. Then spin the mixed stock at 16000 x g at 4 °C. Remove the supernatant and dissolve pellet in the target-buffer from Section 4.3.
5. Reconfiguration of the Accordion Rack from a ‘Free State’ to a ‘Locked State’
- Synthesize the structure without DNA locks for the configuration experiment.
- Prepare DNA lock strands from Section 3.
- Add 2 μL of DNA lock strands of the desired length into 20 μL of the synthesized structure. The DNA lock strands’ concentration is five times higher than the structure.
- Incubate the sample for 0, 10, 25, 50, or 100 minutes to see how fast reconfiguration occurs.
- For the 100 minute incubation, incubate the sample at 50 °C for 30 minutes and slowly cool down to 25 °C at a rate of 0.33 °C/minute.
- For the 50 minute incubation, incubate the sample at 50 °C for 15 minutes and slowly cool down to 25 °C at a rate of 0.66 °C/minute.
- For the 25 minute incubation, incubate the sample at 50 °C for 7.5 minutes and slowly cool down to 25 °C at a rate of 1.32 °C/minute.
- For the 10 minute incubation, incubate the sample at 50 °C for 3 minutes and slowly cool down to 25 °C at a rate of 3.3 °C/minute.
- For the 0-minute incubation, store sample at 4 °C right after DNA locks strands are added.
- Right after the attaching step, rapidly cool down the sample to 4 °C to prevent unwanted denaturation.
6. Reconfiguration of the Accordion Rack from a ‘Locked State’ to a ‘Free State’
- Synthesize the structure with DNA locks of the desired length for the configuration experiment.
- Prepare reverse complementary strands from Section 3.
- Add 2 μL of strands that are reverse complementary to the lock strands of the desired length into 20 μL of the synthesized structure. The DNA lock strands’ concentration is five times higher than the structure.
- Incubate the sample for 0, 12, 60, 120, 240 minutes to see how fast reconfiguration occurs.
- For the 12, 60, 120, 240 minute incubation, rapidly heat the sample to 40 °C and slowly cool down to 20 °C for the time corresponding to each. Right after the detaching step, rapidly cool down the sample to 4 °C to prevent unwanted denaturation.
- For the 0-minute incubation, store sample at 4 °C right after reverse complementary strands are added.
7. TEM Imaging
Note: TEM imaging protocol was adapted from previous literature18,20.
- Prepare 1.25 M NaOH solution by mixing 87.5 μL of nuclease-free water and 12.5 μL of 10 M NaOH solution.
- Add 1 μL of 1.25 M NaOH solution to 50 μL of the 2% uranyl formate solution.
- Vortex the solution for 3 minutes and centrifuge at max speed for 3 minutes. Deposit 3 μL of the purified sample on the glow-discharged TEM grid for 3 minutes and rapidly wash out with filter paper.
- Deposit 7 μL of prepared uranyl formate solution for 30 seconds and rapidly wash out with filter paper.
- Measure the length and angle of the accordion structure imaged by TEM.
8. FRET Analysis
- Use Atto 550 and Atto 647N dye, for which the Förster distance is 6.5 nm. Replace Staple 58 and Staple 117 in Table 1 with fluorescently labelled strands. Then synthesize the structure with fluorescently labelled strands by the method described in Section 3.
- Measure the concentration of the purified sample.
- Normalize the sample to 10 nM and load 50 μL to the 384 microplates well.
- Excite the sample with donor and acceptor dyes at 550 nm and measure the fluorescence spectrum from 570 nm to 800 nm with a fluorometer.
- Measure the fluorescence spectrum of the donor-only sample in the same way.
- Excite the dyes of the sample at 650 nm and the fluorescence spectrum and measure from 670 nm to 800 nm. This is to measure the concentration of the acceptor.
- Obtain the standard deviations by repeating the same experiment with three samples, which are synthesized and purified separately.
- Calculate the FRET efficiency with the ratio A method as described by the equation below21.
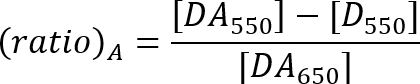
DA550: Acceptor peak fluorescence intensity of the sample with donor and acceptor at 550 nm excitation.
D550: Fluorescence intensity at the acceptor emission range of the donor-only sample at 550 nm excitation.
DA650: Acceptor peak fluorescence intensity of the sample with donor and acceptor at 650 nm excitation.
Representative Results
The designed 6 by 6 DNA accordion rack is simulated from the oxDNA16,17 and the results are shown in Figure 6. From the simulation result, it was confirmed that the intended structure is formed without distortion of the structure.
The TEM images in Figure 7 are images of configured structures with a lock length of 2, 8, 13 and 20 bp. On the image, the angle of the structures (Figure 8A) is decreased as the length of the lock becomes longer. To statistically analyze the tendency, the average and the standard deviation of the angle (Figure 8A) of multiple configured structures were obtained (Figure 8B). Also, the FRET measurement results show structural change and its speed, in addition to the structural tendency shown in the TEM image (Figure 8C).
From these analysis procedures, characterization of the DNA accordion rack such as how many locks are needed for structural configuration and reconfiguration speed was determined.
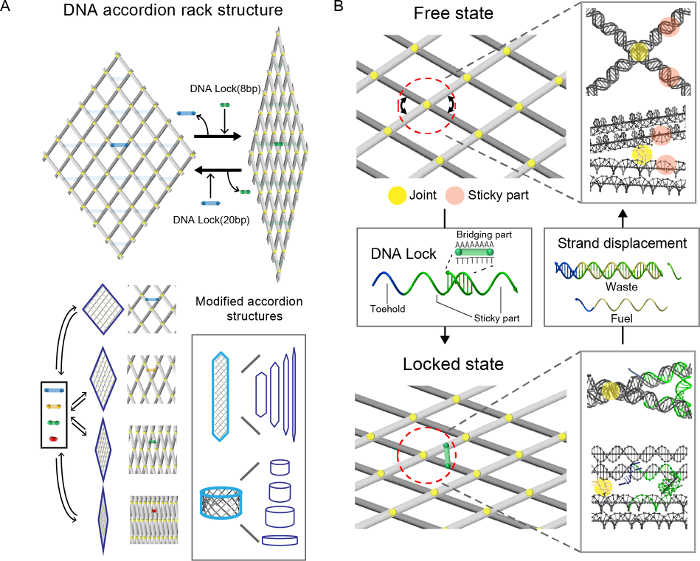
Figure 1. A reconfigurable DNA accordion rack. A. The DNA accordion rack is composed of rigid DNA beams and the multiple flexible joints that connect each beam. The accordion rack is configured differently according to the length of the DNA locks. Various 2D and 3D platforms can be designed by modifying the platform. B. The reconfiguration of the DNA accordion rack is achieved with hybridization of DNA locks with different lengths. In the 'free state' structure, when extra nucleotides are added to the crossovers (or joints, yellow), due to the loose connection between the DNA beams (comprising two helices), the angle can be changed (upper part). Hybridization of the DNA locks with various lengths of the bridge part to the sticky part of both adjacent beams generates the 'locked state' (Lower part). Strand displacement using the toehold parts (blue) of the DNA locks with the fuel strand detach the DNA locks from the locked state of the structure and change the state to the free. By repeating the process with various libraries of the DNA locks, the structure is reconfigured with different angles. This figure has been modified from the previous research11. Please click here to view a larger version of this figure.
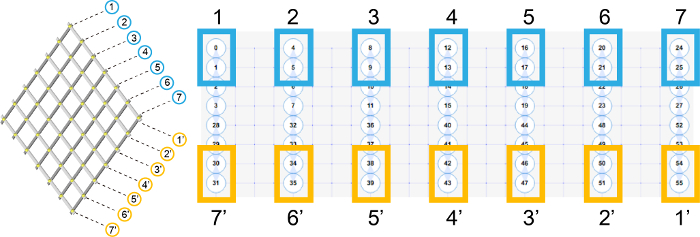
Figure 2. 6 by 6 accordion rack is designed using the caDNAno. The 14 beams of the DNA accordion rack are numbered and drawn on the lattice panel of the caDNAno. This figure has been modified from the previous research11. Please click here to view a larger version of this figure.
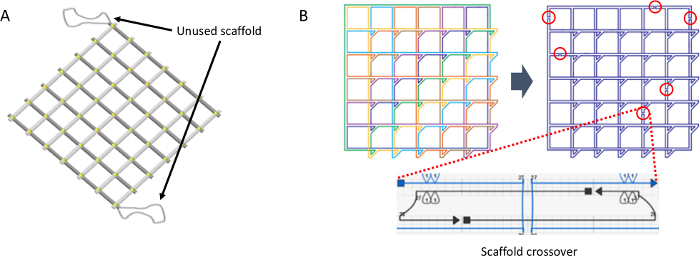
Figure 3. Scaffold routing algorithm. A. Scaffold that are not used in the assembly at the vertices located on opposite sides of the accordion rack. B. Seven closed loops are merged into a single loop by generating scaffold crossovers. At least six scaffold crossovers points are needed and the position of the scaffold crossovers is adapted from previous literature. This figure has been modified from the previous research11. Please click here to view a larger version of this figure.
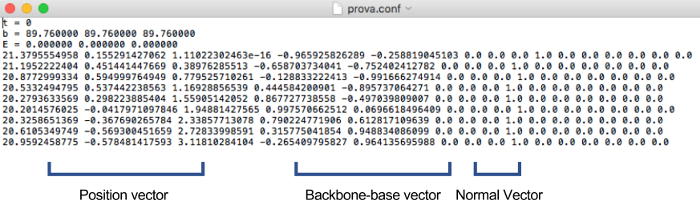
Figure 4. Example of a configuration file. The configuration file includes general information such as the timestep, energy and box size and orientation information. The nine columns on the left in the configuration file are the position vector, backbone-base vector, and normal vector respectively. The six columns on the right are velocities and angular velocities. Please click here to view a larger version of this figure.
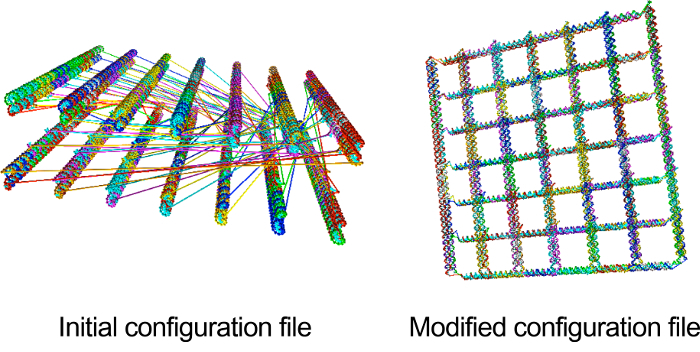
Figure 5. Visualization of the configuration file before and after the modification. Position, backbone-base, and normal vector was modified using rotational transformation to make the information in the configuration file reflect the structural information of the accordion rack. Please click here to view a larger version of this figure.
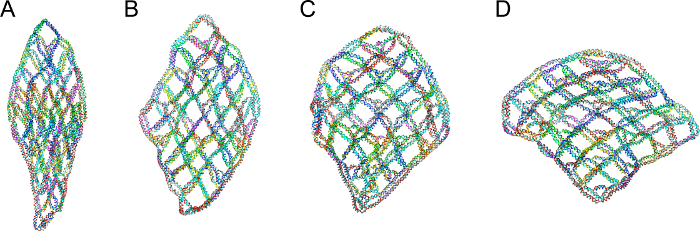
Figure 6. Simulation result from oxDNA. The accordion rack with DNA locks, of which the lengths are 2, 8, 13 and 20 bp, were simulated. 18 locking sites were used. A 6 by 6 accordion rack with a DNA lock of which the length is A 2 bp, B 8 bp, C 13 bp and D 20 bp are simulated andvisualized by cogil. Please click here to view a larger version of this figure.
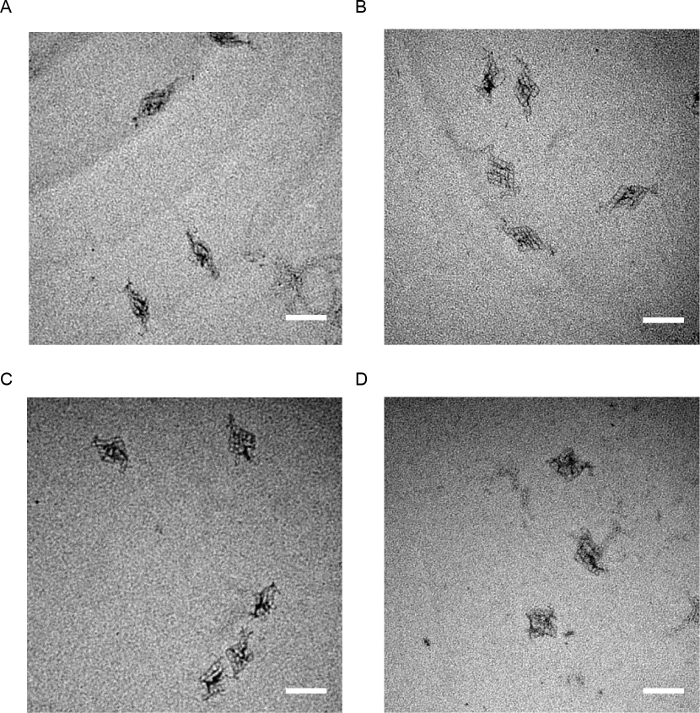
Figure 7. TEM images of configured 6 by 6 DNA accordion racks. Structures with lock lengths of A 2 bp, B 8 bp, C 13 bp and D 20 bp were imaged. 18 locking sites were used for this experiment. (scale bar: 100 nm). This figure has been modified from the previous research11. Please click here to view a larger version of this figure.
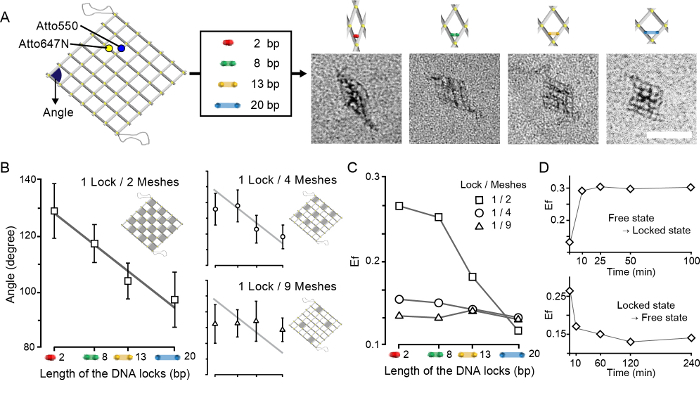
Figure 8. The angle control of the 2D accordion rack. The angle of the accordion rack comprised of 6-by-6 meshes was controlled by the length of the DNA locks. A. By adding DNA locks with lengths of 2, 8, 13 and 20 bp to the 18 locking sites of the accordion rack, the angle (blue) is configured as seen in the representative TEM images. The dye pair, Atto647N, and Atto550 is attached to the structure for FRET efficiency analysis. (scale bar: 100 nm) B. The distribution of the configured angles. The overlaid trend line (grey) is from the graph of the 1 lock per 2 meshes. Additionally, the grid of the x-axis corresponds to the same grid on the graph of the 1 lock per 2 meshes. Bars represent one standard deviation from the mean angle. C. The distribution of FRET efficiencies. Bars represent one standard deviation from the mean efficiency. D. FRET efficiency change while the state transitioned from free to locked or vice versa was measured according to different incubation times. The DNA lock with the length of 2 bp was used. Bars represent one standard deviation from the mean efficiency. This figure has been modified from the previous research11. Please click here to view a larger version of this figure.
| Name | Sequence | Length | |||
| Staple 1 | ATAAGAGGTCATTTTTGCGGATGGATGTTACT | 32 | |||
| Staple 2 | CAAAGTTACCAGAAGGAAACCGAGACATCGGG | 32 | |||
| Staple 3 | ACAATGAAATAGCAATAGCTATCTCAAATAAA | 32 | |||
| Staple 4 | CGCTAATATGAACGGTGTACAGACCAGGCGCATAGGCTGGGAACAAAGTCAGAGGG | 56 | |||
| Staple 5 | CAGAAAACCCGGAATAGGTGTATCACCGTACTCAGGAGGTCCCCCTCAAATGCTTT | 56 | |||
| Staple 6 | AGTGAGAATAGAAAGGTATGATATTCAACCGTTCTAGCTGATAAATTATCAACAGT | 56 | |||
| Staple 7 | ATTTGGGGCGCGAGCTATATGGTT | 24 | |||
| Staple 8 | GCCTTTATTTCAACGCAAGGATAATTAATGGA | 32 | |||
| Staple 9 | TCAATTACCCACTACGAAGGCACCGGTAAAAT | 32 | |||
| Staple 10 | ATTCAGTGAATAAGGCTTGCCCTGAAAACAGA | 32 | |||
| Staple 11 | AGAAACAATAACGGATTCGCCTGATAGCCGAA | 32 | |||
| Staple 12 | ATCAATATCTGGTCAGTTGGCAAAAGTAACAA | 32 | |||
| Staple 13 | CGGAGACAGTCAAATCACCATCAAGGGTTAGA | 32 | |||
| Staple 14 | GAGAATTAACTGAACACGAAACAAAGTACAACGGAGATTTGTATCATCGAAGCGCA | 56 | |||
| Staple 15 | CGAGAGGGTTGATATAAGTATAGCATAGGAAC | 32 | |||
| Staple 16 | CCCTTATTAGCGTTTGCCATCTTTTGGTGCCG | 32 | |||
| Staple 17 | GTTTTCATCGGCATTTATGCCGGA | 24 | |||
| Staple 18 | GTAGCAACTTCCAGTAAGCGTCATGTCTCTGA | 32 | |||
| Staple 19 | AGCTAATGCAGAACGCAATAAACA | 24 | |||
| Staple 20 | CCGTAATGGGATAGGTCACGTTGGGCCTATTT | 32 | |||
| Staple 21 | ACCAGAGCCACCACCGCGAGAGGC | 24 | |||
| Staple 22 | AGGCTCCAAAAGGAGCCGTTTACCAGACGACGATAAAAACCAAAATAGAAAATCTC | 56 | |||
| Staple 23 | GCCGGAAGGAAACGCAATAATAACGGAATACCCAAAAGAACTCACAATTCCACACA | 56 | |||
| Staple 24 | TTTTTCACCGGTGTCTGGAAGTTTCATTCCATATAACAGTACTAAAGGAATTGCGA | 56 | |||
| Staple 25 | ATCAGGTCTTTACCCTCGCATTAAATTTTTGTTAAATCAGCTCATTTTTGACCATA | 56 | |||
| Staple 26 | TGCTGCAAGGCGATTAAGTTGGGTGACATTCA | 32 | |||
| Staple 27 | AATAAAGAAATTGCGTAGATTTTCAGCTGCTC | 32 | |||
| Staple 28 | CCCAATCCTCGCACTCCAGCCAGCTTTCCGGCACCGCTTCAAAATAAACAGCCATA | 56 | |||
| Staple 29 | AAAGACTTCAAATATCATCAAAAGAATAGCCCGAGATAGGGTTGAGTGATTAAGAG | 56 | |||
| Staple 30 | GTAGCGACAGAATCAAGTTTGCCTGATTTAGA | 32 | |||
| Staple 31 | GCCACCCTCAGAACCGCCACCCTCAACAAGAG | 32 | |||
| Staple 32 | AAGCAAGCCGTTTTTATTTTCATCTCCTGATT | 32 | |||
| Staple 33 | GAGGGTAGCTATTTTTGAGAGATCCGTCAGAC | 32 | |||
| Staple 34 | TATGACCCTGTAATACTCCCATCCTAATTTACGAGCATGTAGAAACCAGTTGTACC | 56 | |||
| Staple 35 | GATTAGAGAGTACCTTAACAGTGCCCGTATAAACAGTTAATGCCCCCTAACTCCAA | 56 | |||
| Staple 36 | CGTCTTTCCAGACGTTGTAGGAATCATTACCGCGCCCAATAGCAAGCACGATCTAA | 56 | |||
| Staple 37 | TTTAACGTCAAAAATGAGTGAGACGGGCAACAGCTGATTGCCCTTCACAAGAAACG | 56 | |||
| Staple 38 | TTTAACGGGGTCAGTGCCTTGAGTATAAGGGA | 32 | |||
| Staple 39 | AATTGTGTGCAAAATCCCTTATAAGGTGGTTC | 32 | |||
| Staple 40 | ACCAGTAGCACCATTACCATTAGCTTTCAGGG | 32 | |||
| Staple 41 | AAACATCAAGAAAACAAAATTAATTCACATTA | 32 | |||
| Staple 42 | TTGGCCTTGATATTCATCATCTTT | 24 | |||
| Staple 43 | TGCATCAATCAACAGTTGAAAGGAATTGAGGAAGGTTATCATTATAGTCAGAAGCA | 56 | |||
| Staple 44 | GCCCCAGCAGGCGAAAATCCTGTTAGGGGACG | 32 | |||
| Staple 45 | TCCTCATTAAAGCCAGAATGGAAAAGCAAGAA | 32 | |||
| Staple 46 | GAGCCTAATTTGCCAGGGGGGTAATAGTAAAATGTTTAGACTGGATAGAACGAGCG | 56 | |||
| Staple 47 | TAAGAGCAACACTATCATAACCCTTTAACGTC | 32 | |||
| Staple 48 | ATAGCAAGCCCAATAGGAACCCATGCAAAATC | 32 | |||
| Staple 49 | CATAAAAACTTAGAGCTTAATTGCTGAATATAATGCTGTATAGCAGCCTTTACAGA | 56 | |||
| Staple 50 | ATTTACCGGGCTACAGAGGCTTTGGAACGAGG | 32 | |||
| Staple 51 | GTGCCTAATGAGTGAGAATGCAGATACATAACGCCAAAAGGAATTACGAAGTGTAA | 56 | |||
| Staple 52 | ATACCGATAGTTGCGCATCGGCCAACGCGCGGGGAGAGGCGGTTTGCGTTTCTTAA | 56 | |||
| Staple 53 | ATAACCGATACCGAAGCCCTTTTTAAGAAAAGTAAGCAGAAATGACAACAACCATC | 56 | |||
| Staple 54 | TCAGAGCCGCCACCCTCAGAACCGGCTCAACA | 32 | |||
| Staple 55 | TTTTGCAAAAGAAGTTTTGCCAGAAATCAAAA | 32 | |||
| Staple 56 | GGCCTCTTCGCTATTACGCCAGCTAAGAAACC | 32 | |||
| Staple 57 | ACATGTTCAAAGACACCACGGAATCATATAAA | 32 | |||
| Staple 58 | CTGTGTGAAATTGTTAGAGTAATGTGTAGGTAAAGATTCAAAAGGGTGTGGTCATA | 56 | |||
| Staple 59 | TTTCGAGGCCGGATATTCATTACCCAAATCAACGTAACAAAATTGTATCGGTTTAT | 56 | |||
| Staple 60 | CTTTAGGAGCACTAACAACTAATAAGCCCCAA | 32 | |||
| Staple 61 | AGATTAGTTGCTATTTTACAAAGGCTATCAGGTCATTGCCTGAGAGTCTTGAAGCC | 56 | |||
| Staple 62 | GAGCCGCCGCCAGCATTGACAGGACTGACCTT | 32 | |||
| Staple 63 | GAAACCAGGCAAAGCGCCATTCGCTCATAGCC | 32 | |||
| Staple 64 | CATCAAGAGTAATCTTGACAAGAACACCACCA | 32 | |||
| Staple 65 | AACCTTGCTTCTGTAAATCGTCGCTTATCAAC | 32 | |||
| Staple 66 | TAGTTAGCGACAACTCGTATTAAATCCTTTGCCCGAACGTGTAGCATTCCACAGAC | 56 | |||
| Staple 67 | ACCGATTGAGGGAGGGAAGGTAAAGGGGGATG | 32 | |||
| Staple 68 | TACCAGCGCCAAAGACAAAAGGGCGTTTAGCT | 32 | |||
| Staple 69 | TCATATATTTTAAATGCAATGCCTAAAACGAA | 32 | |||
| Staple 70 | ACATTATCATTTTGCGTATTGACG | 24 | |||
| Staple 71 | CGGAACCTATTATTCTGAAACATGCGGATTGA | 32 | |||
| Staple 72 | CCTGAGAGATATTTTGTTAAAATTTAAATTGT | 32 | |||
| Staple 73 | GAAACCATCGATAGCAAATCAGAT | 24 | |||
| Staple 74 | CTCAGAGCCGCCACCATATTGGGC | 24 | |||
| Staple 75 | CACCGACTTGAGCCATTTGGGAATACACTGAG | 32 | |||
| Staple 76 | TCATGAGGAAGTTTCCATTAAACGTTTGCGGG | 32 | |||
| Staple 77 | AAACAGGAAGATTGTATAAGCAAATAAAATAT | 32 | |||
| Staple 78 | ATCGTCACCCTCAGCAGCGAAAGAAGACTTTT | 32 | |||
| Staple 79 | ATCAACATTAAATGTGTTGTTCCA | 24 | |||
| Staple 80 | TGCAGGGAGTTAAAGGAACGAAAGAGGCAAAAGAATACACTAAAACACTTCGGTCG | 56 | |||
| Staple 81 | ACGTAATGCTGAGCAAAAGAAGATTATTCATT | 32 | |||
| Staple 82 | AATCGATGAACGGTAATCGTAAAATTAGTACC | 32 | |||
| Staple 83 | GACCCCCAGCGATTATACCAAGCGGAGGCAGG | 32 | |||
| Staple 84 | CGAAATCCGCGACCTGGCCTGATA | 24 | |||
| Staple 85 | GCCATCAAAAATAATTCGCGTCTGAGTGCCGT | 32 | |||
| Staple 86 | TCGGCTGTCTTTCCTTATCATTCCATTACCTT | 32 | |||
| Staple 87 | CAGACCGGGGGCGCATCGTAACCGTGCATCTGCCAGTTTGTTTAATTCGAGCTTCA | 56 | |||
| Staple 88 | GTTTGGAACAAGAGTCCACTATTACCTGTAGC | 32 | |||
| Staple 89 | AACAGTACATAAATCAATATATGTCGGGAGAA | 32 | |||
| Staple 90 | GACGTTGGGAAGAAAAATCTACGTCTGGCATG | 32 | |||
| Staple 91 | CAAAATCGCGCAGAGGCGGGAAAC | 24 | |||
| Staple 92 | ATTGCGTTGCGCTCACTGCCCGCTGATGAAAC | 32 | |||
| Staple 93 | CCCCGGTTGATAATCACGTCCAAT | 24 | |||
| Staple 94 | ATGCCTGCAGGTCGACAAGAACGGGTATTAAACCAAGTACCGCACTCACCAGTGCC | 56 | |||
| Staple 95 | GTTTGGATTATACTTCTGAATAATTGATTCCC | 32 | |||
| Staple 96 | ACCGAACTGACCAACTTTGAAAGATAATAAGT | 32 | |||
| Staple 97 | GCCAGGGTGGTTTTTCTTTTCACCCCTCAGAG | 32 | |||
| Staple 98 | CAGTTGAGATTTAGGAATACCACATACATTTA | 32 | |||
| Staple 99 | TAGCCGGAACGAGGCGCAGACGGTTCCTTTTG | 32 | |||
| Staple 100 | AGTATTAGACTTTACAAACAATTCCGTAATCA | 32 | |||
| Staple 101 | CTTGCGGGAGAACCGCCACCCTCAGAGCCACCACCCTCATGAGGCGTTTTAGCGAA | 56 | |||
| Staple 102 | TTAAGAACTGGCTCATATCAATAA | 24 | |||
| Staple 103 | CATACAGGCAAGGCAAAGAATTAGAAGTTTAT | 32 | |||
| Staple 104 | ATTGAGTTAAGCCCAAACATGGCTTTTGATGATACAGGAGTGTACTGGGAGATAAC | 56 | |||
| Staple 105 | ATAGAAGGCTTATCCGGTATTCTACGGAAACG | 32 | |||
| Staple 106 | CTGTCGTGCCAGCTGCATTAATGATTTGAATA | 32 | |||
| Staple 107 | CTTACCAAGATTAGAGCCGTCAATAGATAATACATTTGAGCCCAGCTACAATTTTA | 56 | |||
| Staple 108 | ACCTTGCTGAACCTCAAAAGTATTAAGAGGCTGAGACTCCTCAAGAGAAAAAATCT | 56 | |||
| Staple 109 | TTTGTCACAATCAATAGAAAATTCCAATAAAT | 32 | |||
| Staple 110 | CAATTCTATTCAACTTTAATCATTCTTGAGAT | 32 | |||
| Staple 111 | ACGACAGTATCGGCCTCAGGAAGAGCTGGTTT | 32 | |||
| Staple 112 | ACTGCGGAATCGTCATAAATATTCATGTCAAT | 32 | |||
| Staple 113 | TTTCGTCACCAGTACAAACTACAAATCACCGT | 32 | |||
| Staple 114 | AATTCTGCGAACGAGTAGATTTAGTCCTGATT | 32 | |||
| Staple 115 | GAAATTATTCATTAAAGGTGAATTTTTAAAAG | 32 | |||
| Staple 116 | ACCAGAAGGAGCGGAATTATCATCTCGGTGCG | 32 | |||
| Staple 117 | AATTCGTAACGAGAAACACCAGAACGAGTAGTAAATTGGGGAGGATCCCCGGGTAC | 56 | |||
| Staple 118 | ATTAAGACTCCTTATTACGCAGTACCAGTCAG | 32 | |||
| Staple 119 | GGTTTAATCTAATAGTAGTAGCATGGTGGCAT | 32 | |||
| Staple 120 | AAACCGTCTTAGCGGGGTTTTGCTCAGTACCAGGCGGATATGGACTCCAACGTCAA | 56 | |||
| Staple 121 | TATCAAAATTATTTGCAGAAAGGC | 24 | |||
| Staple 122 | ACAACATTATTACAGGTAGAACCC | 24 | |||
| Staple 123 | CCCGTCGGATTCTCCGTGGGAACAAACCCTCA | 32 | |||
| Staple 124 | ACAATTTCATTTGAATTACCTTTTAGATTCAT | 32 | |||
| Staple 125 | GCTAAACACATTCAGGCTGCGCAACTGTTGGGAAGGGCGAAATGAATTTTCTGTAT | 56 | |||
| Staple 126 | AATAGATAAGTCCTGAACAAGAAAGAGTGAAT | 32 | |||
| Staple 127 | AAAGCTAATGTTAGCAAACGTAGAAAATACATACATAAAGTTAAGCAATAAAGCCT | 56 | |||
| Staple 128 | TGTTTTAAATATGCAACTAAAGTACGCCTCCC | 32 | |||
| Staple 129 | ATACAGTAACAGTACCAGGCATAG | 24 | |||
| Staple 130 | GTAAAACGTTTGACCATTAGATACATTTCGCAAATGGTCACAGGGTTTTCCCAGTC | 56 | |||
| Staple 131 | AGTTGCAGCAAGCGGTCGCCTGGC | 24 | |||
| Staple 132 | ATGGCAATTCATCAATTCGAGAAC | 24 | |||
Table 1. Sequence of the staple strands
| Name | Sequence |
| Toehold_1 | AGAAGTCG |
| Toehold_2 | AGGTATGG |
| Toehold_3 | CTCCACTC |
| Toehold_4 | GTGTCCGA |
| Toehold_5 | AAGTAGAC |
| Toehold_6 | GAGTCGCT |
| Toehold_7 | AGTGTCTA |
| Toehold_8 | GTATCGTG |
| Toehold_9 | CATCACAG |
| Toehold_10 | CGAGACTT |
| Toehold_11 | ACGGACGA |
| Toehold_12 | GCCTACTC |
| Toehold_13 | GATGTGGA |
| Toehold_14 | GCGTGCGT |
| Toehold_15 | GTGGACAA |
| Toehold_16 | TACGACTG |
| Toehold_17 | CAGCCGTG |
| Toehold_18 | TGCCGCAT |
Table 2. Toehold sequence for the strand displacement experiment
Discussion
This protocol introduces the entire process from design, simulation, synthesis, and analysis of the basic 2D DNA accordion rack. The modified design and simulation rules have been described because the design rule differs from that of standard DNA origami, in that the DNA accordion rack has additional nucleotides at the crossovers for flexibility14,15. From this, we expect that the protocol can accelerate various researches using DNA accordion racks. In addition, the described protocol can also be applied to other research using DNA nanostructure rather than standard DNA origami.
For structures other than the basic 2D structure demonstrated in the original research paper, the design can be done with modifications of the described protocol. In brief, a boxing-glove-spring structure is designed by changing the beam number and length from the basic model. 3D nanotubular structures can also be created by connecting both ends of a 2D accordion rack structure. However, for simulating the 3D structures, the initial position of the beams should be considered in 3D space and the initial state of the beams is supposed to be bent. Therefore, more computational transformation is needed. Design and simulation of various 2D and 3D DNA accordion rack structures will be conducted by developing the computer-aided program in future research.
Speed and repeatability of actuation are also important factors in the field of dynamic DNA nanotechnology. Recently dynamic DNA structures that respond to external electrical or magnetic field are proposed and operated with high speed22,23. While the DNA accordion rack has enabled collective actuation of multiple DNA beams, the reconfiguration speed was not significantly improved since actuation is based on DNA strand displacement. For further applications, response to external stimuli should be improved by optimizing DNA lock design or using another external stimuli.
As an application study for the reconfigurable accordion structure, various functional materials such as drug molecules, nanoparticles, or proteins can be attached to the structure. For this, the molecules can be connected to the staple that exists at the intended position. Overall, a variety of application studies can be conducted through this protocol and its modifications.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This research was partially supported by the Global Research Development Center Program through the National Research Foundation of Korea(NRF) funded by the Ministry of Science and ICT (MSIT) (2015K1A4A3047345) and Nano·Material Technology Development Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science and ICT (MSIT) (2012M3A7A9671610). The Institute of Engineering Research at Seoul National University provided research facilities for this work. Authors acknowledge gratitude towards Tae-Young Yoon (Biological Sciences, Seoul National University) regarding the fluorescence spectroscopy for the FRET analysis.
Materials
| M13mp18 Single-stranded DNA | NEB | N4040s | |
| 1M MgCl2 Solution | Biosesang | M2001 | |
| Tris-EDTA buffer | Biosesang | T2142 | |
| Nuclease-Free Water | Qiagen | 129114 | |
| 5M Sodium Chloride solution | Biosesang | s2007 | |
| PEG 8000 | Sigma Aldrich | 1546605 | |
| 10N NaOH | Biosesang | S2038 | |
| Uranyl formate | Thomas Science | C993L42 | |
| Thermal cycler C1000 | Biorad | ||
| Nanodropic 2000 | Thermo Fisher Scientific | ||
| TEM (LIBRA 120) | Carl Zeiss | ||
| Fluorometer Enspire 2300 | Perkin-Elmer | ||
| Centrifuge | Labogene | LZ-1580 |
References
- Andersen, E. S., et al. Self-assembly of a nanoscale DNA box with a controllable lid. Nature. 459 (7243), 73-76 (2009).
- Cha, T. -. G., et al. Design principles of DNA enzyme based walkers: Translocation kinetics and photo-regulation. Journal of the American Chemical Society. 137 (29), 9429-9437 (2015).
- Gerling, T., Wagenbauer, K. F., Neuner, A. M., Dietz, H. Dynamic DNA devices and assemblies formed by shape-complementary, non-base pairing 3D components. Science. 347 (6229), 1446-1452 (2015).
- Pinheiro, A. V., Han, D., Shih, W. M., Yan, H. Challenges and opportunities for structural DNA nanotechnology. Nature nanotechnology. 6 (12), 763-772 (2011).
- Li, J., et al. Exploring the speed limit of toehold exchange with a cartwheeling DNA acrobat. Nature Nanotechnology. 1, (2018).
- Krishnan, Y., Simmel, F. C. Nucleic acid based molecular devices. Angewandte Chemie International Edition. 50 (14), 3124-3156 (2011).
- Liu, M., et al. A DNA tweezer-actuated enzyme nanoreactor. Nature communications. 4, 2127 (2013).
- Douglas, S. M., Bachelet, I., Church, G. M. A logic-gated nanorobot for targeted transport of molecular payloads. Science. 335 (6070), 831-834 (2012).
- Kuzyk, A., et al. Reconfigurable 3D plasmonic metamolecules. Nature Materials. 13 (9), 862-866 (2014).
- Zhou, C., Duan, X., Liu, N. A plasmonic nanorod that walks on DNA origami. Nature communications. 6, 8102 (2015).
- Choi, Y., Choi, H., Lee, A. C., Lee, H., Kwon, S. A Reconfigurable DNA Accordion Rack. Angewandte Chemie International Edition. 57 (11), 2811-2815 (2018).
- Chen, H., et al. Understanding the Mechanical Properties of DNA Origami Tiles and Controlling the Kinetics of their Folding and Unfolding Reconfiguration. Journal of the American Chemical Society. 136 (19), 6995-7005 (2014).
- Han, D., Pal, S., Liu, Y., Yan, H. Folding and cutting DNA into reconfigurable topological nanostructures. Nature Nanotechnology. 5 (10), 712-717 (2010).
- Douglas, S. M., et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucleic Acids Research. 37 (15), 5001-5006 (2009).
- Castro, C. E., et al. A primer to scaffolded DNA origami. Nature methods. 8 (3), 221-229 (2011).
- Ouldridge, T. E., Louis, A. A., Doye, J. P. K. DNA Nanotweezers Studied with a Coarse-Grained Model of DNA. Physical Review Letters. 104 (17), 178101 (2010).
- Snodin, B. E. K., et al. Direct Simulation of the Self-Assembly of a Small DNA Origami. ACS Nano. 10 (2), 1724-1737 (2016).
- Amir, Y., Abu-Horowitz, A., Bachelet, I. Folding and Characterization of a Bio-responsive Robot from DNA Origami. Journal of Visualized Experiments. (106), e51272 (2015).
- Stahl, E., Martin, T. G., Praetorius, F., Dietz, H. Facile and Scalable Preparation of Pure and Dense DNA Origami Solutions. Angewandte Chemie International Edition. 53 (47), 12735-12740 (2014).
- Wei, B., Vhudzijena, M. K., Robaszewski, J., Yin, P. Self-assembly of Complex Two-dimensional Shapes from Single-stranded DNA Tiles. Journal of Visualized Experiments. (99), e52486 (2015).
- Clegg, R. M. Fluorescence resonance energy transfer and nucleic acids. Methods in enzymology. 211, 353-388 (1992).
- Kopperger, E., et al. A self-assembled nanoscale robotic arm controlled by electric fields. Science. 359 (6373), 296-301 (2018).
- Lauback, S., et al. Real-time magnetic actuation of DNA nanodevices via modular integration with stiff micro-levers. Nature Communications. 9 (1), 1446 (2018).

