An Explant System for Time-Lapse Imaging Studies of Olfactory Circuit Assembly in Drosophila
Summary
This protocol describes the dissection procedure, culture condition, and live imaging of an antennae-brain explant system for the study of the olfactory circuit assembly.
Abstract
~Neurons are precisely interconnected to form circuits essential for the proper function of the brain. The Drosophila olfactory system provides an excellent model to investigate this process since 50 types of olfactory receptor neurons (ORNs) from the antennae and maxillary palps project their axons to 50 identifiable glomeruli in the antennal lobe and form synaptic connections with dendrites from 50 types of second-order projection neurons (PNs). Previous studies mainly focused on identifying important molecules that regulate the precise targeting in the olfactory circuit using fixed tissues. Here, an antennae-brain explant system that recapitulates key developmental milestones of olfactory circuit assembly in culture is described. Through dissecting the external cuticle and cleaning opaque fat bodies covering the developing pupal brain, high quality images of single neurons from live brains can be collected using two-photon microscopy. This allows time-lapse imaging of single ORN axon targeting from live tissue. This approach will help reveal important cell biological contexts and functions of previously identified important genes and identify mechanisms underpinning the dynamic process of circuit assembly.
Introduction
Neurons are precisely interconnected to form circuits essential for the proper function of the brain. For over 100 years, neuroscientists have been trying to understand how neurites extend toward their intermediate and final targets with extreme precision. As a result, they have identified important genes that encode guidance cues for developing neuronal processes1. The Drosophila olfactory system provides an excellent model to investigate this process since olfactory receptor neurons (ORNs, the primary sensory neurons) project to 50 identifiable glomeruli with stereotypical size, shape, and relative position, where they form synaptic connections with dendrites from 50 types of second-order projection neurons (PNs), each of which send dendrites to one of the 50 glomeruli2 (Figure 1A). Therefore, it is relatively easy to identify mutant phenotypes at synaptic (glomerular) resolution in the fly olfactory system. This led to discoveries of important genes that regulate olfactory circuit assembly3.
The assembly of the fly olfactory circuit relies on temporally and spatially coordinated developmental processes3. ORNs and PNs acquire distinct cell fates, which set up the program for their wiring specificities. Next, PN dendrites prepattern the antennal lobe (Figure 1B). The axons of ORNs then circumnavigate the ipsilateral antennal lobe and cross the midline of the brain to reach the contralateral antennal lobe. Subsequently, ORN axons invade both ipsi- and contralateral antennal lobes and form synapses with dendrites of their partner PNs in specific glomeruli. This coarse model for olfactory circuit assembly was proposed based on the characterization of fixed samples from intermediate time points during the development. The poor temporal resolution and inability to follow the same neuronal processes across development from fixed tissue limit the mechanistic understanding of the circuit assembly process.
It is technically challenging to live image ORN and PN processes in vivo since the wiring process occurs in the first half of the pupal stage when the antennal lobe is surrounded by opaque fat body inside the pupal case. It is, therefore, impossible to directly image the developing olfactory circuit from intact pupae. Dissected tissues cultured ex vivo can circumvent tissue opacity and have been successfully used to study neural development4,5,6. The challenge of using a similar ex vivo explant culture strategy to study neuronal wiring in the pupal brain is whether it recapitulates the precise neuron targeting in a culture condition. Based on a previously reported ex vivo culture condition for the fly eye-brain complex7, an explant that contains the whole pupal brain, antennae, and the connecting antennal nerves intact has been recently developed, which retains precise targeting of the olfactory circuit and can be subjected to two-photon microscopy-based live imaging for up to 24 h at the frequency of every 20 min8. Here, a detailed protocol of the explant culture and imaging is described. The explant system provides a powerful method to study the assembly of olfactory circuit and potentially other circuits in the central brain.
Protocol
1. Preparation of reagents
NOTE: All the steps in this protocol are carried out at room temperature (20-25 °C) unless explained otherwise.
- To prepare the culture dish for immobilizing the explant during time lapse imaging, lay 0.5 cm thick Sylgard (thoroughly mix two liquid components at 10:1 ratio before use) on the bottom surface of a 60 mm x 15 mm Petri dish and let it cure for 48 h at room temperature (Figure 2A, referred as Sylgard plate in the following text).
- To prepare micro pins for immobilizing explant on this plate, use a pair of forceps to stick multiple micro pins on a tape with the sharp ends aligned on one side (Figure 2B). Use a pair of scissors to cut ~2 mm from the sharp ends of the micro pins (Figure 2B'). Use forceps to hold the cut micro pins and insert into the Sylgard layer of a pre-made Sylgard plate (Figure 2C). Two micro pins are used to immobilize one explant.
- Use a brush to collect white pupae, which form puparium within 1 h, of hsFLP, pebbled-GAL4/+; UAS-FRT100-stop-FRT100-mCD8-GFP8 genotype and transfer them to new vials. Heat shock in 37 °C water bath for 40 min to induce sparse ORN clones from random types. After heat shock, put the vials at 25 °C for 30 h, resulting in pupae aged at 30 h after puparium formation (APF).
- To prepare the culture medium for explant, add 5 mL of Penicillin-Streptomycin (10,000 U/mL) to 500 mL Schneider's Drosophila Medium. Filter the medium and make 45 mL aliquots in 50 mL conical tubes. The medium can be stored at 4 °C for 1-2 months.
- On the day of imaging, take one tube of 45 mL of Schneider's Drosophila medium and add 5 mL of Fetal Bovine Serum (10% v/v), 125 µL of 4 mg/mL human insulin stock solution (10 µg/mL final concentration), 50 µL of 1 mg/mL 20-hydroxyecdysone stock solution dissolved in ethanol (1 µg/mL final concentration). Mix well and transfer 15 mL of full medium into a new 50 mL conical tube. The rest of the full medium can be stored at 4 °C for a week. Fetal Bovine Serum, human insulin stock solution and 20-hydroxyecdysone stock solution are aliquoted and stored at -20 °C.
- Oxygenate the 15 mL full medium by pumping oxygen bubbles from an oxygen cylinder under the liquid surface through a sterile 5 mL pipette tip at the rate of one bubble/s for 20-30 min. Use a paraffin film to cover the opening of the tube during this process.
- Sterilize the dissection well surface and the Sylgard plate (with micro pins inserted on the Sylgard layer, prepared in steps A1 and A2) with 70% ethanol. Let them dry before use.
2. Explant dissection
- Use a brush to transfer 30 h APF (30 h after puparium formation) pupae to a paper tissue and dry the external surface of the pupae for 5 min.
- Put a piece of double-sided tape on a glass slide. Carefully attach the dried pupae on the sticky surface of the tape with the dorsal side facing upward. Gently press the pupae with a brush to help the ventral side of the pupae attach well to the tape (Figure 3A). Do not damage the pupa.
- Use a pair of forceps to remove brown pupal case covering the dorsal side of the head (Figure 3A,B). Insert one sharp tip of the forceps between the brown pupal case and the pupa from the lateral side and carefully break the brown pupal case through a line to the posterior end of the pupa (Figure 3B,C). Open the brown pupal case. Use a pair of forceps to gently hold the pupa and transfer to the dissection well with 1 mL of oxygenated full medium. Submerge floating pupa on the medium surface to help it sink to the bottom of the well (Figure 3D).
NOTE: Do not insert the forceps tip too deeply inside the brown pupal case to prevent injuring the pupa with the forceps. - To dissect the antennae-brain explant from the pupa, use forceps to gently hold the pupa with one hand and use a pair of microscissors to cut a small hole from the posterior side of the pupa with the other hand (Figure 3E). This small hole releases the high pressure inside the pupa.
- Cut through the ventral midline of the pupa from the hole until the neck (the narrow structure that connects the head and the thorax) with the microscissors (Figure 3F). Then, cut through the circumference of the neck to detach the head from the body of the pupa (Figure 3G). Remove the body and place it in a different well.
NOTE: Do not cut the neck directly from the dorsal/ventral side of the pupa, which may squeeze the brain. - Cut the transparent cuticle that covers the dorsal side of the brain (Figure 3H). This will expose the fat body on top of the brain. Keep some cuticle to which the retina and antennae attach. Repeat the same procedure to the ventral side of the brain.
NOTE: Do not insert the blade of the scissors too deeply under the cuticle as this will cause severing of the antennal nerves connecting the antennae and brain (Figure 3H'). - Use a P10 pipette to gently wash out the fat body that covers the brain and antennae by pipetting the medium toward the open regions on the dorsal and ventral sides of the head (Figure 3I).
NOTE: Be very gentle when pipetting the medium as the brain can easily be detached from the cuticle. Make sure all fat body is removed during this step. Arrested development of ORN axons were observed when fat body was not cleaned well, probably due to poor oxygen access from the medium. - To study the interaction of bilateral ORN axons or ORN axons to PN dendrite targeting, sever one or two antennal nerves with the microscissors during this stage8 (Figure 3J). Carefully place the blades of the scissors between the cuticle and the brain and sever interested antennal nerves.
- To transfer the dissected explant to the Sylgard plate, place a droplet of oxygenated full medium (~200 µL) on the Sylgard surface. Coat the inner surface of a 200 µL wide tip pipette tip with the fat body from the dissected trunk (step 1.6) by pipetting the fat body several times, which prevents the explants from sticking the pipette tip during transfer. Then, use this wide tip pipette tip to transfer the explant from the dissection well to the medium droplet on the culture plate (Figure 3K).
- Use forceps to pin the explant on the Sylgard layer in the two optic lobes (Figure 3L). Carefully position the Sylgard plate on the imaging station and immobilize the plate with tapes. Slowly add 10 mL of oxygenated full medium to the Sylgard plate using P1000 pipette.
NOTE: Avoid disrupting the explant when adding the medium to the Sylgard plate.
3. Two-photon microscopy-based live imaging
- To perform time-lapse imaging, use a two-photon microscope, a Ti:Sapphire laser, a 20x water-immersion objective (1.0 NA) and an imaging software. Use the excitation wavelength at 920 nm for imaging GFP proteins. Adjust the pixel dwell time to 10 µs.
- Adjust the imaging station position so that the explants are roughly under the objective. Use 70% ethanol to sterilize the lens before imaging. Slowly lower the objective under the medium close to the explants. Check whether there is any bubble on the lens of objective.
- If so, lift the objective above the medium and repeat this until the bubble is gone. Find the explants using the eyepiece and center one explant in the field.
- To ensure an explant with a few ORNs sparsely labeled for time lapse imaging, dissect ~10 explants each time and align on y axis on the culture plate. Screen all explants by moving the objective along y axis and choose an explant in which a few single ORN axons have just reached the antennal lobe for imaging (Figure 4A).
- Recognize the antennal lobe by its oval shape and the ORN axons that are beginning to circumnavigate it. Image a ~150 µm x 150 µm area in the xy plane (3x zoom with the 20x objective). Estimate the boundary of the two antennal lobes and center them in the imaging area.
- Select an initial imaging region along the z axis by defining the bottom section and top section of scanning. Set up imaging area along the z axis. Set the deepest section with ORN axon signals as the first imaging session and the session 100 µm above (more superficial side) as the last imaging session (Figure 4B).
NOTE: This leaves some sections on top (superficial side) of the ORN axons and avoids shifting of ORN axons upward outside the imaging area due to the growth of the brain during culture.- Image at 2 µm intervals. Set automatic imaging scanning at the frequency of every 20 min using imaging software.
- Shift the imaging region 20 µm upward along the z axis after the first 4 h imaging and another 20 µm upward along the z axis after 16 h imaging. This can be achieved by setting a script and different z stacks in the imaging software.
- Culture the explant for an additional period post imaging (up to 24 h ex vivo) before fixation and staining with N-cadherin, a neuropil marker, to reveal the genetic identity of each single ORN by the glomerulus it targets to.
4. Image processing
- To process z stack images from section series taken at each time point using the Fiji software, open image section series, click on Image | Stacks | Z project [/].
- To correct lateral drift of the sample during culture, install TurboReg Plugin in Fiji.
- Open a z stack image series and a single z stack image from the series. Open Plugins | Registration | TurboReg.
- Select the z stack image series in Source and the single z stack image in Target | Translation. Click on the Batch button to register all the images from the opened z stack image series.
- To maximize the utility of imaged samples, separate sparsely labeled single axons in the vicinity to each other from the z stack images following 3D image sections.
- To extract single ORN axons from a few axons in the same image, open the image section series, click on Plugins | Segmentation | Segmentation Editor [/]. Select the brush tool and mask interested ORN axon in the Segmentation Editor working window using Select "+" or "–" buttons on each image section.
- Click on Process | Image Calculator [/]. Select "Image X" in Image 1, "Multiply" in Operation, "Image X. labels" in Image 2, [/]. This generates a new image series file with the interested axon only. Perform step 4.1 to process the z stack image. Repeat this step for all time points to generate a time series image file.
- To pseudocolor different axons from the same image, first perform step 4.3 to generate the time series image file of each axon separately. Open the time series image files for different single axons from the same raw data image. Click on Image | Color | Merge Channels. Select different time series image files in different color channels and click OK.
Representative Results
ORN axons arrive at the antennal lobe between 18 h and 36 h APF. They then navigate the antennal lobe, cross the midline, and innervate the glomeruli. Video 1 is a representative video showing the entire process for several individually identifiable axons, taken at the frequency of every 20 min for 24 h. Before registration using TurboReg, the axons exhibit some lateral drifting as the brain develops (first half of the video). After registration, the drifting is corrected (second half of the video).
To separate a few ORN axons from the same explant, one example is shown in Figure 5. Following the procedure in step 4.3, VA1d and VA1v axons from explant shown in Figure 5A were extracted to generate a new z stack image with only these two axons (Figure 5B). Similarly, VM2 and VM3 axons (Figure 5B') and DL2 axon (Figure 5B'') were extracted. Figure 5C shows a merge of images in Figure 5B-B'' with pseudocolors. The genetic identities of each ORN axons were revealed by immunostaining of a neuropil marker N-cadherin of the fixed explant (Figure 5D,E).
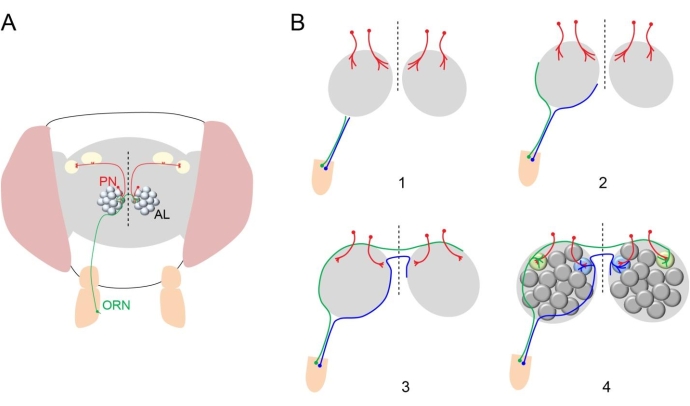
Figure 1: Structure of fly olfactory circuit. (A) An adult fly head is shown with one ORN from the right antennae (green) sending its axon to both antennal lobes (ALs) in the brain and forming synaptic connection in a specific glomerulus with dendrites of PNs (red) in the ipsilateral and contralateral ALs. Dashed vertical line indicates midline in this and subsequent diagrams and images. (B) Diagram showing the olfactory circuit development. (1) PN dendrites first innervate a region in the antennal lobe (red). ORN axons reach the antennal lobes in the brain. (2) ORN axons take either a dorsolateral (green) or ventromedial (blue) trajectory to circumnavigate the antennal lobe. (3) ORN axons cross the midline. (4) ORN axons innervate glomeruli in the antennal lobe. Please click here to view a larger version of this figure.
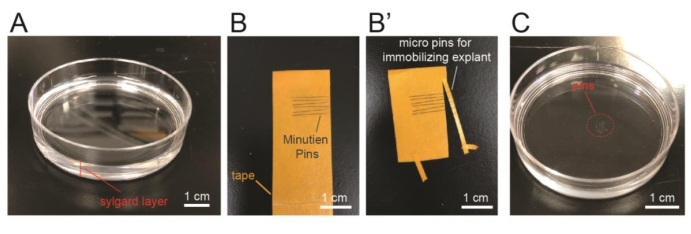

Figure 2: Preparation of the imaging chamber for the explant. (A) Lay a layer of silicone elastomer (~0.5 cm) at the bottom of a 60 mm Petri dish. (B–B') Align pins on a tape and cut to ~2 mm long using a pair of scissors. (C) Pin the micro pins on the silicone elastomer layer of the culture plate. Please click here to view a larger version of this figure.
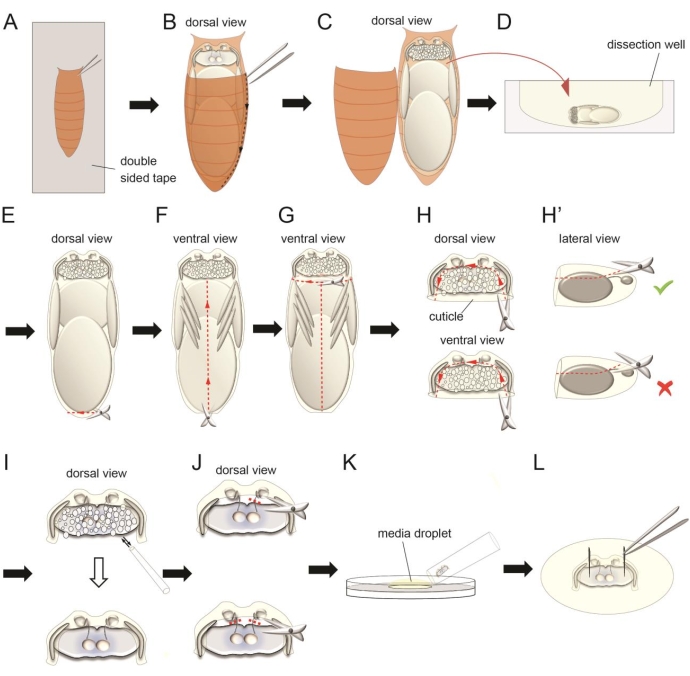
Figure 3: The dissection procedure for the antennae-brain explant. (A) Attach the ventral side of a paper tissue-dried pupa on a double-sided tape on a glass slide. (B–C) Use forceps to cut the external brown cuticle to expose the pupa inside. (D) Transfer the pupa to a dissection well with oxygenated full medium. (E–G) Carefully separate the pupal trunk from the head using microscissors. (H) Cut the pieces of semitransparent cuticle covering the dorsal and ventral sides of the brain. Keep some cuticle on the anterior and lateral sides of the brain to retain connections between the retina, antennae, and brain. (H') Avoid severing the antennal nerve during this step. (I) Clean the fat body covering the brain by gently pipetting. (J) Sever one or two antennal nerve(s) using microscissors in certain experiments. (K) Place a droplet of oxygenated full medium on the surface of the culture plate. Transfer the dissected explant using a wide tip pipette tip. (L) Use forceps to pin the two optic lobes of the explant on the silicone elastomer layer. Please click here to view a larger version of this figure.
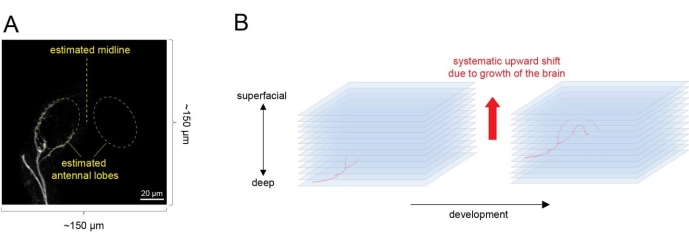
Figure 4: Time-lapse imaging of single ORN axon targeting from an explant. (A) Select an explant with a few ORN axons just reaching the antennal lobe. Estimate the shape of two antennal lobes by the curvature of the axons and center the antennal lobes in the imaging field. (B) Set the imaging region along the z axis. Consider that the antennal lobe will shift upward as the brain grows and develops. Please click here to view a larger version of this figure.
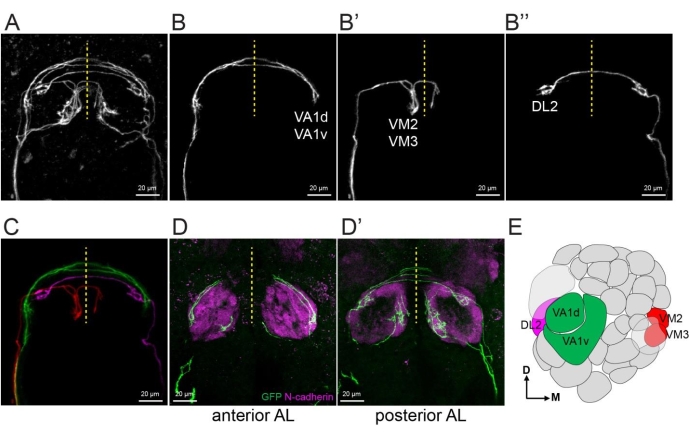
Figure 5: Extract single ORNs and reveal their glomerular identities. (A) A maximum projection image of an explant with 5-10 single ORN axons using two-photon microscopy with 20x objective and 3x zoom in. (B–B'') 1-2 single axons are extracted from (A) by manually creating masks in image sections from the raw image data. (C) Merge the images of (B-B'') with each axon pseudo-colored differently. (D–D') Maximum projection confocal images taken with 40x objective and 1.5x zoom. Explant shown in (A) was fixed followed by staining with anti-GFP and anti-N-cadherin (neuropil marker). Anterior and posterior halves of the antennal lobes are stacks separately in (D) and (D'). (E) Antennal lobe map shows extracted ORN axons in (B,C). Some images shown in this figure are modified from a prior study8. Please click here to view a larger version of this figure.
Video 1: Two-photon microscopy based time-lapse images show targeting of two ORN axons, before and after image registration. Please click here to download this Video.
Discussion
The Drosophila antennae-brain explant retains normal targeting of the olfactory circuit. We did notice that the development is 2 times slower ex vivo compared to in vivo. It is noted that the explant system does not retain maxillary palp, which hosts six types of ORNs. To ensure normal development is recapitulated ex vivo, stretching of the antennal nerves needs to be avoided during explant dissection. During ex vivo culture bacteria growth usually causes arrested development of the olfactory circuit. Therefore, thorough sterilization of the culture dish and pins before imaging and keeping the imaging room clean and isolated is important.
This explant supports long-term two-photon microscopy based time-lapse imaging. Combined with a newly developed reporter for sparse labeling of single ORNs, the explant system allows high-resolution imaging from single axon terminus. This system is powerful to study the cell biological mechanisms underpinning the dynamic process of olfactory circuit assembly8. Although olfactory circuit development was shown here as an example, this system can potentially be expanded to studies of other circuits or other developmental processes in the developing central brain.
The explant maintains normal development in culture for at least 24 h, which can capture the entire process of ORN targeting. It enables researchers to reveal the genetic identity of single ORN axons through counter-staining with a neuropil marker post fixation, as the antennal lobe already develops obvious glomerular structure by the end of the culture. This strategy circumvents the problem of lacking specific genetic drivers for many ORN types at an early developmental stage to achieve imaging of specific types of ORNs using a pan-ORN driver.
To achieve higher spatiotemporal resolution, this explant system can be imaged using more advanced microscopy, the adaptive optics-lattice light-sheet microscopy (AO-LLSM). It has been shown that the AO-LLSM enables visualization of fine structures of axon terminals and scanning frequency at every 30 second per volume8,9,10,11. One advantage of the explant is its compatibility with Janelia Fluorophore dye12,13,14 labeling by incubating the explant expressing Halo-tag in specific neurons with dyes in the medium before imaging. It was noticed that incubating the explant with dye-containing medium results in much stronger labeling than feeding the larvae with the dye. This unique advantage allowed us to image axons at an early developmental stage that was hardly visualized by GFP labeling8.
In addition to time-lapse imaging, the explant system has other advantages. For example, it is possible to sever antennal nerves from the dissected explant unilaterally or bilaterally at specific developmental time points (step 2.8). This allows researchers to probe the requirement of ORN axons in the targeting of any neuron types in the olfactory circuit at distinct developmental steps. In particular unilateral antennal nerve severing assays, which cannot be achieved by traditional genetic manipulation, led to an interesting discovery that interaction between bilateral ORN axons is required for correct contralateral targeting of ORN axons8. Furthermore, the explant is cultured directly in medium instead of being embedded in agarose, therefore allowing fast delivery and wash out of some small molecules or drugs. Compared with genetic manipulation, drug treatment has the advantages of rapid effect and reversibility of the manipulation. It enables researchers to assess some essential processes for cells at later developmental stages by bypassing cell lethality or unhealthy issues due to constitutive disruption through genetic manipulations. It also helps visualization of subtle changes by comparing before and after drug treatment, and after drug washout.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank N. Özel and R. Hiesinger for their advice on the explant culture; M. Wagner for technical help of the two-photon microscopy; D.J. Luginbuhl for generating transgenic flies; D. Friedmann for suggestions of Fiji software analysis; Y. Ge for assistance on fly work; C. McLaughlin and K.K.L. Wong for comments on the manuscript. L.L. is a Howard Hughes Medical Institute investigator. This work was supported by National Institutes of Health grants 1K99DC01883001 (to T.L.) and R01-DC005982 (to L.L.).
Materials
| 20-hydroxyecdysone | Sigma | H5142 | |
| Chameleon Ti:Sapphire laser | Coherent | Coherent MRU X1 | |
| Fetal Bovine Serum | Thermo Fisher Scientific | 10082147 | |
| Human insulin | Thermo Fisher Scientific | 12585014 | |
| Imaging software | Prairie | ||
| Micro Scissors | World Precision Instruments | 501778 | |
| Minutien Pins | Fine Science Tools | 26002-10 | |
| Oxygen cylinder | Praxair | OX M-E | |
| Penicillin-Streptomycin | Thermo Fisher Scientific | 15140122 | |
| Schneider’s Drosophila Medium | Thermo Fisher Scientific | 21720024 | |
| SYLGARD 184 Silicone Elastomer | Thermo Fisher Scientific | NC0162601 | |
| Two-photon microscopy | Bruker | ||
| water immerse objective (20X) | Zeiss | 421452-9800-000 |
References
- Kolodkin, A. L., Tessier-Lavigne, M. Mechanisms and molecules of neuronal wiring: a primer. Cold Spring Harbor Perspective Biology. 3 (6), 001727 (2011).
- Vosshall, L. B., Stocker, R. F. Molecular architecture of smell and taste in Drosophila. Annual Review Neuroscience. 30, 505-533 (2007).
- Hong, W., Luo, L. Genetic control of wiring specificity in the fly olfactory system. 유전학. 196 (1), 17-29 (2014).
- Bentley, D., Caudy, M. Pioneer axons lose directed growth after selective killing of guidepost cells. Nature. 304 (5921), 62-65 (1983).
- Godement, P., Wang, L. C., Mason, C. A. Retinal axon divergence in the optic chiasm: dynamics of growth cone behavior at the midline. Journal of Neuroscience. 14 (11), 7024-7039 (1994).
- Harris, W. A., Holt, C. E., Bonhoeffer, F. Retinal axons with and without their somata, growing to and arborizing in the tectum of Xenopus embryos: a time-lapse video study of single fibres in vivo. Development. 101 (1), 123-133 (1987).
- Ozel, M. N., Langen, M., Hassan, B. A., Hiesinger, P. R. Filopodial dynamics and growth cone stabilization in Drosophila visual circuit development. Elife. 4, 10721 (2015).
- Li, T., et al. Cellular bases of olfactory circuit assembly revealed by systematic time-lapse imaging. Cell. 184, 5107-5121 (2021).
- Chen, B. C., et al. Lattice light-sheet microscopy: Imaging molecules to embryos at high spatiotemporal resolution. Science. 346 (6208), 1257998 (2014).
- Liu, T. L., et al. Observing the cell in its native state: Imaging subcellular dynamics in multicellular organisms. Science. 360 (6386), (2018).
- Wang, K., et al. Rapid adaptive optical recovery of optimal resolution over large volumes. Nature Methods. 11 (6), 625-628 (2014).
- Kohl, J., et al. Ultrafast tissue staining with chemical tags. Proceedings of the National Academy of Science U. S. A. 111 (36), 3805-3814 (2014).
- Sutcliffe, B., et al. Second-Generation Drosophila Chemical Tags: Sensitivity, Versatility, and Speed. 유전학. 205 (4), 1399-1408 (2017).
- Grimm, J. B., Brown, T. A., English, B. P., Lionnet, T., Lavis, L. D. Synthesis of Janelia Fluor HaloTag and SNAP-Tag Ligands and Their Use in Cellular Imaging Experiments. Methods Molecular Biology. 1663, 179-188 (2017).

