Construction of Homozygous Mutants of Migratory Locust Using CRISPR/Cas9 Technology
Summary
This study provides a systematically optimized procedure of CRISPR/Cas9 ribonuclease-based construction of homozygous locust mutants as well as a detailed method for cryopreservation and resuscitation of the locust eggs.
Abstract
The migratory locust, Locusta migratoria, is not only one of the worldwide plague locusts that caused huge economic losses to human beings but also an important research model for insect metamorphosis. The CRISPR/Cas9 system can accurately locate at a specific DNA locus and cleave within the target site, efficiently introducing double-strand breaks to induce target gene knockout or integrate new gene fragments into the specific locus. CRISPR/Cas9-mediated genome editing is a powerful tool for addressing questions encountered in locust research as well as a promising technology for locust control. This study provides a systematic protocol for CRISPR/Cas9-mediated gene knockout with the complex of Cas9 protein and single guide RNAs (sgRNAs) in migratory locusts. The selection of target sites and design of sgRNA are described in detail, followed by in vitro synthesis and verification of the sgRNAs. Subsequent procedures include egg raft collection and tanned-egg separation to achieve successful microinjection with low mortality rate, egg culture, preliminary estimation of the mutation rate, locust breeding as well as detection, preservation, and passage of the mutants to ensure population stability of the edited locusts. This method can be used as a reference for CRISPR/Cas9 based gene editing applications in migratory locusts as well as in other insects.
Introduction
Gene editing technologies could be used to introduce insertions or deletions into a specific genome locus to artificially modify the target gene on purpose1. In the past years, CRISPR/Cas9 technology has developed rapidly and has a growing scope of applications in various fields of life sciences2,3,4,5,6. The CRISPR/Cas9 system was discovered back in 19877, and widely found in bacteria and archaea. Further research indicated that it was a prokaryotic adaptive immune system that depends on the RNA-guided nuclease Cas9 to fight against phages8. The artificially modified CRISPR/Cas9 system mainly consists of two components, a single guide RNA (sgRNA) and the Cas9 protein. The sgRNA is made up of a CRISPR RNA (crRNA) complementary to the target sequence and an auxiliary trans-activating crRNA (tracrRNA), which is relatively conserved. When the CRISPR/Cas9 system is activated, the sgRNA forms a ribonucleoprotein (RNP) with the Cas9 protein and guides Cas9 to its target site via the base pairing of RNA-DNA interactions. Then, the double-strand DNA can be cleaved by the Cas9 protein and as a result, the double-strand break (DSB) emerges near the protospacer adjacent motif (PAM) of the target site9,10,11,12. To mitigate the damage caused by the DSBs, cells would activate comprehensive DNA damage responses to efficiently detect the genomic damages and initiate the repair procedure. There are two distinct repair mechanisms in the cell: non-homologous end joining (NHEJ) and homology-directed repair (HDR). NHEJ is the most common repair pathway that can repair DNA double-strand breaks quickly and prevents cell apoptosis. However, it is error-prone because of leaving small fragments of insertions and deletions (indels) near the DSBs, which usually results in an open reading frame shift and thus can lead to gene knock-out. In contrast, homologous repairment is quite a rare event. On the condition that there is a repair template with sequences homologous to the context of the DSB, cells would occasionally repair the genomic break according to the nearby template. The result of HDR is that the DSB is precisely repaired. Especially, if there is an additional sequence between the homologous sequences in the template, they would be integrated into the genome through HDR, and in this way, the specific gene insertions could be realized13.
With the optimization and development of the sgRNA structures and Cas9 protein variants, the CRISPR/Cas9-based genetic editing system has also been successfully applied in research of insects, including but not limited to Drosophila melanogaster, Aedes aegypti, Bombyx mori, Helicoverpa Armigera, Plutella xylostella, and Locusta migratoria14,15,16,17,18,19. To the best of the authors' knowledge, although RNPs consisting of the Cas9 protein and in vitro transcribed sgRNA have been used for locust genome editing20,21,22, a systematic and detailed protocol for CRISPR/Cas9 ribonuclease mediated construction of homozygous mutants of the migratory locust is still lacking.
The migratory locust is an important agricultural pest that has a global distribution and poses substantial threats to food production, being especially harmful to gramineous plants, such as wheat, maize, rice, and millet23. Gene function analysis based on genome editing technologies can provide novel targets and new strategies for the control of migratory locusts. This study proposes a detailed method for knocking out migratory locust genes via the CRISPR/Cas9 system, including the selection of target sites and design of sgRNAs, in vitro synthesis and verification of the sgRNAs, microinjection and culture of eggs, estimation of mutation rate at the embryonic stage, detection of mutants as well as passage and preservation of the mutants. This protocol could be used as a basal reference for manipulation of the vast majority of locust genes and can provide valuable references for genome editing of other insects.
Protocol
1. Target site selection and sgRNA design
- Collect as much sequence information as possible for the gene of interest through literature research and/or searching for the mRNA or coding DNA sequence (CDS) of the gene at NCBI and locust database24.
- Compare the sequence of the interested gene to its genomic DNA sequence to distinguish the exon and intron regions.
- Select a candidate region for target site design based on the research purpose. Design primer pairs to amplify the candidate region fragment and verify its wild-type sequence by sequencing analysis of the PCR product.
NOTE: There are different strategies for the selection of the candidate target region. For example, in most gene/protein function research, use the exon fragment close to its start codon as the candidate region to ensure that the whole protein/RNA function is lost after the genome editing (i.e., gene knock-out). While for the function analysis of a specific domain, select the candidate region at the beginning (or both ends) of the domain. - Use online resources such as the E-CRISP Design25 to search for potential target sites in the candidate target region.
- Select "Drosophila melanogaster BDGP6" (or any other insect) in the drop-down list of reference genomes and choose "Input is FASTA sequence" followed by pasting the sequence of target region into the textbox (in FASTA format).
- Start the application by pressing "Start sgRNA search" with "medium" and "single design" to get the possible target sites. Select 1-3 potential targets from the results based on their predicted scores and design the corresponding sgRNAs accordingly.
NOTE: There are many more online platforms for CRISPR design, such as CRISPOR, CHOPCHOP, CRISPRdirect, ZiFiT, and so on (see Table of Materials). Some of them have the function of specificity check for the species whose genomic sequence data is in their databases. However, the genomic sequence of locusts has not been found on any online website. It is better to use more than one online tool for the CRISPR design in locusts. Comprehensively, consider the results from different websites and select the sgRNA from the results on the principle of highest specificity, highest efficiency, and least mismatch rate (Figure 1A,B).
2. Synthesis and verification of the sgRNA in vitro
- Synthesize the sgRNA using sgRNA synthesis kits according to the manufacturer´s manual (Table of Materials). This procedure usually includes three steps: DNA template amplification, in vitro transcription, and sgRNA purification21. Dilute the synthesized sgRNA with nuclease-free water to a storage concentration of 300 ng/µL for further use.
- Amplify and purify the target gene fragment to serve as the substrate of the in vitro cleavage assay of the CRISPR/Cas9 system. Perform these steps following the manufacturer's instructions.
- Dilute the purchased Cas9 protein to 300 ng/µL with nuclease-free water. Incubate 1 µL of sgRNA (300 ng/µL) and 1 µL of Cas9 protein (300 ng/µL) with 200 ng of purified target fragment in the cleavage buffer at 37 °C for 1 h (10 µL of total volume; see Table 1). Estimate the efficiency of sgRNA induced Cas9 cleavage by agarose gel electrophoresis (Figure 1C). Select the sgRNAs with high activity for the following microinjection.
NOTE: The results of agarose gel electrophoresis can be converted to grayscale images with image processing software and the cleavage efficiency is estimated according to the grayscale of the bands of speculated sizes.
3. Microinjection and culture of the eggs
- Prepare the injection needles by pulling glass capillaries with a micropipette puller (Table of Materials). Set the parameters as follows: Heat to 588, Pull to 90, Velocity to 60, and Time to 40. Grind the tip of the injection needle with a micro grinder (Table of Materials).
NOTE: Ideal needle tips are open and sharp-edged (Figure 2A). It is highly recommended to prepare additional needles for the experiments because needles are sometimes blocked or accidentally broken during injections. - Mix 1 µL of Cas9 protein (300 ng/µL) with 1 µL of the verified sgRNA (300 ng/µL) together to obtain the RNPs for injection. Add 8 µL of RNase-free sterile water to make the final volume of the mixture to 10 µL. Mix the solution thoroughly and keep it on ice.
NOTE: The final concentrations of the Cas9 protein and sgRNAs can be optimized according to the editing results. Avoid repetitive freezing and thawing of the prepared RNP solution and immediate use is recommended. - Rear the male and female adult locusts together at 30 °C with a 16 (light):8 (dark) photoperiod and supply them sufficient fresh wheat seedlings as food. Observe these locusts daily and put an oviposition pot (a plastic flowerpot or cup filled with wet sterile sand) into the rearing cage for oviposition once the locusts mated.
NOTE: Usually, 100 pairs of adult locusts reared in a rearing cage (40 cm x 40 cm x 40 cm) are enough to generate eggs for knocking out a single gene. Culture additional locust pairs if more eggs are needed. - Collect the freshly-laid egg pods from the oviposition pot and wait for about 30 min for egg tanning. Gently isolate the eggs from the egg pods in water using a fine brush and wash them with sterile water three times. Keep the eggs in a Petri dish and add sterile water to keep them moist.
NOTE: Tanning of the newly-laid eggs can significantly improve the mutant efficiency21. - Fill an injection needle with the prepared RNP solution and load it to the micromanipulator (Table of Materials).
- Set the microinjection parameters as follows: 300 hPa of the injection pressure (pi), 0.5 s of the injection time (ti), and 25 hPa of the compensation pressure (pc).
- Press the pedal to evaluate the volume of the injected solution. Adjust the injection pressure and injection time to ensure that the injection volume is about 50-100 nL.
NOTE: Exhaust the residual air in the needle to make the injection solution continuous and controllable as much as possible. The connection of the needle and the micromanipulator need to be tight.
- Arrange the sterile eggs regularly on the injection pad (Figure 2B,C) and place the pad on the object table of the microscope (Figure 3A). Adjust the magnification of the microscope until the eggs are observed. Adjust the microinjection needle under the microscope until the injection tip can be seen and position it near the egg to be injected.
- Start the injection at a suitable height and an angle of 30-45 degrees. Insert the tip into the egg carefully near the micropyles of the egg (Figure 3B) and press the pedal to accomplish the injection. Retract the needle quickly and move the injection pad for injection of the next egg.
NOTE: A slight expansion of the egg during the microinjection should be observed. A small amount of cytoplasmic leakage at the pinhole is acceptable. Replace the injection needle with a new one or adjust the injection angle if the outflow of fluid is too much. - Transfer the injected eggs to a culture dish (a Petri dish with a piece of moist filter paper), and place them in an incubator at 30 °C.
NOTE: It will take about 13-14 days till nymphs hatch from these injected eggs (on condition that mutation of the target gene does not affect the development of the embryos) (Figure 3C). Inject at least 100 eggs to ensure a sufficient hatching and eclosion amount for knocking out a specific gene.
4. Mutation rate estimation and the screening of mutants
- Check the development of the injected eggs every day after the injection. Transfer injected eggs to a new culture dish every 24 h in the first 5 days.
- On the 6th day (or later) after injection, randomly pick and transfer 10 eggs into a tube and adequately grind the eggs with two steel balls at 60 Hz for 6 min using a grinder (see Table of Materials). Resuspend the debris with 1 mL of PBS. Transfer 5 µL of the mixture into 45 µL of 50 mM NaOH and lyse at 95 °C for 5 min. Add 5 µL of 1 M Tris-HCl (pH=8.0) into the lysis system to terminate the alkaline lysis reaction.
NOTE: Eggs can be picked out for alkaline lysis on any day after the last transfer and multiple eggs can be used as one sample depending on their development situation (e.g., five 6-day-old embryos can be used as one sample, and two 10-day-old embryos can be used as one sample). Sometimes, it is not feasible to get the genomic DNA of eggs using the alkaline lysis method. Commercial genomic DNA extraction kits can be used as an alternative method for isolating the genomic DNA from the eggs. - Take 1 µL of the lysis product as the PCR template to amplify the target gene fragment (Table 2 and Table 3) and send the PCR products for sequencing. Compare the sequencing results with the wild-type sequence to preliminarily evaluate whether the CRISPR/Cas9 system cleaved the target gene in vivo (Figure 4A). Allow the rest of the eggs for subsequent development on the condition that indels are detected at the embryonic stage.
NOTE: In this way, the mutation rate can be preliminarily estimated at the embryo stage. If no indels are found at this step, prepare some new sgRNAs for the target gene and repeat the knock-out protocol from step 2.1. - Transfer the hatched nymphs to a rearing cage and culture them as described in step 3.3. When the nymphs developed to the fifth instar stage, separate them with plastic culture cups (1 nymph/cup).
NOTE: It usually takes about 25-35 days for the nymphs to develop to their fifth instar and the most obvious phenotype is that the wing buds extend to the fourth or fifth abdominal segments. Moreover, it is recommended to record the phenotypes of dead nymphs to predict the relationship between the target gene and phenotypes in further research. - Cut off about 2 mm length of the antennae with dissecting scissors and lyse it using the alkaline lysis method (step 4.2). Analyze the target gene fragment sequence of these nymphs as described above (step 4.3) to identify the G0 mutants (Figure 4B).
NOTE: The antennae cut for lysis can be a little longer and grinding or mincing the antennae is helpful for lysis although the antennae can be directly digested. Individuals with multiple peaks near the target site in the sequencing result are identified as positive mutants and allowed for subsequent development and mating.
5. Establishment and passaging of mutant lines
- Perform cross breeding using the G0 mutants and wild-type locusts (Figure 4B). Collect the oocysts and incubate these oocysts separately at 30 °C. Use 3-6 developed eggs in each oocyst to detect mutations as described in steps 4.2 and 4.3. Keep mutation-positive oocysts for the subsequent development and abandon the mutation-negative oocysts.
NOTE: Mutation rate estimation at the embryonic stage can greatly accelerate the screening of G1 mutants. Usually, 3-6 developed eggs can be mixed as one sample for PCR-mediated genotyping as described above. - Rear the G1 nymphs as described in step 4.4. Cut off about 2 mm length of the antennae to perform PCR-based genotyping as described in step 4.5 for detecting G1 heterozygotes. Perform TA-cloning according to the manufacturer's instructions (see Table of Materials) to identify their mutations. Perform in-cross using G1 heterozygotes with the same mutations to obtain G2 nymphs.
NOTE: Sanger sequencing of the PCR products can only provide information for finding out the heterozygotes, but not enough information for identifying the exact mutations. Thus, TA cloning using the PCR products is required to clearly identify the mutations and can promote the establishment of stable mutant lines. - Rear the G2 nymphs till their fifth instar. Use the PCR-based genotyping described in step 5.2 to identify the homozygotes and/or heterozygotes that are suitable for further research and stable passaging (Figure 4B).
NOTE: Pay attention to avoid mixing of homozygotes and heterozygotes. It is recommended to perform PCR-based genotyping in every generation to confirm the homozygosity and/or heterozygosity of each population. The number of locusts used for this check depends on the population size. Moreover, the population of locusts can be expanded by the in-cross strategy.
6. Egg cryopreservation and resuscitation
- Wash the eggs to be cryopreserved with sterile water and incubate them in a culture dish as described above (step 3.8) for 5-6 days. Gather these eggs together in the culture dish and cover them with filter paper fragments. Wrap the entire culture dish with a paraffin film.
- Keep the dish at 25 °C for 2 days followed by another 2 days at a relatively lower temperature (13-16 °C). Then, transfer the dish to a 6 °C refrigerator. Add water into it every 2 weeks to provide a moist environment for the eggs (Figure 5A).
NOTE: The embryos are speculated to be at the katatrepsis stage after being developed for 5-6 days at 30 °C26. Gradient cooling is helpful for cryopreservation of eggs (Table 5). - For resuscitating the cryopreserved eggs, take the Petri dish out from the refrigerator and keep it at 25 °C for 2 days. Place these eggs in a 30 °C incubator until the nymphs hatch (Figure 5B).
NOTE: It is necessary to put the cryopreserved eggs at 25 °C before transferring them to a 30 °C incubator. The embryos can be cryopreserved for at least five months although the hatching rate and eclosion rate can decrease because of the cryopreservation (Table 5).
Representative Results
This protocol contains the detailed steps for generating homozygous mutants of the migratory locusts with the RNP consisting of Cas9 protein and in vitro synthesized sgRNA. The following are some representative results of CRISPR/Cas9-mediated target gene knockout in locusts, including target selection, sgRNA synthesis and verification (Figure 1A), egg collection and injection, mutant screening and passaging, cryopreservation, and resuscitation of the homozygous eggs.
In this study, the target site for CRISPR/Cas9 system is selected according to the results of three online programs (the E-CRISP, CRISPOR, and ZiFit) and located in the first exon (Figure 1B). According to the Cas9 cleavage assay in vitro (Table 1), the CRISPR/Cas9 RNP could digest the PCR fragment (containing the target site) with a cleavage rate of about 55% (Figure 1C). Then, this RNP was microinjected into 120 fertilized eggs at their single-cell stage using the standard microinjection system (Figure 2 and Figure 3). The sequencing results for embryonic stage mutation rate estimation suggested efficient genome editing at the target site (Figure 4A). Further, this study resulted in a 52.73% nymph hatching rate and 66.7% of the G0 adults were mosaic mutants (Table 4).
Further, the G0 chimeras were crossed with wild-type locusts to obtain heterozygous G1 individuals and the G1 heterozygotes with the same mutations (screened by TA-cloning) were in-crossed to generate the G2 animals. PCR-based phenotyping was used for detecting mutants and the obtained homozygotes were in-crossed to establish stable mutant lines (Figure 4B).
Meanwhile, the excess homozygous eggs were cryopreserved to improve the utilization rate of the homozygotes. Although the hatching rate of the cryopreserved eggs was reduced with the extension of cryopreservation time (Table 5), the remedial actions including applying filter paper fragments for keeping eggs wet and recovering these preserved eggs with a gradient of rising temperature were both helpful for the resuscitation (Figure 5 and Table 5). Finally, the homozygous population of mutants was successfully kept for subsequent research.
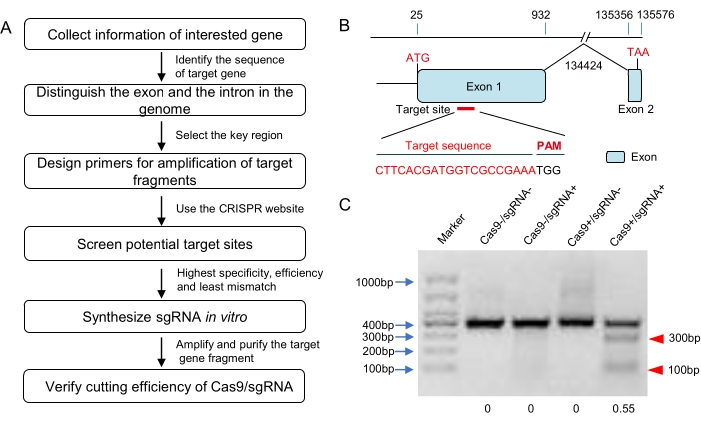
Figure 1: Design of the target and in vitro verification of the sgRNA. (A) The flow diagram for target selection, sgRNA synthesis, and bioactivity verification in vitro (including but not limited to locusts). (B) Schematic diagram of the gene structure and target site. The exons of this target gene are shown as blue domains and the target site was selected downstream of the start codon in exon 1. The target sequence is highlighted in red. (C) The agarose gel electrophoresis result of in vitro Cas9 cleavage assay. A DNA fragment harboring the target sequence (about 400 bp in length) was amplified and used as the substrate for Cas9 digestion. The expected small bands (about 100 bp and 300 bp here; marked with red triangles) suggested that the synthesized sgRNA could induce effective Cas9 cleavage. The cleavage rate was about 55% according to the grayscale analysis. Please click here to view a larger version of this figure.
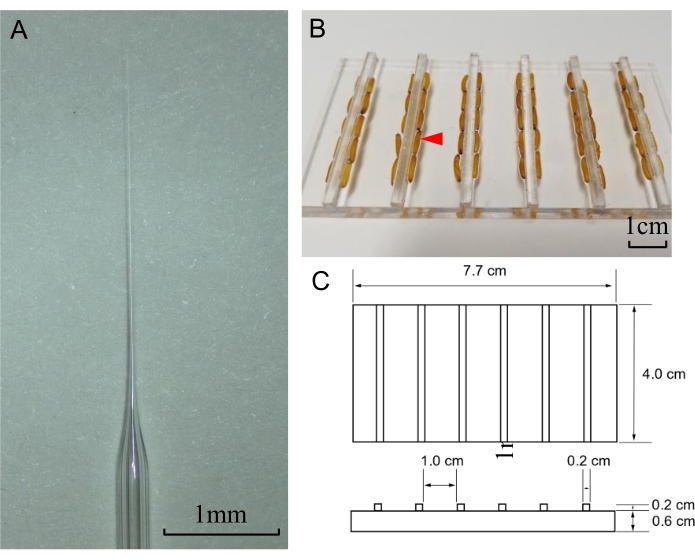
Figure 2: Needle preparation and the microinjection pad for locust eggs. (A) Tip of a prepared needle for microinjection of locust eggs. Scale bar: 1 mm. (B) Picture of a microinjection pad with eggs (indicated with a red triangle) arranged on it. The scale bar represents 1 cm. (C) The size of the microinjection pad. Please click here to view a larger version of this figure.
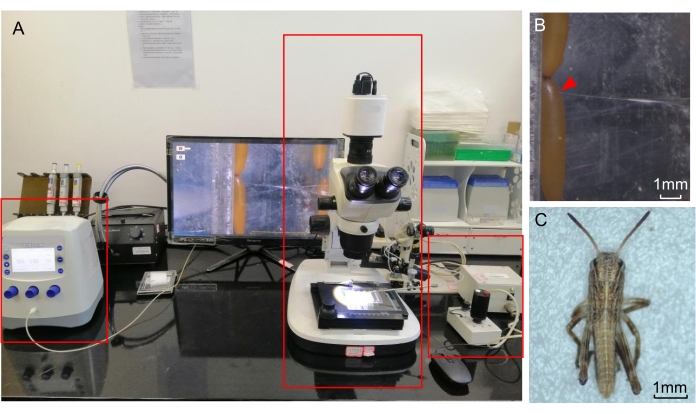

Figure 3: Egg microinjections. (A) A representative microinjection system labeled with red boxes indicating a microscope (middle) and a micromanipulator (right) connected to a microinjector (left). The computer is used for data storage and for providing auxiliary observation. (B) Injection of the locust egg. The injection site is marked with a red triangle. (C) A hatched nymph under the microscope. The scale bars represent 1 mm. Please click here to view a larger version of this figure.
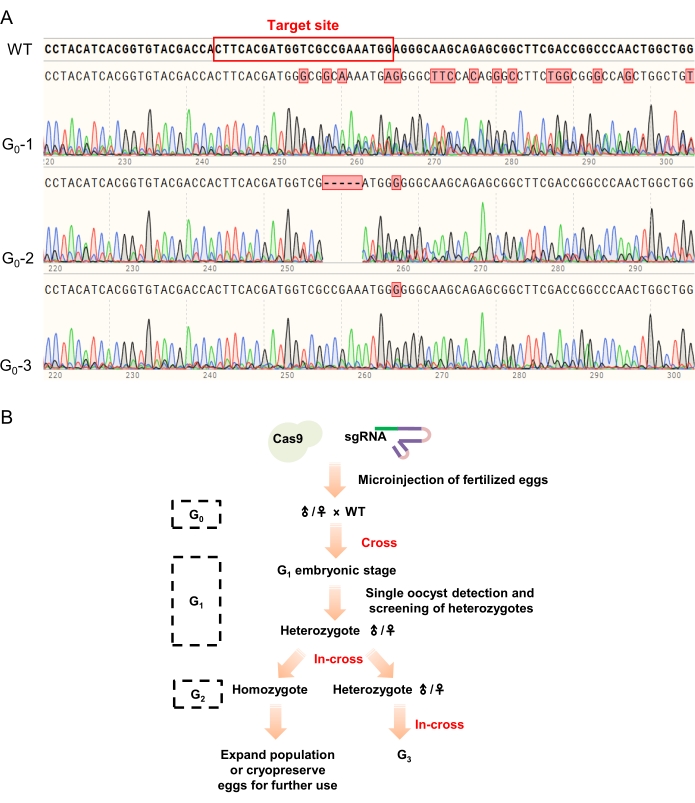
Figure 4: The sequencing results of G0 animals and passaging strategy of the mutant lines. (A) The typical sequencing results in the G0 screening. Multiple peaks near the target site (marked with a red rectangle in the wild-type sequence) indicated that the tested egg/locust was successfully edited by CRISPR/Cas9 system (as shown in G0-1 and G0-2), while the individual without any change in the sequencing result was considered as not edited and abandoned (e.g., G0-3). (B) The passaging strategy of the mutant lines. The G0 animals were firstly screened by PCR-mediated genotyping and those with multiple peaks in their sequencing results were crossed with wild-type locusts to generate the G1 animals. PCR-mediated genotyping and TA cloning were used to identify the G1 heterozygotes. Then, heterozygotes with the same mutations were in-crossed to generate the G2 animals (homozygotes and heterozygotes) for further research and passaging. Please click here to view a larger version of this figure.
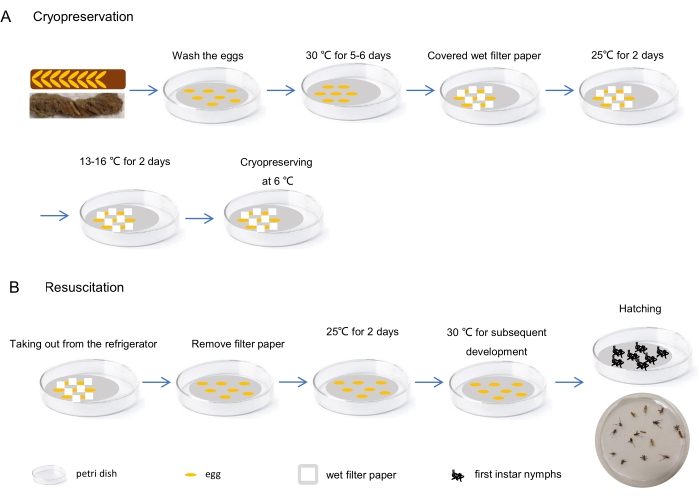
Figure 5: Egg cryopreservation and resuscitation. (A) The procedure of cryopreservation: Isolate the eggs from egg pods and incubate them at 30 °C for 5-6 days after washing with sterile water. Then, gather the developed eggs together in the Petri dish and cover them with small scraps of moist filter paper. Wrap the entire Petri dish with a paraffin film and keep it at 25 °C for 2 days, followed by another 2 days at a relatively lower temperature (e.g., 13-16 °C). Finally, refrigerate it at 6 °C. (B) The procedure of resuscitation: Take the Petri dishes with cryopreserved eggs out from the refrigerator and keep them at 25 °C for 2 days after removing the filter paper scraps. Then, the eggs can be cultured for subsequent development at 30 °C until the nymphs hatch. Please click here to view a larger version of this figure.
| Reagent | Volume (μL) |
| Cas9 protein (300 ng/μL) | 1 |
| sgRNA (300 ng/μL) | 1 |
| PCR product (200 ng/μL) | 1 |
| 10×NEBuffer r3.1 | 1 |
| Nuclease-free water | 6 |
Table 1: In vitro digestion system. 200 ng of purified target fragment (PCR product) was mixed with the CRISPR/Cas9 system (300 ng of each component) for testing the bio-activity of the synthesized sgRNAs. A commercial buffer was used and nuclease-free water was added to make up the total volume to 10 µL.
| Reagent | Volume (μL) |
| 2xEs Taq MasterMix | 12.5 |
| Forward primer | 0.5 |
| Reverse primer | 0.5 |
| Template (lysis product) | 1 |
| Nuclease-free water | 10.5 |
Table 2: PCR system for target gene fragment amplification. To amplify the genomic fragment of the target gene, a commercial Taq mix was used and primers were added according to the manufacturer's instructions. 1 µL of the lysis product was used as the template and nuclease-free water was used to make the total volume to 25 µL.
| Temperature (°C) | Time | Cycles |
| 95 | 5 min | 1 |
| 95 | 30 s | 35 |
| 55 | 30 s | |
| 72 | 40 s | |
| 72 | 10 min | 1 |
Table 3: PCR program for target gene amplification. The PCR program for target gene amplification was: 95 °C for 5 min; 35 cycles of 95 °C for 30 s, 55 °C for 30 s, 72 °C for 40 s; 72 °C for 10 min.
| Items | Data |
| No. of injected eggs | 120 |
| Tested embryos | 10 |
| No. of hatched nymphs | 58 |
| Hatching rate | 52.73% |
| No. of G0 adult | 12 |
| No. of G0 mutants | 8 |
| Mutation efficiency in G0 adults | 66.67% |
Table 4: Summary of editing efficiency. 120 eggs were injected to knock out the target gene and 10 eggs were taken for testing during the embryo stage. 58 nymphs hatched from the rest embryos (the hatching rate was 52.73%). Finally, 12 G0 locusts developed to the adult stage and 8 of them were successfully edited at the target site. The mutation rate in G0 adults was 66.67%.
| Control group | Group 1 | Group 2 | Group 3 | Group 4 | |
| No. of eggs | 120 | 120 | 120 | 120 | 120 |
| Temperature treatment for storage | 30 °C | 30°C (5-6 d)→6 °C | 30°C (5-6 d)→25°C (2 d)→13-16°C (2 d)→6 °C | 30°C (5-6 d)→25°C (2 d)→13-16°C (2 d)→6 °C | 30°C (5-6 d)→25°C (2 d)→13-16°C (2 d)→6 °C |
| Storage time | 14 days | 14 days | 1 month | 3 months | 5 months |
| Temperature treatment for resuscitation | 30 °C | 6°C→30 °C | 6°C→25°C (2 d)→30 °C | 6°C→25°C (2 d)→30 °C | 6°C→25°C (2 d)→30 °C |
| No. of hatched locusts | 108 | 0 | 96 | 86 | 78 |
| Hatching rate | 90.00% | 0 | 80.00% | 71.67% | 65.00% |
| No. of adult locusts | 81 | 0 | 60 | 46 | 31 |
| Eclosion rate | 75.00% | 0 | 62.50% | 53.49% | 39.74% |
Table 5: Summary of the cryopreservation and resuscitation of eggs. 600 eggs were divided into five groups for cryopreservation and resuscitation study. The first group (control) of 120 eggs was always incubated at 30 °C and stored for 14 days. The second group (Group 1) of 120 eggs was transferred to a refrigerator after incubation at 30 °C for 5-6 days and stored at 6 °C for 14 days. Then, these eggs were transferred to 30 °C for resuscitation. The other groups (group 2, group 3, and group 4, each group contained 120 eggs) experienced a gradient cooling and recovering temperature treatment as described in the protocol (steps 6.1-6.3) with a different storage time (1 month for group 2, 3 months for group 3, and 5 months for group 4). At last, 108 nymphs hatched from the control group (at a hatching rate of 90%) and 81 of them developed to the adult stage (75% of the eclosion rate). No locusts hatched in the second group (Group 1). The hatching rate and eclosion rate of the other groups were lower compared to the control group and declined with the storage time. 96 nymphs hatched in group 2 and 60 of them developed to the adult stage. 86 nymphs hatched in group 3 and 46 of them developed to the adult stage. In group 4, 78 nymphs hatched and 31 of them developed to the adult stage.
Discussion
Locusts have been among the most devastating pests to agriculture since the civilization of human beings23. CRISPR/Cas9-based genome editing technology is a powerful tool for providing better knowledge of the biological mechanisms in locusts as well as a promising pest control strategy. Thus, it is of great benefit to develop an efficient and easy-to-use method of CRISPR/Cas9-mediated construction of homozygous locust mutants. Although some great works have been reported and provided some basic workflow for genome editing in locusts18,20,21,22, a systematic optimization of the whole procedure is still lacking. In general, the homozygous locust mutant generated by embryonic injection of CRISPR/Cas9 system can be divided into three major steps: a) sgRNA design, synthesis, and screening in vitro, b) egg collection, preparation, and injection; and c) mutant screening and homozygote maintaining. This protocol provides an optimized workflow and the steps of target site selection, in vitro sgRNA screening, microinjection of the eggs as well as detecting mutants are especially critical for effectively generating homozygous mutant locusts with the CRISPR/Cas9 system.
Firstly, the target site, the form of corresponding sgRNA, and the concentration of sgRNA have been suggested as the critical factors of genome editing efficiency in insects18,27,28. To solve this problem, it is recommended to first analyze the gene structure and design more than one target site on exon 1 or near the start codon using the online resources. Then, synthesize sgRNA in vitro and mix it with Cas9 protein at different concentrations to optimize the cutting efficiency of RNP. This procedure will help to identify a sgRNA with high bioactivity and optimize the composition of the RNP complex for the candidate target gene.
Secondly, collecting as many fresh eggs as possible in a limited time and improving the hatching rate as well as mutation rate are also big challenges for genome editing operations. The gregarious adult locusts reared at the condition of 16 (light):8 (dark) photoperiod are used for collecting enough eggs in a short period during 9:00-11:00 am. It is important to gently apply downward pressure between the eggs using a paintbrush or tweezers to carefully isolate the eggs from the pod. With enough practice, multiple users can reliably separate individual eggs from the pod. After each egg is isolated and washed with sterile water, the eggs are aligned on the designed injection pads (Figure 2B,C) to stabilize the eggs during microinjection. The egg is injected with the optimized RNP near the micropyles. To limit the mechanical damage and minimize the pollution of eggs, fresh eggs are allowed to stay in the pods for about 30 min to make sure that the eggshell tanning is ideal for microinjection21, owing to its condensed structure and high defense ability on invasion after enough tanning29,30. Meanwhile, a published report indicated that the syncytial division31 of locust eggs occurs at some point between 0 h and 4 h after laying. With the current standard microinjection protocol, injection of about 100 eggs can be accomplished within 40 min. Taken together, the procedure of tanning and microinjection of 100 fertilized eggs can be completed within 2 h. Thus, there is at least 2 h left for CRISRP/Cas9-mediated cleavage and the repair of target genes to achieve efficient mutation in the injected eggs and future adult locusts.
Lastly, effective strategies for passaging and mutant identification are important for generating homozygotes in most mutant animals. In locusts, for genes that are nonlethal after editing, the cross strategy suggested here is that G0 mutants cross with wide-type locusts and further in-crosses can be performed using the G1 heterozygotes with the same mutations to generate homozygous mutants in the following generation. To verify the mutation in each locust during the mutant inheriting process, it is recommended to use the alkaline lysis method to digest a short segment of antennae as the template for PCR-based genotyping. Finally, use the obtained homozygous adults for further in-cross to expand the population. Limited by the population size and development stage differences between homozygous locusts, the excess eggs are recommended for cryopreservation at 6 °C and resuscitation by gradient temperature rise (Figure 5), which is based on the principle of low-temperature diapause and high-temperature diapause release of migratory locust eggs32,33.
In short, the CRISPR/Cas9 system has been proven to be a reliable tool for facilitating the study of functional genomics of migratory locusts. This detailed protocol can be used as a reference for CRISPR/Cas9 based gene editing applications in migratory locusts as well as in other insects.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (32070502, 31601697, 32072419 and the China Postdoctoral Science Foundation (2020M672205).
Materials
| 10×NEBuffer r3.1 | New England Biolabs | B7030S | The buffer of in vitro Cas9 cleavage assays |
| 2xEs Taq MasterMix (Dye) | Cwbio | CW0690 | For gene amplification |
| 2xPfu MasterMix (Dye) | Cwbio | CW0686 | For gene amplification |
| CHOPCHOP | Online website for designing sgRNAs, http://chopchop.cbu.uib.no/. | ||
| CRISPOR | Online website for designing sgRNAs, http://crispor.org. | ||
| CRISPRdirect | Online website for designing sgRNAs, http://crispr.dbcls.jp/. | ||
| Electrophoresis power supply | LIUYI BIOLOGY | DYY-6D | Separation of nucleic acid molecules of different sizes |
| Eppendorf Tube | Eppendorf | 30125177 | For sample collection, etc. |
| Fine brushes | Annigoni | 1235 | For cleaning and isolating eggs. Purchased online. |
| Flaming/brown micropipette puller | Sutter Instrument | P97 | For making the microinjection needles |
| Gel Extraction Kit | Cwbio | CW2302 | DNA recovery and purification |
| Gel Imaging Analysis System | OLYMPUS | Gel Doc XR | Observe the electrophoresis results |
| GeneTouch Plus | Bioer | B-48DA | For gene amplification |
| Glass electrode capillary | Gairdner | GD-102 | For making injection needles with a micropipette Puller |
| Incubator | MEMMERT | INplus55 | For migratory locust embryo culture |
| Metal bath | TIANGEN | AJ-800 | For heating the sample |
| Micro autoinjector | Eppendorf | 5253000068 | Microinjection of embryos early in development |
| Micro centrifuge | Allsheng | MTV-1 | Used for mixing reagents |
| Microgrinder | NARISHIGE | EG-401 | To ground the tip of injection needle |
| Microloader | Eppendorf | 5242956003 | For loading solutions into the injection needles. |
| Micromanipulation system | Eppendorf | TransferMan 4r | An altinative manipulation system for microinjection |
| Microscope | cnoptec | SZ780 | For microinjection |
| Motor-drive Manipulator | NARISHIGE | MM-94 | For controling the position of the micropipette during the microinjection precedure |
| Multi-Sample Tissue Grinder | jingxin | Tissuelyser-64 | Grind and homogenize the eggs |
| ovipisition pot | ChangShengYuanYi | CS-11 | Filled with wet sterile sand for locust ovipositing in it. The oocysts are collected from this container. Purchased online. |
| Parafilm | ParafilmM | PM996 | For wrapping the petri dishes. |
| pEASY-T3 Cloning Kit | TransGen Biotech | CT301-01 | For TA cloning |
| Petri dish | NEST | 752001 | For culture and preservation of the eggs. |
| Pipettor | Eppendorf | Research®plus | For sample loading |
| plastic culture cup | For rearing locusts seperately and any plastic cup big enough (not less than 1000 mL in volume) will do. Purchased online. | ||
| Precision gRNA Synthesis Kit | Thermo | A29377 | For sgRNA synthesis |
| Primer Premier | PREMIER Biosoft | Primer Premier 5.00 | For primer design |
| SnapGene | Insightful Science | SnapGene®4.2.4 | For analyzing sequences |
| Steel balls | HuaXinGangQiu | HXGQ60 | For sample grinding.Purchased online. |
| Tips | bioleaf | D781349 | For sample loading |
| Trans DNA Marker II | TransGen Biotech | BM411-01 | Used to determine gene size |
| TrueCut Cas9 Protein v2 | Thermo | A36496 | Cas9 protein |
| UniversalGen DNA Kit | Cwbio | CWY004 | For genomic DNA extraction |
| VANNAS Scissors | Electron Microscopy Sciences | 72932-01 | For cutting off the antennae |
| Wheat | To generate wheat seedlings as the food for locusts. Bought from local farmers. | ||
| ZiFiT | Online website for designing sgRNAs, http://zifit.partners.org/ZiFiT/ChoiceMenu.aspx. |
References
- Doudna, J. A. The promise and challenge of therapeutic genome editing. Nature. 578 (7794), 229-236 (2020).
- van Haasteren, J., Li, J., Scheideler, O. J., Murthy, N., Schaffer, D. V. The delivery challenge: fulfilling the promise of therapeutic genome editing. Nature Biotechnology. 38 (7), 845-855 (2020).
- McCarty, N. S., Graham, A. E., Studena, L., Ledesma-Amaro, R. Multiplexed CRISPR technologies for gene editing and transcriptional regulation. Nature Communications. 11 (1), 1281 (2020).
- Manghwar, H., et al. CRISPR/Cas systems in genome editing: Methodologies and tools for sgRNA design, off-target evaluation, and strategies to mitigate off-target effects. Advanced Science. 7 (6), 1902312 (2020).
- Anzalone, A. V., Koblan, L. W., Liu, D. R. Genome editing with CRISPR-Cas nucleases, base editors, transposases and prime editors. Nature Biotechnology. 38 (7), 824-844 (2020).
- Rees, H. A., Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nature Reviews Genetics. 19 (12), 770-788 (2018).
- Ishino, Y., Shinagawa, H., Makino, K., Amemura, M., Nakata, A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. Journal of Bacteriology. 169 (12), 5429-5433 (1987).
- Barrangou, R., et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science. 315 (5819), 1709-1712 (2007).
- Mali, P., et al. RNA-guided human genome engineering via Cas9. Science. 339 (6121), 823-826 (2013).
- Jinek, M., et al. RNA-programmed genome editing in human cells. eLife. 2, 00471 (2013).
- Cong, L., et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 339 (6121), 819-823 (2013).
- Jinek, M., et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 337 (6096), 816-821 (2012).
- Scully, R., Panday, A., Elango, R., Willis, N. A. DNA double-strand break repair-pathway choice in somatic mammalian cells. Nature Reviews Molecular Cell Biology. 20 (11), 698-714 (2019).
- Ai, D., et al. Embryo microinjection and knockout mutant identification of CRISPR/Cas9 genome-edited Helicoverpa Armigera (Hubner). Journal of Visualized Experiments: JoVE1. (173), e62068 (2021).
- Huang, Y., et al. CRISPR/Cas9 mediated knockout of the abdominal-A homeotic gene in the global pest, diamondback moth (Plutella xylostella). Insect Biochemistry and Molecular Biology. 75, 98-106 (2016).
- Gratz, S. J., et al. Highly specific and efficient CRISPR/Cas9-catalyzed homology-directed repair in Drosophila. 유전학. 196 (4), 961-971 (2014).
- Kistler, K. E., Vosshall, L. B., Matthews, B. J. Genome engineering with CRISPR-Cas9 in the mosquito Aedes aegypti. Cell Reports. 11 (1), 51-60 (2015).
- Li, Y., et al. CRISPR/Cas9 in locusts: Successful establishment of an olfactory deficiency line by targeting the mutagenesis of an odorant receptor co-receptor (Orco). Insect Biochemistry and Molecular Biology. 79, 27-35 (2016).
- Xu, X., et al. BmHpo mutation induces smaller body size and late stage larval lethality in the silkworm, Bombyx mori. Insect Science. 25 (6), 1006-1016 (2018).
- Chen, D., et al. CRISPR/Cas9-mediated genome editing induces exon skipping by complete or stochastic altering splicing in the migratory locust. BMC Biotechnology. 18 (1), 60 (2018).
- Zhang, T., et al. Egg tanning improves the efficiency of CRISPR/Cas9-mediated mutant locust production by enhancing defense ability after microinjection. Journal of Integrative Agriculture. 20 (10), 2716-2726 (2021).
- Guo, X., et al. 4-Vinylanisole is an aggregation pheromone in locusts. Nature. 584 (7822), 584-588 (2020).
- Zhang, L., Lecoq, M., Latchininsky, A., Hunter, D. Locust and grasshopper management. Annual Review of Entomology. 64, 15-34 (2019).
- Pétavy, G. Origin and development of the vitellophags during embryogenesis of the migratory locust, Locusta migratoria L. (Orthoptera : Acrididae). International Journal of Insect Morphology and Embryology. 14 (6), 361-379 (1985).
- Barry, S. K., et al. Injecting Gryllus bimaculatus eggs. Journal of Visualized Experiments: JoVE. (150), e59726 (2019).
- Watanabe, T., Noji, S., Mito, T. Genome editing in the cricket, Gryllus bimaculatus. Methods in Molecular Biology. 1630, 219-233 (2017).
- Du, M. H., et al. Suppression of Laccase 2 severely impairs cuticle tanning and pathogen resistance during the pupal metamorphosis of Anopheles sinensis (Diptera: Culicidae). Parasites & Vectors. 10 (1), 171 (2017).
- Eisner, T., Shepherd, J., Happ, G. M. Tanning of grasshopper eggs by an exocrine secretion. Science. 152 (3718), 95-97 (1966).
- Ho, K., Dunin-Borkowski, O. M., Akam, M. Cellularization in locust embryos occurs before blastoderm formation. Development. 124 (14), 2761-2768 (1997).
- Wang, X., et al. Interactive effect of photoperiod and temperature on the induction and termination of embryonic diapause in the migratory locust. Pest Managment Science. 77 (6), 2854-2862 (2021).
- Jarwar, A. R., et al. Comparative transcriptomic analysis reveals molecular profiles of central nervous system in maternal diapause induction of Locusta migratoria. G3-Genes Genomes Genetics. 9 (10), 3287-3296 (2019).

