Differentiated Mouse Adipocytes in Primary Culture: A Model of Insulin Resistance
Summary
This protocol describes the isolation of mouse preadipocytes from subcutaneous fat, their differentiation into mature adipocytes, and the induction of insulin resistance. Insulin action is evaluated by the phosphorylation/activation of members of the insulin signaling pathway through western blot. This method allows direct determination of insulin resistance/sensitivity in primary adipocytes.
Abstract
Insulin resistance is a reduced effect of insulin on its target cells, usually derived from decreased insulin receptor signaling. Insulin resistance contributes to the development of type 2 diabetes (T2D) and other obesity-derived diseases of high prevalence worldwide. Therefore, understanding the mechanisms underlying insulin resistance is of great relevance. Several models have been used to study insulin resistance both in vivo and in vitro; primary adipocytes represent an attractive option to study the mechanisms of insulin resistance and identify molecules that counteract this condition and the molecular targets of insulin-sensitizing drugs. Here, we have established an insulin resistance model using primary adipocytes in culture treated with tumor necrosis factor-α (TNF-α).
Adipocyte precursor cells (APCs), isolated from collagenase-digested mouse subcutaneous adipose tissue by magnetic cell separation technology, are differentiated into primary adipocytes. Insulin resistance is then induced by treatment with TNF-α, a proinflammatory cytokine that reduces the tyrosine phosphorylation/activation of members of the insulin signaling cascade. Decreased phosphorylation of insulin receptor (IR), insulin receptor substrate (IRS-1), and protein kinase B (AKT) are quantified by western blot. This method provides an excellent tool to study the mechanisms mediating insulin resistance in adipose tissue.
Introduction
Insulin is an anabolic hormone produced by pancreatic islet β-cells and the key regulator of glucose and lipid metabolism. Among its many functions, insulin regulates glucose uptake, glycogen synthesis, gluconeogenesis, protein synthesis, lipogenesis, and lipolysis1. The initial molecular signal after insulin interaction with its receptor (IR) is the activation of the intrinsic tyrosine protein kinase activity of IR2, resulting in its autophosphorylation3 and the subsequent activation of a family of proteins known as insulin receptor substrates (IRS), which binds to adaptor proteins leading to activation of a cascade of protein kinases4. Insulin activates two main signaling pathways: phosphatidylinositol 3-kinase (PI3K)-protein kinase B (AKT) and Ras-mitogen-activated protein kinase (MAPK). The former constitutes a major branch point or node4,5 for the activation of numerous downstream effectors involved in diverse physiological functions, including the regulation of fuel homeostasis, whereas the latter regulates cell growth and differentiation4,6. Insulin actions ultimately depend on the cell type and physiological context7.
One of the main insulin-responsive metabolic tissues is the adipose tissue. White adipose tissue is the most abundant type of fat in humans and rodents, distributed within subcutaneous fat (between the skin and muscles) and visceral fat (around the organs in the abdominal cavity). Given their large volume, adipocytes or fat cells are the most abundant cell type in adipose tissue. These fat cells can be brown/beige (thermogenic), pink (in the mammary gland), and white8,9. White adipocytes keep the main energy reserves in the body in the form of triglycerides, an insulin-dependent process. Insulin promotes glucose transport and lipogenesis, while it inhibits lipolysis or lipid breakdown7,10. It also facilitates the differentiation of preadipocytes into adipocytes — the mature fat-storing cells11.
Insulin resistance occurs when a normal insulin level produces an attenuated biological response, resulting in compensatory hyperinsulinemia12. Insulin resistance is a condition associated with overweight and obesity5, that when combined leads to type 2 diabetes (T2D) and other metabolic diseases13. Hyperinsulinemia compensates insulin resistance in peripheral tissues to maintain normal blood glucose levels14. However, eventual β-cell loss or exhaustion, together with exacerbated insulin resistance, leads to elevated blood glucose levels consistent with T2D5. Therefore, insulin resistance and hyperinsulinemia can contribute to the development of obesity-derived metabolic diseases15. Furthermore, obesity may cause chronic low-grade local inflammation promoting insulin resistance in adipose tissue15,16,17. In addition, obesity-derived alterations in adipose tissue, such as fibrosis, inflammation, and reduced angiogenesis and adipogenesis, lead to lower adiponectin serum levels (an insulin sensitizer) and increased secretion of factors such as plasminogen activator inhibitor 1 (PAI-1), free fatty acids, and exosomes into the bloodstream, exacerbating insulin resistance17.
Many aspects underlying insulin resistance remain unknown. In vitro and in vivo models have been developed to study the mechanisms mediating insulin resistance in major target tissues, including adipose tissue. The advantage of in vitro models is that researchers have more control of the environmental conditions and can evaluate insulin resistance in specific cell types. Particularly, adipocyte precursor cells (APCs) have the individual phenotype of the donor tissue, which might reflect physiology better than adipocyte cell lines. A main factor inducing insulin resistance in vitro is tumor necrosis factor-α (TNF-α). TNF-α is a proinflammatory cytokine secreted by adipocytes and macrophages in adipose tissue18. While it is required for proper adipose tissue remodeling and expansion19, long-term exposure to TNF-α induces insulin resistance in adipose tissue in vivo and in adipocytes in vitro20. Chronic TNF-α treatment of several cell types leads to increased serine phosphorylation of both IR and IRS-1, thereby promoting decreased tyrosine phosphorylation21. Increased phosphorylation of IRS-1 on serine residues inhibits the IR tyrosine kinase activity and may be one of the key mechanisms by which chronic TNF-α treatment impairs insulin action22,23. TNF-α activates pathways involving the serine/threonine kinase inhibitor of nuclear factor ĸB kinase β (IKKβ) and c-Jun N terminal kinase (JNK)24. JNK induces a complex proinflammatory transcriptional program but also directly phosphorylates IRS-16.
Understanding the pathogenesis of insulin resistance has become increasingly important to guide the development of future therapies against T2D. APCs have proven to be an excellent model for the study of fat cell biology, including the sensitivity and resistance to insulin, and for identifying the intrinsic properties of adipocytes independent of the systemic environment. APCs can be easily obtained from different adipose depots, and under the appropriate conditions, differentiated into mature adipocytes. With this method, direct effects on insulin resistance/sensitivity can be evaluated in adipocytes.
Protocol
All rodent experiments were approved by the Bioethics Committee of the Institute of Neurobiology of the UNAM, protocol number 075.
1. Isolation of mouse adipocyte precursor cells
- Euthanize 8-10-week-old male C57BL/6 mice (e.g., by CO2 inhalation and subsequent cervical dislocation) (four animals per isolation). Disinfect the mice by rubbing with 70% ethanol and obtain the adipose tissue immediately after sacrifice.
NOTE: After the euthanasia of rodents, isolate the adipose tissue immediately and perform all steps inside a sterile laminar flow hood. Mice are kept under 12 h light/dark cycles with free access to food and water. - Dissect the inguinal subcutaneous adipose tissue from each mouse. Remove only the adipose tissue, avoiding the removal of muscle, skin, or any other tissue, and collect it in a 50 mL conical tube containing 15 mL of type 1 collagenase solution (1.5 mg/mL) on ice. Dissolve type 1 collagenase in Dulbecco's Modified Eagle Medium (DMEM) high glucose-1% bovine serum albumin (BSA) and filter through 0.2 µm syringe filters.
- Cut the adipose tissue into small pieces with sterile surgical scissors and digest the samples by incubation with type 1 collagenase solution at 37 °C in an orbital shaker (150 rpm) for 30 min (place the tube containing fat and collagenase in a horizontal position for a greater shaking surface). Check the digestion every 10 min to make sure it works (tissue degradation is visible) and to prevent overdigestion (see Supplementary Figure S1).
- Filter using a 200 µm mesh-syringe (previously autoclaved) to eliminate tissue not digested with collagenase. Pass the filter over the edge of the tube to drain as many cells as possible into the solution. Add 15 mL of cold DMEM-1% BSA to stop digestion and centrifuge at 400 × g for 10 min at 4 °C.
- Aspirate the top layer containing mature adipocytes and most of the liquid layer; avoid touching or disturbing the pellet. Add 20 mL of cold phosphate buffered saline (PBS)-2% fetal bovine serum (FBS) and resuspend the pellet. Centrifuge at 400 × g for 5 min at 4 °C.
- Remove the supernatant, aspirating first the upper layer to eliminate the remaining adipocytes and fat. Resuspend the pellet in 1 mL of ammonium chloride potassium (ACK) lysing buffer (to lyse red blood cells) and incubate on ice for 5 min. Add 10 mL of PBS-2% FBS, mix, and centrifuge at 400 × g for 5 min at 4 °C.
- Aspirate the supernatant and resuspend the pellet in 200 µL of anti-Fc solution (purified rat anti-mouse CD16/CD32 diluted in PBS-2% FBS [1:150]) to reduce Fc receptor-mediated binding by antibodies of interest. Incubate on ice for 5 min. Transfer the cell suspension to a 5 mL tube that fits into the prechilled racks of a magnetic cell separator and centrifuge at 400 × g for 5 min at 4 °C.
- Eliminate the supernatant, add 200 µL of the mixture of CD31(PECAM-1) monoclonal antibody (390)-biotin (1:100) and CD45 monoclonal antibody (30-F11)-biotin (1:100), mix well, and incubate on ice for 15 min (prepare antibody dilutions in anti-Fc solution; see Table 1). Add 400 µL of PBS-2% FBS and centrifuge at 400 × g for 5 min at 4 °C.
NOTE: CD31 and CD45 are markers of endothelial cells and hematopoietic cells, respectively. - Aspirate the supernatant and incubate in 100 µL of anti-biotin microbeads (1:5) for 15 min on ice (prepare antibody dilutions in anti-Fc solution, see Table 1). Add 400 µL of PBS-2% FBS and centrifuge at 400 × g for 5 min at 4 °C.
- Discard the supernatant and resuspend the pellet in 350 µL of PBS-2% FBS. Remove cell aggregates or large particles from the single-cell suspensions with a preseparation filter (70 µm). First, activate the filter with 100 µL of PBS-2% FBS, pass the cell suspension through the filter (collect in a clean tube), and finally wash the filter with 100 µL of PBS-2% FBS.
- Perform magnetic separation of cells using the negative separation strategy with a magnetic cell separator.
- Place the sample in position A of the chill rack (ensure that the rack is precooled) and two empty tubes in position B and C to recover the non-labeled and labeled cells, respectively.
- Load the washing buffer and running buffer into the corresponding bottles before turning on the equipment. Program the equipment to perform the magnetic separation of cells with the DEPLETE protocol.
NOTE: This protocol is for depletion in standard mode for the removal of cells with normal antigen expression (in this case, cells labeled with anti-CD31 and anti-CD45), if recovery is the highest priority. - In the separation section, select the number of samples to be separated and select the DEPLETE protocol. Press Run and start the separation. At the end of the program, recover the unlabeled cells and centrifuge at 400 × g for 5 min at 4 °C.
- Remove the supernatant and resuspend the pellet in 500 µL of growth medium (Table 1).
- Seed the APCs in one well of a 12-well plate previously coated with 2.5% basement membrane matrix (approximately 50,000-75,000 cells are obtained). Add 400 µL of 2.5% basement membrane matrix to cover the entire surface of each well. Immediately after removing the excess solution and leaving only a small layer, let the plate dry (without a lid) inside the laminar flow hood for at least 1 h. Incubate at 37 °C in a 5% CO2 atmosphere. Change the medium every 48 h until the cells reach 80% confluence.
- At 80% confluence, remove the medium completely and wash with 350 µL of PBS-2% FBS. Harvest the cells with 350 µL of 0.05% trypsin-EDTA for 2 min at 37 °C. Add 2 mL of growth medium and collect the cells in a new 50 mL conical tube, then centrifuge at 400 × g for 5 min.
- Passage the APCs to 12-well plates (10,000-20,000 cells per well) previously coated with 2.5% basement membrane matrix. Incubate at 37 °C with 5% CO2. Change the medium every 48 h until cells reach 80% confluence.
NOTE: APCs can be passaged once. Subsequently, they lose their ability to proliferate and differentiate. See the workflow for the APCs isolation process in Supplementary Figure S2.
2. Adipocyte differentiation and induction of insulin resistance
NOTE: Maintain cells at 37 °C in 5% CO2 and perform steps involving change of medium and treatments with TNF-α and insulin inside a sterile hood.
- At 80% confluence, begin the differentiation process; aspirate off any growth medium and replace it with 500 µL of differentiation medium (Table 1) containing 3.3 nM bone morphogenetic protein (BMP4) in each well.
- After 48 h, replace the medium with the differentiation medium (500 µL per well) and the differentiation cocktail (Table 1).
- Remove the medium 72 h later and add 100 nM of insulin in 500 µL of fresh differentiation medium.
NOTE: Cells begin to show a round appearance with small lipid droplets inside. - Aspirate off any differentiation medium and replace it with 500 µL of simple medium-2% FBS (Table 1) 48 h later. Induce insulin resistance with 4 ng/mL of TNF-α. Include control wells without TNF-α treatment.
- After 24 h of incubation, add 4 ng/mL TNF-α in simple medium-0% FBS (Table 1). Maintain control wells without TNF-α treatment.
- Activate the insulin signaling pathway 24 h later by adding 100 nM insulin and incubate for 15 min at 37 °C in a 5% CO2 atmosphere. Remove the medium and wash with 500 µL of PBS. Include control wells without insulin treatment.
3. Evaluation of insulin signaling pathway by western blot
- Protein extraction and quantification
- Wash each well of cells with 500 µL of PBS and keep on ice to lyse the cells in 50 µL of RIPA buffer (Table 1) containing 1% protease inhibitor cocktail added just prior to sample isolation. Scrape the entire surface of the well with a tip and transfer the lysate to a 0.6 mL microcentrifuge tube (keep on ice).
- Incubate for 1 h at 4 °C in a rotating shaker, mix by vortexing, and centrifuge at 8,000 × g for 15 min at 4 °C and recover the supernatant (keep on ice).
- Determine the protein concentration using the Bradford assay (do not dilute the sample to quantify).
- Take the necessary volume corresponding to 40-60 µg and denature with 6x Laemmli buffer (Table 1) by incubating at 97 °C for 15 min while shaking at 550 rpm. Freeze until use.
- SDS-polyacrylamide gel electrophoresis
- Perform SDS-polyacrylamide gel electrophoresis (SDS-PAGE) with a 7.5% polyacrylamide gel of 1.5 mm thickness. Insert the gel into the tank and fill with running buffer (Table 1).
- Add 3 µL of prestained protein standard to the first well of the gel and load each sample into subsequent wells.
- Run the gel at 80 V for approximately 15 min to ensure the sample enters the polyacrylamide concentrator gel matrix. Thereafter, increase the voltage to 90 V for 2.5 h or until the 37 kDa molecular weight marker can be seen at the bottom edge of the gel.
- Transfer the proteins from the gel to a nitrocellulose membrane in a semi-dry chamber for 35 min at 25 V. Beforehand, submerge the gel, membrane, and filters in transfer buffer (Table 1) for a few minutes.
- Western blot
- After transfer, wash the membrane with Tris-buffered saline (TBS) containing 0.1% Tween 20 (TBS-T 0.1%) (Table 1) for 3 min with shaking at 30 rpm.
- Block the membrane with blocking solution (Table 1) at 4 °C for 1 h in constant rotation.
- Cut the membrane into three parts according to the molecular weight marker to incubate overnight with different primary antibodies (diluted in blocking solution) at 4 °C in constant rotation: Phospho-IRS1 (Tyr608) antibody (1:1,000), expected band at 180 kDa; Phospho-Insulin receptor β (Tyr1150/1151) antibody (1:1,000), expected band at 95 kDa; Phospho-Akt (Ser473) antibody (1:1,000), expected band at 60 kDa.
- Wash the membranes 5x for 5 min each with TBS-T 0.1%.
- Incubate the membranes with the corresponding peroxidase-coupled secondary antibody (diluted in blocking solution) for 2 h at room temperature in constant rotation: peroxidase affiniPure donkey anti-rabbit IgG (H+L) (1:5,000).
- Wash the membranes 5x for 5 min each with TBS-T 0.1%.
- Detect the proteins using the chemiluminescence detection system. Prepare the substrate according to the manufacturer's instructions.
- Place the blot in a plastic bag and add 200 µL of chemiluminescent substrate to completely cover the membrane. Drain excess substrate and remove any air bubbles.
- Expose each blot for 30-60 s in a high-performance imaging system compatible with chemiluminescence.
- Strip the membranes (steps 10-13 of the western blot section) to break the antibody-antigen interactions and thus allow nitrocellulose membranes to be re-blotted. Recover the membranes and wash 3x for 5 min each with TBS-T 1% at 50 rpm.
- Incubate for 7 min with 10 mL of 0.2 M NaOH at 50 rpm.
- Wash 3x for 5 min each with tap water at 50 rpm.
- Wash for 10 min with TBS-T 1% at 50 rpm.
- Repeat steps 2-9 of the western blot section to incubate the membrane with anti-beta tubulin (55 kDa) as a loading control using 1:1,000 dilution.
- Perform densitometric analysis25 for each protein using ImageJ software (https://imagej.nih.gov/).
Representative Results
Over the last few years, the increased prevalence of obesity and T2D has prompted an intense search for the mechanisms mediating insulin resistance in adipose tissue. With the protocol described here, APCs can be differentiated into mature adipocytes to evaluate insulin resistance and sensitivity. Once the APCs reach confluence, it takes 10 days to complete their differentiation into mature adipocytes and their TNF-α-mediated induction of insulin resistance (Figure 1).
APCs show a fibroblast-like morphology, characterized by their flat and elongated shape and their adhesion to the plate completed 48 h after seeding (Figure 2A). APCs take 2-5 days to reach 80% confluence (depending on the number of seeded cells), a time at which the differentiation process is started by exposure to the differentiating cocktail (Figure 2A). Mature adipocytes begin to appear in the following 6-8 days. Differentiated adipocytes are characterized by a round morphology and the intracellular accumulation of lipid droplets. Up to 80% of cells are differentiated at the end of the differentiation process (Figure 2B).
Insulin resistance is induced in primary adipocytes by TNF-α treatment (4 ng/mL) for 48 h; this concentration of TNF-α does not alter cell viability (Supplementary Figure S3). Subsequently, the insulin signaling pathway is activated by adding 100 nM insulin for 15 min, and the phosphorylation of main signaling molecules (IR, IRS-1, and AKT) is measured by western blot. Insulin stimulates the phosphorylation of these three molecules (Figure 3) compared to control, non-treated differentiated adipocytes. TNF-α reduces the insulin-induced phosphorylation of IR (30%), IRS-1 (20%), and AKT (45%) (Figure 3). Insulin-resistant cells show less activation of the insulin signaling pathway compared to insulin-sensitive cells. Moreover, TNF-α treatment decreases the mRNA expression of known insulin-sensitivity markers: Insr, Irs2, Glut4, and Adipoq (Supplementary Figure S4). These findings confirm that TNF-α reduces the action of insulin in primary adipocytes, and thereby induces insulin resistance.
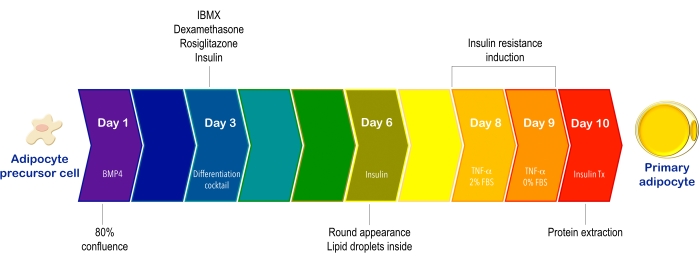
Figure 1: Schematic timeline representing the process of adipocyte precursor cell differentiation and the induction of insulin resistance. APCs are isolated from subcutaneous adipose tissue using collagenase treatment and magnetic cell separation technology. Then, they undergo a 7 day differentiation process to obtain primary adipocytes. Subsequently, insulin resistance is induced with TNF-α for 48 h, the first 24 h in medium containing 2% FBS and the next 24 h with serum-free medium to prevent the activation of the insulin signaling pathway. The signaling cascade is activated with insulin and protein is extracted for western blot analysis 10 days after starting the differentiation process to evaluate the phosphorylation/activation of IR, IRS-1, and AKT. Abbreviations: APCs = adipocyte precursor cells; BMP4 = Bone morphogenetic protein; IBMX = 3-isobutyl-1-methylxanthine; TNF-α = tumor necrosis factor-α; FBS = fetal bovine serum; IR = insulin receptor; IRS = insulin receptor substrate; AKT = protein kinase B; Tx = treatment. Please click here to view a larger version of this figure.
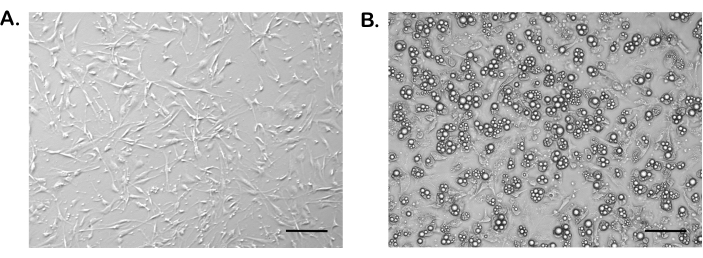
Figure 2: Morphology of adipocyte precursor cells and mature adipocytes. (A) Subcutaneous APCs at 80% confluence prior to inducing differentiation in 12-well culture plates. (B) Subcutaneous primary adipocytes after 7 days of inducing differentiation in 12-well culture plates. Images were taken at 10x magnification. Scale bars = 100 µm. Abbreviation: APCs = adipocyte precursor cells. Please click here to view a larger version of this figure.
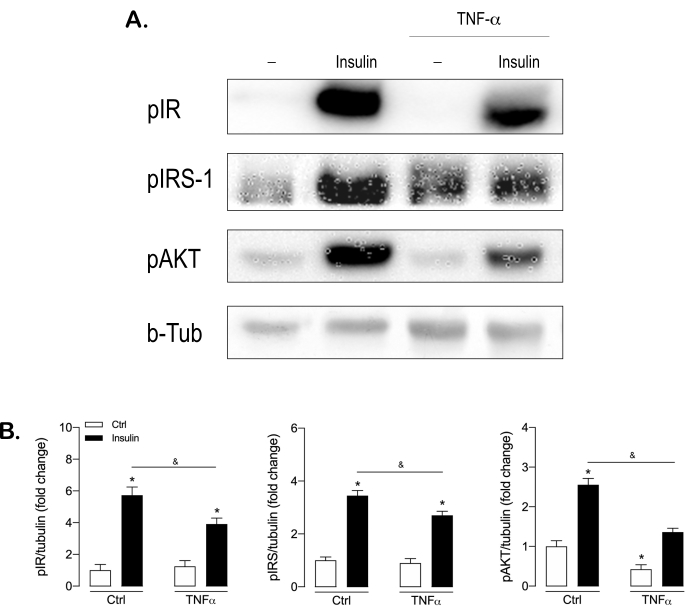
Figure 3: Insulin resistance induced by TNF-α in subcutaneous primary adipocytes demonstrated by decreased insulin-induced phosphorylation of IR, IRS-1, and AKT. Subcutaneous adipocytes were treated with TNF-α (4 ng/mL) for 48 h and serum-starved for the last 24 h. Following the stimulation with insulin (100 nM) for 15 min, 40 µg of total protein was loaded on 7.5% gels and subjected to SDS-PAGE, transferred to nitrocellulose membranes, and blocked in 4% non-fat milk in TBS-T 0.1%. The membrane was probed with anti-phosphorylated IR (pIR), anti-phosphorylated IRS-1 (pIRS-1), and anti-phosphorylated AKT (pAKT), as well as with an anti-rabbit HRP secondary antibody. The signal was visualized with chemiluminescence detection and anti-β tubulin was used as a loading control. (A) Representative blots and (B) quantification from three independent experiments. Data are mean ± SEM; *, p < .05 versus control. Abbreviations: TNF-α = tumor necrosis factor-α; IR = insulin receptor; IRS = insulin receptor substrate; pX = phosphorylated form of X; b-Tub = beta-tubulin; HRP = horseradish peroxidase; Ctrl = control. Please click here to view a larger version of this figure.
| Anti-Fc solution | purified rat anti-mouse CD16 / CD32 diluted in PBS–2% FBS [1:150] | |||
| TBS-T 0.1% | 0.01 M Tris-HCl (pH 8), 0.15 M NaCl, 0.1% Tween 20 | |||
| Blocking solution | 4% nonfat dry milk diluted in TBS-T 0.1% | |||
| Growth medium | 60% DMEM low glucose, 40% MCDB 201 medium, 1x penicillin-streptomycin, 1 nM dexamethasone, 0.1 mM L-ascorbic acid 2-phosphate, 1x insulin, transferrin, sodium, selenite (ITS) liquid media supplement, 1x linoleic acid-albumin from BSA, 10% FBS, 10 ng/mL epidermal growth factor (EGF), 10 ng/mL leukemia inhibitory factor (LIF), 10 ng/mL platelet-derived growth factor BB (PDGF-BB), 5 ng/mL fibroblast growth factor-basic (bFGF), and 50 µg/mL normocin | |||
| Differentiation medium | 60% DMEM low glucose, 40% MCDB 201 medium, 1x penicillin-streptomycin, 1 nM dexamethasone, 0.1 mM L-ascorbic acid 2-phosphate, 1x ITS liquid media supplement, 1x linoleic acid-albumin from BSA, and 2% FBS | |||
| Differentiation cocktail | 0.5 µM 3-isobutyl-1-methylxanthine [IBMX], 1 µM dexamethasone, 10 µM rosiglitazone, and 100 nM insulin | |||
| Simple medium-2% FBS | 60% DMEM low glucose, 40% MCDB 201 medium, 1x penicillin-streptomycin, and 2% FBS | |||
| Simple medium–0% FBS | 60% DMEM low glucose, 40% MCDB 201 medium, 1x penicillin-streptomycin | |||
| RIPA buffer | 50 mM Tris-HCl, 1 mM EGTA, 1 mM EDTA, 1% octylphenoxy poly(ethyleneoxy)ethanol, 1 mM Na3VO4, 48.8 mM NaF, 8.2 mM Na4P2O7, and 0.26 M saccharose | |||
| 6x Laemmli buffer | 1.2 g SDS, 6 mg bromophenol blue, 4.7 mL glycerol, 1.2 mL Tris base 0.5 M pH 6.8, 845 µL 2- mercaptoethanol, and 2.1 mL H2O | |||
| Running buffer | 25 mM Tris base, 192 mM glycine, 1% SDS | |||
| Transfer buffer | 25 mM Tris-base, 192 mM glycine, 20% methanol | |||
Table 1: Solutions used in this protocol.
Supplementary Figure S1: Subcutaneous adipose tissue digested with collagenase. Once the adipose tissue is removed, it is cut into small pieces with scissors to start the digestion process (A), then it is incubated for 30 min with collagenase type 1 at 37 °C, 150 rpm (B). Please click here to download this File.
Supplementary Figure S2: Scheme of the adipocyte precursor cell isolation process. Dissect the inguinal subcutaneous adipose tissue and digest the samples with collagenase. Eliminate mature adipocytes by centrifugation and wash the pellet to remove excess fat. Lyse the red blood cells and block to reduce nonspecific binding. Label the endothelial cells and macrophages with antibodies coupled to magnetic particles. Perform the magnetic separation of cells using the negative separation strategy with a magnetic cell separator. Seed APCs (unlabeled cells) in plates covered with basement membrane matrix. Change the medium every 48 h until the cells reach 80% confluence. Abbreviations: APCs = adipocyte precursor cells; BSA = bovine serum albumin; FBS = fetal bovine serum. Created with Biorender.com. Please click here to download this File.
Supplementary Figure S3: Treatment with TNF-α to induce insulin resistance does not alter the viability of adipocytes. The viability of mature adipocytes was measured by the MTT assay after treatment with 4 ng/mL of TNF-α for 48 h. Data are mean ± SEM. Abbreviations: TNF-α = tumor necrosis factor-α; Ctrl = control. Please click here to download this File.
Supplementary Figure S4: TNF-α treatment in adipocytes decreases the expression of insulin sensitivity markers. The mRNA expression of Insr, Irs, Glut4, and Adipoq was measured by real-time PCR after treatment with 4 ng/mL of TNF-α for 48 h. Data are mean ± SEM; *, p < .05 versus control. Abbreviations: TNF-α = tumor necrosis factor-α; Ctrl = control. Please click here to download this File.
Discussion
This paper provides a method for studying insulin resistance that uses primary adipocytes in culture treated with TNF-α. This model has the advantage that primary adipocytes can be cultured under defined conditions for long periods of time with a tight control of cellular environmental factors26. The assay duration is 15-20 days, although variations in the percentage of differentiated adipocytes can occur between experiments. Primary adipocytes have advantages over cell lines since they have not been continuously expanded in culture and more closely capture the diversity of the tissue from which they are derived27. Furthermore, primary adipocytes allow studying donors under different physio-pathological contexts, such as lean versus obese, male versus female, young versus old, and cells from different fat depots. Another advantage of this method is that cell lines from CRE and knockout mice can be generated after APCs isolation.
One possible problem during APCs isolation and differentiation is that these cells may lose the ability to differentiate, which would likely occur when 80% confluence is exceeded at the start of the differentiation process. The medium should also be changed very gently, especially after lipid accumulation, since differentiated adipocytes easily detach from the culture dish. Moreover, each lot of collagenase must be tested for digestion efficiency, cell yield, and cytotoxicity26. Techniques have been developed to identify and isolate APCs from the stromovascular fraction of white adipose tissue, using negative markers such as CD31 and CD45, as well as positive stem cell population markers such as CD34 and Sca128,29. In this protocol, we only perform a negative separation, eliminating endothelial cells and macrophages from the stromovascular fraction, thus avoiding the loss of preadipocytes in the positive separation.
Although primary cultures allow the study of donor variability and complexity27,30, setting the experimental model introduces limitations. For example, adipocytes isolated from old animals will have a lower differentiation capacity and a higher rate of lipotoxicity compared to those from young animals31,32. The protocol could also be adapted to use different cell separation techniques. If an automatic magnetic cell separator is not available, manual separation of the cells can be achieved with columns or FACS sorting. The method described here for APCs isolation using a magnetic cell separator is an adapted and simplified version of the FACS sorting protocol described by Macotela et al.28.
Several inflammatory cytokines have been used to induce insulin resistance in fat cells, including TNF-α, IL-1β, and IL-618,33,34,35,36. Cultured 3T3-L1 adipocytes exposed to TNF-α become insulin-resistant within several days, as assessed by the reduced ability of insulin to stimulate glucose uptake37. These results show that TNF-α decreases insulin-induced phosphorylation of IR, IRS-1, and AKT21,23, measured by the western blot detection of total protein extracted from primary adipocytes. However, proteases and phosphatases released during cell lysis could influence western blot detection. The active state of proteins should be preserved by the addition of protease and phosphatase inhibitors to the lysis buffer and after protein quantification, by mixing the sample with Laemmli buffer. In conclusion, this method provides a useful tool to study the mechanisms mediating insulin resistance in adipocytes differentiated from mouse APCs.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank Daniel Mondragón, Antonio Prado, Fernando López-Barrera, Martín García-Servín, Alejandra Castilla, and María Antonieta Carbajo for their technical assistance, and Jessica Gonzalez Norris for critically editing the manuscript. This protocol was supported by Consejo Nacional de Ciencia y Tecnología de México (CONACYT), Fondo Sectorial de Investigación para la Educación Grant 284771 (to Y.M.).
Materials
| 1. Isolation mouse adipocyte precursor cells | |||
| ACK lysing buffer | LONZA | 10-548E | |
| Anti-Biotin Microbeads | Miltenyi | 130-090-485 | |
| Anti-CD31 | eBioscience | 13-0311-85 | |
| AutoMACS Pro Separator | Miltenyi | ||
| Basement membrane matrix (matrigel) | Corning | 354234 | |
| bFGF | Sigma | F0291 | Growth factor |
| BSA | Equitech-Bio, Inc. | BAC63-1000 | |
| CD45 Monoclonal Antibody (30-F11) – Biotin | eBioscience | 13-0451-85 | |
| Collagenase, Type 1 | Worthington Biochem | LS004197 | |
| Dexamethasone | Sigma | D1756 | |
| DMEM | GIBCO | 12800017 | |
| DMEM low glucose | GIBCO | 31600-034 | |
| EGF | Peprotech | 315-09 | Growth factor |
| FBS | GIBCO | 26140-079 | |
| ITS mix | Sigma | I3146 | |
| L-ascorbic acid 2-phosphate | Sigma | A8960 | |
| LIF | Millipore | ESG1107 | Growth factor |
| Linoleic acid-albumin | Sigma | L9530 | |
| MCDB 201 medium | Sigma | M6770 | |
| Normocin | InvivoGen | ant-nr-2 | |
| PDGF-BB | Peprotech | 315-18 | Growth factor |
| Peniciline-Streptomycine | BioWest | L0022-100 | |
| Pre-Separation Filters (70 µm) | Miltenyi | 130-095-823 | |
| Purified Rat Anti-Mouse CD16 / CD32 | BD Pharmingen | 553142 | |
| Trypsin-EDTA | GIBCO | 25300062 | |
| 2. Adipocyte differentiation and insulin resistance induction | |||
| 3-Isobutyl-1-methylxanthine [IBMX] | Sigma | I5879 | Differentiation cocktail |
| BMP4 | R&D Systems | 5020-BP-010 | Differentiation cocktail |
| Dexamethasone | Sigma | D1756 | Differentiation cocktail |
| Insulin | Sigma | I9278 | |
| Rosiglitazone | Cayman | 71742 | Differentiation cocktail |
| TNFα | R&D Systems | 210-TA-005 | |
| 3. Evaluation of insulin signaling pathway by western blot | |||
| Anti-beta tubulin antibody | Abcam | ab6046 | |
| Bromophenol blue | BioRad | 161-0404 | Laemmli buffer |
| EDTA | Sigma | E5134 | RIPA buffer |
| EGTA | Sigma | E4378 | RIPA buffer |
| FluorChem E system | ProteinSimple | ||
| Glycerol | Sigma | G6279 | Laemmli buffer |
| Glycine | Sigma | G7126 | Running and Transfer buffer |
| Igepal | Sigma | I3021 | RIPA buffer |
| 2- mercaptoethanol | Sigma | M3148 | Laemmli buffer |
| Methanol | JT Baker | 907007 | Transfer buffer |
| NaCl | JT Baker | 3624-05 | TBS-T |
| NaF | Sigma | 77F-0379 | RIPA buffer |
| NaOH | JT Baker | 3722-19 | |
| Na4P2O7 | Sigma | 114F-0762 | RIPA buffer |
| Na3VO4 | Sigma | S6508 | RIPA buffer |
| Nitrocellulose membrane | BioRad | 1620112 | |
| Nonfat dry milk | BioRad | 1706404 | Blocking solution |
| Prestained protein standard | BioRad | 1610395 | |
| Protease inhibitor cocktail | Sigma | P8340-5ML | |
| Peroxidase AffiniPure Donkey Anti-Rabbit IgG (H+L) | Jackson ImmunoResearch | 711-035-132 | |
| Phospho- Insulin Receptor β | Cell signaling | 3024 | |
| Phospho-Akt (Ser473) Antibody | Cell signaling | 9271 | |
| Phospho-IRS1 (Tyr608) antibody | Millipore | 9432 | |
| Saccharose | JT Baker | 407205 | RIPA buffer |
| SDS | BioRad | 1610302 | Running and laemmli buffer |
| SuperSignal West Pico PLUS Chemiluminescent Substrate | Thermo Scientific | 34577 | |
| Tris-base | Promega | H5135 | Running, transfer and laemmli buffer |
| Tris-HCl | JT Baker | 4103-02 | RIPA buffer – TBS |
| Tween 20 | Sigma | P1379 | TBS-T |
References
- Elmus, G. B. Insulin signaling and insulin resistance. Journal of Investigative Medicine. 61 (1), 11-14 (2013).
- Kasuga, M., Zick, Y., Blithe, D. L., Crettaz, M., Kahn, C. R. Insulin stimulates tyrosine phosphorylation of the insulin receptor in a cell-free system. Nature. 298 (5875), 667-669 (1982).
- Kasuga, M., Karlsson, F. A., Kahn, C. R. Insulin stimulates the phosphorylation of the 95,000-dalton subunit of its own receptor. Science. 215 (4529), 185-187 (1982).
- Taniguchi, C. M., Emanuelli, B., Kahn, C. R. Critical nodes in signalling pathways: insights into insulin action. Nature Reviews Molecular Cell Biology. 7 (2), 85-96 (2006).
- Insulin Resistance. StatPearls Available from: https://www.ncbi.nlm.nih.gov/books/NBK507839/ (2021)
- Petersen, M. C., Shulm, G. I. Mechanisms of insulin action and insulin resistance. Physiological Reviews. 98 (4), 2133-2223 (2018).
- Batista, T. M., Haider, N., Kahn, C. R. Defining the underlying defect in insulin action in type 2 diabetes. Diabetologia. 64 (5), 994-1006 (2021).
- Unamuno, X., Frühbeck, G., Catalán, V. Adipose Tissue. Encyclopedia of Endocrine Diseases. Second edition. , 370-384 (2019).
- Cinti, S. Pink adipocytes. Trends in Endocrinology & Metabolism. 29 (9), 651-666 (2018).
- Kahn, B. B., Flier, J. S. Obesity and insulin resistance. The Journal of Clinical Investigation. 106 (4), 473-481 (2000).
- Klemm, D. J., et al. Insulin-induced adipocyte differentiation. The Journal of Biological Chemistry. 276 (30), 28430-28435 (2001).
- Wilcox, G. Insulin and insulin resistance. The Clinical Biochemist. Reviews. 26 (2), 19-39 (2005).
- Yohannes, T. W. Obesity, insulin resistance, and type 2 diabetes: associations and therapeutic implications. Diabetes, Metabolic Syndrome and Obesity: Targets and Therapy. 13, 3611-3616 (2020).
- James, D. E., Stöckli, J., Birnbaum, M. J. The etiology and molecular landscape of insulin resistance. Nature Reviews Molecular Cell Biology. 22 (11), 751-771 (2021).
- Wondmkun, Y. T. Obesity, insulin resistance, and type 2 diabetes: associations and therapeutic implications. Diabetes, Metabolic Syndrime and Obesity: Targets and Therapy. 9 (13), 3611-3616 (2020).
- Hardy, O. T., Czech, M. P., Corvera, S. What causes the insulin resistance underlying obesity. Current Opinion in Endocrinology, Diabetes, and Obesity. 19 (2), 81-87 (2012).
- Klein, S., Gastaldelli, A., Yki-Järvinen, H., Scherer, P. E. Why does obesity cause diabetes. Cell Metabolism. 34 (1), 11-20 (2022).
- Lo, K., et al. Analysis of in vitro insulin-resistance models and their physiological relevance to in vivo diet-induced adipose insulin resistance. Cell Reports. 5 (1), 259-270 (2013).
- Asterholm, I. W., et al. Adipocyte inflammation is essential for healthy adipose tissue expansion and remodeling. Cell Metabolism. 20 (1), 103-118 (2014).
- Hotamisligil, G. S., Murray, D. L., Choy, L. N., Spiegelman, B. M. Tumor necrosis factor alpha inhibits signaling from the insulin receptor. Proceedings of the National Academy of Sciences. 91 (11), 4854-4858 (1994).
- Ruan, H., Hacohen, N., Golub, T. R., Parijs, L. V., Lodish, H. F. Tumor necrosis factor-α suppresses adipocyte-specific genes and activates expression of preadipocyte genes in 3T3-L1 adipocytes. Nuclear factor-κB activation by TNF-α is obligatory. Diabetes. 51 (5), 1319-1336 (2002).
- Boucher, J., Kleinridders, A., Kahn, C. R. Insulin receptor signaling in normal and insulin-resistant states. Cold Spring Harbor Perspectives in Biology. 6 (1), 009191 (2014).
- Hotamisligil, G. S., et al. IRS-1-mediated inhibition of insulin receptor tyrosine kinase activity in TNF-alpha- and obesity-induced insulin resistance. Science. 271 (5249), 665-668 (1996).
- Copps, K. D., White, M. F. Regulation of insulin sensitivity by serine/threonine phosphorylation of insulin receptor substrate proteins IRS1 and IRS2. Diabetologia. 55 (10), 2565-2582 (2012).
- Gassmann, M., Grenacher, B., Rohde, B., Vogel, J. Quantifying western blots: pitfalls of densitometry. Electrophoresis. 30 (11), 1845-1855 (2009).
- Skurk, T., Hauner, H. Primary culture of human adipocyte precursor cells: expansion and differentiation. Methods in Molecular Biology. 806, 215-226 (2012).
- Hausman, D. B., Park, H. J., Hausman, G. J. Isolation and culture of preadipocytes from rodent white adipose tissue. Methods in Molecular Biology. 456, 201-219 (2008).
- Macotela, Y., et al. Intrinsic differences in adipocyte precursor cells from different white fat depots. Diabetes. 61 (7), 1691-1699 (2012).
- Rodeheffer, M. S., Birsoy, K., Friedman, J. M. Identification of white adipocyte progenitor cells in vivo. Cell. 135 (2), 240-249 (2008).
- Hausman, D. B., DiGirolamo, M., Bartness, T. J., Hausman, G. J., Martin, R. J. The biology of white adipocyte proliferation. Obesity Reviews. 2 (4), 239-254 (2001).
- Kirkland, I. M., Tchkonia, T., Pirtskhalava, T., Han, J., Karagiannides, I. Adipogenesis and aging: does aging make fat go MAD. Experimental Geronotology. 37 (6), 757-767 (2002).
- Guo, W., et al. Aging results in paradoxical susceptibility of fat cell progenitors to lipotoxicity. American Journal of Physiology. Endocrinology and Metabolism. 292 (4), 1041-1051 (2007).
- Ruan, H., Hacohen, N., Golub, T. R., Van Parijs, L., Lodish, H. F.Tumor necrosis factor-a suppresses adipocyte-specific genes and activatesexpression of preadipocyte genes in 3T3-L1 adipocytes: nuclear factor-kappaB activation by TNF-a is obligatory. Diabetes. 51 (5), 1319-1336 (2002).
- Jager, J., Gre ́meaux, T., Cormont, M., Le Marchand-Brustel, Y., Tanti, J. F. Interleukin-1b-induced insulin resistance in adipocytes throughdown-regulation of insulin receptor substrate-1 expression. Endocrinology. 148 (1), 241-251 (2007).
- Rotter, V., Nagaev, I., Smith, U. Interleukin-6 (IL-6) induces insulinresistance in 3T3-L1 adipocytes and is, like IL-8 and tumor necrosis factor-a,overexpressed in human fat cells from insulin-resistant subjects. The Journal of Biological Chemistry. 278 (46), 45777-45784 (2003).
- Isidor, M. S., et al. Insulin resistance rewires the metabolic gene program and glucose utilization in human white adipocytes. International Journal of Obesity. 46 (3), 535-543 (2021).
- Houstis, N., Rosen, E. D., Lander, E. S. Reactive oxygen species have a causal role in multiple forms of insulin resistance. Nature. 440 (7086), 944-948 (2006).

