RNA In situ Hybridization in Whole Mount Embryos and Cell Histology Adapted for Marine Elasmobranchs
Summary
By combining methods for RNA whole mount in situ hybridization and histology, gene expression can be linked with cell fate decisions in the developing embryo. These methods have been adapted to marine elasmobranchs and facilitate the use of these animals as model organisms for biomedical, toxicology and comparative studies.
Abstract
Marine elasmobranchs are valued animal models for biomedical and genomic studies as they are the most primitive vertebrates to have adaptive immunity and have unique mechanisms for osmoregulation 1-3. As the most primitive living jawed-vertebrates with paired appendages, elasmobranchs are an evolutionarily important model, especially for studies in evolution and development. Marine elasmobranchs have also been used to study aquatic toxicology and stress physiology in relationship to climate change 4. Thus, development and adaptation of methodologies is needed to facilitate and expand the use of these primitive vertebrates to multiple biological disciplines. Here I present the successful adaptation of RNA whole mount in situ hybridization and histological techniques to study gene expression and cell histology in elasmobranchs.
Monitoring gene expression is a hallmark tool of developmental biologists, and is widely used to investigate developmental processes 5. RNA whole mount in situ hybridization allows for the visualization and localization of specific gene transcripts in tissues of the developing embryo. The expression pattern of a gene’s message can provide insight into what developmental processes and cell fate decisions a gene may control. By comparing the expression pattern of a gene at different developmental stages, insight can be gained into how the role of a gene changes during development.
While whole mount in situ‘s provides a means to localize gene expression to tissue, histological techniques allow for the identification of differentiated cell types and tissues. Histological stains have varied functions. General stains are used to highlight cell morphology, for example hematoxylin and eosin for general staining of nuclei and cytoplasm, respectively. Other stains can highlight specific cell types. For example, the alcian blue stain reported in this paper is a widely used cationic stain to identify mucosaccharides. Staining of the digestive tract with alcian blue can identify the distribution of goblet cells that produce mucosaccharides. Variations in mucosaccharide constituents on short peptides distinguish goblet cells by function within the digestive tract 6. By using RNA whole mount in situ‘s and histochemical methods concurrently, cell fate decisions can be linked to gene-specific expression.
Although RNA in situ‘s and histochemistry are widely used by researchers, their adaptation and use in marine elasmobranchs have met limited and varied success. Here I present protocols developed for elasmobranchs and used on a regular basis in my laboratory. Although further modification of the RNA in situ‘s hybridization method may be needed to adapt to different species, the protocols described here provide a strong starting point for researchers wanting to adapt the use of marine elasmobranchs to their scientific inquiries.
Protocol
I. RNA Whole Mount In situ Hybridization in Marine Elasmobranchs
1. Embryo Fixation and Preparation
- Skate embryos can be staged according to Ballard 7. Optimal stages for detection of gene expression depends on the tissue of interest. To track gene expression in the skate digestive tract, stages 27-30 are optimal 7.
- Dissect embryos into PBS, separating the embryos from the yolk sac.
- Fix in 30 ml of freshly made 4% paraformaldehyde (PFA) in a 50 ml conical tube at 4 °C with gentle rocking. Fix for 24 hr.
- Wash embryos in PBT, twice for 15 min, rotating at room temperature. All solution recipes for RNA whole mount in situ‘s are included (Table 1).
- Dehydrate embryos by washing in a methanol series of 25%, 50% and 75% methanol:PBT rocking at room temperature. The duration of the washes depends on the stage of the embryos. For pre-neurulation staged embryos, 5 min of each wash in the methanol series is sufficient. For embryos older than stage 30, washes can last up to 30 min. To determine if embryos are sufficiently penetrated and acclimated to each wash, hold the tube upright in your hand. If the embryos sink to the bottom of the tube, then you are ready for the next wash. If the embryos float to the top of the solution, change the solution and repeat that particular dehydration step.
- Wash twice in 100% methanol for 10 min gently rocking.
- Once dehydrated to 100% methanol, embryos should be stored at -20 °C for at least 24 hr before proceeding with the protocol. Embryos have been successfully stored at -20 °C and used for RNA whole mounts for up to 1 year.
2. Synthesis of RNA Probe
- Generate a linear fragment of the gene whose expression you want to detect. This can be done either by PCR amplification or linearizing a plasmid with your gene of interest. To amplify by PCR, choose primers whose binding sites flank the gene insert and retain RNA polymerase binding sites in the amplified region. For commonly used plasmids such as pBS or pGEM, M15 forward and reverse primers are ideal. Alternatively, linearize a plasmid by digesting 15 μl of QIAGEN miniprep DNA with an enzyme that cleaves at the 5′ end of the gene. Gel extract and purify using the appropriate QIAquick kit.
- Reverse transcribe the RNA probe using 8 μl of linear plasmid or 100 ng of PCR product as a template using the appropriate RNA polymerase (see Table 2 for transcription reaction).
- Incubate reverse transcription reaction for 2 hr at 37 °C.
- To confirm reaction, run 1 μl of the transcription reaction on a 1% agarose/TAE gel. Run gel at 100 mVolts until the dye has just run into the gel. A single prominent band should be visible. The linear DNA template fragment may be faintly visible at a higher molecular weight.
- Add 1 μl of RNAse-free DNAse I to the transcription reaction and incubate at 37 °C for 15 min.
- To precipitate, add 100 μl TE pH 8, 10 μl 4 M LiCl, 300 μl 100% EtOH and incubate at -20 °C for at least 60 min or overnight.
- Spin for 10 min in a 4 °C microfuge at 14,000 rpm.
- Wash pellet with 70% EtOH and air dry.
- Resuspend RNA probe in 20 μl of dH2O. Add 80 μl of hybridization buffer (hybridization buffer should be pre-warmed to 70 °C). The probe can be stored in 80% hybridization buffer for several months at -20 °C or longer at -80 °C.
3. Embryo Pre-treatment and Hybridization
- Remove embryos stored in 100% methanol from -20 °C. For younger embryos proceed with step (3.2). For later staged embryos (stage 29/30 <) you may see an epidermis layer that has ‘lifted’ off the body of the embryo. Under the microscope, carefully dissect off the epidermal layer to maximize probe penetration.
- Place embryos in clean glass scintillation vials. Rehydrate the embryos in the reverse methanol series describe above: (75%, 50%, 25% methanol:PBT) by rocking gently each solution for 5-30 min each at room temperature. Complete rehydration with two washes of PBT for 10 min each, gently rocking.
- To remove any pigment, bleach embryos with 6% hydrogen peroxide in PBT for 1 hr at room temperature, rocking.
- Remove hydrogen peroxide by washing three times in PBT, (rocking at room temperature, 10 min each).
- Treat embryos with proteinase K to allow for penetration of probe during hybridization. The amount of proteinase K and the duration of the treatment depends on the tissue you want to examine and the age of the embryo. More superficial tissue and younger embryos require shorter and less proteinase K, while deeper tissues and older embryos require a higher concentration and longer treatment time to fully permeabilize the embryo for probe hybridization. For example, to detect Shh in the gut endoderm of stage 29 skate embryos, 30 μg/ml of proteinase K/PBT is incubated for 20 min at room temperature.
- Quickly rinse embryos in PBT to wash away proteinase K.
- To deactivate proteinase K, postfix embryos with 4% PFA, 0.2% glutaraldehyde in PBT for 20 min, gently rocking.
- Wash twice for 10 min with PBT.
-
- For pre-hybridization, add 2-3 ml of pre-heated hybridization solution to the embryos, just enough to cover them. Rock gently in a heated water bath at 70 °C for 1 hr.
- To prepare hybridization solution, preheat RNA probe to 70 °C for 10 min and cool on ice. Dilute 15-20 μl of RNA probe to 2 ml of fresh preheated hybridization solution, to obtain a final concentration of 0.5-1.0 μg/ml. Remove pre-hybe and add diluted RNA probe to embryos. Embryos should be just covered by solution, add more hybridization solution if necessary. Tightly secure lid of scintillation vial. Hybridize by rocking gently in a heated water bath at 70 °C overnight.
4. Post Hybridization Washes and Antibody Hybridization
- Remove hybridization solution with probe from embryos and place in a fresh scintillation vial or Eppendorf tube. The probe can be stored at -20 °C and reused in the future. For post-hybridization washes, place embryos in a Netwell seated in a 6-well tissue culture plate.
- Wash 3 times for 30 min each in pre-warmed Solution #1, rocking at 70 °C.
- Wash 3 times for 30 min each in pre-warmed Solution #2, rocking at 70 °C.
- Wash 3 times for 5 min each in TBST, rocking at room temperature.
- Pre-block with 10% heat-inactivated sheep serum in TBST for 1 hr, rocking at room temperature.
- Transfer embryos from Netwell to a fresh glass scintillation vial. Add anti-DIG Fab Fragments (1:5,000) in 1% heat-inactivated sheep serum/TBST to embryos. Rock overnight at 4 °C.
5. Post Antibody Hybridization Washes
- Remove antibody and discard. Place embryos back into Netwell for washes.
- Wash 3 times for 5 min each in TBST, rocking at room temperature
- Wash at least 6 times for 1 hr each, rocking at room temperature.
- Wash overnight in TBST, rocking at 4 °C. As the post antibody washes help cut down on background staining, maximizing the number and duration of washes is preferable. Embryos are routinely washed in TBST at 4 °C, over the weekend.
6. Detection of Probe
- Make up fresh NTMT solution and filter sterilize.
- Discard TBST wash solution and place embryos in a clean glass scintillation vial. Wash embryos 3 times in fresh NTMT for 10 min each at room temperature, rocking.
- Remove NTMT wash and add reaction mix of 1 x NBT / 1 x BCIP in NTMT. As these reactants are light sensitive, cover scintillation vial in foil and rock at room temperature.
- Monitor color reaction every 15 min until desired gene expression is clearly visible. Strong probes may take as little as 10 min to develop. Weak probes may take 1-2 hr. Do not let reaction go for too long or background staining will begin to appear (the whole embryo starts to turn a dull purple or brown color).
- Wash 2 times for 10 min each in PBT at room temperature, rocking.
- Postfix embryos in 4% paraformaldehyde, 0.1% glutaraldehyde for 1 hr at room temperature. Embryos can be store indefinitely in fixative at 4 °C.
- Visualize embryos with a dissecting stereomicroscope. To produce a light blue background for pictures, place embryos in a dish with pre-hardened 1% agarose/PBS.
7. Representative Results for I. RNA Whole Mount in situ hybridization in marine elasmobranchs
RNA whole mount in situ‘s depicting expression of Sonic hedgehog (Shh) and Hoxa13 in skate embryos are shown in Figure 1. Expression of Shh in higher vertebrates is found in the notochord and gut endoderm and this expression pattern is conserved in the skate (Figure 1a) 8,9 . Marine elasmobranchs have a unique method of osmoregulation that uses the rectal gland to secrete salts. Hoxa13 expression is high in the developing rectal gland (Figure 1b) 10. Hoxa13 gene product role in patterning the rectal gland remains unknown.
II. Paraffin Embedding and Sectioning Elasmobranch Tissue
1. Harvest and Preparation of Tissue
- Dissect desired tissue and fix in 10% formalin for 24-48 hr, rocking at room temperature. 4% paraformaldehyde (PFA) can also be used as a fixative (see Discussion).
- Wash out fixative by washing in PBS, 3 times for 30 min each. Depending on the size of the tissue, washing times can be decreased to 10 min or increased to 1 hr.
- Dehydrate embryos by washing in an ethanol series of 25%, 50% and 75% ethanol:PBS rocking at room temperature. Again, the time for each wash will depend on the size of the tissue. For 0.5 cm3 tissue, incubate each wash for 15 min. Increase time for larger pieces. If long term storage of tissue is desired, wash an additional two times in 75% ethanol and store in -20 °C for up to one year. Otherwise proceed to next step.
- Further dehydrate by washing 3 times in 100% ethanol for 15 min each, rocking.
2. Paraffin Embedding and Sectioning
- Clarify tissue by washing 2 times in xylene for 5 min each in glass scintillation vials with screw cap lids. Xylene will render the tissue brittle, so do not exceed a total of 10 min for washes. The tissue should look translucent when you hold it up to a light. If the tissue is still very opaque after the two washes, do an additional wash in xylene for 5 min and then proceed with protocol. Only handle xylene inside a fume hood with gloves.
- Remove xylene and fill the scintillation vial 2/3 full with melted paraffin. Incubate in a 60 °C oven for 30 min. When putting paraffin into the vial, you will notice the paraffin solidify along the sides of the vial. Place the vial in the oven and allow the paraffin to re-dissolve for a few minutes. Take the vial out of the oven and give the solution a quick swirl to mix and replace in oven for remainder of incubation.
- Repeat paraffin incubations two more times for 30 min.
- Incubate in paraffin twice for an hour each. Change paraffin a final time and incubate overnight in the 60 °C oven.
- The following day, place the tissue in a heated paraffin bath chamber and place under vacuum for 2-3 hr. This will facilitate infiltration of paraffin into the tissue and remove any air bubbles. For some types of thin tissue, a vacuum step may not be required, but it is necessary for whole embryos or digestive tract and other organs with luminal compartments.
- After vacuum-facilitated infiltration of tissue, place tissue in a metal mold with melted paraffin, and cool on an ice plate. Leave on ice or at 4 °C overnight before trying to remove the tissue from the mold. Paraffin blocks of tissue can be stored indefinitely at 4 °C.
- Place paraffin block onto a microtome and cut 8 μm sections. Sections should be floated on a water bath heated to 48 °C and mounted onto glass Superfrost*Plus slides.
- Dry slides overnight in a 37 °C oven.
- Store sections at 4 °C until needed.
III. Alcian Blue / Nuclear Fast Red Stain of Elasmobranch Tissue
1. Alcian Blue Stain for Mucins
- Place slides with sections facing upward on a slide warmer. Melt for 45 min.
- Place slides into a slide holder. All consequent solutions are contained in tubs inside a fume hood. Slides are washed by transferring them from one tub with a solution to another. Make sure that the solution levels in the tubs covers the slides; the tissues on the slides are completely submerged. Dissolve paraffin by washing slides 2 times in xylene for 5 min each.
- Wash slides two times with 100% ethanol for 1 min each.
- Rehydrate slides in an ethanol series (75%, 50%, 25% ethanol) washing in each solution for 1 min. Wash in dH2O for 1 min.
- Stain in Alcian Blue Solution, pH 2.5 for 15 min.
- Develop stain by washing in running tap water for 3 min. Dip once in dH2O.
- Counterstain in Nuclear Fast Red Counterstain solution for 10 min.
- Wash in running tap water for 3 min and dip in dH2O one time.
- Dehydrate in an ethanol series (25%, 50%, 75%, 100%, 100%) for 20 dips each, or 1 min each. Wash 2 times in xylene for 5 min each. Mount with DPX mounting medium and coverslips. Allow mounting medium to dry and harden for 48 hr in the fume hood before manipulating slides.
Representative Results
Examples of alcian blue staining in different regions of the L. erinacea digestive tract are shown in Figure 2. Acid mucin containing globlet cells are clearly visible by the alcian blue stain throughout the digestive tract. The distribution of acid mucins differs in the different regions of the digestive tract, thus reflecting differences in function. Acidic mucins are sparsely produced in the spiral intestine and cloaca, while a high concentration of acid mucins are detected in the distal intestine (compare Figure 2a and 2c with 2b).
| PBT | 1 x PBS |
| 0.1% Tween-20 | |
| Filter-sterilize in Nalgene-type tissue culture filter units. | |
| 20 x SSC, pH 4.5 | 3 M NaCl |
| 0.3 M NaCitrate | |
| Adjust pH with citric acid. | |
| Hybridization solution | 50% formamide |
| 1.3 x SSC, pH 4.5 | |
| 5 mM EDTA, pH 8 | |
| 50 mg/ml tRNA | |
| 0.2% Tween-20 | |
| 0.5% Chaps | |
| 100 mg/ml heparin | |
| Aliquot and store at -20 °C for up to one year. | |
| Proteinase K | 10 mg/ml stock solution |
| 20 μl aliquots into 0.5 μl Eppendorf-like tubes and store at -20 °C. | |
| Solution #1 | 50% formamide |
| 1.3 x SSC, pH 4.5 | |
| 0.2% Tween-20 | |
| Store at -20 °C | |
| Solution #2 | 50% formamide |
| 1 x SSC, pH 4.5 | |
| 0.2% Tween-20 | |
| Store at -20 °C | |
| Sheep serum | Inactivate by heating to 55 °C for 1 hr, and store in aliquots at -20 °C. |
| 10 x TBS | 80 g NaCl |
| 2 g KCl | |
| 250 ml, 1 M Tris-HCl, pH 7.5 | |
| Add water to 1 L | |
| 1 x TBST | 1 x TBS |
| 0.1% Tween-20 | |
| NTMT | 100 mM NaCl |
| 100 mM Tris-HCl, pH 9.5 | |
| 50 mM MgCl2 | |
| 0.1% Tween-20 | |
| Make approximately 100 ml fresh before using and filter. | |
| 200 x NBT stock | 50 mg/ml NBT in 70% dimethyl formamide |
| Store in -20 °C in 1 ml aliquots. | |
| 200 x BCIP stock | 25 mg/ml BCIP in water |
| Store in -20 °C in 1 ml aliquots. |
Table 1. RNA whole mount in situ solutions.
| Linearized plasmid | 8 μl |
| 10 x transcription buffer | 4 μl |
| DIG-UTP nucleotide mix | 2 μl |
| RNAse inhibitor | 0.5 μl |
| RNA polymerase (SP6, T3 or T7) | 1 μl |
| RNAse-free sterile H20 | 4.5 μl |
| Total volume | 20 μl |
Table 2. Transcription reaction to generate RNA probe.
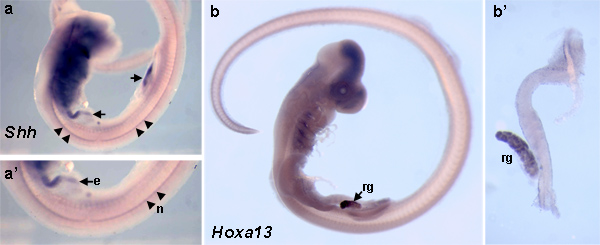
Figure 1. Shh and Hoxa13 expression in Leucoraja erinacea embryos are visualized by RNA whole mount in situ hybridization. (a) Shh expression is detected in a stage 29 skate embryo. (a’) Higher magnification of (a) reveals Shh expression in the notochord (arrowheads) and gut endoderm (arrows). (b) Hoxa13 expression is localized to the rectal gland (arrow) of a stage 29 skate embryo. (b’) Dissected digestive tract from the embryo in (b) demonstrates the specificity of Hoxa13 transcript expression to the rectal gland. e, endoderm; n, notochord; rg, rectal gland. Click here to view larger figure.
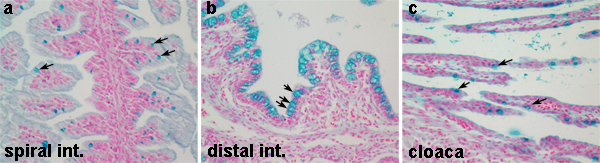
Figure 2. Distribution of acidic mucin-producing goblet cells in the skate digestive tract. (a, c) Low numbers of acidic mucin-producing goblet cells are detected by the alcian blue stain in the spiral intestine and cloaca, respectively (arrows). (b) The distal intestine contains a higher density of acidic mucin goblet cells (arrows). In all panels, nuclei are clearly visible by the nuclear fast red stain. Click here to view larger figure.
Discussion
The protocols presented are classic methods for monitoring gene expression and identifying differentiated cell types, and have been adapted for use in marine elasmobranchs. Further modifications of these protocols may be needed to adapt to different elasmobranch species.
The most common concern regarding RNA whole mount in situ‘s is the risk of RNase contamination and thereby the degradation of the RNA probe and endogenous messages. Two aspects need to be considered: the synthesis of the riboprobe and its hybridization to embryonic messages, and general techniques for handling RNA. In general, contamination by ribonucleases can be avoided by using plasticware from unopened containers, and previously unused glassware. If new glassware is unavailable, treat all glassware with RNase-AWAY (VWR#17810-491). Washes are done by gently pouring off solutions through a perforated spoon and adding fresh solution to the vial. Alternatively, very small embryos or organs/tissues can be placed in a Netwell that fits into a 6-well tissue culture dish. The tissue can be moved from one solution to the next by simply lifting the Netwell from one well to the next. This helps prevent the loss of any tissue by pouring, and also preserves the architecture of the tissue. Additional suggestions for handling RNA and preventing RNase contamination can be found in Molecular Cloning; A Laboratory Manual, and on several pharmaceutical websites, including Roche (http://www.roche-applied-science.com/labfaqs/p2_1.htm) 11.
Generation of the RNA probe and pre-treatment and hybridization of embryos are the most vulnerable steps to contamination by ribonucleases. Antisense RNA probes that complement native mRNA transcript are synthesized by in vitro reverse transcription in the presence of digoxigenin-UTP. Briefly, cDNA plasmids are linearized with a unique restriction enzyme whose site is located at the 5′ end of the gene. This allows for the RNA polymerase to fall off at the end of the gene. Choose a RNA polymerase by identifying the RNA polymerase promoter available in the plasmid, just 3′ to the stop codon. While T3 and T7 RNA polymerases are often used for pBluescript, SP6 is the choice for genes subcloned into the directional pGEM plasmids. In addition to RNase inhibitor, DEPC-treated water is used in the transcription reaction and purification of the probe. Hybridization solution and steps using PBT in the pre-treatment of embryos should all be made with DEPC water. To DEPC treat water, add 1 ml of DEPC to 1 L of water in a large flask and mix well. Leave overnight in a hood and autoclave the next day. Solutions such as 10 x PBS and SSC may also be made with DEPC-treated water. After hybridization with the riboprobe, the use of RNase-free solutions and similar precautions are no longer necessary.
Additional variables worth considering are the concentration and length of proteinase K treatment, which can affect the penetration of the probe. A careful titration of the proteinase K stock is recommended to optimize treatment time for different embryonic stages. The protocol described here has been routinely and successfully used with embryos at stages 20 – 31 according to Ballard 7. For embryos that are very young or for tissue that is particularly “sticky”, the detergent Tween-20 can be replaced with Triton-X. In addition, gene-specific modifications to the protocol may be needed depending on the abundance of transcript. To reduce background, 10% dimethylformamide has been routinely added to the development solution (NTMT) by several groups 12.
In contrast to the sensitive nature of RNA in situ‘s, the joy of histology is that it is quick and pretty foolproof! Any variability in staining is likely due to inadequate fixation. To ensure that tissue is fully fixed, it is recommended that very large tissues (such as the digestive tract of an adult animal) be dissected into smaller pieces and fresh fixative solution replaced every 10 hr over a 48 hr period. It may be necessary to optimize conditions for fixation, both under and over-fixed tissues can result in poor sectioning or uneven staining. In addition, different histochemical stains work optimally with different fixatives. Thus, it is worth modifying the fixative according to the histochemical stain used 13. 4% paraformaldehyde is recommended when fixing soft tissue organs or young embryos. For older embryos or whole animals, 10% formalin is preferred. Once tissue is paraffin embedded and sectioned, it may be used for multiple purposes, including RNA in situ‘s. This allows for resolution of gene expression on the cellular level. Alternatively, sections may be stained with different histochemical stains to identify differentiated cell types.
The alcian blue and nuclear fast red staining has been successfully performed in the skate (Leucoraja erinacea), dogfish (Squalas acanthias), hagfish (Myxine glutinosa) and lamprey (Petromyzon marinus), (unpublished results) 10,14 . In addition to the alcian blue pH 2.5 stain described here, modifying the pH of the alcian blue solution can distinguish different mucosaccharides (i.e. sialo- versus sulfomucins) 13. Different mucosaccharide constituents can distinguish goblet cells by function and localization within the digestive tract 15,16 . Alcian blue stain in the digestive tract has been used to diagnose conditions such as Barrett’s esophagus and to enhance staining of pancreatic beta-cell granules 17,18 .
In summary, histochemistry and RNA whole mount in situ‘s when used together can lead to a better understanding of the link between where a gene is expressed during development and the resulting differentiated cell type. Expression of posterior Hox genes has been linked to specification of acid mucin goblet cells in the development digestive tract 10,19 . Here I provide an example of the expression of Shh and a Hox gene along with the presence of acid mucin-producing goblet cells in the L. erinacea digestive tract. These techniques can be widely applied to different species of elasmobranchs, different developmental patterning genes and different histological stains. The continued application and adaptation of techniques to marine elasmobranchs is important to keep pace with the broad use of these animals for biomedical research models in physiology, endocrinology, toxicology and genomics.
Declarações
The authors have nothing to disclose.
Acknowledgements
I wish to thank the many undergraduate students who have worked in my laboratory and contributed to the evolution of these protocols. NAT has received support from the Skidmore-Union Network, a project established with a NSF ADVANCE PAID grant.
Materials
| Name of the reagent | Company | Catalogue number | Comments |
| 10 x transcription buffer | Roche | 11-465-384-001 | |
| DIG-RNA labeling mix | Roche | 11-277-073-910 | |
| RNAse inhibitor | Roche | 03-335-399-001 | |
| RNA polymerase – SP6 | Roche | 10-810-274-001 | |
| DNAseI, RNAse-free | Roche | 10-776-785-001 | |
| Yeast RNA | Invitrogen | 15401-029 | |
| CHAPS | EMD-Millipore | 220201 | |
| heparin | Sigma-Aldrich | H4784 | |
| DEPC (diethyl pyrocarbonate) | Research Organics | 2106D | |
| Moria Perforated Spoon | Fine Science Tools | 10370-17 | |
| Netwell inserts | Electron Microscopy Sciences | 64713-00 | Netwells for use in 6-well tissue culture dishes |
| 6-well tissue culture plate | Corning | 3516 | |
| Glass scintillation vials with screw-cap lids | Weaton Science Products | 986540 | |
| formamide | Fisher | BP227500 | |
| Proteinase K | Invitrogen | 59895 (AM2542) | |
| NBT | 11585029001 | ||
| BCIP | Roche | 11585002001 | |
| Hydrogen peroxide, 30% | EMD | HX0635-1 | |
| Sheep serum | VWR | 101301-478 | |
| glutaraldehyde | Sigma-Aldrich | G5882 | |
| tRNA | Roche | 10-109-541-001 | |
| Anti-DIG Fab Fragments | Roche | 1137-6623 | |
| Table 3. Reagents and equipment for RNA whole mount in situ‘s. | |||
| 1% Alcian Blue 8GS, pH 2.5 | Electron Microscopy Sciences | 26323-01 | |
| Nuclear Fast Red | Electron Microscopy Sciences | 26078-05 | |
| DPX Mountant | Electron Microscopy Sciences | 13510 | |
| Paraffin (Paraplast X-tra) | McCormick Scientific | 39503002 | |
| 10% Formalin, NBF | VWR | 95042-908 | |
| Glass scintillation vials with screw-cap lids | Weaton Science Products | 986540 | |
| Stainless steal base molds | Tissue-Tek | 4161-4165 | Multiple sizes available. |
| Cassettes | Tissue-Tek | 4170 | |
| Slide warmer | Fisher-Scientific | 12-594Q | |
| Tissue Embedder | Leica Microsystems | EG1160 | |
| Microtome, rotary | Leica Microsystems | RM2235 | |
| Tissue-Tek Slide Staining Set | Electron Microscopy Sciences | 62540-01 | |
| Tissue-Tek 24-Slide Holder | Electron Microscopy Sciences | 62543-06 | |
| Superfrost*Plus slides | Fisherbrand | 12-550-17 | |
| Table 4. Reagents and equipment for Alcian Blue stain. |
Referências
- Forester, R., Goldstein, L. Intracellular osmoregulatory role of amino acids and urea in marine elasmobranchs. Am. J. Physiol. 230, 925-931 (1976).
- Shuttleworth, T., Shuttleworth, R. . Physiology of elasmobranch fishes. , 171-194 (1988).
- Yancey, P. H., Clark, M. E., Hand, S. C., Bowlus, R. D., Somero, G. N. Living with water stress: evolution of osmolyte systems. Science. 217, 1214-1222 (1982).
- Ballatori, N., Villalobos, A. Defining the molecular and cellular basis of toxicity using comparative models. Toxicl. Appl. Pharmacol. 183, 207-220 (2002).
- Nieto, M. A., Patel, K., Wilkinson, D. G. In situ hybridization analysis of chick embryos in whole mount and tissue sections. Methods Cell Biol. 51, 219-235 (1996).
- Jass, J. R., Walsh, M. D. Altered mucin expression in the gastrointestinal tract: a review. J Cell Mol Med. 5, 327-351 (2001).
- Ballard, W. W., Mellinger, J., Lechenault, H. A series of normal stages for development of Scyliorhinus canicula, the lesser spotted dogfish (Chondrichthyes; Scyliorhinidae). J. Exp. Zool. 267, 318-336 (1993).
- Echelard, Y., et al. Sonic hedgehog, a member of a family of putative signaling molecules, is implicated in the regulation of CNS polarity. Cell. 75, 1417-1430 (1993).
- Riddle, R. D., Johnson, R. L., Laufer, E., Tabin, C. Sonic hedgehog mediates the polarizing activity of the ZPA. Cell. 75, 1401-1416 (1993).
- Theodosiou, N. A., Hall, D. A., Jowdry, A. L. Comparison of acid mucin goblet cell distribution and Hox13 expression patterns in the developing vertebrate digestive tract. J. Exp. Zool. B. Mol. Dev. Evol. 308, 442-453 (2007).
- Sambrook, J., Russell, D. W. Ch. 7. Molecular Cloning; A Laboratory Manual. 1, 7.82 (2001).
- Zhang, G., Miyamoto, M. M., Cohn, M. J. Lamprey type II collagen and Sox9 reveal an ancient origin of the vertebrate collagenous skeleton. Proc. Natl. Acad. Sci. U.S.A. 103, 3180-3185 (2006).
- Sheehan, D. C., Hrapchak, B. B. . Theory and practice of histotechnology. , (1980).
- Theodosiou, N. A., Simeone, A. Evidence of a rudimentary colon in the elasmobranch, Leucoraja erinacea. Dev. Genes Evol. 222, 237-243 (2012).
- Filipe, M. I. Mucins in the human gastrointestinal epithelium: a review. Invest. Cell Pathol. 2, 195-216 (1979).
- Corfield, A. P., Wagner, S. A., Clamp, J. R., Kriaris, M. S., Hoskins, L. C. Mucin degradation in the human colon: production of sialidase, sialate O-acetylesterase, N-acetylneuraminate lyase, arylesterase, and glycosulfatase activities by strains of fecal bacteria. Infect. Immun. 60, 3971-3978 (1992).
- Reid, B. J., et al. Flow-cytometric and histological progression to malignancy in Barrett’s esophagus: prospective endoscopic surveillance of a cohort. Gastroenterology. 102, 1212-1219 (1992).
- Mowry, R. Selective staining of pancreatic beta-cell granules. Evolution and present status. Arch. Pathol. Lab Med. 107, 464-468 (1983).
- Roberts, D. J., Smith, D. M., Goff, D. J., Tabin, C. J. Epithelial-mesenchymal signaling during the regionalization of the chick gut. Development. 125, 2791-2801 (1998).

