Organelle Transport in Cultured Drosophila Cells: S2 Cell Line and Primary Neurons.
Summary
Drosophila S2 cells and cultured neurons are great systems for imaging of motor-driven organelle transport in vivo. Here we describe detailed protocols for culturing both cell types, their imaging and analysis of transport.
Abstract
Drosophila S2 cells plated on a coverslip in the presence of any actin-depolymerizing drug form long unbranched processes filled with uniformly polarized microtubules. Organelles move along these processes by microtubule motors. Easy maintenance, high sensitivity to RNAi-mediated protein knock-down and efficient procedure for creating stable cell lines make Drosophila S2 cells an ideal model system to study cargo transport by live imaging. The results obtained with S2 cells can be further applied to a more physiologically relevant system: axonal transport in primary neurons cultured from dissociated Drosophila embryos. Cultured neurons grow long neurites filled with bundled microtubules, very similar to S2 processes. Like in S2 cells, organelles in cultured neurons can be visualized by either organelle-specific fluorescent dyes or by using fluorescent organelle markers encoded by DNA injected into early embryos or expressed in transgenic flies. Therefore, organelle transport can be easily recorded in neurons cultured on glass coverslips using living imaging. Here we describe procedures for culturing and visualizing cargo transport in Drosophila S2 cells and primary neurons. We believe that these protocols make both systems accessible for labs studying cargo transport.
Introduction
Drosophila S2 cells have gained increasing popularity for studying many cellular processes. These cells were originally obtained from trypsinized late stage embryos of OregonR Drosophila melanogaster and maintain macrophage-like features1. Drosophila S2 cells offer many advantages over other cell lines. They are cultured at 25 °C without CO2, and can be imaged for many hours at room temperature without the need of heating or gas exchange. More importantly, Drosophila S2 cells are very sensitive to RNAi treatment allowing high efficiency knockdown of proteins of interest. S2 cells are highly transfectable and stable cell lines can be selected in less than four weeks to allow stable ectopic expression of proteins of interest. S2 cells normally grow either loosely attached but not spread on a flask or in suspension. They can be induced to spread on coverslips precoated with a plant lectin concanavalin A (ConA). The periphery of the spread cells is especially thin and therefore is ideal for live cell microscopy. Detailed protocols for S2 cells culture, RNAi treatment, live light imaging and immunostaining can be found in the literature2,3. Our lab has adopted S2 cells as a model system to study microtubule-based cargo transport. Drosophila S2 cells treated with any drug that depolymerizes or severs F-actin, such as cytochalasin D (CytoD), grow long processes filled with uniformly polarized microtubules4-10, which makes it an ideal system to study cargo movement along microtubule arrays. Different parameters of transport in these processes (i.e. trajectories lengths, velocities, directionality etc.) can be easily measured using automated particle-tracking software, DiaTrack (Semasopht: http://www.microscopy.info/Organization/Details/4423; Dr. Pascal Vallotton: http://www.csiro.au/en/Organisation-Structure/Divisions/Mathematics-Informatics-and-Statistics/PascalVallotton.aspx#a4). These properties make S2 cells an appealing system for studying cargo transport along microtubules in tissue culture cells.
Pilot experiments in S2 cells can be extended to a physiologically important model of cargo transport studies in Drosophila primary neurons. Similar to S2 cells, neurons cultured from dissociated embryos are permeable to small molecules such as dyes and inhibitors, and high-resolution live imaging of organelle transport can be performed in neurons at room temperature. Using Drosophila with different genetic backgrounds, we can examine gene function in primary neurons by knockout (genetic loss-of-function mutations), knockdown (dsRNA injection or expression) or ectopic expression (transgenic flies). Specifically, fragmentation of actin filaments using CytoD treatment does not prevent neurite outgrowth; instead they grow faster, and thus we can study microtubule-dependent organelle transport in CytoD-treated neurons without the influence of actin filaments11. Therefore, primary neuronal cultures combine the advantages of tissue culture cells and fly genetics, which makes it a great system to study cargo transport in a physiological relevant system10,12. Since several familial neurodegenerative diseases could be caused by or associated with cargo transport defects (e.g. Parkinson's disease13, Huntington's disease14, Alzheimer's Disease15,16, Amyotrophic Lateral Sclerosis/ALS17, HMN7B and Perry syndrome18,19, and Spinocerebellar ataxia type 5/SCA520), primary neuronal cultures can be useful in analysis of the effects of specific mutations causing these diseases on microtubule-dependent transport.
Finally, important properties of microtubule-dependent transport are well conserved between mammals and flies, but Drosophila genetic and cell culture is less expensive and time-consuming, justifying the use of cultured Drosophila cells for studying essential aspects of organelle transport in normal and pathological conditions.
Protocol
1. Drosophila S2 cells
- Preparation of ConA coated coverslips
- Place coverslips in a ceramic rack and wash them by chromic acid immersion for 1 hr. [CAUTION: chromic acid is corrosive and can cause irritation of eyes, nose, throat, and skin].
- Rinse coverslips thoroughly with constantly running dH2O for 30 min until the acid is completely washed out.
- Let coverslips air dry and coat them with ConA (0.5 mg/ml solution in dH2O) for 30 min.
- Rinse coverslips with dH2O for 15 min and let them air dry. ConA coated coverslips can be stored up to 1 month.
- Plating the cells
- Allow S2 cells to exponentially grow in T25 or T75 cm2 flasks depending on the number of cells required for the experiment.
- Count the cell density using a hemocytometer. Add 1 ml growth medium (for example Insect-Xpress) into a 35 mm tissue culture dish with ConA-coated coverslip, and gently transfer ~1 x 105 cells to the dish. This cell density is optimal for imaging because processes from different cells do not overlap.
- In order to induce formation of processes, immediately after plating add 1ml medium with 5 µM CytoD to the dish (to the final concentration of 2.5 µM CytoD). Allow full development of processes by incubating the cells at least for 2 hr at 25 ºC before imaging.
2. Drosophila Primary Neuron Culture
- Preparation for neuron culture
- Prepare Drosophila neuronal growth medium (Schneider's medium supplemented with: 20% fetal bovine serum, heat inactivated at 55 ºC for 30 min; 5 µg/ml insulin; 100 µg/ml penicillin, 100 µg/ml streptomycin; 10 µg/ml tetracycline). This supplemented Schneider's medium can be stored at 4 ºC for 6 months.
- Prepare apple juice agar vials for collecting Drosophila embryos (22.5 g agar, 12.0 g sucrose and 750 ml H2O in a 2-liter flask; autoclave for 20-25 min; add 250 ml apple juice and pour ~10 ml into each Drosophila vial; after solidification, plug vials with cotton balls, and wrap vials with plastic food wrap to avoid over drying of the food). The apple juice in agar vials can be stored at 4 ºC in well-sealed plastic bags for 4 weeks.
- Coat acid-washed 25 mm circle coverslips with ConA (see details in the Protocol for Drosophila S2 cells) and sterilize them under UV light for 10-15 min. These coverslips can be stored for a month.
- Drosophila embryonic neuron culture
- Keep young adult flies in apple juice agar vials supplemented with dry yeast for 1-2 days before collecting embryos for neuron preparation.
- On the collecting day, in order to synchronize the collected embryos, transfer adult flies to a fresh apple juice agar vial for 1 hr precollection in light and discard these embryos.
- Transfer flies to a new apple juice agar vial, collect embryos for 1 hr, and transfer the flies to another vial for another hour, both in dark to increase the embryo yield.
- Remove the adult flies from the apple juice agar vial and let the embryos develop another 4 hr to stages 9-11 at room temperature (4-6 hr-old embryos are between stage 9-11).
- Remove the embryos by squirting water into vials and loosening the embryos from the apple juice agar with a small paintbrush (at least 10 embryos are required for one good neuron prep).
- Transfer the embryos to a 100 µm nylon mesh cell strainer with a transfer pipette to drain water (prerinse the transfer pipette in Drosophila embryo wash buffer, 0.4% NaCl, 0.03% Triton X-100).
- Collect the embryos from the strainer using a small paintbrush, and transfer them into a 1.5 ml microtube filled with 1 ml Drosophila embryo wash (Note: clean any agar or cotton fibers using the paintbrush or pipette tips). In this protocol, the 1.5 ml microtube comes with a matching disposable pellet pestle.
- Wash embryos in Drosophila embryo wash 3x.
- Dechorionate the embryos in a fresh 1:1 solution of commercial bleach and 95% ethanol for 10 min [CAUTION: bleach can cause irritation to eyes, nose, throat, and skin].
- Move to the tissue culture hood, and wash embryos 2x in supplemented Schneider's medium.
- Leave 200-300 µl medium in the tube with the embryos, and gently mechanically dissociate dechorionated embryos using Kimble Chase disposable pellet pestles that have been sterilized by 70% ethanol (grind 10-15x).
- Centrifuge the homogenized mixture at 20 x g for 2 min, and transfer the supernatant to a fresh tube. [This step is optional; it helps to remove embryo debris]
- Centrifuge the supernatant at 550 x g for 2 min to pellet cells.
- Resuspend the pellet in 500 µl supplemented Schneider's medium and spin cells down again at 550 x g for 2 min; repeat the resuspension-spinning cycle once.
- Finally resuspend the pellet in ~100 µl supplemented Schneider's medium and plate the cells onto a 25 mm ConA-coated coverslip in a sterile 35 mm Petri dish. Incubate for 10 min to allow cell attachment.
- Add 2 ml of supplemented Schneider's medium, with 5 µM CytoD to submerge the coverslip with attached cells.
- Incubate the dish in a humidified 25 ºC incubator overnight before imaging.
Representative Results
Drosophila S2 cells attached on the ConA-coated coverslip spread into a flat round “pancake” shape (Figures 1A and 1C). However, when S2 cells plated on coverslips in the presence of CytoD and allowed to spread for 2-3 hr, they form long thin processes filled by parallel microtubules (Figures 1B and 1D). These microtubules have uniform polarity with plus-ends out7. Time-lapse fluorescence imaging of Drosophila S2 stably expressing mCherry-tubulin shows that processes are very dynamic, especially during the first 60 min after attachment (Figure 2). After 1 hr incubation, their growth slows down and 2-3 hr incubation ensures that majority of cells have fully grown processes. Here we show two representative examples of organelle transport along microtubules in S2 cells, lysosomes (labeled with LysoTracker, 200 nM; Figures 3A-B) and peroxisomes (labeled with peroxisome-targeted GFP, GFP-SKL)7 (stable transfection with pAC-GFP-SKL; Figures 3D-E). Time-lapse images of the cells were taken every 1 sec for 61 frames using a Nikon TU-2000 inverted microscope equipped with a Perfect Focus system (Nikon) and Coolsnap CCD camera (Roper Scientific) driven by Metamorph software. A 100-W halogen light source was used for fluorescence excitation to minimize photobleaching and phototoxicity. Each of the 61 frames in sequence was color coded according to the bar at upper right and images were superimposed to generate color-coded tracks. Thus moving particles generated rainbow tracks, while stationary particles appeared white10. ImageJ plug-in Temporal Color-Code (http://fiji.sc/Temporal-Color_Code) was used for this processing. For quantitative analysis of organelle movement we used DiaTrack software (Semasopht Inc.), we only select organelles that are localized in processes because they can be easily tracked and the polarity of microtubule tracks is known. Figures 3C and 3F show typical distributions of retrograde and anterograde velocities obtained by lysosomes and peroxisomes tracking, respectively.
Similar techniques are used in our lab for analysis of organelle transport in primary cultured neurons. Neurons are a predominant cell type in mixed cultures obtained from stage 9-11 embryos. After overnight culturing they are easy to recognize by characteristic cell shape with neurites that are over 50 µm long (Figure 4A). These cells are also positive for the pan-neuronal marker, Elav21 (Figures 4B and 4B'). Processes in the neurons are filled with microtubules (Figures 4B and 4B"). We can track organelle transport along the neurites using the techniques described above for S2 cells. Mitochondria can be labeled with mitochondrial GFP (Mito-GFP) under control of a motor neuron-specific promoter D4222 (Figures 5A and 5A'), and peroxisomes can be marked by injection of DNA encoding a peroxisome-targeted mCherry7, into syncytial blastoderm stage embryos (Figures 5B and 5B').
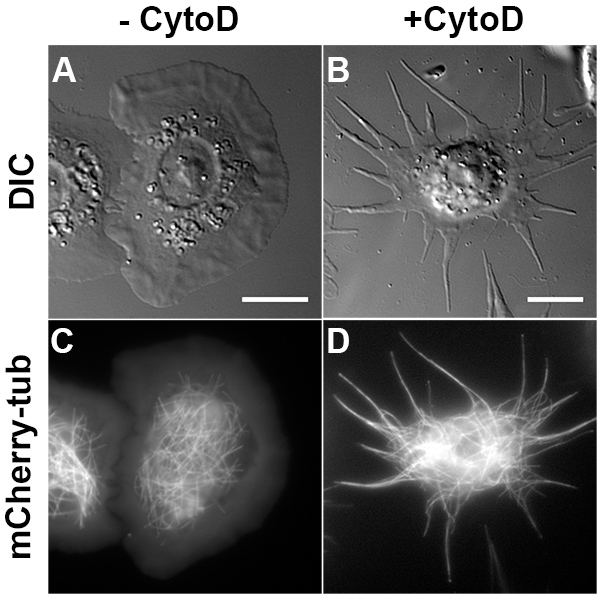
Figure 1. Drosophila S2 process formation induced by CytoD. (A, C) Drosophila S2 cells attach and spread into flatten and round shapes when plated in ConA-precoated coverslips. (B, D) S2 cells plated in the presence of 2.5 µm CytoD form long processes filled with microtubules after 3 hr incubation. A-B and C-D are transmitted light and fluorescent mCherry-tagged tubulin images, respectively. Scale bar, 10 µm. Click here to view larger image.

Figure 2. Time lapse of processes growth in a Drosophila S2 cell. Fluorescence time-lapse of a Drosophila S2 cell expressing mCherry-tubulin in the presence of 2.5 µM CytoD. The numbers indicate the time after the cell attach to the coverslip. Scale bar, 5 µm. Click here to view larger image.
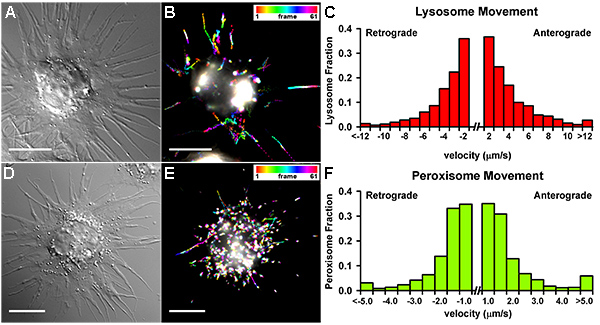
Figure 3. Organelle transport in Drosophila S2 cells. (A-C) Lysosome movement in Drosophila S2 cell. A representative 1 min-lysosome transport (labeled with 200 nM Lysotracker) in one cell (A, DIC image) represented by the artificial Temporal-Code (created by FIJI) (B). Velocities obtained from the analysis of lysosome movement tracked in processes (DiaTrack; 663 particles in 23 cells) (C). (D-F) Peroxisome movement in Drosophila S2 cell. A representative 1 min-peroxisome transport (labeled with pMT-GFP-SKL) in one cell (D, DIC image) represented by the artificial Temporal-Code (created by FIJI) (E). Velocities obtained from the analysis of peroxisome movement tracked in processes (DiaTrack; 228 particles in 20 cells) (F). Note that lysosomes move with longer and faster trajectories than peroxisomes. Scale bar, 5 µm. Click here to view larger image.
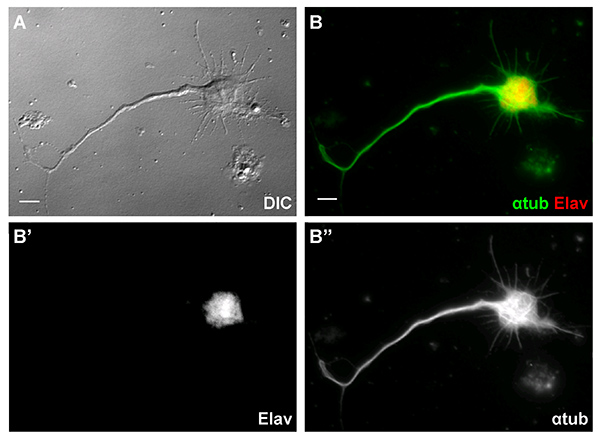
Figure 4. Primary Drosophila neurons from overnight culture. (A) An overnight cultured Drosophila neuron has characteristic neuronal morphology with long neurites. (B) The neuron expressed a pan-neuronal marker, Elav (red in B and white in B', DSHB, 7E8A10, 1:100) and with long neurites filled with microtubules (green in B and white in B'', DM1α, 1:1000). Scale Bar, 5 µm. Click here to view larger image.
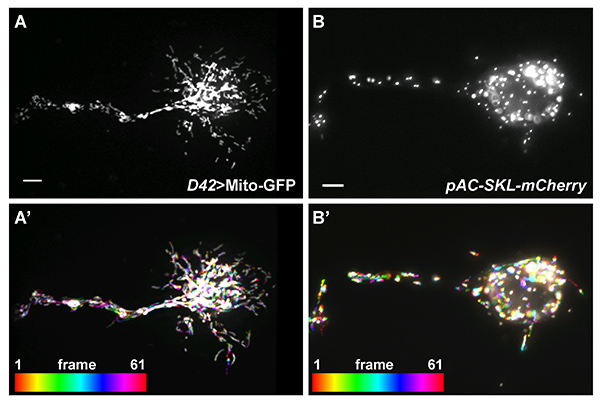
Figure 5. Organelle transport in cultured Drosophila neurons. (A) Mitochondria in a cultured Drosophila neuron marked by Mito-GFP under control of a motor neuron-specific promoter, D42. (A') 5 min-mitochondrial movement in (A) is represented by the artificial Temporal-Color code (created by FIJI). (B) Peroxisomes in a cultured Drosophila neuron marked by pAC-SKL-mCherry, injected into early syncytial blastoderm stage embryos. (B') 2 min-peroxisome movement in (B) is represented by the artificial Temporal-Color code (created by FIJI). Scale bar, 5 µm. Click here to view larger image.
| Organelle | Description |
| Peroxisomes | Peroxisomal targeting signal 1 tripeptide (SKL) tagged with a FP25 |
| Mitochondria | Mitochondrial targeting sequence of cytochrome c oxidase subunit VIII tagged with a FP26 |
| Mitochondria | Mitotracker (Dye which accumulates in active mitochondria)27 |
| Lysosomes | Lysotracker (Dye which accumulates in cellular compartments with low internal pH)28 |
Table 1. Most common organelle labeling strategies.
This table summarizes the most common organelle labeling strategies in tissue culture cells, including labeling of peroxisomes, mitochondria, and lysosomes.
Discussion
Here we present detailed protocols for studying microtubule-dependent cargo transport in Drosophila S2 cells and primary neuron culture. Both S2 processes and neurites are structures filled by bundled microtubules arrays that serve as tracks for organelle transport.
Two strategies are typically used to label organelles: transfection of vectors encoding a fluorescent protein with an organelle-targeting sequence or staining with fluorescent dyes added directly to cultures. Common examples of cargo labels are listed in Table 1. Transient transfection or RNAi treatments of Drosophila S2 cells if necessary is performed in suspension4,6,7,9,10 before cells are plated on coverslips for imaging.
In order to get reliable results in S2 cell experiments, it is important to keep the following points in mind:
- Plate cells at optimal density for imaging. High density will result in overlapping of processes from different cells, while at low density cells have poor viability and are unable to form processes.
- Allow S2 cells to completely develop processes. We routinely incubate cells for 2 hr after plating but incubation up to 24 hr can be performed if necessary.
- S2 cells must be treated with CytoD before they attach to ConA coated coverslips. Adding the drug after cells are attached will not induce processes formation.
- When using dyes, change samples for imaging every 20-25 min; long-term incubation of cells with either Mitotracker or Lysotracker is toxic to the cells.
- We usually track cargo transport in S2 cells by imaging for 1 min at 1 frame/sec. Under this condition cells are not photo-damaged.
- Only analyze organelle movement in cell processes. Organelle density in the cell body is too high for reliable tracking and polarity of the movement could not be easily determined.
The Drosophila primary neuron culture preparation procedure was modified from the protocols by the Mahowald lab23 and Lieu lab24. This protocol allows you to obtain primary neuron culture within a day and it is ready for imaging and tracking the next day.
In this protocol, you may need to pay attentions to several key steps listed below:
- Stage embryos correctly. The collected embryos need to be synchronized and staged to stages 9-11, when neuroblast delamination from the ectoderm occurs. Staging embryos too early or too late may result in fewer neuroblasts in each embryo and thus failure of harvesting neurons out of the dissociated embryos.
- Dechorionate embryos to the right extent (all embryos change color from completely white to more transparent). Under-dechorionation could lead to difficulty in embryo dissociation, while over-dechorionation could damage the embryos. Dechorionated embryos become very sticky and soft, thus try to avoid pipetting or touching the embryos with pipette tips during the washing steps before dissociation.
- Pay special attention to embryos dissociation. We find that plastic disposable pellet pestles and matching microtubes give better dissociation than a glass dounce homogenizer, especially for small number of embryos. Under-dissociation will leave large cell clusters, while over-dissociation will lead to cells breakdown and massive accumulation of cell debris. Adjust the dissociation level according to the embryo numbers.
- Plating neuron on the glass coverslip to a reasonable density. Crowded neurons usually develop very short neurites, while sparse neurons often fail to survive.
Additionally, we noticed that primary cell cultures often contain muscle cells, and blood cells. Overnight treatment with CytoD helps to reduce the number of muscle cell and blood cell, and promotes neurite outgrowth.
Declarações
The authors have nothing to disclose.
Acknowledgements
We thank the members of the Gelfand lab who contributed to the development of these protocols. Research reported in this publication was supported by the National Institute of General Medical Science of the National Institutes of Health under award number R01GM052111 to VIG and by Basque Government Department of Education, Universities and Research under award number BFI-2011-295 to UdC.
Materials
| Powder Insect cell medium (Schneider’s) | Sigma | S9895 |
| Insect –Xpress medium | Fisher | BW12730Q |
| Insulin | Sigma | I6634 |
| Penicillin | Research Products International | P93000 |
| Streptomycin | Research Products International | S62000 |
| Tetracycline hydrochloride | Sigma | T7660-5G |
| Fetal bovine serum | Atlanta Biologicals | S11150 |
| Concanavalin A | Sigma | C2010 |
| Cytochalasin D | VWR | IC15077105 |
| LysoTracker Red DND-99 | Molecular Probes | L7528 |
| 100 µm Cell strainer | Fisher Scientific | 22363549 |
| 25 mm Circle cover glass | VWR | 48380 080 |
| Drosophila Vials | VWR | 89092-730 |
| Disposable pellet pestles with matching microtubes | Kimble Chase | 749520-0000 |
Referências
- Schneider, I. Cell lines derived from late embryonic stages of Drosophila melanogaster. J. Embryol. Exp. Morphol. 27, 353-365 (1972).
- Rogers, S. L., Rogers, G. C. Culture of Drosophila S2 cells and their use for RNAi-mediated loss-of-function studies and immunofluorescence. 3, 606-611 (2008).
- Buster, D. W., Nye, J., Klebba, J. E., Rogers, G. C. Preparation of Drosophila S2 cells for light microscopy. J. Vis. Exp. (40), e1982 (2010).
- Ling, S. C., Fahrner, P. S., Greenough, W. T., Gelfand, V. I. Transport of Drosophila fragile X mental retardation protein-containing ribonucleoprotein granules by kinesin-1 and cytoplasmic dynein. Proc. Natl. Acad. Sci. U.S.A. 101, 17428-17433 (2004).
- Kural, C., et al. Kinesin and dynein move a peroxisome in vivo: a tug-of-war or coordinated movement. Science. 308, 1469-1472 (2005).
- Kim, H., et al. Microtubule binding by dynactin is required for microtubule organization but not cargo transport. J. Cell. Biol. 176, 641-651 (2007).
- Ally, S., Larson, A. G., Barlan, K., Rice, S. E., Gelfand, V. I. Opposite-polarity motors activate one another to trigger cargo transport in live cells. J. Cell. Biol. 187, 1071-1082 (2009).
- Bensenor, L. B., Barlan, K., Rice, S. E., Fehon, R. G., Gelfand, V. I. Microtubule-mediated transport of the tumor-suppressor protein Merlin and its mutants. Proc. Natl. Acad. Sci. U.S.A. 107, 7311-7316 (2010).
- Jolly, A. L., et al. Kinesin-1 heavy chain mediates microtubule sliding to drive changes in cell shape. Proc. Natl. Acad. Sci. U.S.A. 107, 12151-12156 (2010).
- Barlan, K., Lu, W., Gelfand, V. I. The microtubule-binding protein ensconsin is an essential cofactor of Kinesin-1. Curr. Biol. 23, 317-322 (2013).
- Lu, W., Fox, P., Lakonishok, M., Davidson, M. W., Gelfand, V. I. Initial Neurite Outgrowth in Drosophila Neurons Is Driven by Kinesin-Powered Microtubule Sliding. Curr. Biol. 23, 1018-1023 (2013).
- Pathak, D., Sepp, K. J., Hollenbeck, P. J. Evidence that myosin activity opposes microtubule-based axonal transport of mitochondria. J. Neurosci. 30, 8984-8992 (2010).
- Clark, I. E., et al. Drosophila pink1 is required for mitochondrial function and interacts genetically with parkin. Nature. 441, 1162-1166 (2006).
- Gunawardena, S., et al. Disruption of axonal transport by loss of huntingtin or expression of pathogenic polyQ proteins in Drosophila. Neuron. 40, 25-40 (2003).
- Koo, E. H., et al. Precursor of amyloid protein in Alzheimer disease undergoes fast anterograde axonal transport. Proc. Natl. Acad. Sci. U.S.A. 87, 1561-1565 (1990).
- Stokin, G. B., Goldstein, L. S. Axonal transport and Alzheimer’s disease. Annu. Rev. Biochem. 75, 607-627 (2006).
- Bilsland, L. G., et al. Deficits in axonal transport precede ALS symptoms in vivo. Proc. Natl. Acad. Sci. U.S.A. 107, 20523-20528 (2010).
- Lloyd, T. E., et al. The p150(Glued) CAP-Gly domain regulates initiation of retrograde transport at synaptic termini. Neuron. 74, 344-360 (2012).
- Moughamian, A. J., Holzbaur, E. L. Dynactin is required for transport initiation from the distal axon. Neuron. 74, 331-343 (2012).
- Lorenzo, D. N., et al. Spectrin mutations that cause spinocerebellar ataxia type 5 impair axonal transport and induce neurodegeneration in Drosophila. J. Cell. Biol. 189, 143-158 (2010).
- Robinow, S., White, K. The locus elav of Drosophila melanogaster is expressed in neurons at all developmental stages. Dev. Biol. 126, 294-303 (1988).
- Pilling, A. D., Horiuchi, D., Lively, C. M., Saxton, W. M. Kinesin-1 and Dynein are the primary motors for fast transport of mitochondria in Drosophila motor axons. Mol. Biol. Cell. 17, 2057-2068 (2006).
- Furst, A., Mahowald, A. P. Differentiation of primary embryonic neuroblasts in purified neural cell cultures from Drosophila. Dev. Biol. 109, 184-192 (1985).
- Salvaterra, P. M., Bournias-Vardiabasis, N., Nair, T., Hou, G., Lieu, C. In vitro neuronal differentiation of Drosophila embryo cells. J. Neurosci. 7, 10-22 (1987).
- Wiemer, E. A., Wenzel, T., Deerinck, T. J., Ellisman, M. H., Subramani, S. Visualization of the peroxisomal compartment in living mammalian cells: dynamic behavior and association with microtubules. J. Cell. Biol. 136, 71-80 (1997).
- Rizzuto, R., et al. A gene specifying subunit VIII of human cytochrome c oxidase is localized to chromosome 11 and is expressed in both muscle and non-muscle tissues. J. Biol. Chem. 264, 10595-10600 (1989).
- Poot, M., et al. Analysis of mitochondrial morphology and function with novel fixable fluorescent stains. J. Histochem. Cytochem. 44, 1363-1372 (1996).
- Haller, T., Dietl, P., Deetjen, P., Volkl, H. The lysosomal compartment as intracellular calcium store in MDCK cells: a possible involvement in InsP3-mediated Ca2+ release. Cell Calcium. 19, 157-165 (1996).

