Aplysia Ganglia Preparation for Electrophysiological and Molecular Analyses of Single Neurons
Summary
Marine snail Aplysia californica has been widely used as a neurobiology model for the studies on cellular and molecular basis of behavior. Here a methodology is described for exploring the nervous system of Aplysia for the electrophysiological and molecular analyses of single neurons of identified neural circuitry.
Abstract
A major challenge in neurobiology is to understand the molecular underpinnings of neural circuitry that govern a specific behavior. Once the specific molecular mechanisms are identified, new therapeutic strategies can be developed to treat abnormalities in specific behaviors caused by degenerative diseases or aging of the nervous system. The marine snail Aplysia californica is well suited for the investigations of cellular and molecular basis of behavior because neural circuitry underlying a specific behavior could be easily determined and the individual components of the circuitry could be easily manipulated. These advantages of Aplysia have led to several fundamental discoveries of neurobiology of learning and memory. Here we describe a preparation of the Aplysia nervous system for the electrophysiological and molecular analyses of individual neurons. Briefly, ganglion dissected from the nervous system is exposed to protease to remove the ganglion sheath such that neurons are exposed but retain neuronal activity as in the intact animal. This preparation is used to carry out electrophysiological measurements of single or multiple neurons. Importantly, following the recording using a simple methodology, the neurons could be isolated directly from the ganglia for gene expression analysis. These protocols were used to carry out simultaneous electrophysiological recordings from L7 and R15 neurons, study their response to acetylcholine and quantitating expression of CREB1 gene in isolated single L7, L11, R15, and R2 neurons of Aplysia.
Introduction
The human brain is extraordinarily complex with almost 100 billion neurons and trillions of synaptic connections. There are almost an equal number of nonneuronal cells that interact with neurons and regulate their function in the brain. Neurons are organized into circuits that regulate specific behaviors. Despite the advances in our understanding of brain functions and neural circuits, little is known about the identity of circuitry components that control a specific behavior. Knowledge of the identities of various components of a circuitry will greatly facilitate our understanding of both cellular and molecular basis of behavior and aid in developing novel therapeutic strategies for neuropsychiatric disorders.
The marine snail Aplysia californica has been a workhorse for determining neuronal circuits underlying specific behaviors1-14. The Aplysia nervous system contains approximately 20,000 neurons that are organized into 9 different ganglia. The neurons of Aplysia are large and can be easily identified based on their size, electrical properties, and position in the ganglia. Aplysia has a rich repertoire of behaviors that can be studied. One of the well-studied behaviors is the gill-withdrawal reflex (GWR). The central components of this reflex are situated in abdominal ganglia. Components of the GWR circuitry have been mapped and contributions of various components determined. Importantly, GWR circuitry undergoes associative and nonassociative learning5,6,15-19. Decades of study on this reflex have also identified several signaling pathways that have a key role in learning and memory20-24.
Several different preparations of Aplysia were used to study cellular and molecular basis of memory storage. These include the intact animal2,3, semi-intact preparation1,7,13,14,16 and reconstitution of major components of neural circuitry25-29. A reduced preparation for exploring Aplysia ganglia for the electrophysiological and molecular analyses of identified neuronal circuits is described here. The following four identified neurons were studied. R15, a bursting neuron, L7 and L11, two different motor neurons and R2, a cholinergic neuron were studied. R2 is the largest neuron described in the invertebrate nervous system. Briefly, this methodology involves protease treatment of ganglia, electrophysiological measurements before and after pharmacological treatments, and isolation of single neurons for quantitative analysis of gene expression. This methodology enables us to combine molecular analyses with simultaneous recording from multiple neurons. This methodology was successfully used to study responses of R15 and L7 neurons to acetylcholine (Ach) by paired intracellular recordings. Following electrophysiological measurements R15 and L7 and other identified neurons such as L11 and R2 were isolated for quantitative polymerase chain reaction (qPCR) analysis of expression of CREB1, a transcription factor important for memory storage.
Protocol
1. Preparation of Abdominal Ganglia, Electrophysiological Measurements, and Isolation of Single Identified Neurons from Abdominal Ganglion of Aplysia californica
- Maintain Aplysia in the laboratory aquarium with circulating artificial seawater (ASW) at 16 °C under 12:12 light:dark conditions.
- Isolation of abdominal ganglion.
- Anesthetize animals by injecting 380 mM MgCl2 solution for 5-10 min (equivalent to 30-35% of the animal's body weight).
- Identify the abdominal ganglion based on its position in the central nervous system24,28.
- Remove the ganglia by surgical operation and immediately store in artificial seawater consisting of 450 mM NaCl, 10 mM KCl, 10 mM CaCl2, 55 mM MgCl2, 2.5 mM NaHCO3, and 10 mM HEPES (pH 7.4).
- Protease treatment.
- Incubate abdominal ganglion for 30 min at 34 °C±0.5 in a Petri dish with 0.1% protease (dispase) diluted in ASW described above.
NOTE: The amount of the protease used needs to be standardized with the age and weight of the animals. In general, older and bigger animals would require more protease.
- Incubate abdominal ganglion for 30 min at 34 °C±0.5 in a Petri dish with 0.1% protease (dispase) diluted in ASW described above.
- Removal of ganglia sheath.
- After the enzyme treatment, pin down the ganglia to the Sylgard Silicone base of a cell chamber (Ø= 1 cm; vol. = 0.3 ml) and perfuse with ASW (flow rate: 150 μl/min) at room temperature (18 °C±2 ).
NOTE: In order to increase the visibility of identified neurons, place the ganglion on the top of a polycarbonate glass cylinder (Figure 1) (diameter Ø= 2 mm modified cell chamber) that can be easily illuminated from the bottom using a binocular stereomicroscope with 20X and 40X magnification. - Carefully remove the sheath from the ganglion by using forceps and microscissors.
- After the enzyme treatment, pin down the ganglia to the Sylgard Silicone base of a cell chamber (Ø= 1 cm; vol. = 0.3 ml) and perfuse with ASW (flow rate: 150 μl/min) at room temperature (18 °C±2 ).
- Simultaneous electrophysiological recording from two neurons.
- Identify neuron of interest.
- Impale the neuron with a single intracellular sharp microelectrode with resistance 10-15 MΩ filled with 3 M KCl solution.
- Record neuronal activity (10 kHz band pass) using an intracellular recording system.
- Process the recording data using electrophysiology software such as Axograph.
NOTE: One can easily carry out recording from multiple neurons. Neurons L7, L11 or R15 and R2 are easily identified based on their position. Two amplifiers and two micromanipulators are required for simultaneously recording from two L7 and R15 neurons (Figure 2).
- Drug application.
- Apply drug of choice into the cell chamber by gentle pipetting or inject directly into neurons.
- Carry out electrophysiological measurements.
NOTE: Depending on the experimental objective, one could measure different electrophysiological parameters of neurons such as the changes in membrane potential (MP) or action potential (AP) before, during and after the drug application.
- Isolation of single neurons.
- At the conclusion of the electrophysiological experiments, stop the perfusion of ASW and wash the ganglion with 100% ethanol. Exposure to ethanol will petrify the neurons and, using fine forceps, make it easy to remove individual neurons without damaging other neurons from the ganglion.
- Remove single neurons under a stereomicroscope and transfer to a small Eppendorf tube containing ice cold 500 μl RNA extraction reagent. Isolated single neurons can be stored in RNA extraction reagent at -80 °C for future analysis.
2. RNA Isolation from Single Neurons and Gene Expression Analysis by Quantitative Real Time PCR (qPCR)
- Precautions to minimize RNA degradation:
Ribonucleases (RNases) degrade RNA and are very stable and active enzymes. Take the following steps during handling RNAs.- Clean the working area and instruments with RNase deactivating agents.
- Wear gloves at all times while handling RNA and change gloves often.
- Use only RNase free plastics to handle RNAs.
- Isolating Total RNA from Single neurons:
- Standard Trizol protocol for RNA isolation can be used to isolate RNAs from single neurons. Briefly add 20% v/v of chloroform to Trizol, mix well by brief vortexing.
- Place the tubes on rotator for 10 min at 4 °C. Spin the Trizol-Chloroform mix at 12,000 x g for 15 min at 4 °C.
- Collect the aqueous phase and transfer to a fresh tube.
- Add sodium acetate solution (pH 5.5) to a final concentration of 0.3 M, 100 ng/ml of the coprecipitant GlycoBlue, and 2.5 volumes of 100% ethanol to precipitate RNA.
- Mix the samples well and incubate at -80 °C overnight.
- Spin the samples at 12,000 x g for 20 min at 4 °C and a small blue pellet will be visible at the bottom of the tube.
- Remove the supernatant and wash the pellet with 850 μl of 75% ice-cold ethanol at 12,000 x g for 10 min at 4 °C.
- Remove the supernatant carefully and air-dry the RNA pellet for 7-10 min, maximum.
NOTE: Make sure that RNA is not left for long to dry as over–drying the RNA pellet makes solubilization very difficult. - Once dried, resuspend the RNA pellet in 10 μl of RNAse free water and determine RNA concentration.
NOTE: Determine RNA concentration using a spectrophotometer. When not using right away, store RNA at -80 °C.
- aRNA synthesis from single neurons:
- Amplify the RNA (one or two rounds depending on experimental objective) using commercially available linear RNA amplification system.
NOTE: The total RNA from single neurons ranged from ~10-60 ng. Expected yield of first round of amplification is ~100-300 ng and the second round of amplification is ~0.8-1.5 μg of RNA.
- Amplify the RNA (one or two rounds depending on experimental objective) using commercially available linear RNA amplification system.
- Reverse transcription of RNA:
- To generate cDNA from aRNA, use 1.0 μg of aRNA in 20 μl of a reverse transcription reaction. Several kits are commercially available for this purpose.
NOTE: cDNA was synthesized according to the manufacturer's protocol using the following settings on thermal cycler.
5 min at 25 °C
30 min at 42 °C
5 min at 85 °C
Hold at 4 °C - After the reverse transcription step, store the cDNA at -20 °C for long-term storage.
- To generate cDNA from aRNA, use 1.0 μg of aRNA in 20 μl of a reverse transcription reaction. Several kits are commercially available for this purpose.
- Primer design and standardization:
Several excellent software programs are available, free of charge, for designing primers for real time amplifications.- Select 18-24 mer oligonucleotides that produce 70-110 bp amplicons and synthesize the primers using commercial sources.
NOTE: Two to three primer pairs should be designed for each gene of interest. Each primer set needs to standardized by qPCR. The forward and reverse primer pairs that were used for gene CREB1 (Figure 4) are below:
Primer set 1
Forward: 5'-TGATTCTGACGCAAAGAAAAGA-3'
Reverse: 5'-ACCGTGAGCAGTCAGTTGTAGA-3'
Primer set 2
Forward: 5'-AGGGAATGTCGATGAAAGAAGA-3'
Reverse: 5'-GACACACAGGAAGTATGCCAAA-3' - To select the best primer to be used for the downstream analyses, monitor Ct value and the shape of the amplification curve in qPCR.
NOTE: Primers that give Ct (Cycle threshold) values between 10-35 in the 40 round amplification and smooth curves during the exponential phase of the PCR followed by a smooth plateau are optimal for gene expression analyses.
- Select 18-24 mer oligonucleotides that produce 70-110 bp amplicons and synthesize the primers using commercial sources.
- Quantitative Real Time PCR (qPCR):
Note: Use appropriate tubes or plates that are compatible with the qPCR machine.- Dilute cDNA to 5x with nuclease free water.
- To 2 μl of diluted cDNA, add 8 μl of a qPCR master mix containing 2 μl of H2O, 5 μl of 2x SYBR Green master mix, and 1.0 μl of 10 μM (each) forward and reverse primer.
- Close the tubes and seal the plate after pipetting cDNA and master mix. Mix the contents by gentle tapping and spinning down at 1,500 rpm for 30 sec.
- Proceed to setting up the qPCR machine.
Representative Results
The weights of animals that were used in this study ranged from 100-200 g. Following the described protocols, we conducted electrophysiological measurements and molecular analysis of neurons of abdominal ganglia isolated from animals ranging from 2-5 g to 200-300 g.
Standardization of protease treatment is important for successful electrophysiological measurements of neurons in the ganglia. Initially, multiple protease (Dispase) concentrations and durations were used and bursting action potentials from R15 neuron were recorded26,31. These measurements were compared to direct (without protease treatment) recording from the intact ganglia. Concentration of the protease (0.1% protease, see step 1.3) that produced bursting potentials that were comparable to those obtained from exposed ganglion without protease were used in the subsequent experiments.
Using the protocol described, action potentials (AP) in response to Ach exposure to R15, a cholinergic neuron and L7, a motor neuron that does not respond to Ach were recorded. As expected, simultaneous electrophysiological recordings show that Ach induce depolarization in R15 whereas there was no change in firing of L7. Figure 2 shows the application of Acetylcholine (Ach) directly into the cell chamber. 1 M Ach (volume 0.3 μl) was directly applied into the chamber using micropipette in order to obtain a final 1 mM concentration inside the chamber.
Ach was washed off from the bath by perfusion and recorded again from R15 neurons. Figure 3 shows waveforms suggesting that Ach effect is reversible. Analysis of the parameters AP in control, during depolarization and after wash is shown. During depolarization amplitude is decreased and duration of AP increased significantly.
Following electrophysiological measurements, R15 and L7 neurons were isolated and RNAs were prepared for qPCR analyses. A linear T7 RNA Polymerase—driven transcription using MessageAmp II aRNA Amplification Kit from Ambion was used for amplifying RNAs from single neurons. For comparison, identified neurons R2 and L11 and the whole abdominal ganglia from adult (6-9 months old) and young animals (1-2 months old) were also studied. qPCR amplification were carried out in a 7900HT Fast Real-Time PCR System under the following conditions: 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 sec, 60 °C for 1 min. There were four biological replicates in the experiment. Each biological replicate had four technical replicates in the qPCR set up. Ct values from four replicates were averaged and 1/Ct was calculated to find the absolute expression level of CREB1 in each sample (Figure 4). An estimation of the absolute expression levels of CREB1 gene in young and adult Aplysia ganglia, and in four identified single neurons was monitored by the Ct value obtained from qPCR measurements.
The starting mRNA concentration was the same in both young and adult abdominal ganglia. The same forward and reverse CREB1 primer set were used in all qPCRs, both in studies using ganglia and single neurons. The Ct values are very similar between young and adult abdominal ganglia. Similarly, Ct values were measured in the case of single neurons where the starting total mRNA concentrations were low and two rounds of amplification yielded a good amount of RNAs for cDNA synthesis. The Ct values of CREB1 amplification suggested that CREB1 expression in R15 and L7 neurons are comparable whereas R2 and L11 showed significant differences in expression when compared to L7 or R15 (Student's T test, p<0.05) (Figure 4).
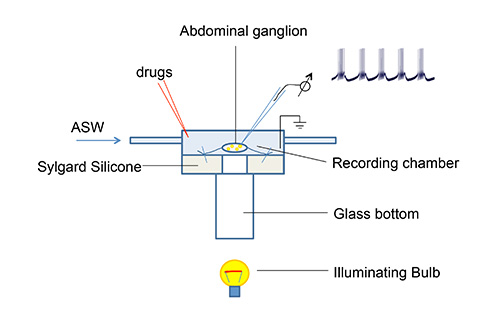
Figure 1. Microscope stage and perfusion system for ganglia electrophysiology. Perfusion system and stage were made using Plexiglas and are transparent. This setup is mounted on to a metal block (not shown) to hold position. The metal block is designed such that samples can be illuminated from the bottom. A sample electrophysiological bursting pattern recorded from R15 neuron is shown.
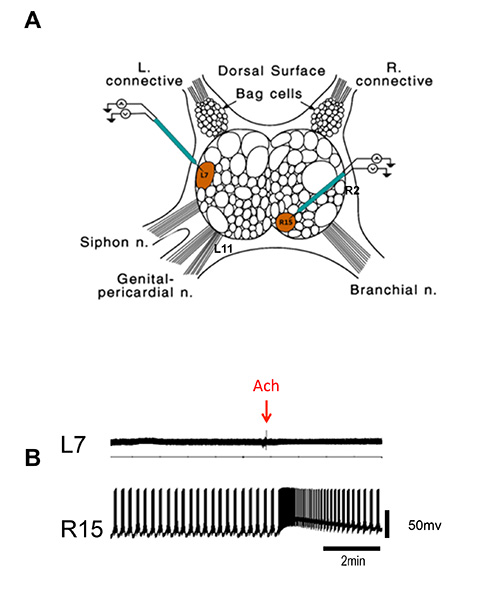
Figure 2. Simultaneous recording from L7 and R15 neuron of abdominal ganglia. (A) Cartoon of abdominal ganglia showing the position of L7 and R15 neurons (modified from Kandel 2001, reprinted with permission from AAAS). Also shown are the positions of R2 and L11 in the abdominal ganglia. (B) Representative intracellular recordings from L7 and R15. Arrow indicates time of Ach application. One could continue these measurements for 8-10 hr.
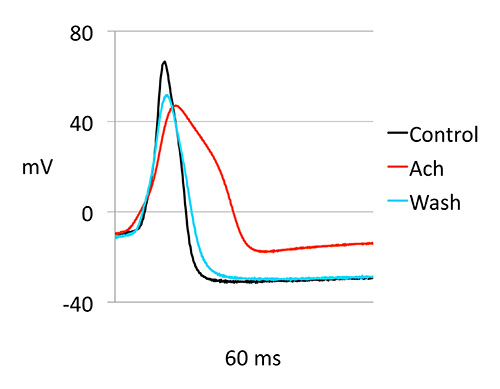
Figure 3. Ach induced changes in action potential (AP) of R15. AP waveforms were analyzed before, during and after Ach exposure. Representative waveforms are shown. ms: millisecond, mV: millivolt.
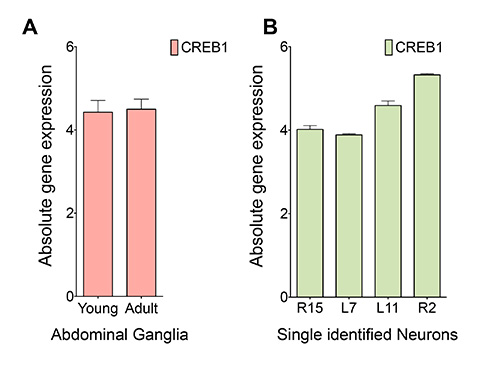
Figure 4. qPCR analysis of expression of CREB1 in abdominal ganglia and single identified neurons. (A) Absolute quantification of CREB1 transcript in the young and adult abdominal ganglia of Aplysia. Total RNA was isolated using standard Trizol protocol as described in the methods. 1 μg of total RNA was used to synthesize cDNA, followed by real time PCR amplification using CREB1 primers. (B) Absolute quantification of CREB1 transcript in identified single neurons of old Aplysia. Total RNA was subjected to two rounds of in vitro amplification before cDNA synthesis. 1 μg of aRNA was used to synthesize cDNA, followed by real time PCR amplification using CREB1 primers.
Discussion
The neuron R15 is involved in regulating cardiovascular, digestive, respiratory, and reproductive systems30. A regularly rhythmic bursting activity of the AP is a feature of R15. As shown in the results section, paired recording of R15 and L7 show that the ganglia preparation has preserved activity of R15 neurons. R15 and L7 neurons responded appropriately to Ach. This ganglia preparation could be maintained up to 8-10 hr and electrophysiological activity could be continuously monitored. Thus, one could study the electrophysiological effects of brief neurotransmitter exposure (exposed locally to a specific neuron or to the bath) for a long period of time. Furthermore, one could study the effects of exposure to multiple neurotransmitters (simultaneously or in tandem) on the same neuron on firing properties of itself or the follower neuron or both.
An example of gene expression measurements by qPCR is shown in Figure 4. Since the RNA content of a single neuron is low, in vitro amplification is necessary to obtain a sufficient amount of RNA for quantitating expression of multiple genes. Commercial kits are available for the in vitro transcription process. The amplification of the CREB1 transcript was quantified directly by the number of cycles required to obtain a standardized fluorescence signal, the Ct-method31. The amplified RNA from a single neuron is sufficient for the studies to determine gene expression changes in a large number of genes using microarray32 and for the entire transcriptome by RNAseq analyses33.
A limitation of this technique is that this preparation may not preserve all the synaptic connections of the neurons of the abdominal ganglia. Several of the neurons form synapses with neurons in other ganglia and in the ganglia preparation these long-distance connections will be missed. However, the methodology described here could be easily modified to include the entire CNS or could be easily integrated to semi-intact preparation13,14,16 that was described to study cellular and electrophysiological changes in gill and siphon withdrawal reflex.
Recent advances in genomics techniques now allow us to obtain a deep insight into the dynamics of expression of coding and noncoding RNAs34. The reduced preparation that is described here is ideal for studying temporal changes of gene expression in conjunction with electrophysiological measurements. For example, one could apply pharmacological agents and measure their effects on electrical properties of individual neurons or multiple neurons that are part of the same neural circuitry and isolate individual neurons at different times for RNA isolation. The effect of manipulation of a single neuron (such as electrical stimulation, or exposures to pharmacological agents) on the entire identified circuitry could be studied at the level of changes in expression of specific genes in single neurons. Thus this methodology facilitates successful integration of electrophysiological and genomic methodologies to study neural circuitry in Aplysia.
This approach could be easily extended to the study of other invertebrate models such as nudibranch mollusc Tritonia, pond snail Lymnaea, and terrestrial snail Helix whose nervous systems contain several identified neurons35-57. These models are also used extensively for the study of neural control of behaviors such as escape swimming, magnetic field orientation, synapse formation, learning and memory storage. The methodology we described is a simple integration of genomics analysis with physiological measurements of neuronal activity at single cell level. A deeper insight into molecular and physiological basis of neural control of behavior could be obtained from such studies.
Declarações
The authors have nothing to disclose.
Acknowledgements
We sincerely thank the Whitehall Foundation for their funding support and startup funds from The Scripps Research Institute for carrying out this work.
Materials
| Aplysia | National Aplysia Resource Facility, University of Miami | |
| NaCl | SIGMA | S 3014-1KG |
| KCl | SIGMA | P 9333-500G |
| CaCl2•2H2O | SIGMA | C5080- 500G |
| MgCl2•6H2O | Fisher Scientific | BP 214-501 |
| NaHCO4 | SIGMA | S 6297-250G |
| HEPES | SIGMA | H 3375-500G |
| Protease | GIBCO | 17105-042 |
| Trizol | Ambion | 15596-026 |
| Chloroform | MP Biomedicals | 2194002 |
| 100% Ethanol | ACROS | 64-17-5 |
| GlycoBlue | Ambion | AM9515 |
| 3 M NaOAc, pH 5.5 | Ambion | AM9740 |
| Nuclease free water | Ambion | AM9737 |
| MessageAmp II aRNA Amplification Kit | Ambion | AM1751 |
| qScript cDNA SuperMix | Quanta Biosciences | 95048-100 |
| Power SYBR Green PCR Master Mix | Applied Biosystems | 4367659 |
| Forceps | Fine Science Tools | 11252-20 |
| Scissors | Fine Science Tools | 15000-08 |
| Stainless Steel Minutien Pins | Fine Science Tools | 26002-10 or |
| 26002-20 | ||
| Veriti Thermal Cycler | Applied Biosystems | Veriti Thermal Cycler |
| 5430R Centrifuge | Eppendorf | 5430R Centrifuge |
| 7900HT Fast Real-Time PCR | Applied Biosystems | 7900HT Fast Real-Time PCR |
| Amplifier | BRAMP-01R | NPI Electronics |
| Digidata Converter | Instrutech ITC-18 | HEKA ELEKTRONIK |
| Micro Manipulator | Patch Star | Scientifica |
Referências
- Cleary, L. J., Byrne, J. H., Frost, W. N. Role of interneurons in defensive withdrawal reflexes in Aplysia. Learn. Mem. 2, 133-151 (1995).
- Elliott, C. J., Susswein, A. J. Comparative neuroethology of feeding control in molluscs. The J. Exp. Biol. 205, 877-896 (2002).
- Nargeot, R., Simmers, J. Functional organization and adaptability of a decision-making network in Aplysia. Front. Neurosci. 6, 113 (2012).
- Baxter, D. A., Byrne, J. H. Feeding behavior of Aplysia: a model system for comparing cellular mechanisms of classical and operant conditioning. Learn. Mem. 13, 669-680 (2006).
- Castellucci, V., Pinsker, H., Kupfermann, I., Kandel, E. R. Neuronal mechanisms of habituation and dishabituation of the gill-withdrawal reflex in Aplysia. Science. 167, 1745-1748 (1970).
- Castellucci, V. F., Carew, T. J., Kandel, E. R. Cellular analysis of long-term habituation of the gill-withdrawal reflex of Aplysia californica. Science. 202, 1306-1308 (1978).
- Dembrow, N. C., et al. A newly identified buccal interneuron initiates and modulates feeding motor programs in Aplysia. J. Neurophysiol. 90, 2190-2204 (2003).
- Fredman, S. M., Jahan-Parwar, B. Command neurons for locomotion in Aplysia. J. Neurophysiol. 49, 1092-1117 (1983).
- Jing, J., Vilim, F. S., Cropper, E. C., Weiss, K. R. Neural analog of arousal: persistent conditional activation of a feeding modulator by serotonergic initiators of locomotion. J. Neurosci. 28, 12349-12361 (2008).
- McManus, J. M., Lu, H., Chiel, H. J. An in vitro preparation for eliciting and recording feeding motor programs with physiological movements in Aplysia californica. J. Vis. Exp. (4320), (2012).
- McPherson, D. R., Blankenship, J. E. Neuronal modulation of foot and body-wall contractions in Aplysia californica. J. Neurophysiol. 67, 23-28 (1992).
- Miller, N., Saada, R., Fishman, S., Hurwitz, I., Susswein, A. J. Neurons controlling Aplysia feeding inhibit themselves by continuous NO production. PloS one. 6, (2011).
- Perrins, R., Weiss, K. R. A cerebral central pattern generator in Aplysia and its connections with buccal feeding circuitry. J. Neurosci. 16, 7030-7045 (1996).
- Xin, Y., Weiss, K. R., Kupfermann, I. An identified interneuron contributes to aspects of six different behaviors in Aplysia. J. Neurosci. 16, 5266-5279 (1996).
- Carew, T. J., Castellucci, V. F., Byrne, J. H., Kandel, E. R. Quantitative analysis of relative contribution of central and peripheral neurons to gill-withdrawal reflex in Aplysia californica. J. Neurophysiol. 42, 497-509 (1979).
- Cohen, T. E., Kaplan, S. W., Kandel, E. R., Hawkins, R. D. A simplified preparation for relating cellular events to behavior: mechanisms contributing to habituation, dishabituation, and sensitization of the Aplysia gill-withdrawal reflex. J. Neurosci. 17, 2886-2899 (1997).
- Frost, L., et al. A simplified preparation for relating cellular events to behavior: contribution of LE and unidentified siphon sensory neurons to mediation and habituation of the Aplysia gill- and siphon-withdrawal reflex. J. Neurosci. 17, 2900-2913 (1997).
- Frost, W. N., Castellucci, V. F., Hawkins, R. D., Kandel, E. R. Monosynaptic connections made by the sensory neurons of the gill- and siphon-withdrawal reflex in Aplysia participate in the storage of long-term memory for sensitization. Proc. Natl. Acad. Sci. U.S.A. 82, 8266-8269 (1985).
- Hawkins, R. D., Greene, W., Kandel, E. R. Classical conditioning, differential conditioning, and second-order conditioning of the Aplysia gill-withdrawal reflex in a simplified mantle organ preparation. Behav. Neurosci. 112, 636-645 (1998).
- Hawkins, R. D., Clark, G. A., Kandel, E. R. Operant conditioning of gill withdrawal in Aplysia. J. Neurosci. 26, 2443-2448 (2006).
- Cai, D., Chen, S., Glanzman, D. L. Postsynaptic regulation of long-term facilitation in Aplysia. Curr. Biol. 18, 920-925 (2008).
- Ho, V. M., Lee, J. A., Martin, K. C. The cell biology of synaptic plasticity. Science. 334, 623-628 (2011).
- Kandel, E. R. The molecular biology of memory storage: a dialogue between genes and synapses. Science. 294, 1030-1038 (2001).
- Wan, Q., Abrams, T. W. Trans-synaptic plasticity: presynaptic initiation, postsynaptic memory. Curr. Biol. 18, 220-223 (2008).
- Bao, J. X., Kandel, E. R., Hawkins, R. D. Involvement of presynaptic and postsynaptic mechanisms in a cellular analog of classical conditioning at Aplysia sensory-motor neuron synapses in isolated cell culture. J. Neurosci. 18, 458-466 (1998).
- Lorenzetti, F. D., Baxter, D. A., Byrne, J. H. Classical conditioning analog enhanced acetylcholine responses but reduced excitability of an identified neuron. J. Neurosci. 31, 14789-14793 (2011).
- Martin, K. C., et al. Synapse-Specific, Long-Term Facilitation of Aplysia Sensory to Motor Synapses: A Function for Local Protein Synthesis in Memory Storage. Cell. 91, 927-938 (1997).
- Montarolo, P. G., et al. A critical period for macromolecular synthesis in long-term heterosynaptic facilitation in Aplysia. Science. 234, 1249-1254 (1986).
- Mozzachiodi, R., Lorenzetti, F. D., Baxter, D. A., Byrne, J. H. Changes in neuronal excitability serve as a mechanism of long-term memory for operant conditioning. Nat. Neurosci. 11, 1146-1148 (2008).
- Alevizos, A., Weiss, K. R., Koester, J. Synaptic actions of identified peptidergic neuron R15 in Aplysia. I. Activation of respiratory pumping. J. Neurosci. 11, 1263-1274 (1991).
- Heid, C. A., Stevens, J., Livak, K. J., Williams, P. M. Real time quantitative PCR. Genome. Res. 6, 986-994 (1996).
- Moroz, L. L., et al. Neuronal transcriptome of Aplysia: neuronal compartments and circuitry. Cell. 127, 1453-1467 (2006).
- Moroz, L. L., Kohn, A. B. Do different neurons age differently? Direct genome-wide analysis of aging in single identified cholinergic neurons. Front. Aging Neurosci. 2, (2010).
- Kadakkuzha, B. M., Puthanveettil, S. V. Genomics and proteomics in solving brain complexity. Mol. BioSyst. , (2013).
- Clemens, S., Katz, P. S. Identified serotonergic neurons in the Tritonia swim CPG activate both ionotropic and metabotropic receptors. J. Neurophysiol. 85, 476-479 (2001).
- Murray, J. A., Hewes, R. S., Willows, A. O. Water-flow sensitive pedal neurons in Tritonia: role in rheotaxis. J. Comp. Physiol. 171, 373-385 (1992).
- Katz, P. S., Frost, W. N. Intrinsic neuromodulation in the Tritonia swim CPG: the serotonergic dorsal swim interneurons act presynaptically to enhance transmitter release from interneuron C2. J. Neurosci. 15, 6035-6045 (1995).
- Brown, G. D., Frost, W. N., Getting, P. A. Habituation and iterative enhancement of multiple components of the Tritonia swim response. Behav. Neurosci. 110, 478-485 (1996).
- Popescu, I. R., Frost, W. N. Highly dissimilar behaviors mediated by a multifunctional network in the marine mollusk Tritonia diomedea. J. Neurosci. 22, 1985-1993 (2002).
- Megalou, E. V., Brandon, C. J., Frost, W. N. Evidence that the swim afferent neurons of tritonia diomedea are glutamatergic. Biol. Bull. 216, 103-112 (2009).
- Hill, E. S., Vasireddi, S. K., Bruno, A. M., Wang, J., Frost, W. N. Variable neuronal participation in stereotypic motor programs. PloS one. 7, (2012).
- Yeoman, M. S., Patel, B. A., Arundell, M., Parker, K., O’Hare, D. Synapse-specific changes in serotonin signalling contribute to age-related changes in the feeding behaviour of the pond snail. Lymnaea. J. Neurochem. 106, 1699-1709 (2008).
- Moroz, L. L., Dahlgren, R. L., Boudko, D., Sweedler, J. V., Lovell, P. Direct single cell determination of nitric oxide synthase related metabolites in identified nitrergic neurons. J. Inorg. Biochem. 99, 929-939 (2005).
- Alania, M., Sakharov, D. A., Elliott, C. J. Multilevel inhibition of feeding by a peptidergic pleural interneuron in the mollusc Lymnaea stagnalis. J. Comp. Physiol. 190, 379-390 (2004).
- Straub, V. A., Benjamin, P. R. Extrinsic modulation and motor pattern generation in a feeding network: a cellular study. J. Neurosci. 21, 1767-1778 (2001).
- Vehovszky, A., Elliott, C. J. The octopamine-containing buccal neurons are a new group of feeding interneurons in the pond snail Lymnaea stagnalis. Acta Biol. Hungarica. 51, 165-176 (2000).
- Jansen, R. F., Pieneman, A. W., ter Maat, A. Spontaneous switching between ortho- and antidromic spiking as the normal mode of firing in the cerebral giant neurons of freely behaving Lymnaea stagnalis. J. Neurophysiol. 76, 4206-4209 (1996).
- McCrohan, C. R., Benjamin, P. R. Synaptic relationships of the cerebral giant cells with motoneurones in the feeding system of Lymnaea stagnalis. J. Exp. Biol. 85, 169-186 (1980).
- Malyshev, A. Y., Balaban, P. M. Buccal neurons activate ciliary beating in the foregut of the pteropod mollusk Clione limacina. J. Exp. Biol. 212, 2969-2976 (2009).
- Ierusalimsky, V. N., Balaban, P. M. Primary sensory neurons containing command neuron peptide constitute a morphologically distinct class of sensory neurons in the terrestrial snail. Cell Tissue Res. 330, 169-177 (2007).
- Malyshev, A. Y., Balaban, P. M. Identification of mechanoafferent neurons in terrestrial snail: response properties and synaptic connections. J. Neurophysiol. 87, 2364-2371 (2002).
- Balaban, P. M., et al. A single serotonergic modulatory cell can mediate reinforcement in the withdrawal network of the terrestrial snail. Neurobiol. Learn. Mem. 75, 30-50 (2001).
- Ierusalimsky, V. N., Zakharov, I. S., Palikhova, T. A., Balaban, P. M. Nervous system and neural maps in gastropod Helix lucorum. 24, 13-22 (1994).
- Kharchenko, O. A., Grinkevich, V. V., Vorobiova, O. V., Grinkevich, L. N. Learning-induced lateralized activation of the MAPK/ERK cascade in identified neurons of the food-aversion network in the mollusk Helix lucorum. Neurobiol. Learn. Mem. 94, 158-166 (2010).
- Ivanova, J. L., et al. Intracellular localization of the HCS2 gene products in identified snail neurons in vivo and in vitro. Cell. Mol. Neurobiol.. 26, 127-144 (2006).
- Kiss, T. Evidence for a persistent Na-conductance in identified command neurones of the snail, Helix pomatia. Brain Res. 989, 16-25 (2003).
- Balaban, P. M. Cellular mechanisms of behavioral plasticity in terrestrial snail. Neurosci. Biobehav. Rev. 26, 597-630 (2002).

