Characterization of MLKL-mediated Plasma Membrane Rupture in Necroptosis
Summary
We report methods for characterization of MLKL-mediated plasma membrane rupture in necroptosis including conventional and confocal live-cell microscopy imaging, scanning electron microscopy, and NMR-based lipid binding.
Abstract
Necroptosis is a programmed cell death pathway triggered by activation of receptor interacting protein kinase 3 (RIPK3), which phosphorylates and activates the mixed lineage kinase-like domain pseudokinase, MLKL, to rupture or permeabilize the plasma membrane. Necroptosis is an inflammatory pathway associated with multiple pathologies including autoimmunity, infectious and cardiovascular diseases, stroke, neurodegeneration, and cancer. Here, we describe protocols that can be used to characterize MLKL as the executioner of plasma membrane rupture in necroptosis. We visualize the process of necroptosis in cells using live-cell imaging with conventional and confocal fluorescence microscopy, and in fixed cells using electron microscopy, which together revealed the redistribution of MLKL from the cytosol to the plasma membrane prior to induction of large holes in the plasma membrane. We present in vitro nuclear magnetic resonance (NMR) analysis using lipids to identify putative modulators of MLKL-mediated necroptosis. Based on this method, we identified quantitative lipid-binding preferences and phosphatidyl-inositol phosphates (PIPs) as critical binders of MLKL that are required for plasma membrane targeting and permeabilization in necroptosis.
Introduction
Identifying genetic components of necroptosis has facilitated the use of animal models to test the implication of necroptosis in physiology and disease1,2,3,4,5. Knockout of RIPK3 or MLKL in mice had minimal implication in development and adult homeostasis suggesting that necroptosis is not essential for life3,6. Moreover, certain species do not contain either RIPK3 or MLKL genes, supporting the non-essential role of necroptosis in animals7,8. On the other hand, challenging knockout animal models with various pathologies induced in the laboratory has revealed an important role of necroptosis in inflammation, innate immunity, and viral infection9,10,11,12.
Necroptosis can be activated in several ways by signaling through different innate immunity sensors, all of which result in the activation of RIPK31,13,14. Active RIPK3 in turn phosphorylates and activates MLKL3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18. The most studied, and perhaps the most complex way, leading to activation of RIPK3 involves death receptor ligation, which bifurcates based on the downstream composition of the signaling complexes to either induce apoptosis or necroptosis1. Necroptosis ensues when signaling through RIPK1 is favored and results in engagement of RIPK319,20. This outcome is easily favored upon pharmacological inhibition or genetic deletion of caspase 8, a putative endogenous inhibitor of necroptosis that keeps necroptosis at bay. RIPK1 binds to and activates RIPK3. Another way to activate necroptosis is through Toll-like receptors TLR3/TLR4 signaling, which engages and activates RIPK3 through TIR-domain-containing adapter-inducing interferon-β (TRIF)21. Yet another way to die by necroptosis is by activation of the DNA sensor DAI, which directly engages and activates RIPK322.
MLKL is a cytosolic protein comprised of an N-terminal helix bundle (NB) domain and a C-terminal pseudokinase domain (psKD) linked by a regulatory brace region3. In normal cells, MLKL is found in the cytosol where it is thought to be in an inactive complex with RIPK314. Activation of necroptosis triggers RIPK3 phosphorylation of MLKL in the activation loop of the psKD, and potentially additional sites in the NB and brace3,15,23. Phosphorylation induces a conformational change in MLKL that results in dissociation from RIPK314. Poorly understood conformational changes release the brace from the psKD24. The brace, which contains 2 helices, mediates oligomerization of MLKL into a putative trimer through the C-terminal helix25. The N-terminal helix of the brace inhibits the NB domain, which is essential for membrane permeabilization24,26. In isolation, NB domain is sufficient to induce plasma membrane permeabilization and necroptosis16,24,27. The pro-necroptotic activity of NB was reconstituted in mouse embryonic fibroblasts deficient in MLKL (mlkl-/- MEFs). NB is a lipid binding domain that preferentially engages the phospholipid phosphatidylinositol 4,5 diphosphate (PIP2). We proposed a stepwise mechanism of activation of MLKL, wherein brace oligomerization facilitates recruitment of MLKL to the plasma membrane via weak interactions of NB with the PIP2 polar head group24. At the membrane, the NB undergoes regulated exposure of an additional high-affinity binding site for PIP2, which is masked by the brace in inactive MLKL. Overall, the multiple interactions of NB with PIP2 destabilize the plasma membrane leading to its rupture, although the molecular mechanism of these events have not been elucidated.
Here we illustrate specific methods used to characterize the function of MLKL as executioner of necroptosis24. In particular, we focus on the most minimal domain of MLKL, the NB and brace (NBB), which is regulated by brace inhibition and can be activated through enforced dimerization to induce plasma membrane rupture and necroptosis. We describe our inducible expression system combined with enforced drug-induced FKBP-mediated dimerization for live-cell imaging, and electron microscopy of cells undergoing necroptosis. Additionally, we illustrate our in vitro NMR analysis of the interactions of NBB with phosphatidylinositols (PIPs).
Protocol
1. Cloning and Cell Line Generation
- PCR amplify the NBB region, corresponding to amino acid residues 1-140 (NBB140), from human MLKL cDNA for in frame standard restriction enzyme-based cloning with the oligomerization domain 2x FK506 binding protein (2xFKBP or 2xFV) and Venus fluorescent protein into the Doxycycline (Dox)-inducible retroviral vector pRetroX-TRE3G to obtain NBB140-2xFV-Venus (Table 1, Figures 1A-B).
- Immortalize primary mlkl-/- or ripk3-/- mlkl-/- mouse embryonic fibroblasts (MEFs) obtained from the respective mice available in the Green laboratory by transient transfection with SV40-large antigen expressing plasmid. Select immortalized cells in ~1–2 weeks, and maintain in DMEM supplemented with 10% fetal bovine serum (FBS), 2 mM L-glutamine, 100 U/mL penicillin and streptomycin, 1 mM sodium pyruvate, and nonessential amino acids at 37 °C and 5% CO2 in standard tissue culture incubators.
- Transduce SV40-immortalized mlkl-/- MEFs with the reverse tetracycline controlled transactivator (rtTA)-containing retrovirus expressed from a plasmid containing the blasticidin resistance gene (our modified plasmid) and treat for 1 week with 5 μg/mL blasticidin to select for cells stably expressing rtTA28.
- Transduce the rtTA-expressing MEFs with the Dox-inducible retroviral vector containing the chimeric MLKL (pRetroX-NBB140-2xFV-Venus-puro) and select using 4 μg/mL puromycin for one week, 2 days after transduction.
2. Live-Cell Microscopy Imaging of MLKL-mediated Necroptosis
- Plate mlkl-/- MEFs expressing NBB140-2xFV-Venus at a density of 50,000 cells per well (1 mL/well) in 24 well plates, and incubate overnight (Figure 2A). Use live-cell staining for automated cell counting (see Table of Materials).
- Treat cells for 12 h with Dox (0.5 μg/mL) to induce the expression of NBB140-2xFV-Venus.
- Induce necroptosis with 25 nM FKBP dimerizer (Dim) in addition to 25 nM membrane-impermeable green fluorescent dye (see Table of Materials) in standard DMEM (see 1.2). Cells undergoing necroptosis accumulate the dye as their plasma membrane integrity is compromised by NBB140-mediated rupture. Perform assays in triplicate or quadruplicate.
- To monitor necroptosis using single- or dual-color imaging systems (see Table of Materials), place the vessels in the respective imaging unit housed in a mammalian tissue culture incubator.
- Open the software (see Table of Materials) and select the tab Schedule Upcoming Scans under the Tasks List.
NOTE: This protocol is for imaging using single-color systems, but a similar software is available for dual-color systems. - Select the "Tray, Vessel, Scan Types", and the "Scan Pattern" in the "Physical Layout" tab and "Fast Fluorescence" in the Properties tab.
- Add the "Scan Time" by clicking on the "Timeline" window and the total number of time points and frequency of acquisition (typically 1 acquisition every 0.5 h to 1 h for 6 h to 12 h, or every 5 min for fast-kinetics necroptosis) and start image acquisition by selecting "Apply" (red button).
- Quantify necroptosis using the image analysis software (see Table of Materials) by selecting from the "Tasks Pane", "Object Counting New Analysis with Fixed Segmentation Threshold above the background level", which can be determined by hovering the mouse cursor over background regions in several images used in the analysis.
- Examine fluorescent objects by selecting Previewing using current image and refine the Threshold levels as needed.
- Integrate green fluorescence counts by opening the "Launch New Analysis" tab, choose the "Time Range", select the wells to be analyzed, enter the "Analysis Job Name", and press "OK".
- Access analyzed data from the "Analysis Jobs" tab by double-clicking on the respective job name and exporting from the Graph/Export window for additional analysis in external graphing software (see Table of Materials).
- Quantitate data as green fluorescence positive (+ve) counts/mm2 or normalize the counts by confluence and express the data as green fluorescence +ve counts/confluence.
- Optionally, at the experimental endpoint, stain cells with 100 nM of the membrane-permeant green fluorescent dye (see Table of Materials). Normalize data to % necroptotic cells as the ratio of green fluorescence/green fluorescence at endpoint.
3. Live-cell Confocal Microscopy Imaging of Plasma Membrane Recruitment and Permeabilization by MLKL
- Plate mlkl-/- MEFs expressing NBB140-2xFV-Venus at a density of 20,000 cells per well (0.5 mL/well) in a glass bottom 4-well microscopy chamber pretreated with 50 µg/mL fibronectin in PBS at 37 °C for 30 min and incubate for ~12 h (Figure 2B).
- Treat the cells with Dox (0.5 μg/mL) for ~12 h to induce the expression of NBB140-2xFV-Venus.
- Induce necroptosis by incubating with fresh medium (1 mL/well) containing 25 nM Dim to induce NBB140-2xFV-Venus activation and translocation to the plasma membrane.
- Place the chamber on a spinning disc laser scanning confocal microscope built on an inverted stand equipped with environmental control and a 63 1.4 NA objective and begin imaging acquisition using software of choice (see Table of Materials).
- Initialize the software, select "Focus" icon and YFP filter configuration (excitation wavelength 515 nm).
- Locate the field of view and focal plane, and engage the focus maintenance system using the "Definite Focus" tab.
- To begin acquisition, select the "Capture" tab followed by assignment of filter configuration, appropriate EMCCD camera exposure time and intensification gain, and time-lapse acquisition parameters including interval and length of acquisition.
- Monitor cellular redistribution of the fluorescent NBB140-2xFV-Venus protein during necroptosis by acquiring images every 10 s by a EMCCD camera using a 515 nm laser (Movie 1).
NOTE: Photobleaching is a concern with fluorescent protein imaging. Venus is one of the more resistant fluorescent proteins to photobleaching29. If fluorescent proteins that are more susceptible to photobleaching are used, image every 30 s or longer to allow recovery of signal between imaging time points. - To monitor plasma membrane association of NBB140-2xFV-Venus during necroptosis, image every 10 s the sample prepared in 3.3 by total internal reflection fluorescence (TIRF) microscopy using a microscope equipped with an automated TIRF slider and a 100X 1.4 NA objective and an EMCCD camera (Movie 2). Follow steps 3.5–3.7, but adjust the TIRF angle within the TIRF focus tab according to sample and bottom glass thickness of the microscopy chamber.
NOTE: Plasma membrane markers such as LCK-C-RFP may be used as controls for plasma membrane localization in TIRF analysis. - Perform image processing to enhance quality by convolution with the negative normalized second derivative of a Gaussian function (Marr algorithm).
4. Electron Microscopy
- Plate mlkl-/- MEFs expressing NBB140-2xFV-Venus at a density of 5 million cells in a plasma-coated 150 mm cell culture dish and incubate for 12 h (Figure 3).
- Treat the cells with Doxycycline (0.5 μg/mL) to induce the expression of NB1-140-2xFV-Venus for 12 h.
- Induce NBB140-2xFV-Venus activation and translocation to the plasma membrane with 25 nM of homodimerizer for 5 min. This time is established from the kinetics of necroptosis observed in live-cell imaging.
- Remove the media and fix cells using 10 mL of 2.5% glutaraldehyde, 2% paraformaldehyde in 0.1 M sodium cacodylate buffer pH 7.4 (cacodylate buffer) pre-warmed at 37 °C.
- Collect the sample by scraping and spin the samples for 10 min at 500 x g. Discard the supernatant.
- Postfix the sample for 1.5 h in reduced 2% osmium tetroxide with 1.5% potassium ferrocyanide in 0.1 M sodium cacodylate buffer. Osmium tetroxide is a staining agent widely used in TEM to provide contrast. We used TEM as preliminary analysis prior to SEM of cell undergoing necroptosis.
- Rinse the sample 5 times in ultrapure water for 5 min each at 500 x g, followed by rinses in buffer (see 4.6), water, and ethanol on an automatic processor. Discard the supernatant.
- For the SEM, stain the previously fixed samples with 1% uranyl acetate and lead aspartate heavy-metal stain31.
- Dehydrate the samples through a graded series of alcohol and propylene oxide solutions.
- Embed the sample in hard resin32 to obtain a resin block.
- Cut thick sections of 0.5 μm to determine the correct area of analysis.
- Coat the samples with an ultra-thin film of electrically-conducting metal (Iridium).
- Image and analyze the samples (Figure 3). For quantification, a low-magnification image of the resin block surface with exposed cells is scanned at 5 keV using an electron microscope.
- Visualize cells across individual images captured by SEM or imaging software may be utilized to join multiple fields-of-view into one contiguous image30. Roughly 100 cells may be used to evaluate the fraction of cells undergoing necroptosis in this manner by generating a high-resolution montage in software of choice (see Table of Materials).
NOTE: Scoring necrotic morphology by SEM compares normal cell features with the progression of pre-necrotic or necrotic phenotypes. These features include plasma membrane alterations including microvilli disappearance, flattening, ruptures in cells ranging from 10 nm to 1 µm in size, or loss of organelle structure across at least 30–40% of the cytosolic cell area.
5. Lipid Binding of MLKL by Nuclear Magnetic Resonance (NMR) Spectroscopy
- Prepare 15N-labeled NBB1-156 by standard protein expression and purification as previously described24.
- Dissolve 500 µg of 1,2-distearoyl-sn-glycero-3-phosphoinositol ammonium salt (18:0 PI) and 1,2-dioleoyl-sn-glycero-3-phospho-(1'-myo-inositol) ammonium salt (18:1 PI) in 500 µL of ice-cold CHCl3 each for final concentrations of 1 mg/mL.
- Sonicate 1 mg/mL 18:0 and 18:1 PI in a sonication water bath for 1 min at room temperature or up to 5 min in an ice-water mixture.
- Aliquot 1 mg/mL 18:0 and 18:1 PI into clean glass vials for individual NMR titration series (Figure 4). Transfer lipids in organic solvents using glass air-tight syringes.
- Aliquot 132 µL of 1 mg/mL 18:1 PI (molar mass = 880.137 g/mol) in CHCl3 for a final resuspension volume of 1,200 µL at 125 µM concentration.
NOTE: For 5 mm NMR tubes, prepare serial dilution volumes of >1,000 µL and final NMR sample volumes of >500 µL. For 3 mm NMR tubes, prepare serial dilution volumes of >300 µL and final NMR sample volumes of >150 µL.
- Aliquot 132 µL of 1 mg/mL 18:1 PI (molar mass = 880.137 g/mol) in CHCl3 for a final resuspension volume of 1,200 µL at 125 µM concentration.
- Aliquot 1 mg/mL porcine brain L-α-phosphatidylinositol-4,5-bisphosphate ammonium salt in 20:9:1 CHCl3/MeOH/H2O (brain PI(4,5)P2) into clean glass vials.
- Place vials under a gaseous stream of argon (Ar) or nitrogen (N2) with moderate flow to evaporate organic solvent without splashing against the glass vial walls. Five to thirty min is typically sufficient for volumes of 10–200 µL organic solvent.
- Arrange glass vial(s) at bottom of a Büchner or vacuum flask using large tweezers, seal with a rubber stopper, and attach plastic tubing connected to a vacuum line. Evacuate flask under mild vacuum overnight to remove residual organics.
- Overlay lipid film aliquots with inert gas, seal with an airtight lid, and store at -20 °C for use within one month or -80 °C for longer-term storage.
- Prepare NMR sample buffers without detergent (-Det) containing 20 mM NaxHyPO4 pH 6.8, 10% D2O and with detergent (+Det) containing 20 mM NaxHyPO4 pH 6.8, 10% D2O, 0.34 mM n-dodecyl-β-D-maltopyranoside (DDM).
- Add +Det buffer to glass vial with lipid film to achieve desired concentration and sonicate in water bath for up to 30 min, alternating sonication for 10 min followed by visual inspection.
NOTE: Sonication may warm sample. Allow to cool to room temperature for 15 min before final protein sample reconstitution. - Perform serial dilution of lipid-detergent micelles in +Det buffer using sonication of 1:1 mixtures for the desired concentration series. Starting at 125 µM lipid concentration, three repetitions will yield 62.5 µM, 31.3 µM, and 15.6 µM lipid in 0.34 mM DDM.
- Prepare control and reference NMR samples of protein and additives in -Det and +Det buffers, respectively. Add protein and additives to each dilution of lipid in +Det buffer.
NOTE: For 15N NBB156, protein is added to a final concentration of 40 µM and supplemented with 2 mM deuterated dithiothreitol to keep Cys residues reduced. - Transfer the appropriate volume of samples to 5 mm or 3 mm NMR tubes (see note in step 5.4).
- Manually load samples in NMR instrument or arrange in an automation platform such as the SampleCase loader.
- Acquire composite NMR spectra using standard pulse programs for 1H proton spectra as quality control followed by 2D 1H-15N correlation spectra either as transverse relaxation optimized spectroscopy (TROSY) or heteronuclear multiple quantum coherence (SOFAST-HMQC)33.
NOTE: Usage of 3 mm NMR tubes requires 64 scans for 1H-15N TROSY (~3 h) or 1H-15N SOFAST-HMQC (~90 min). Scan numbers are halved for 5 mm NMR tubes. - Processed 2D NMR spectra may be added to a CARA repository34 for analysis or other software of choice to the user.
- Analyze software by annotating 2D NMR resonances for three well-dispersed and resolved peaks for each protein state. Record the peak amplitudes for each of the six peaks for each buffer condition containing NBB156 (Figure 5A).
- Use CARA to prepare a batch-integration list (Click Main Menu heading "Spectrum | Setup Batch List"…) of all spectra collected and analyze using a single peak assignment file (Click Main Menu heading "Peaks | Import Peaklist" …and choose a user-defined .peaks file with 6 total peaks, 3 for each protein state, defined within).
- Tune the settings ("Integrator | Tune Peak Model" …) of X-width 0.06 ppm and Y-width 0.30 ppm (Click Main Menu heading "Integrator | Tune Peak Model" …) before recording the final output in two menu selections (1. Click Main Menu heading "Integrator | Integrate Batch List") (2. Click Main Menu heading "Peaks | Export Integration Table") (Figure 5B).
NOTE: The backbone amide resonances for residues M21, V53, and L105 have been assigned for the NBB156 closed-brace state. The open-brace state was assigned to backbone amide resonances of residues G130 and A141 and the epsilon proton side-chain resonance of W133 using 3D NMR experiments24.
- Normalize samples for direct comparison from different magnets or sample conditions by averaging the three open-brace amplitudes of the reference samples and calculating a normalization factor relating the two references. Calculate the scaled amplitudes for NBB156 resonances using the normalization factor and setting negative amplitudes to zero (Table 2).
Examples: 1) Comparing samples from different magnets: Magnet A, NBB156+DDM and Magnet B, NBB156+DDM. 2) Detergent comparison: 0.34 mM DDM and 1.7 mM DDM. - Calculate for each resonance the scaled fraction of closed- or open-brace by dividing with the range of minimum to maximum amplitude of all spectra collected containing appropriate references of free NBB156 and fully open-brace NBB156 in detergent.
- Plot the normalized NMR amplitudes as a function of lipid concentration and compare the fraction of open-brace NBB156 in different conditions (Figure. 5C).
NOTE: Alternatively, analysis of the fraction of closed-brace conformation against lipid concentration can be performed to compare lipid binding to NBB156. We prefer the former analysis because it is quicker and better reports on the fraction fully-engaged by lipids.
Representative Results
Visualizing regulated necroptosis execution in live cells has been possible through inducible expression of a minimal truncated MLKL construct, NBB140-2xFV-Venus. This construct maintains the ability to induce plasma membrane permeabilization and is activated through Dim-induced oligomerization of the FKBP cassette (2xFV). We observe and quantify necroptosis by live-cell microscopy imaging, monitoring kinetically (every 5 min) the uptake of a cell impermeable green fluorescence DNA binding dye (Figure 6A). This inducible system is very robust, and complete necroptosis of mlkl-/- MEFs can be observed within ~1 h upon Dim-mediated oligomerization of Dox-preincubated cells that express NBB140-2xFV-Venus (Figure 1A). We refer to these conditions as fast-kinetics necroptosis. Expression of Venus alone ± Dim or NBB140-2xFV-Venus in the absence of Dim did not induce necroptosis (Figure 6A). We usually perform at least 3 replicate imaging experiments in triplicate or quadruplicate in 24-, 96-, or 384-well plates.
The fast-kinetics necroptosis induced by enforced dimerization of NBB140-2xFV-Venus (Figure 6A) supports the role of MLKL as the putative executioner of plasma membrane rupture. To visualize the redistribution of NBB140-2xFV-Venus to the plasma membrane during necroptosis, live-cell confocal microscopy is used to monitor Venus fluorescence. During fast-kinetics necroptosis, within 2–3 min of incubation with Dim, Venus coating of the cell periphery is observed, followed by gradual cell rounding (Movie 1). NBB140-2xFV-Venus accumulation at the plasma membrane is visualized by TIRF confocal microscopy, which focuses on the cell volume at the cytosol-plasma membrane-glass interface. Plasma membrane-associated Venus puncta are visible within 1–2 min of incubation with Dim (Movie 2). Thus, enforced oligomerization of NBB140-2xFV-Venus induces its rapid redistribution to plasma membrane.
Scanning electron microscopy (SEM) is a powerful tool to reveal the morphological changes in cells undergoing necroptosis. Under fast necroptosis induced by oligomerization of NBB140-2xFV-Venus, cells change morphology from normal elongated shapes (time 0 min) to rounded and swelled (5 min) to partially ruptured (10 min) and extensively dismantled where the cytosol has vanished (20 min) (Figure 6B). SEM complements the previous observations from live-cell microscopy correlating MLKL localization to the plasma membrane with membrane rupture. Overall the microscopy techniques presented herein offer complementary means to monitor, evaluate and quantify necroptosis at cellular level and implicate NBB140-2xFV-Venus in the execution of plasma membrane rupture.
To determine if MLKL can directly contact the plasma membrane we perform in vitro lipid binding experiments monitored by NMR spectroscopy using recombinant NBB156. Serial dilutions of lipid-detergent micelles, using a constant concentration of detergent (vehicle for lipid presentation) and variable lipid concentrations, are tested for binding to 15N-labeled NBB156 by 2D NMR spectroscopy, which monitors binding-induced changes in the protein. NBB156 undergoes major structural changes upon lipid binding to displace the inhibitory brace region (amino acids 132-156) from the closed and helical (Closed Brace) to the open and intrinsically disordered conformation (Open Brace) (Figure 7A). Two-dimensional NMR spectroscopy provides per residue information on mixtures of both conformers (Figures 7B-7C). We previously assigned the backbone amides of both conformers24. We can easily monitor the percentages of closed and open conformers in a given sample as described in Figures 5A-C. Using this binding assay, we explore NBB156 binding to PIPs in DDM detergent, which is inert to NBB156 binding (Figure 5B). In this assay, PIP2 is the best NBB156 ligand, which induces full opening of the inhibitory brace at 125 µM (Figure 5C). In contrast, saturated (18:0) PI and unsaturated (18:1) PI are poor NBB156 ligands inducing partial brace opening under the same conditions (Figure 5C). Our NMR-based lipid binding assay provides supporting evidence for the interaction of MLKL NBB156 with phospholipids and suggests a direct link between MLKL and the plasma membrane.
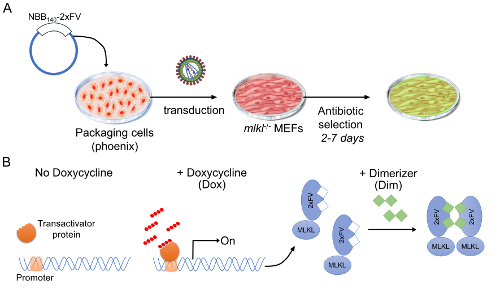
Figure 1: Stable cell line harboring a modified Tet-On 3G inducible system. (A) Stable cell lines were generated by retroviral transduction of mlkl-/- MEFs with the pTREX-rtTA-blast followed by the pRetroX-TRE3G-NBB140-2xFV-Venus-puro. Each transduction was followed by antibiotic selection for up to 1 week before moving on to subsequent analyses. (B) Schematic representation of inducible MLKL expression system. This system is based on 2 drug-inducible regulatory steps: i) MLKL gene expression and protein production (+Dox) and ii) activation by oligomerization (+Dim). When protein production is induced in advance, +Dox, fast-kinetics necroptosis can be triggered, +Dim. Please click here to view a larger version of this figure.
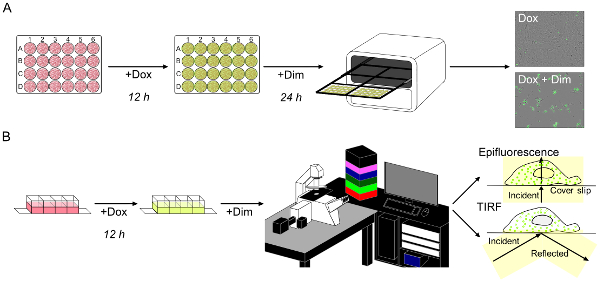
Figure 2: Live-cell imaging of fast-kinetics necroptosis reveals rapid membrane relocalization of MLKL. (A) Fast-kinetics necroptosis induced as in Figure 1B can be monitored in an imaging system. Necroptosis is scored by uptake of the cell-impermeable green fluorescent dye. (B) Schematic representation of live-cell imaging of fast-kinetics necroptosis with epifluorescence and total internal reflection fluorescence (TIRF) microscopy analysis. Please click here to view a larger version of this figure.
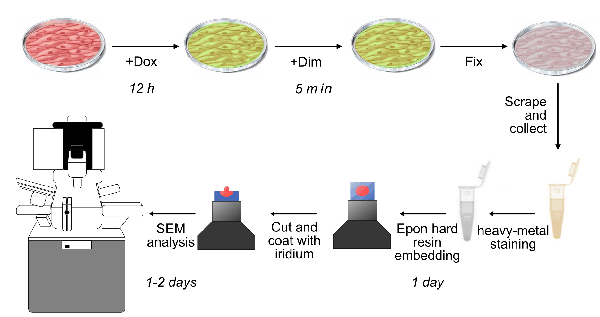
Figure 3: Schematic representation of scanning electron microscopy (SEM) sample preparation and analysis. Cells from fast-kinetics necroptosis induced as described in Figure 1 are fixed on the plate at different time points after addition of Dim. The cells are then prepared for SEM analysis by scraping and pooling, heavy metal staining, resin embedding, and iridium coating as described in section 4. Please click here to view a larger version of this figure.
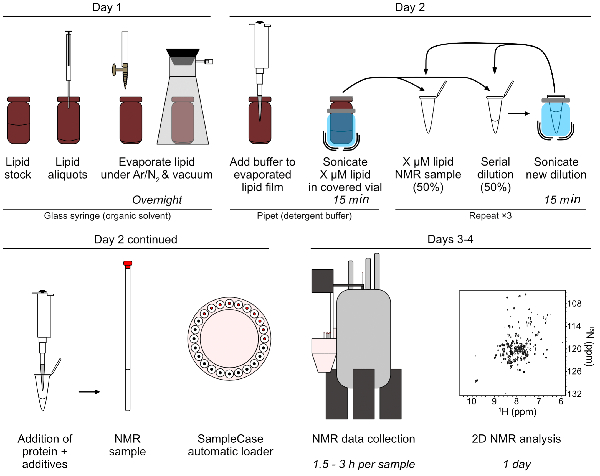
Figure 4: Sample preparation and data collection for NMR titrations of 15N NBB156 and lipid-detergent micelles. This assay and analysis is typically performed in 3–4 days: During day 1, lipid aliquots are dispensed and dried overnight. During day 2, the lipid films are rehydrated in assay buffer ± detergents, sonicated, and serially diluted (two-fold) by mixing equal volumes of the rehydrated lipid solution with lipid-free buffer solution. Each dilution is sonicated to ensure homogeneous mixing and distribution of lipids in detergent micelles prior to subsequent dilutions. Protein samples are mixed in the serial lipid-detergent micelle dilutions, loaded in the appropriate NMR tubes, placed in a SampleCase loader (or loaded manually), and automatic NMR data collection is started. During subsequent days, NMR data collection is completed and followed by 2D NMR analysis. Please click here to view a larger version of this figure.
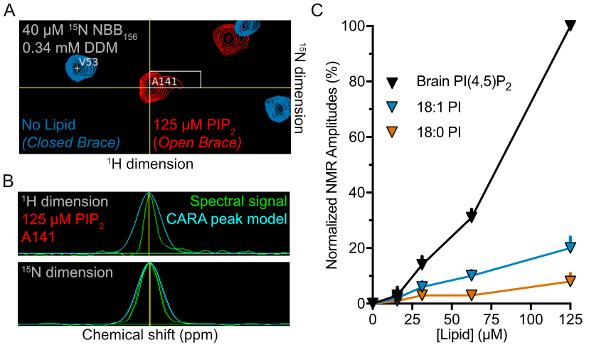
Figure 5: Lipid-binding preferences of NBB156 measured using 2D NMR. (A) Superimposed 1H-15N TROSY spectra for 15N-NBB156 in the presence (red) or absence (blue) of 125 µM PI(4,5)P2 in 0.34 mM DDM visualized in CARA. (B) One-dimensional slices of resonances used for amplitude measurements and normalization. The raw spectrum (green) is overlaid with the boundary for amplitude and integration by the program CARA. (C) Normalized amplitudes of NBB156 in the presence of phosphoinositide-DDM micelles plotted against lipid concentration. PIP2 induces full opening of the brace. Please click here to view a larger version of this figure.
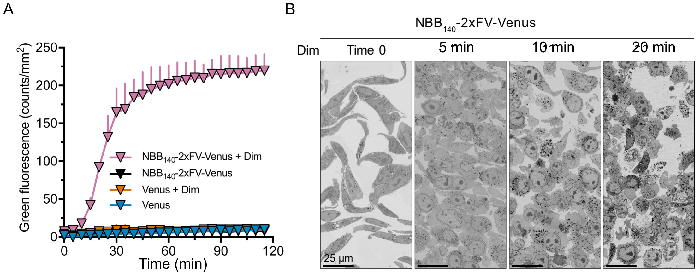
Figure 6: Fast-kinetics necroptosis induced by oligomerization of NBB140-2xFV-Venus in mlkl-/- MEFs. (A) Necroptosis quantification by high-throughput fluorescence imaging of uptake of cell-impermeable, DNA-binding green fluorescent dye. Robust necroptosis induction is only observed upon Dim-induced oligomerization of NBB140-2xFV-Venus, but not in the absence of Dim. Venus-only control experiments are also performed. All conditions are done in triplicate and are plotted as average and SD from one representative replicate. (B) Fast-kinetics necroptosis induced in the presence of Dim visualized with scanning electron microscopy. The time course reveals morphological changes underlying necroptosis including cell rounding and swelling (5 min), rupture of plasma membrane (10 min), and complete extravasation of the cytosol (20 min). Each sample had similar number of cells at time 0 min. From left to right, the zoomed in area contains 26, 32, 23, and 36 cells with nuclei. Please click here to view a larger version of this figure.
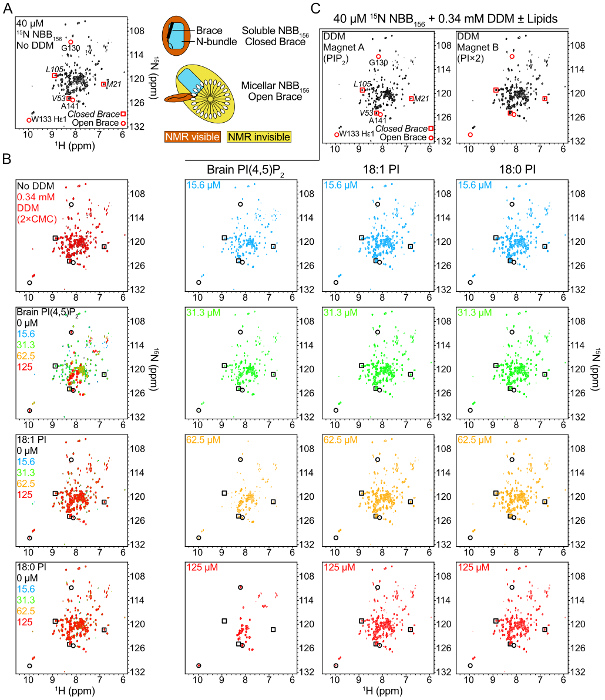
Figure 7: Binding of NBB156 to phosphoinositides monitored by NMR spectroscopy. (A) 1H-15N TROSY spectrum of 15N-NBB156. The chemical shifts of resonances used for normalization of the closed brace spectrum or quantification of the open state are highlighted with squares or circles, respectively. Right, schematic of NMR signatures detected in the absence or presence of lipid-detergent micelles. NBB156 does not bind DDM detergent micelles in the absence of phosphoinositides. (B) Superimposed 1H-15N TROSY spectra of 15N-NBB156 in the presence of the respective lipid-detergent micelles. (C) Single 1H-15N TROSY spectra of 15N-NBB156 in the presence of the respective lipid-detergent micelles from panel B. By detecting the open and closed conformations unambiguously in mixtures of the two conformations we can quantify the percentages of the two conformers in a given sample. PIP2 is the preferred lipid ligand of NBB156 providing a direct link between MLKL and plasma membrane phospholipids. Please click here to view a larger version of this figure.
| PCR buffer (10x) | 5.0 µL |
| DNA template (100 ng/µL) | 1.0 µL |
| cDNAs for MLKL, 2x FKBP, Venus) | |
| dNTPs (25 mM each NTP) | 0.5 µL |
| PCR primers forward (F) (100 ng/µL) | 1.3 µL |
| PCR primers forward (FR (100 ng/µL) | 1.3 µL |
| NBB140 F | ATAATCGATACCATGGAAAATTTGAAGCATATT |
| NBB140 R | TATGCGGCCGCATCCTGCTGATCTTCCTGTGC |
| 2xFKBP F | ATAGCGGCCGCAGGCGTCCAAGTCGAAACCATT |
| 2xFKBP R | TATGCGGCCGCTTCCAGTTTTAGAAGCTCCAC |
| Venus F | ATAGGGCCCACCATGGTGAGCAAGGGCGAG |
| Venus R | TATGAATTCTTACTTGTACAGCTCGTC |
| DNA Polymerase (2.5 U/µL) | 1.0 µL |
| Distilled deionized water (ddH2O) | 39.9 µL |
| Total reaction volume | 50.0 µL |
| PCR cycling parameters | |
| 1 cycle | 94-98 °C; 45 s |
| 25–30 cycles | 94-98 °C; 45 s |
| 58 °C; 45 s | |
| 72 °C; 1-2 min | |
| 1 cycle | 72 °C; 10 min |
Table 1: PCR reaction of NB140-2xFV-Venus for restriction enzyme-based cloning in pRetroX-TRE3G.
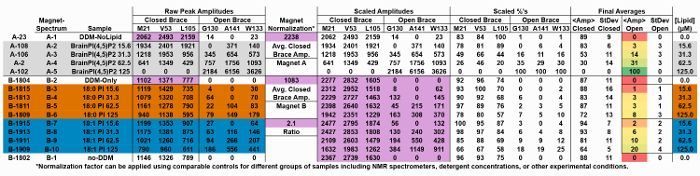
Table 2: Raw data for NMR spectra presented in Figure 5 illustrating the normalized transformation of data collected from a second NMR magnet to the calculated average open brace state. Please click here to view a larger version of this table.
Movie 1: Fast-kinetics necroptosis induced by oligomerization of NBB140-2xFV-Venus in mlkl-/- MEFs. Live confocal microscopy showing the rapid translocation of NBB140-2xFV-Venus from the cytosol to the plasma membrane after enforced oligomerization of the FKBP domain. The cells were treated as described in Figure 1. Yellow: NBB140-2xFV-Venus. Please click here to view this video. (Right-click to download.)
Movie 2: TIRF microscopy of fast-kinetics necroptosis. TIRF microscopy showing fast (~2 min) relocalization and aggregation of MLKL on the plasma membrane upon addition of Dim to mlkl-/- MEFs expressing NBB140-2xFV-Venus. Please click here to view this video. (Right-click to download.)
Discussion
We provide protocols for techniques that we combined to implicated MLKL as the putative executioner of plasma membrane rupture24. In addition to deciphering the regulatory network that regulates MLKL-mediated necroptosis, these techniques can be used independently to characterize other suitable biological systems. Practically speaking, these techniques are medium- to low-throughput discovery tools.
We have routinely used live-cell imaging of NBB140-2xFV-Venus-mediate necroptosis, and that induced by other MLKL constructs, to mechanistically dissect the regulation of MLKL in necroptosis through mutagenesis and by using small molecule chemical probes. In particular, we and other groups have demonstrated that human and mouse constructs of MLKL that contain the minimal region of NBB can mediate necroptosis in human and mouse cell lines. One reported caveat governing MLKL-mediated necroptosis is species specificity of RIPK3 and MLKL interaction, which prevents cross-reactivity among species observed upon upstream stimulation by combination of TNF, smac mimetics, and caspase inhibition25,35,36. Accordingly, human and mouse RIPK3 and MLKL proteins do not cross-react under these conditions25,35,36. To bypass the inter-species limitations, we introduce mutations that activate human MLKL or use oligomerization cassettes as shown here for NBB140-2xFV-Venus24. Human NBB140 is a construct that maintains the inhibitory region and therefore remains inactive in the context of NBB140-2xFV-Venus. Dimerization of 2xFV is thought to activate NBB140 by releasing the brace. In contrast, human NBB182 induces necroptosis even in the absence of oligomerization, because this construct is able to spontaneously oligomerize24. Moreover, point mutants (R30A, R30E, E136A, and E136R) activating the NB region in the context of full-length human MLKL overcome the need for activation by RIPK3 phosphorylation24. Furthermore, we reconstituted dimerizable human RIPK3 (Cerulean-2xFV-RIPK3) and Dox-inducible full-length human MLKL-Venus, which are compatible and robustly induce necroptosis in ripk3-/- mlkl-/- MEFs24. Others recently revealed additional mutations or human-mouse domain swapped MLKL constructs that may overcome species specificity barriers25.
We typically optimize the necroptosis assay conditions by performing several range-finding experiments in 96-well plates. We then follow up with optimized experiments in 24-well plates, which provide sufficient cells for multiplexing with the complementary fluorescent activated cell sorting (FACS) and western blotting analyses performed subsequently on the same samples immediately after imaging the final time point. Currently, live-cell imaging is based on detection of green and red fluorescence, limiting the labeling possibilities for many applications. A common alternative to green fluorescent dyes is the cell-impermeable DNA-binding fluorescent dye propidium iodide (PI). Dual color imaging instruments offer the option of using these dyes with complementary green or red fluorescent proteins to enhance the utility and information content of necroptosis imaging.
The ability of MLKL to translocate to the plasma membrane after its activation, makes live microscopy the technique of choice to study the biology of this protein and to follow in real time the morphological changes underlying necroptosis. TIRF microscopy unequivocally resolves MLKL protein aggregates on the plasma membrane, especially when executed in the presence of markers that co-localize to the plasma membrane. We used LCK-C-RFP as a marker for plasma membrane co-localization24. The ability to tag proteins of interest with different fluorophores, also allows to study concurrently the activity of several proteins involved in the same process. The next generation of super-resolution microscopy partially overcomes the diffraction limit problem in standard confocal microscopy and has the potential to reveal additional features of MLKL-mediated necroptosis.
One of the hallmarks of necroptosis is plasma membrane permeabilization. Even if several techniques can indirectly quantify or indicate membrane ruptures, only EM can visualize discontinuities in the plasma membrane integrity induced by MLKL. While TEM has the highest power of resolution, SEM has the ability to map larger specimen areas. This feature, combined with the development of new and powerful software able to manage greater volume of data, has enhanced the utility of SEM in cell morphology characterization. Moreover, SEM is also able to scan consecutive sample sections allowing 3D reconstruction when coupled to other systems, such as Focused Ion Beam. Some of the drawbacks of EM analysis include the cost of the instrumentation and maintenance, necessity of highly specialized staff, and significant time investment in sample processing and analysis.
Dot blot assays have been used originally to make direct connections between MLKL and plasma membrane phospholipids16,27. Our NMR-based lipid binding assay definitively implicates MLKL in specific phospholipid binding, highlighting PIP2 as the MLKL ligand of choice. Our results demonstrate a deleterious effect of acyl chain saturation on NBB156 binding to phosphatidylinositol. Nonetheless, how MLKL binding to PIP2, and potentially other phospholipids, results in plasma membrane rupture remains unknown. Using this protocol any other phospholipid may be tested for binding to MLKL. Our assay serves as a great tool for testing emerging models of MLKL-mediate necroptosis, as it can be used with mutants of MLKL and various ligands to pinpoint the specific contributions to lipid binding. Additionally, it can be used for any other membrane associated systems to interrogate their lipid binding profiles.
Declarações
The authors have nothing to disclose.
Acknowledgements
None.
Materials
| Cloning and cell line generation | |||
| pRetroX-TRE3G | Clontech | 631188 | |
| Tet-On transactivator plasmid | Llambi et al., 2016 | ||
| Mouse Embryonic Fibroblasts (MEFs) mlkl-/- | Dillon et al., 2014 | ||
| Blasticidin S Hydrochloride | Thermo Fisher Scientific | BP2647100 CAS#3513-03-9 | |
| Cell death quantification and live-cell microscopy | |||
| Doxycycline | Clontech | 631311 CAS# 24390-14-5 | |
| B/B Homodimerizer AP20187 | Takara | 635059 CAS# 195514-80-8 | |
| SYTOX Green | Thermo Fisher Scientific | S7020 | |
| Syto16 | Thermo Fisher Scientific | S7578 | |
| NMR | |||
| 15N Ammonium Chloride | Cambridge Isotope Laboratories | NLM-467-10 CAS# 12125-02-9 | |
| Deuterated DTT | Cambridge Isotope Laboratories | DLM-2622-1 | |
| Deuterium Oxide | Sigma Aldrich | 617385-1 CAS# 7789-20-0 | |
| n-Dodecyl-β-D-Maltopyranoside | Anatrace | D310 CAS# 69227-93-6 | |
| L-α-phosphatidylinositol-4,5-bisphosphate (Brain, Porcine) (ammonium salt) | Avanti Polar Lipids | 840046X CAS# 383907-42-4 | |
| 1,2-distearoyl-sn-glycero-3-phosphoinositol (ammonium salt) (18:0 PI) | Avanti Polar Lipids | 850143 CAS# 849412-67-5 | |
| 1,2-dioleoyl-sn-glycero-3-phospho-(1'-myo-inositol) (ammonium salt) (18:1) | Avanti Polar Lipids | 850149 CAS# 799268-53-4 | |
| Specialized Equipment | |||
| IncuCyte FLR or ZOOM | Essen BioScience, Inc. | Live-cell microscopy imaging | |
| Helios NanoLab 660 DualBeam | Thermo Fisher Scientific | Electron microscope | |
| Software | |||
| IncuCyte 2011A Rev2 v20111.3.4288 (FLR) | Essen BioScience, Inc. | http://www.essenbioscience.com | Imaging analysis |
| FEI MAPS | Thermo Fisher Scientific | https://www.fei.com/software/maps/ | EM analysis |
| TopSpin v3.2 | Bruker BioSpin | http://www.bruker.com | NMR data collection |
| CARA v1.9.1.7 | http://cara.nmr.ch/ | NMR data analysis | |
| Slidebook | 3i (Intelligent Imaging Innovations) | https://www.intelligent-imaging.com/slidebook | Confocal microscopy |
Referências
- Weinlich, R., Oberst, A., Beere, H. M., Green, D. R. Necroptosis in development, inflammation and disease. Nature Reviews Molecular Cell Biology. 18 (2), 127-136 (2017).
- Kaiser, W. J., et al. RIP3 mediates the embryonic lethality of caspase-8-deficient mice. Nature. 471 (7338), 368-372 (2011).
- Murphy, J. M., et al. The pseudokinase MLKL mediates necroptosis via a molecular switch mechanism. Immunity. 39 (3), 443-453 (2013).
- Oberst, A., et al. Catalytic activity of the caspase-8-FLIP(L) complex inhibits RIPK3-dependent necrosis. Nature. 471 (7338), 363-367 (2011).
- Zhang, H., et al. Functional complementation between FADD and RIP1 in embryos and lymphocytes. Nature. 471 (7338), 373-376 (2011).
- He, S., et al. Receptor interacting protein kinase-3 determines cellular necrotic response to TNF-alpha. Cell. 137 (6), 1100-1111 (2009).
- Dondelinger, Y., Hulpiau, P., Saeys, Y., Bertrand, M. J. M., Vandenabeele, P. An evolutionary perspective on the necroptotic pathway. Trends in Cell Biology. 26 (10), 721-732 (2016).
- Newton, K., Manning, G. Necroptosis and Inflammation. Annual Review of Biochemistry. 85, 743-763 (2016).
- Kaiser, W. J., Upton, J. W., Mocarski, E. S. Viral modulation of programmed necrosis. Current Opinion in Virology. 3 (3), 296-306 (2013).
- Newton, K., et al. RIPK3 deficiency or catalytically inactive RIPK1 provides greater benefit than MLKL deficiency in mouse models of inflammation and tissue injury. Cell Death & Differentiation. 23 (9), 1565-1576 (2016).
- Kearney, C. J., Martin, S. J. An Inflammatory Perspective on Necroptosis. Molecular Cell. 65 (6), 965-973 (2017).
- Pasparakis, M., Vandenabeele, P. Necroptosis and its role in inflammation. Nature. 517 (7534), 311-320 (2015).
- Sun, L., Wang, X. A new kind of cell suicide: mechanisms and functions of programmed necrosis. Trends in Biochemical Sciences. 39 (12), 587-593 (2014).
- Grootjans, S., Vanden Berghe, T., Vandenabeele, P. Initiation and execution mechanisms of necroptosis: an overview. Cell Death & Differentiation. 24 (7), 1184-1195 (2017).
- Sun, L., et al. Mixed lineage kinase domain-like protein mediates necrosis signaling downstream of RIP3 kinase. Cell. 148 (1-2), 213-227 (2012).
- Wang, H., et al. Mixed lineage kinase domain-like protein MLKL causes necrotic membrane disruption upon phosphorylation by RIP3. Molecular Cell. 54 (1), 133-146 (2014).
- Cai, Z., et al. Plasma membrane translocation of trimerized MLKL protein is required for TNF-induced necroptosis. Nature Cell Biology. 16 (1), 55-65 (2014).
- Chen, X., et al. Translocation of mixed lineage kinase domain-like protein to plasma membrane leads to necrotic cell death. Cell Research. 24 (1), 105-121 (2014).
- Dillon, C. P., et al. RIPK1 blocks early postnatal lethality mediated by caspase-8 and RIPK3. Cell. 157 (5), 1189-1202 (2014).
- Rickard, J. A., et al. RIPK1 regulates RIPK3-MLKL-driven systemic inflammation and emergency hematopoiesis. Cell. 157 (5), 1175-1188 (2014).
- Kaiser, W. J., et al. Toll-like receptor 3-mediated necrosis via TRIF, RIP3, and MLKL. Journal of Biological Chemistry. 288 (43), 31268-31279 (2013).
- Upton, J. W., Kaiser, W. J. DAI Another Way: Necroptotic Control of Viral Infection. Cell Host & Microbe. 21 (3), 290-293 (2017).
- Tanzer, M. C., et al. Necroptosis signalling is tuned by phosphorylation of MLKL residues outside the pseudokinase domain activation loop. Biochemical Journal. 471 (2), 255-265 (2015).
- Quarato, G., et al. Sequential Engagement of Distinct MLKL Phosphatidylinositol-Binding Sites Executes Necroptosis. Molecular Cell. 61 (4), 589-601 (2016).
- Davies, K. A., et al. The brace helices of MLKL mediate interdomain communication and oligomerisation to regulate cell death by necroptosis. Cell Death & Differentiation. , (2018).
- Su, L., et al. A plug release mechanism for membrane permeation by MLKL. Structure. 22 (10), 1489-1500 (2014).
- Dondelinger, Y., et al. MLKL compromises plasma membrane integrity by binding to phosphatidylinositol phosphates. Cell Reports. 7 (4), 971-981 (2014).
- Llambi, F., et al. BOK Is a Non-canonical BCL-2 Family Effector of Apoptosis Regulated by ER-Associated Degradation. Cell. 165 (2), 421-433 (2016).
- Malkani, N., Schmid, J. A. Some secrets of fluorescent proteins: distinct bleaching in various mounting fluids and photoactivation of cyan fluorescent proteins at YFP-excitation. PLoS One. 6 (4), 18586 (2011).
- Perez, A. J., et al. A workflow for the automatic segmentation of organelles in electron microscopy image stacks. Frontiers in Neuroanatomy. 8, 126 (2014).
- Walton, J. Lead aspartate, an en bloc contrast stain particularly useful for ultrastructural enzymology. Journal of Histochemistry & Cytochemistry. 27 (10), 1337-1342 (1979).
- Denk, W., Horstmann, H. Serial block-face scanning electron microscopy to reconstruct three-dimensional tissue nanostructure. PLoS Biology. 2 (11), 329 (2004).
- Rossi, P., Xia, Y., Khanra, N., Veglia, G., Kalodimos, C. G. 15N and 13C- SOFAST-HMQC editing enhances 3D-NOESY sensitivity in highly deuterated, selectively [1H,13C]-labeled proteins. Journal of Biomolecular NMR. 66 (4), 259-271 (2016).
- Keller, R. The computer aided resonance assignment tutorial. Cantina Verlag. , (2004).
- Chen, W., et al. Diverse sequence determinants control human and mouse receptor interacting protein 3 (RIP3) and mixed lineage kinase domain-like (MLKL) interaction in necroptotic signaling. Journal of Biological Chemistry. 288 (23), 16247-16261 (2013).
- Tanzer, M. C., et al. Evolutionary divergence of the necroptosis effector MLKL. Cell Death & Differentiation. 23 (7), 1185-1197 (2016).

