Primary Culture of Porcine Retinal Pigment Epithelial Cells
Summary
Here, an easy-to-follow method to culture primary porcine retinal pigment epithelial cells in vitro is presented.
Abstract
The retinal pigment epithelium (RPE) is a monolayer of polarized pigmented epithelial cells, located between the choroid and neuroretina in the retina. Multiple functions, including phagocytosis, nutrient/metabolite transportation, vitamin A metabolism, etc., are conducted by the RPE on a daily basis. RPE cells are terminally differentiated epithelial cells with little or no regenerative capacity. Loss of RPE cells results in multiple eye diseases leading to visual impairment, such as age-related macular degeneration. Therefore, the establishment of an in vitro culture model of primary RPE cells, which more closely resembles the RPE in vivo than cell lines, is critical for the characteristic and mechanistic studies of RPE cells. Considering the fact that the source of human eyeballs is limited, we create a protocol to culture primary porcine RPE cells. By using this protocol, RPE cells can be easily dissociated from adult porcine eyeballs. Subsequently, these dissociated cells attach to culture dishes/inserts, proliferate to form a confluent monolayer, and quickly re-establish key features of epithelial tissue in vivo within 2 wks. By qRT-PCR, it is demonstrated that primary porcine RPE cells express multiple signature genes at comparable levels with native RPE tissue, while the expressions of most of these genes are lost/highly reduced in human RPE-like cells, ARPE-19. Moreover, the immunofluorescence staining shows the distribution of tight junction, tissue polarity, and cytoskeleton proteins, as well as the presence of RPE65, an isomerase critical for vitamin A metabolism, in cultured primary cells. Altogether, we have developed an easy-to-follow approach to culture primary porcine RPE cells with high purity and native RPE features, which could serve as a good model to understand RPE physiology, study cell toxicities, and facilitate drug screenings.
Introduction
The retinal pigment epithelium (RPE) is located between photoreceptors and choriocapillaris in the outer layer of the retina1 with multiple functions, including forming the blood-retinal barrier, transporting and exchanging nutrients and retinal metabolites, recycling vitamin A to maintain a normal visual cycle, and phagocytosis and clearance of shed photoreceptor outer segments (POSs)2,3. Since POSs require constant self-renewal to generate vision, the RPE cells need to continuously engulf detached POSs to maintain retinal homeostasis4. Therefore, RPE dysfunction results in many blinding eye diseases, such as age-related macular degeneration (AMD)4, retinitis pigmentosa (RP)5, Leber congenital amaurosis6, diabetic retinopathy7, etc. Till now, the exact pathogenesis of most of these diseases remains elusive. As a result, RPE cell culture is established to study RPE cell biology, pathological changes, and underlying mechanisms.
As the simplest model to study cell biology, the culture of RPE cells was started as early as the 1920s8. Although ARPE-19 is widely used as RPE cells, loss of pigmentation, cobblestone morphology, and, especially, the barrier functions in this cell line raise lots of concerns9. In comparison, the culture of primary human RPE cells offers a more realistic scenario for physiological and pathological studies9. However, the relatively limited availability restricts their usage and ethical issues always exist. In addition, several groups used mouse models to culture RPE cells. However, the size of the mouse eye is small, and a single culture usually requires many mice, which is not convenient9. Recently, scientists have developed new methods to use human embryonic stem cells or induced pluripotent stem cells to derive RPE cells. Although this technique has particular potential for the treatment of inherited RPE disorders, it is time-consuming and usually requires several months to generate mature RPE cells10. To overcome these problems, here we introduce an easy-to-follow protocol to isolate and culture high-purity RPE cells in the laboratory routinely. Under suitable culture conditions, these cells can display typical RPE functions and exhibit typical RPE morphologies. Therefore, this culture method can provide a good model to understand RPE physiology, study cytotoxicity, investigate pathological mechanisms of related ocular diseases, and conduct drug screenings.
Protocol
The use of experimental animals complied with the regulations of the Association for Research in Vision and Ophthalmology (ARVO) and was approved by the Ethics Committee of Experimental Animal Management of Xiamen University.
1. Preparation of experimental surgical devices, tissue digestion enzyme, and cell culture buffer
- Prepare the experimental surgical devices, by cleaning and autoclaving two pairs of ophthalmic surgical scissors and forceps the day before the eyeball dissection, and afterward drying the box with surgical devices in a general protocol oven at 65 °C overnight.
- Culture media preparation.
- Prepare DMEM/Basic media supplemented with 10% (v/v) fetal bovine serum, 2 mM L-glutamine, and 1% (v/v) penicillin (100 U/mL) and streptomycin (100 U/mL).
- Prepare DMEM/F12 media supplemented with 1% (v/v) FBS, and 1% (v/v) penicillin (100 U/mL) and streptomycin (100 U/mL).
- Prepare MEM-Nic11, by supplementing MEM alpha with 2 mM L-glutamine, 1% FBS, 1% (v/v) penicillin (100 U/mL) and streptomycin (100 U/mL), 0.1 mM NEAA, 1% (v/v) N1 supplement, taurine (0.25 mg/mL), hydrocortisone (20 ng/mL), triiodo-thyronin (0.013 ng/mL), and 10 mM nicotinamide.
NOTE: To improve cell viability and proliferation, the percentage of FBS can be increased to 20% to neutralize digestion enzymes and seed the dissociated cells. For long-term cell culture, the percentage of FBS can be decreased to as low as 1%12.
- Thaw the tissue digestion enzyme aliquot (0.25% (w/v) Trypsin/EDTA solution supplemented with 0.91 mM EDTA, hereafter referred as Trypsin/EDTA solution) during the experiment.
NOTE: Fresh Trypsin/EDTA solution should be used every time to obtain optimum results. Aliquot 10 mL of fresh Trypsin/EDTA solution into each 15 mL sterile centrifuge tube and freeze the aliquots at -20 °C refrigerator until use. - Dissection solution.
- Sterilize 1x Phosphate Buffered Saline (PBS) (pH 7.2) supplemented with 2% (v/v) penicillin (100 U/mL) and streptomycin (100 U/mL) by filtering the solution through a 0.22 µm syringe filter unit.
- Coat the culture plates and transwell inserts.
- Wash the wells and transwell inserts with 1x PBS. Remove the PBS from wells and transwells with a pipette, and then add 1 mL of fresh 1x PBS to the lower chamber and 600 µL of 10 µg/mL of laminin solution to the upper chamber.
- Incubate the plates/transwell inserts in the cell culture incubator overnight at 37 °C and 5% CO2. Remove the laminin solution from the upper chamber and wash with 1 mL of cold 1x PBS twice before seeding the cells.
2. Dissection of porcine eyeball RPE cells
- Preparation of the instruments.
- Clean the laminar flow hood with 75% ethanol and UV light. Prepare three 50 mL sterile centrifuge tubes containing ~25 mL of 75% ethanol, three 50 mL sterile centrifuge tubes containing ~25 mL of 1x PBS, and three 10 cm sterile cell culture dishes containing ~15 mL of 1x PBS.
NOTE: These settings are usually used for dissecting four porcine eyeballs. Please scale up when more eyeballs are used. To improve the cell viability, precool 1x PBS buffer on ice for at least 30 min. Both 75% ethanol and UV light are necessary to ensure that experimental instruments and cells are not contaminated. Turn on the UV lamp in advance to sterilize the entire operation table for at least 15 min.
- Clean the laminar flow hood with 75% ethanol and UV light. Prepare three 50 mL sterile centrifuge tubes containing ~25 mL of 75% ethanol, three 50 mL sterile centrifuge tubes containing ~25 mL of 1x PBS, and three 10 cm sterile cell culture dishes containing ~15 mL of 1x PBS.
- Clean the porcine eyeballs.
- Obtain fresh eyeballs from a slaughterhouse and keep them on ice until dissection. Soak four porcine eyeballs into ~15 mL of 75% ethanol in a 10 cm Petri dish and use a pair of scissors and forceps to cut off all the residual connective tissues and muscles.
NOTE: Remove as much tissue as possible from outside of the sclera to reduce contaminations. Do not cut the optic nerve at this time to facilitate the transfer of eyeballs during the decontamination and wash step. After this step, all the procedures should be performed in a laminar flow hood.
- Obtain fresh eyeballs from a slaughterhouse and keep them on ice until dissection. Soak four porcine eyeballs into ~15 mL of 75% ethanol in a 10 cm Petri dish and use a pair of scissors and forceps to cut off all the residual connective tissues and muscles.
- Decontaminate the porcine eyeballs by soaking and washing four porcine eyeballs in three 50 mL sterile centrifuge tubes filled with 75% ethanol in a sequential manner; each eyeball is dipped for at least 5 min in each tube. Next, wash the porcine eyeballs in three 50 mL sterile centrifuge tubes filled with 1x PBS in a sequential manner; wash each eyeball for at least 5 min in each tube (Figure 1A). Invert the tubes every minute to wash the eyeballs thoroughly.
- Dissection of porcine eyeballs.
- Move the four porcine eyeballs into a 10 cm sterile cell culture dish containing 1x PBS. Trim the outer surface of each eyeball again to remove the optic nerve and small debris (Figure 1B). Use scissors to make a small cut at the intersection of the limbus and sclera, and then remove the cornea, iris, lens, vitreous body, and neural retina (for detailed structures of these tissues, please refer to Figure 1).
- Transfer the RPE-choroid-sclera complex into a new 10 cm cell culture dish and make four cuts to flat the eyecup in the shape of a four-leaf clover (Figure 1C).
- Perform Trypsin/EDTA solution digestion by placing the four RPE-choroid-sclera complexes into a new 10 cm dish. Pour 20 mL of fresh Trypsin/EDTA solution to merge the RPE-choroid-sclera complexes and put the dish into the cell culture incubator at 37 °C for ~30 min.
NOTE: Use fresh Trypsin/EDTA solution to dissociate the RPE cells. Carefully control the time of Trypsin/EDTA solution digestion, as a longer incubation time (more than 30 min) may increase the contamination of other types of cells.
3. Isolation and culture of porcine eyeball RPE cells
- RPE dissociation.
- Take the dish out of the incubator after 30 min of incubation with Trypsin/EDTA solution and add 20 mL of prewarmed culture media (with 10% FBS) to the dish to neutralize the Trypsin/EDTA solution.
NOTE: At this step, only DMEM/Basic media is used. When cells reach confluency, three culture medias can be used depending on the experimental conditions: DMEM/Basic media, DMEM/F12 media, and MEM-Nic media. - Use a 5 mL transfer pipette to dissociate the RPE cells by gently pipetting several times. Collect cell suspensions into 15 mL centrifuge tubes. Use another 10 mL of fresh culture media (10% FBS) to wash the RPE-choroid-sclera complexes on the dish by gently pipetting to obtain as many RPE cells as possible.
NOTE: Do not triturate the cells too vigorously. Ensure to fill and empty the pipette at a rate of about 3 mL/s. Avoid bubbling the cell suspension.
- Take the dish out of the incubator after 30 min of incubation with Trypsin/EDTA solution and add 20 mL of prewarmed culture media (with 10% FBS) to the dish to neutralize the Trypsin/EDTA solution.
- Collect RPE cells by centrifuging the tubes at 200 x g for 5 min at room temperature. Aspirate the supernatant using a pipette and add another 5 mL of culture media (10% FBS) to resuspend the cells and centrifuge at 200 x g for another 5 min. Decant the supernatant and resuspend the cells with 12 mL of culture media.
NOTE: When aspirating the supernatant, leave about 1 mL of media to avoid the breaking of cell clumps. - Seeding RPE cells.
- Seed ~1-2 x 105 cells/well into 12-well culture plates or transwell inserts. Usually, about 1.5 x 106 RPE cells are obtained from four porcine eyeballs. Change the culture media every 2 days and reduce the serum concentration to 1% when the cells fully cover the surface of the wells/inserts.
NOTE: Do not move the cell culture dish too frequently before cells attach to the culture dish/inserts.
- Seed ~1-2 x 105 cells/well into 12-well culture plates or transwell inserts. Usually, about 1.5 x 106 RPE cells are obtained from four porcine eyeballs. Change the culture media every 2 days and reduce the serum concentration to 1% when the cells fully cover the surface of the wells/inserts.
- Change the culture media every 2 days until the cells are harvested.
NOTE: The concentration of FBS can be reduced after cells reach confluency, and cells can be cultured for up to several months to allow full differentiation and maturation.
4. Characterization of primary porcine RPE cells
- Culture the cells at confluency for 1 or 2 wks, and then harvest the cells for further analysis.
- Harvest cells for quantitative real-time PCR (qRT-PCR) analysis.
- Remove the culture media, wash the cells with 1 mL of cold 1x PBS, and then add 500 µL of RNA extraction solution into each well to collect cell lysate from about 1 x 106 cells.
- Extract mRNA13; approximately 4 µg of RNA is extracted from each sample. Use 100 ng of mRNA to perform reverse transcription14; dilute the cDNA 40-fold for quantitative real-time PCR analysis. Primers are listed in Table 1.
- Harvest cells for immunofluorescence staining.
- Remove the culture media, wash the cells with 1 mL of cold 1x PBS, and then add 500 µL of 4% (w/v) paraformaldehyde in 1x PBS to fix the cells (about 1 x 106 cells/well) for 30 min at room temperature. Then, wash the cells with 1x PBS twice and perform immunofluorescence staining with primary antibodies (1:100 in 1x PBS supplemented with 1% (w/v) bovine serum albumin) at 4 °C overnight and secondary antibodies (1:200 in 1x PBS supplemented with 1% (w/v) bovine serum albumin) at room temperature for 2 h according to online protocols15.
- The immunofluorescence images are acquired by a confocal microscope following the online manual16.
- Harvest cells for Western blot analysis.
- Remove the culture media, wash the cells with cold 1x PBS twice, and then add 80 µL of RIPA buffer (with 1x Protease inhibitor) into each well and use cell scrapers to scratch about 1 x 106 cells off the dishes/inserts.
- Collect the cell lysate from each well/insert into a 1.5 mL microcentrifuge tube. Boil the cell lysates for 10 min and then measure protein concentrations using BCA kits; the protein concentration of each sample is about 2 mg/mL. Use 25 µg of proteins of each sample for Western blot analysis according to online protocols17.
- For Western blot, incubate the membranes in 5 mL of primary antibody solution (1:1,000 dilution of primary antibodies in 1x TBST solution supplemented with 5% (w/v) bovine serum albumin) at 4 °C overnight on a rotating multipurpose shaker.
- Then, incubate the membrane with about 15 mL of anti-rabbit or anti-mouse secondary antibody solution (1:5,000 dilution of secondary antibodies in 1x TBST solution supplemented with 5% (w/v) non-fat milk) at room temperature for 2 h on an orbital shaker, according to the source of primary antibody used.
- Transepithelial resistance (TER) measurement.
- Wash the chopstick electrodes with 70% ethanol and sterile 1x PBS in a sequential way. Then place the shorter end of the electrode in the top chamber and the longer end in the bottom chamber of the transwell inserts and click the button on the epithelial voltmeter for the measurements. Record the TER measurements of each well in triplicates.
Representative Results
The primary porcine RPE (pRPE) cells were cultured in DMEM/Basic media with 10% FBS, and cell morphology under light microscope was photographed at 2 days (Figure 2A), 6 days (Figure 2B) and 10 days (Figure 2C) after seeding. After 1 wk, a confluent monolayer of pigmented pRPE cells with cobblestone morphologies was observed.
To better characterize the primary pRPE cells, primary human RPE cells (hRPE) at Passage 3 (P3)18 and ARPE-19 cells were cultured in DMEM/Basic media with 1% FBS for another wk, and then the total mRNA and protein of the cultured cells, as well as porcine RPE/choroid tissues, were harvested. The expression levels of key characteristic genes19 and protein markers of mature RPE were evaluated by qRT-PCR and Western blot. Compared with ARPE-19 cells, pRPE cells retained significantly higher expression levels of genes functioning in native RPE secretion (Figure 3A(c)), phagocytosis (Figure 3A(i)), transportation (Figure 3A(a),(d)), tight junctions (Figure 3A(j)), barrier formation (Figure 3A(b)), as well as visual cycle (Figure 3A(e),B). However, the expression levels of Krt8 and Krt18, two cytoskeleton marker genes of the RPE, were significantly lower in primary pRPE cells in comparison with primary hRPE and ARPE-19 cells (Figure 3A(g),(h)). Moreover, itgav, which participates in phagocytosis, was lower in primary pRPE cells as well (Figure 3A(f)). Since the expression levels of these three genes in pRPE cells were similar with the porcine RPE/choroid tissue, this may indicate the gene expression differences between the species. Moreover, the expression level of Tyr, which is responsible for melanin production, was highly reduced in both primary pRPE and hRPE cells (Figure 3A(k)), which may explain the loss of pigmentation in long-term cultured cells. The dissimilarities in the qRT-PCR results observed in the data of human pRPE and porcine pRPE cells might be because of the longer storage and higher passage number (P3) of human pRPE, thus indicating a loss of RPE characters on sub-culturing. In addition, Western blot showed that RPE65 protein, which is a key enzyme in the visual cycle featuring RPE cells, was expressed in pRPE cells, while its expression level was significantly reduced in primary hRPE cells and ARPE-19 cells (Figure 3B,C).
To further characterize the polarity and barrier functions of RPE cells, the cultured confluent monolayers of pRPE and ARPE-19 cells were cultured in transwell inserts with DMEM/Basic, DMEM/F12, and MEM-Nic media for 1 wk (Figure 4). Under these culture conditions, Na+-K+-ATPase (Figure 4A–F) and ZO-1 (Figure 4G–L) fluorescent staining results revealed that pRPE cells had higher expression levels of both Na+-K+-ATPase and ZO-1 than ARPE-19 cells. The equal distribution of Na+-K+-ATPase at the apical and basal surface of pRPE cells indicated that a longer culture time was required to restore cell polarities in vitro.
The RPE forms tight junctions near its apical surface to tightly regulate the exchange of metabolites between the inner retina and choroids20. ZO-1 staining suggested that tight junction proteins were normally localized at the plasma membrane of primary pRPE cells with regular cobblestone morphologies, but not of ARPE-19 cells. Among the three culture medias, the best ZO-1 staining patterns were observed in cells cultured in DMEM/Basic, while DMEM/F12 failed to maintain the cobblestone morphology of pRPE cells. A higher TER value serves as an indicator of tight junction formation and better barrier functions of RPE cells21. To confirm the function of tight junctions, transepithelial resistance (TER) was measured. However, only slightly higher TER was detected in comparison with empty transwell inserts (Figure 4M,N).
In the retina, Bruch's membrane exists between the RPE and the choroid. The main components of Bruch's membrane are collagen type IV, proteoglycans, and laminin22. In order to simulate the supporting effect of Bruch's membrane on RPE, laminin was spread on the surface of the transwell membrane to facilitate the maturation of cultured RPE cells. Based on the results obtained from Figure 4, the confluent pRPE cells were cultured on transwell inserts in DMEM/Basic media with 1% FBS for 2 wks. Immunofluorescent staining results showed that RPE-specific proteins RPE65 were expressed in all pRPE cells (Figure 5A,B). Na+-K+-ATPase was distributed on the apical surface of pRPE cells, suggesting the re-establishment of cell polarities (Figure 5C,D). Both ZO-1 staining and TER results indicated that tight junctions were well formed in primary pRPE cells when they were cultured on laminin (Figure 5E,F,H). Figure 5G depicts cells stained with Hoechst dye.
Moreover, Western blot demonstrated that both pRPE and ARPE-19 cells produced growth factors, including pigment epithelium-derived factor (PEDF) and vascular endothelial growth factor (VEGF) (Figure 6A). After confluent monolayers of pRPE were cultured in transwell inserts with DMEM/Basic, DMEM/F12, and MEM-Nic media for 1 wk, culture media from the top and bottom chambers of the transwell inserts were collected, and the secreted VEGF was quantified by ELISA. When DMEM/Basic and MEM-Nic media were used for cell culture, pPRE cells secreted more VEGF than cells in DMEM/F12 media (Figure 6B). However, no differences could be detected in VEGF amounts between the top and bottom chambers of the transwell inserts in all tested conditions (Figure 6B). In addition, after primary pRPE cells were cultured on the inserts coated with laminin and in DMEM/Basic media for 2 wks, the media from the top and bottom chambers were collected. Western blot analysis showed higher levels of PEDF in the top chamber than the bottom chamber, while the protein levels of secreted VEGF were similar at both chambers (Figure 6C). These results further supported the re-establishment of cell polarity when primary pRPE cells were cultured for another 2 wks after they reached confluency on laminin.
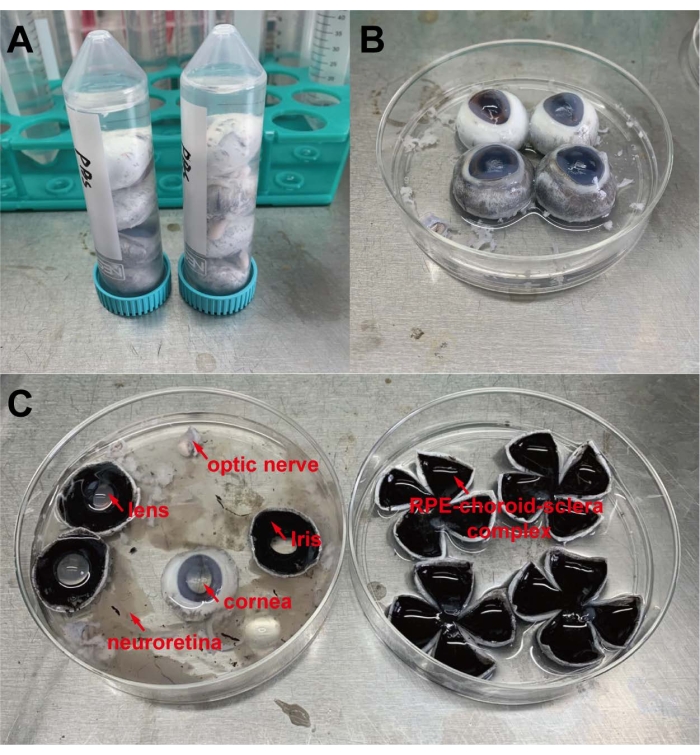

Figure 1: Basic steps of pRPE cell isolation. (A) Wash porcine eyeballs with 1x PBS in 50 mL sterile centrifuge tubes. (B,C) Prepare RPE-choroid-sclera complexes for enzymatic digestion. Please click here to view a larger version of this figure.
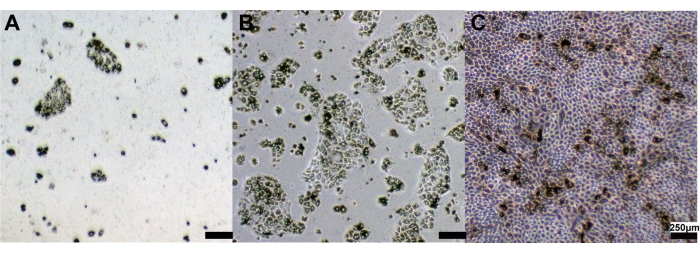
Figure 2: Primary pRPE cell morphologies after cell culture. Representative images of primary pRPE cells at (A) Day 2, (B) Day 6, and (C) Day 10. Scale bar 250 µm. Please click here to view a larger version of this figure.
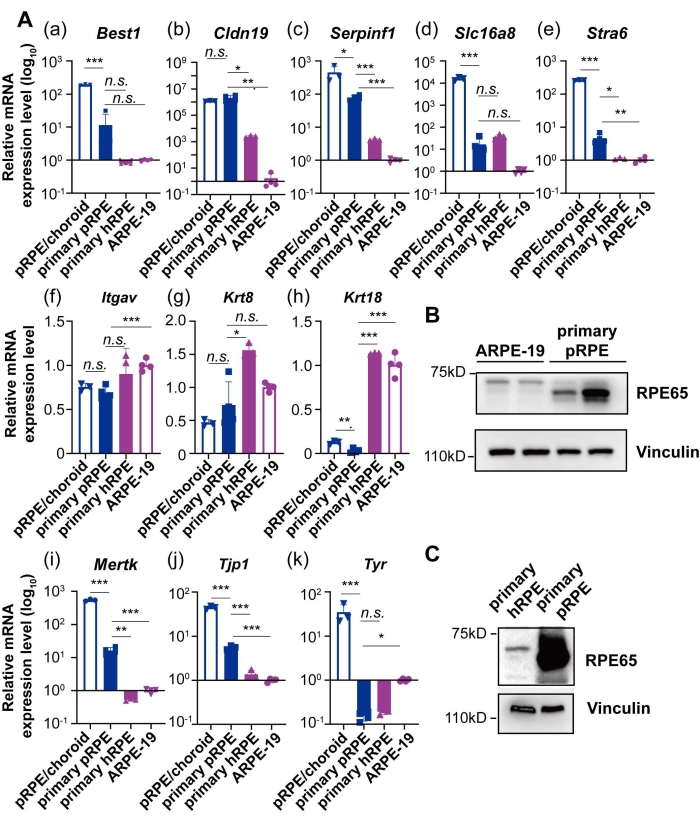
Figure 3: Expression levels of key signature genes and proteins in cultured RPE cells. (A) qRT-PCR analysis of mRNA levels of signature genes in primary pRPE cells, primary hRPE cells, ARPE-19 cells, and pRPE/choroid tissue. Gapdh was used as the house keeping gene for qRT-PCR. Four biological replicates were used for primary pRPE cells and ARPE-19 cells, while three biological replicates were used for primary hRPE cells and pRPE/choroid tissue. The gene expression levels in ARPE-19 cells were set as controls. (B) Western blot analysis of RPE65 proteins in ARPE-19 and pRPE cells. Vinculin was used as a loading control. (C) Western blot analysis of RPE65 proteins in primary hRPE cells and pRPE/choroid tissue. Vinculin was used as a loading control. For comparison, student t-test was used, *p < 0.05, **p < 0.01, ***p < 0.001. Please click here to view a larger version of this figure.
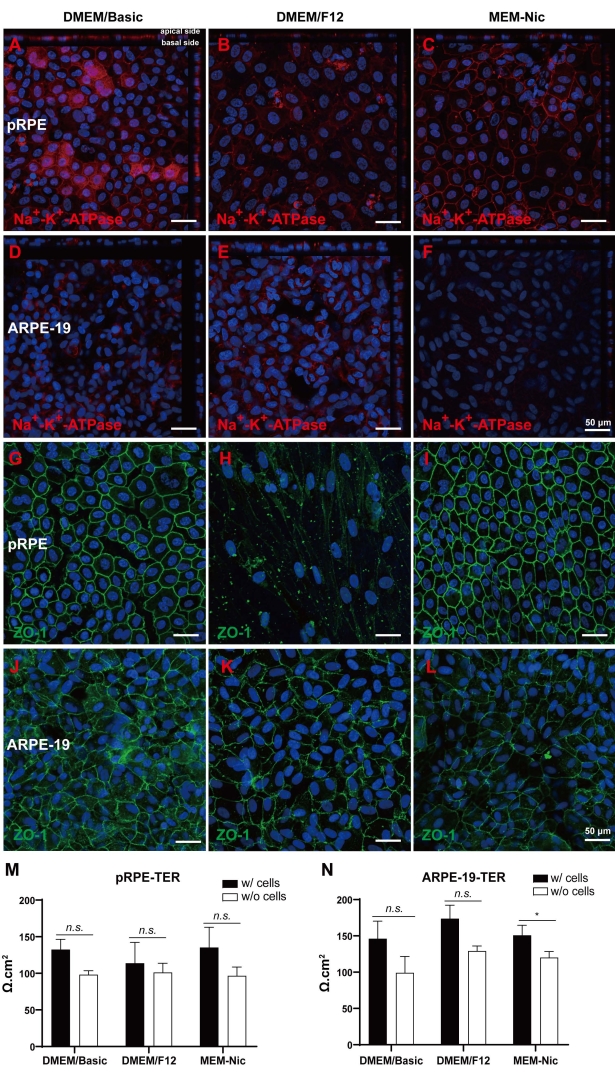
Figure 4: One-wk culture of primary pRPE and ARPE-19 cells in DMEM/Basic, DMEM/F12, and MEM-Nic media with 1% FBS. (A) Na+-K+-ATPase fluorescent staining (Red) of primary pRPE cells in DMEM/Basic, (B) DMEM/F12, and (C) MEM-Nic media. (D) Na+-K+-ATPase fluorescent staining (Red) of ARPE-19 cells in DMEM/Basic media, (E) DMEM/F12 media, and (F) MEM-Nic media. (G) ZO-1 fluorescent staining (Green) of primary pRPE cells in DMEM/Basic media, (H) DMEM/F12 media, and (I) MEM-Nic media. (J) ZO-1 fluorescent staining (Green) in ARPE-19 cells in DMEM/Basic media, (K) DMEM/F12 media, and (L) MEM-Nic media. TER measurements after primary pRPE (M) and ARPE-19 (N) cells were cultured in DMEM/Basic, DMEM/F12, and MEM-Nic media for 1 wk. For each type of cells, data were obtained from at least three different wells. For comparison, student t-test was used, *p < 0.05, **p < 0.01, ***p < 0.001. Scale bar 50 µm. Please click here to view a larger version of this figure.
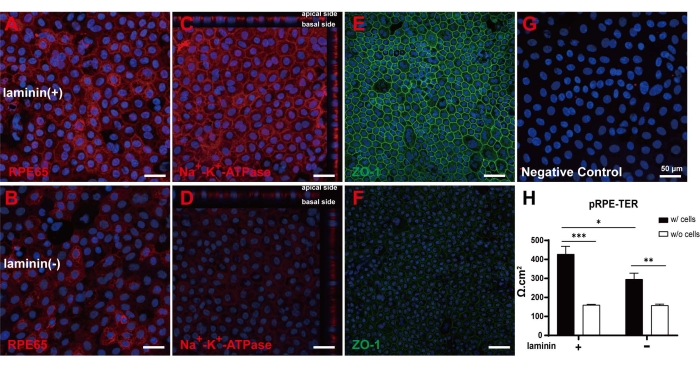
Figure 5: Two-wk culture of pRPE cells in DMEM/Basic media with 1% FBS. (A–F) Cells cultured with or without laminin were stained with RPE65 (A,B, Red), Na+-K+-ATPase (C,D, Red), ZO-1 (E,F, Green), and (G) Hoechst (Blue) . (H) TER measurements in cell sheets cultured with or without laminin. For different treatments, data were obtained from four different wells. For comparison, student t-test was used, *p < 0.05, **p < 0.01, ***p < 0.001. Scale bar = 50 µm. Please click here to view a larger version of this figure.
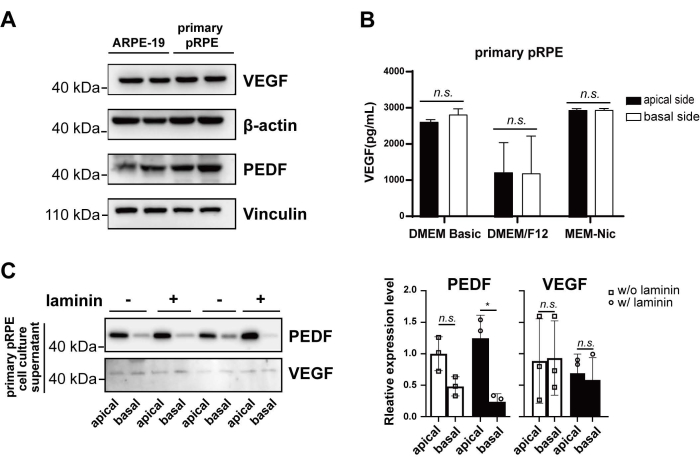
Figure 6: Secretory functions of cultured primary pRPE and ARPE-19 cells. (A) Western blot analysis of VEGF and PEDF in primary pRPE and ARPE-19 cells. (B) Secreted VEGF in the top and bottom chamber of transwell inserts with primary pRPE cells when confluent cells were cultured in DMEM/Basic, DMEM/F12, and MEM-Nic media for 1 wk. (C) Secreted PEDF and VEGF at the top and bottom chamber of transwell inserts with primary pRPE in DMEM/Basic media for 2 wks. For different treatments, data were obtained from three different wells. For comparison, student t-test was used, *p < 0.05, **p < 0.01, ***p < 0.001. Please click here to view a larger version of this figure.
| Homo sapiens | Sus scrofa | |
| Best1 | F: GAAGGCAAGGACGAGCAAG | F: GGACACCTGTATGCCTACGA |
| R: TCCAACTGCTTGTGTTCTGC | R: GGAACGTGAAGAGGGGTACA | |
| Cldn19 | F: GGTGACCCAGGAGTTCTTCA | F: TCGTGACCCAGGAGTTCTTC |
| R: CTGTTGGGTCTCTCTGGCTC | R: GCTGCTGTTGGATCGCTC | |
| Itgav | F: CGCAGTCCCATCTCAAATCC | F: GCTTTCTTCAGGACGGAACA |
| R: GGCCCTGTATAAGATAGCTCGA | R: GAAATGAGCTGACCTTGCCA | |
| Krt8 | F: GGAGCAGATCAAGACCCTCA | F: CCCAGGAGAAGGAGCAGATC |
| R: GCCGCCTAAGGTTGTTGATG | R: ATGTTGTCGATGTTGCTCCG | |
| Krt18 | F: CTTGGAGAAGAAGGGACCCC | F: GCTGATAATCGGAGGCTGGA |
| R: GGCCAGCTCTGTCTCATACT | R: GAAGTCATCAGCAGCGAGAC | |
| Mertk | F: GTGTGCAGCGTTCAGACAAT | F: GCGGCTATTTCTTGGTGGAA |
| R: AAAATGTTGACGGGCTCAGG | R: ACGTAGATGGGGTCAGACAC | |
| Serpinf1 | F: CAGATGAAAGGGAAGCTCGC | F: AAGACGTCGCTGGAGGATTT |
| R: TTAGGGTCCGACATCATGGG | R: GGTCACTTTCAGAGGCAGGA | |
| Slc16a8 | F: GGGTGTCCTCCATCATGCTA | F: CAGTTCGAGGTGCTCATGG |
| R: GTCAGGTAGAGCTCCAGGAG | R: GACAGCCATGAAGACACCAG | |
| Stra6 | F: CTTTGCAGGAAGAAGCTGGG | F: CTAGCCGTGTTGTCGATCCT |
| R: TAAATGGCCGTCCCTGTCAG | R: GACCATGAAGACAGCAGCAG | |
| Tjp1 | F: ACAGGAAAATGACCGAGTTGC | F: AAGACTTGTCAGCTCAGCCA |
| R: TGGTTCAGGATCAGGACGAC | R: CCAGCATCTCGAGGTTCACT | |
| Tyr | F: ACTCAGCCCAGCATCATTCT | F: ATCTACTCAGCCCAGCATCC |
| R: ACATCAGCTGAAGAGGGGAG | R: GAGCCTTGGAGTCCTGGATT | |
| Gapdh | F: CAGCCTCAAGATCATCAGCA | F: CATCCTGGGCTACACTGAGG |
| R: ATGATGTTCTGGAGAGCCCC | R: GGGGCTCTTACTCCTTGGAG |
Table 1: Primers for qRT-PCR.
Discussion
Here, a detailed and optimized protocol for the isolation, culture, and characterization of RPE cells from porcine eyeballs, which generates a good model for in vitro characterization of RPE cells and RPE-related disorder studies has been described. Methods for the isolation of the RPE from human, mouse, and rat eyes have been described previously23,24,25. However, it is difficult to obtain human eyeballs in some laboratories, and it usually raises ethical issues. The RPE tissues of mice and rats are relatively small, the cells are easily damaged during separation, and only a few cells can be obtained from each animal. In contrast, porcine eyeballs are much easier to obtain and handle, which could stably generate a relatively large number of primary cells from a single eyeball. In some studies, tweezers are used to mechanically separate RPE, which can easily lead to cell death. Therefore, hyaluronic acid, dispase, and Trypsin/EDTA solution have been employed to dissociate RPE cells previously. Previous studies have shown that Trypsin/EDTA solution has the ability to help RPE detach from the choroid21. Therefore, Trypsin/EDTA solution was used to digest the porcine tissues, which generate optimum results. After Trypsin/EDTA neuralization, RPE tissues could be dissociated into single cell suspension by gently pipetting with a pipette tip. One trick is that fresh Trypsin/EDTA solution is recommended for each experiment to fully dissociate RPE cells. Due to the large area of the RPE tissues of porcine eyeballs, the half bowl-shaped RPE-choroid-sclera complex needs to be cut into the shape of a four-leaf clover, which is conducive to the full contact between the RPE tissues and the Trypsin/EDTA solution so as to facilitate their separation from the choroid. However, the digestion time should not exceed half an hour. Within this time, other tissues except RPE will not be digested, which could effectively reduce the contaminations from other types of cells.
Laminin was spread on the surface of the transwell membrane to simulate the supportive effect of Bruch's membrane on RPE. The results showed that laminin was beneficial for the growth of pRPE cells, which stimulated the expression of transporter and tight junction proteins. However, the results also indicated that a longer culture time, of about 2 wks, was required for primary pRPE cells to restore certain features of native RPE tissues, including cell polarity and tight junctions.
One shortcoming for primary RPE culture is that it is difficult to maintain visual cycle enzyme expressions in vitro. Therefore, it is critical to examine whether cultured RPE cells could express RPE-specific proteins RPE65. Compared with ARPE-19 cells, pRPE cells expressed significantly more RPE65 protein. In contrast, only a weak band of RPE65 protein was observed in ARPE-19 cell lysates in Western blot. It is interesting to observe that the molecular weight of RPE65 was slightly different between human and pigs (Figure 3B,C). The loss of RPE65 expression in cultured RPE cells remains as a mystery. Even in cultured primary cells, RPE65 was gradually lost with passages (data not shown). Till now, how to maintain the expression of RPE65 in vitro requires further investigation, considering the fact that visual cycle is a critical function for native RPE tissues. Secondly, a major limitation of this protocol is that in vitro cultured RPE cells cannot be passaged. With prolonged culture, pRPE cells tend to lose their hexagonal shape and TER as well as pigmentation, so that only passage 0 cells can be used. Another limitation of this culture protocol is that genetic mutations, which lead to inherited retinal diseases, are difficult to reproduce in these cells. Therefore, virus-mediated gene editing may be employed to study hereditary RPE-disorders such as Leger congenital amaurosis in the future.
Declarações
The authors have nothing to disclose.
Acknowledgements
The authors would like to show their gratitude and respect to all animals contributing their cells in this study. This study was supported in part by grants from the National Key R&D Program of China (2019YFA0111200, Yi Liao & Yuan Gao and Grant nos. 2018YFA0107301, Wei Li). The authors thank Jingru Huang and Xiang You from Central Lab, School of Medicine, Xiamen University for technical support in confocal imaging.
Materials
| ARPE-19 cells | CCTCC | GDC0323 | |
| Bovine serum albumin | Yeasen | 36101ES60 | |
| Confocal microscopy | Zeiss | LSM 880 with Airyscan | |
| ChemiDoc Touch | Bio-Rad | 1708370 | |
| Cell scraper | Sangon | F619301 | |
| 10 cm culture dish | NEST | 121621EH01 | |
| 12-well culture plate | NEST | 29821075P | |
| DMEM F12 Medium | Gibco | C11330500BT | |
| DMEM basic Medium | Gibco | C11995500BT | |
| EVOM2 | World Precision Instruments | EVOM2 | For TER measurement |
| Fetal bovine serum | ExCell Bio | FSP500 | |
| Goat anti-Rabbit IgG (H+L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor 488 | ThermoFisher Scientific | A-11034 | |
| Goat anti-Rabbit IgG (H+L) Cross-Adsorbed Secondary Antibody, Alexa Fluor 594 | ThermoFisher Scientific | A-11012 | |
| Goat anti Mouse IgG (H/L):HRP | Bio-Rad | 0300-0108P | |
| Goat anti Rabbit IgG (H/L):HRP | Bio-Rad | 5196-2504 | |
| hydrocortisone | MCE | HY-N0583/CS-2226 | |
| Hoechst 33342 solution (20 mM) | ThermoFisher Scientific | 62249 | |
| LightCycler 96 Instrument | Roche | 5815916001 | |
| Liothyronine | MCE | HY-A0070A/CS-4141 | |
| laminin | Sigma-Aldrich | L2020-1MG | |
| MEM(1X)+GlutaMAX Medium | Gibco | 10566-016 | |
| MEM NEAA(100X) | Gibco | 11140-050 | |
| Millex-GP syringe filter unit | Millipore | SLGPR33RB | |
| N1 | Sigma-Aldrich | SLCF4683 | |
| NcmECL Ultra | New Cell&Molecular Biotech | P10300 | |
| Non-fat Powdered Milk | Solarbio | D8340 | |
| Nicotinamide | SparkJade | SJ-MV0061 | |
| Na+-K+ ATPase antibody | Abcam | ab76020 | Recognize both human and porcine proteins |
| PAGE Gel Fast Preparation Kit(10%) | Epizyme | PG112 | |
| primary Human RPE cells | – | – | Generous gift from Shoubi Wang lab |
| Pierce BCA Protein Assay Kit | ThermoFisher Scientific | 23225 | |
| Prism | GraphPad by Dotmatics | version 8.0 | |
| Protease Inhibitor Cocktails | APExBIO | K1024 | |
| PRE65 antibody | Proteintech | 17939-1-AP | Recognize both human and porcine proteins |
| PEDF antibody | Santa Cruz Biotechnology | sc-390172 | Recognize both human and porcine proteins |
| 100 x penicillin/streptomycin | Biological Industries | 03-031-1BCS | |
| Phosphate buffered saline (PBS) | RARBIO | RA-9005 | |
| ReverTra Ace qPCR RT Master Mix | Toyobo | FSQ-201 | |
| RIPA buffer | ThermoFisher Scientific | 89900 | |
| 15 mL sterile centrifuge tubes | NEST | 601052 | |
| 50 mL sterile centrifuge tubes | NEST | 602052 | |
| 0.25% Trypsin-EDTA | Gibco | 25200-056 | |
| Taurine | Damas-beta | 107-35-7 | |
| Trizol | Thermo-Fisher | 15596026 | RNA extraction solution |
| TB Green Fast qPCR Mix | Takara | RR430A | |
| 12-well transwell inserts | Labselect | 14212 | |
| VEGF antibody | Proteintech | 19003-1-AP | Recognize both human and porcine proteins |
| VEGF ELISA kit | Novusbio | VAL106 | |
| ZO-1 antibody | ABclonal | A0659 | Recognize both human and porcine proteins |
Referências
- Tan, L. X., Germer, C. J., La Cunza, N., Lakkaraju, A. Complement activation, lipid metabolism, and mitochondrial injury: Converging pathways in age-related macular degeneration. Redox Biology. 37, 101781 (2020).
- Caceres, P. S., Rodriguez-Boulan, E. Retinal pigment epithelium polarity in health and blinding diseases. Current Opinion in Cell Biology. 62, 37-45 (2020).
- Lakkaraju, A., et al. The cell biology of the retinal pigment epithelium. Progress in Retinal and Eye Research. 78, 100846 (2020).
- Somasundaran, S., Constable, I. J., Mellough, C. B., Carvalho, L. S. Retinal pigment epithelium and age-related macular degeneration: A review of major disease mechanisms. Clinical & Experimental Ophthalmology. 48 (8), 1043-1056 (2020).
- Ducloyer, J. B., Le Meur, G., Cronin, T., Adjali, O., Weber, M. Gene therapy for retinitis pigmentosa. Medecine Sciences. 36 (6-7), 607-615 (2020).
- den Hollander, A. I., Roepman, R., Koenekoop, R. K., Cremers, F. P. Leber congenital amaurosis: genes, proteins and disease mechanisms. Progress in Retinal and Eye Research. 27 (4), 391-419 (2008).
- Samuels, I. S., Bell, B. A., Pereira, A., Saxon, J., Peachey, N. S. Early retinal pigment epithelium dysfunction is concomitant with hyperglycemia in mouse models of type 1 and type 2 diabetes. Journal of Neurophysiology. 113 (4), 1085-1099 (2015).
- Smith, D. T. Melanin pigment in the pigmented epithelium of the retina of the embryo chick eye studied in vivo and in vino. The Anatomical Record. 18, 260-261 (1920).
- Schnichels, S., et al. Retina in a dish: Cell cultures, retinal explants and animal models for common diseases of the retina. Progress in Retinal and Eye Research. 81, 100880 (2021).
- D’Antonio-Chronowska, A., D’Antonio, M., Frazer, K. A. In vitro differentiation of human iPSC-derived retinal pigment epithelium cells (iPSC-RPE). Bio-Protocol. 9 (24), 3469 (2019).
- Hazim, R. A., Volland, S., Yen, A., Burgess, B. L., Williams, D. S. Rapid differentiation of the human RPE cell line, ARPE-19, induced by nicotinamide. Experimental Eye Research. 179, 18-24 (2019).
- Dunn, K. C., Aotaki-Keen, A. E., Putkey, F. R., Hjelmeland, L. M. ARPE-19, a human retinal pigment epithelial cell line with differentiated properties. Experimental Eye Research. 62 (2), 155-169 (1996).
- . Scientific, T Available from: https://www.thermofisher.cn/document-connect/document (2022)
- . Toyota Available from: https://www.toyoboglobal.com/seihin/xr/likescience/support/manual/FSQ-201.pdf (2022)
- . Abcam Available from: https://www.abcam.cn/protocols/immunocytochemistry-immunofluorescence-protocol (2022)
- . Zeiss Available from: https://www.zeiss.com/microscopy/en/products/software/zeiss-zen-lite.html#manuals (2022)
- . Cell Signal Technology Available from: https://www.cellsignal.cn/learn-and-support/protocols/protocol-western (2022)
- Wang, S., et al. Reversed senescence of retinal pigment epithelial cell by coculture with embryonic stem cell via the TGFbeta and PI3K pathways. Frontiers in Cell and Developmental Biology. 8, 588050 (2020).
- Pfeffer, B. A., Philp, N. J. Cell culture of retinal pigment epithelium: Special Issue. Experimental Eye Research. 126, 1-4 (2014).
- Lehmann, G. L., Benedicto, I., Philp, N. J., Rodriguez-Boulan, E. Plasma membrane protein polarity and trafficking in RPE cells: past, present and future. Experimental Eye Research. 126, 5-15 (2014).
- Anderson, J. M., Van Itallie, C. M. Physiology and function of the tight junction. Cold Spring Harbor Perspectives in Biology. 1 (2), 002584 (2009).
- Nita, M., Strzalka-Mrozik, B., Grzybowski, A., Mazurek, U., Romaniuk, W. Age-related macular degeneration and changes in the extracellular matrix. Medical Science Monitor. 20, 1003-1016 (2014).
- Fernandez-Godino, R., Garland, D. L., Pierce, E. A. Isolation, culture and characterization of primary mouse RPE cells. Nature Protocols. 11 (7), 1206-1218 (2016).
- Langenfeld, A., Julien, S., Schraermeyer, U. An improved method for the isolation and culture of retinal pigment epithelial cells from adult rats. Graefe’s Archive for Clinical and Experimental Ophthalmology. 253 (9), 1493-1502 (2015).
- Sonoda, S. A protocol for the culture and differentiation of highly polarized human retinal pigment epithelial cells. Nature Protocols. 4 (5), 662-673 (2009).

