IDG-SW3 Cell Culture in a Three-Dimensional Extracellular Matrix
Summary
Here, we present a protocol for culturing IDG-SW3 cells in a three-dimensional (3D) extracellular matrix.
Abstract
Osteocytes are considered to be nonproliferative cells that are terminally differentiated from osteoblasts. Osteoblasts embedded in the bone extracellular matrix (osteoid) express the Pdpn gene to form cellular dendrites and transform into preosteocytes. Later, preosteocytes express the Dmp1 gene to promote matrix mineralization and thereby transform into mature osteocytes.This process is called osteocytogenesis. IDG-SW3 is a well-known cell line for in vitro studies of osteocytogenesis. Many previous methods have used collagen I as the main or the only component of the culturing matrix. However, in addition to collagen I, the osteoid also contains a ground substance, which is an important component in promoting cellular growth, adhesion, and migration. In addition, the matrix substance is transparent, which increases the transparency of the collagen I-formed gel and, thus, aids the exploration of dendrite formation through imaging techniques. Thus, this paper details a protocol to establish a 3D gel using an extracellular matrix along with collagen I for IDG-SW3 survival. In this work, dendrite formation and gene expression were analyzed during osteocytogenesis. After 7 days of osteogenic culture, an extensive dendrite network was clearly observed under a fluorescence confocal microscope. Real-time PCR showed that the mRNA levels of Pdpn and Dmp1 continually increased for 3 weeks. At week 4, the stereomicroscope revealed an opaque gel filled with mineral particles, consistent with the X-ray fluorescence (XRF) assay. These results indicate that this culture matrix successfully facilitates the transition from osteoblasts to mature osteocytes.
Introduction
Osteocytes are terminally differentiated cells derived from osteoblasts1,2. Once the osteoblast is buried by the osteoid, it undergoes osteocytogenesis and expresses the Pdpn gene to form preosteocytes, the Dmp1gene to mineralize the osteoid, and the Sost and Fgf23 genes to function as a mature osteocyte in bone tissue3. Here, a 3D culturing system is introduced to identify dendrite extension and marker gene expression in the osteocytogenesis process.
IDG-SW3 cells are an immortalized primary cell line derived from transgenic mice and can expand or replicate osteoblast to late osteocyte differentiation when cultured in different media4. Compared with MLO-A5, MLO-Y4, and other cell lines, the expression profile of functional proteins, the ability to perform calcium salt deposition, and the responses to various hormones in IDG-SW3 cells are more likely to be the same as those of primary osteocytes in the bone tissue4.
Compared to 2D systems, 3D culturing systems are more capable of mimicking the in vivo cellular growth environment, including the nutrient gradient, low mechanical stiffness, and surrounding mechanical range (Table 1). Most of the previous methods for culturing osteoblastic cells in a 3D system used collagen I as the unique component in formulations4,5,6, because collagen I fibers serve as the site of calcium and phosphorus deposition. However, an indispensable constituent in the osteoid, the extracellular matrix, contains a large group of cellular factors that promote cellular growth, adhesion, and migration7,8 and is transparent and convenient for imaging observation. Thus, this protocol uses Matrigel (hereafter referred to as basement membrane matrix) as a secondary component for the osteocytogenesis study.
Protocol
This protocol is suitable for culturing cells in four wells of 24-well plates. If preparing multiple samples or plates, the amounts of the reagents should be increased accordingly.
1. Preparation of the collagen I mixture
NOTE: Collagen I and basement membrane matrix gel quickly at room temperature. Therefore, collagen should be handled on ice (2 °C to 8 °C). All the tips and tubes used must be prechilled unless otherwise indicated. All the procedures should be performed in a safety hood.
- Place all the reagents and centrifuge tubes on ice.
- Slowly pipette 0.48 mL of collagen I (5 mg/mL) and 0.1 mL of 10x minimum essential medium (MEM) (with phenol red) into a sterile 15 mL centrifuge tube kept on ice. Mix well by pipetting up and down gently.
- According to the color palette of the phenol red indicator, use an appropriate volume of 7.5% (w/v) NaHCO3 (approximately 0.2 mL) to adjust the phenol red indicator to orange, which indicates that the pH value is in the range of 7.0-7.4.
NOTE: Depending on the pH of the medium, a volume of NaHCO3 is used to bring the pH to approximately 7.0-7.4. Additionally, NaOH is used for pH neutralization. - Bring the final volume of the mixture up to 1 mL by adding an appropriate volume of ddH2O. Mix well by pipetting gently up and down to prepare for use.
NOTE: The final volume of ddH2O will depend on the amount of NaHCO3 or NaOH used in step 1.3.
2. Preparation of the cell-matrix mixture
- Slowly pipette 0.9 mL of basement membrane matrix into a new 15 mL centrifuge tube on the ice.
- Culture the IDG-SW3 cells in a T25 flask with 4 mL of complete medium (alpha-MEM containing 10% fetal bovine serum [FBS]) supplemented with 50 U/mL INF-γ at 33 °C.
- Once the IDG-SW3 cells reach 90% confluency, remove the medium, and wash the cells with phosphate-buffered saline (PBS, pH 7.4 throughout the protocol) with a pipette. Digest the cells with 0.5 mL of 0.25% trypsin-ethylenediaminetetraacetic acid (EDTA) at 37 °C for 30 s.
- Inactivate the trypsin by adding 3.5 mL of complete medium (alpha-MEM containing 10% fetal bovine serum [FBS]). Transfer the 4 mL cell suspension to a 15 mL centrifuge tube.
- Spin down the cell suspension at 300 × g for 5 min at 4 °C. Discard the supernatant and resuspend the cell pellet in 1 mL of the complete medium on ice.
- Count the cells using a hemocytometer and adjust the final cell density with the complete medium to 4 × 105 cells/mL. Add 0.1 mL of the prepared cell suspension to 0.9 mL of the extracellular matrix from step 2.1. Mix well by pipetting gently up and down.
NOTE: If a cell-free control is needed, add 0.1 mL of the complete medium to 0.9 mL of extracellular matrix as a negative control.
3. Plating the cell-gel mixture
- Gently, mix well the two mixtures from step 2.6 and step 1.4 to 2 mL by pipetting up and down on the ice. Pipette 0.5 mL of the final mixture into each well (four wells of a 24-well plate). Incubate at 37 °C for 1 h to form a cell-gel mixture.
- Add 0.5 mL of osteogenic medium (complete medium containing 50 µg/mL ascorbic acid and 4 mM β-glycerophosphate, also known as osteogenic induction medium)4 to the cell-gel mixture in each well to start the osteogenic differentiation at 37 °C. Consider this as Day 0.
- Every 2 days, replace half the medium with fresh osteogenic medium maintained at 37 °C. Continue the culturing for 35 days.
4. Identifying the cell viability and cellular dendrites using a confocal microscope
- Cell staining
NOTE: Calcein acetoxymethyl ester (calcein AM) is a cell-permeant dye that can be used to determine cell viability. Ethidium homodimer-I (EthD-1) does not cross the membrane of intact/viable cells9. Thus, calcein AM/EthD-1 can be used to identify cell viability and cellular dendrites.- On Day 1 and Day 7 of the culture, remove the calcein AM stock solution (4 mM in DMSO) and EthD-1 stock solution (2 mM in DMSO/H2O 1:4 [v/v]) from the freezer, and allow them to warm to room temperature.
NOTE: The mineralized matrix has a strong autofluorescence signal, so the preosteocyte stage without Dmp1 expression within 7 days of culturing4 is more suitable for the observation of cell staining. - Add 4 µL of the 2 mM EthD-1 stock solution and 5 µL of the 4 mM calcein AM stock solution to 2 mL of Dulbecco's phosphate-buffered saline (D-PBS), and mix well. This gives approximately 2 µM calcein AM and 4 µM EthD-1 as working solutions.
- Pipette 0.5 mL of the working solution into the wells in the 24-well plate. Incubate for 30-45 min at room temperature to stain the cell-gel matrix.
- After incubation, add approximately 0.5 mL of fresh D-PBS to a new 35 mm glass-bottom culture dish. Cover the dish to prevent contamination or drying of the samples.
- Using elbow-tipped forceps, carefully transfer the stained cell-gel matrix from step 4.1.3 into the 35 mm culture dish from step 4.1.4. Avoid damaging or shearing the cell-gel matrix.
- View the labeled cells under a laser confocal fluorescence microscope.
- On Day 1 and Day 7 of the culture, remove the calcein AM stock solution (4 mM in DMSO) and EthD-1 stock solution (2 mM in DMSO/H2O 1:4 [v/v]) from the freezer, and allow them to warm to room temperature.
- Microscope scanning set
- Place the 35 mm dish on the stage. Select the regions of the cell-gel matrix to be scanned through the eyepiece using a 10x objective. Higher-magnification objectives are not recommended due to the size of the extended dendrites.
- Select the optical filters. Calcein can be viewed as green with a standard fluorescein bandpass filter, and EthD-1 can be viewed as red with filters for propidium iodide or Texas Red dye. Select the "line mode" for scanning and set the "pinhole" to 2 µm.
- Select a data depth of 8 bit data depth and an image resolution of 1,024 pixels x 1,024 pixels.
- Image using sequential line scanning with the optimal laser and detector settings (keep a minimal laser energy to avoid potential photobleaching).
NOTE: The optimal detector setting depends on the final fluorescence signals.
- Collecting a "z-stack" of images
- According to the green channel signal, define the top and bottom positions of the cell-gel matrix to be scanned by focusing through the sample while continuously scanning, .
- Once the top and bottom of the sample have been specified, select the desired scanning frame. Start scanning. The sample will be scanned from the top to the bottom with the selected settings from step 4.2, generating a gallery of images.
- Cell viability identification.
- Select one slice from step 4.3. The green and red channels indicate live and dead cells, respectively.
- Cell dendrite identification.
- Select the single green channel to generate 3D reconstructions with the image acquisition software. The depth information is displayed as a series of rainbow pseudocolor images. The dendrites located at the bottom of the gel are displayed in red, whereas those at the top are displayed in blue.
5. Identifying appearance using a stereomicroscope
- On Day 1, Day 7, Day 21, and Day 35 of the culture, remove the culturing medium, and wash the gels in the plate twice with D-PBS. Add 0.5 mL of 4% (v/v) paraformaldehyde in D-PBS to the well to fix the cell-gel matrix in the plate for 10 min at room temperature.
- Remove the paraformaldehyde in the well and wash the cell-gel matrix two more times with D-PBS in the plate. Leave 0.5 mL of D-PBS in each well for imaging.
- Place the plate on the stage and select the optimal position through the eyepiece using a 0.5x objective. Image the full-field view of the cell-gel matrix in the plate individually using "automatic exposure" under the bright field.
6. Identifying mineral deposition with an XRF assay
- On Day 35 of the culture, check whether the liquid nitrogen is sufficient. Start the XRF system. Open the sample room and transfer the gels with elbow-tipped forceps from the plate to the middle of the sample stage of the instrument. Close the sample room and wait for 30 min to allow the instrument to cool before use.
- Set the "Exposure time" to 35 ms, the "Spectrum range" to 0-40 keV, and the "Electric current" to 770 µA for acquisition setup.
- Choose three to five scanning sites for analysis by moving the sample stage.
- Select the elements (Ca and P) for detection. Start the detection and export the results.
NOTE: Ca and P are the most abundant elements in hydroxyapatite.
7. Identifying functional gene expression
- On Day 1, Day 7, Day 21, Day 28, and Day 35 of the culture, remove the culturing medium and wash the cell-gel matrix twice with ice-cold D-PBS in the plate.
- Transfer the gels with elbow-tipped forceps from the plate to a 2.0 mL tube (according to the culture time, the cell-gel matrix will gradually shrink to approximately 0.1 mL on Day 35). Make sure one gel is placed in one tube. Add 1 mL of phenol-chloroform and two sterile stainless-steel beating beads (4 mm, as a lysing matrix) into each tube. Place the tubes symmetrically into the prechilled sample slot of the mechanical beating homogenizer.
- Set the beating to a frequency of 55 Hz and an amplitude of 1 cm in each 1 min of mechanical beating with a 10 s pause, with six cycles in total. Start the bead-beating.
- Extract the total RNA from the cell-gel matrix using the phenol-chloroform-isoamylachohol method. Reverse-transcribe 2 µg of total RNA to cDNA with random hexamer primers.
- Design primers targeting β-actin, Pdpn, Dmp1, Sost, and Fgf23 for real-time PCR (Table 2). β-Actin is the housekeeping gene used for normalization. The Pdpn10 and Dmp111 genes are mainly expressed in pre-osteocytes and mineralized osteocytes, whereas Sost12 and Fgf2313 are mainly expressed in mature osteocytes.
- Place the tube on a thermocycler with the heated lid set to 105 °C and perform PCR amplification using the following PCR cycling settings: 95 °C for 5 s and 60 °C for 30 s for 35 cycles with an initial denaturation step at 95 °C for 5 min and a final extension step at 60 °C for 30 s. Analyze the fold changes with the ΔΔCt method.
Representative Results
After live/dead cell staining, the cells were visualized using a confocal laser microscope. All the cells were calcein AM-positive (green color), and there were almost no EthD-1-positive cells (red color) in the field, indicating that the gel system made by this method is highly suitable for osteocytogenesis (Figure 1A, left). To better determine the spatial distribution of the cells, a pseudocolor image was chosen to display the cell dendrites at different depths of the gel; red shows the dendrites at the bottom of the gel, and blue shows the dendrites at the top. The results indicated that the IDG-SW3 cells grew well in this cell-gel matrix, and the cellular dendrites gradually extended into a network in the osteogenic medium on Day 7 (Figure 1A, right, and Figure 1B).
Under a stereomicroscope, the transparency of the gel matrix continued to decline until it became opaque at Day 35, unlike the cell-free gel matrix (Figure 2A). The XRF spectrum of the opaque gel at Day 35 indicated that the gel was completely filled with calcium and phosphorus deposits (Figure 2B).
On Day 1, Day 7, Day 21, Day 28, and Day 35 of the culture, the expression of several marker genes was analyzed by real-time PCR. The results showed that the mRNA levels of Pdpn and Dmp1 continually increased from Day 1 until Day 21, whereas the mRNA level of Pdpn decreased after Day 21 (Figure 3). The mRNA levels of Fgf23 and Sost continually increased during all stages (Figure 3).
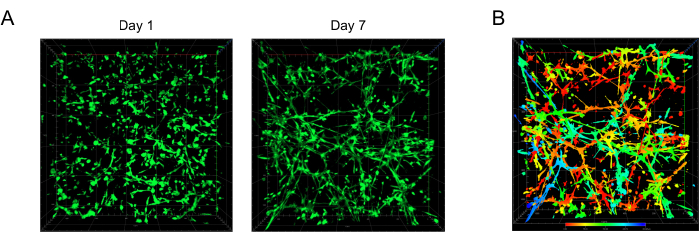
Figure 1: The cellular dendrite network of IDG-SW3 visualized by confocal microscopy. (A) Representative images of a partial area of the extensive dendrite network in the gel. The green color indicates the live cells and the extended cellular dendrite network. (B) Pseudocolor of the image in A (right, Day 7). The red (blue) color indicates the dendrites located at the bottom (top) of the gel. Scale bar = 200 µm. Please click here to view a larger version of this figure.
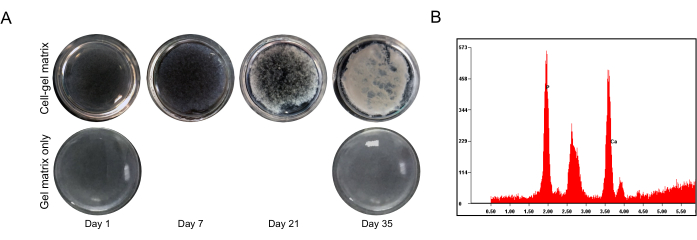
Figure 2: Mineral deposition of the IDG-SW3 cells after osteogenic culturing. (A) Representative full-field images of the gel with (top) and without (bottom) cells in a 24-well plate at the indicated times, as visualized by a stereomicroscope. (B) The calcium and phosphorus content in the cell-gel matrix on Day 35, as analyzed by the XRF assay. Please click here to view a larger version of this figure.
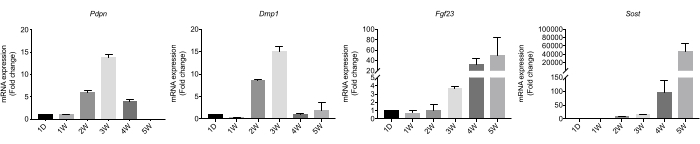
Figure 3: The functional gene expression of the IDG-SW3 cells after osteogenic culturing. Changes in the functional genes in the IDG-SW3 cells cultured with an osteogenic medium at the indicated times. Abbreviations: D = day; W = week. Data are represented as the mean ± SEM. Fold changes were analyzed with the Ct method, referring to the β-actin gene. Please click here to view a larger version of this figure.
| 2D | 3D | |
| Difficulty of preparation | easy | medium |
| Gradient of nutrient | absent | present |
| Arrangement of collagen | monolayer | multiaxial discrete |
| Mechanics stiffness of ECM | high | low |
| Mechanical range of ECM | unilateral | surround |
| Cell motility | flat surface | free |
| Intercellular communication | insufficient | sufficient |
Table 1: Comparison between 2D and 3D culture systems for IDG-SW3 cells.
| Names | Sequences (5’- 3’) |
| Pdpn-for | GGAGGGCTTAATGAATCTACTGG |
| Pdpn-rev | GGTTGTACTCTCGTGTTCTCTG |
| Dmp1-for | CCCAGTTGCCAGATACCAC |
| Dmp1-rev | CACTATTTGCCTGTCCCTCTG |
| Sost-for | ACAACCAGACCATGAACCG |
| Sost-rev | CAGGAAGCGGGTGTAGTG |
| Fgf23-for | GGTGATAACAGGAGCCATGAC |
| Fgf23-rev | TGCTTCTGCGACAAGTAGAC |
| β-actin-for | ACCTTCTACAATGAGCTGCG |
| β-actin-rev | CTGGATGGCTACGTACATGG |
Table 2: Primers used in the real-time PCR assay.
Discussion
A critical point in this protocol is that steps 1 and step 2 must be performed on ice to prevent spontaneous coagulation. In this method, the final concentration of collagen I was 1.2 mg/mL. Thus, an optimal ddH2O volume should be calculated to match the different collagens from various manufacturers.
In vivo, osteocytogenesis involves a polar movement of osteoblasts from the surface to the interior of trabecular bone14. This protocol frees cells from growth in the matrix without directional polarity. Of course, seeding IDG-SW3 cells on the cell-free gel matrix surface is optional. However, the collagen concentration provided in this protocol is not enough to form a theoretically flat surface for osteoblast infiltration. Different ratios of collagen I and extracellular matrix should be tested for this purpose.
Compared with that of the osteoid, the stiffness of this gel is far from sufficient. Some articles report increasing the strength of gels by adding several chemical materials to better mimic the stiffness of the osteoid in vivo6,15. The benefit of this method is that the gel formed can provide a medium with good transparency for imaging and tracing cells in situ rather than using histological methods15. Nevertheless, it is difficult to satisfy the transparency of the material for imaging while also ensuring that the in vitro physical and chemical properties can mimic the osteoid. Therefore, more culture models need to be explored to address any specific scientific questions.
Additionally, the physicochemical properties of this IDG-SW3 mineralized gel, including the rheological properties, were introduced in a previous study16. Thus, this biogenic mineralized gel formed by this method could be useful as a biomedical material for future orthopedics research.
Declarações
The authors have nothing to disclose.
Acknowledgements
We thank Dr. Lynda F. Bonewald for gifting the IDG-SW3 cell line. This work was supported by the National Natural Science Foundation of China (82070902, 82100935, and 81700778) and the Shanghai "Science and Technology Innovation" Sailing Project (21YF1442000).
Materials
| 0.25% Trypsin-ethylenediaminetetraacetic acid | Hyclone | SH30042.01 | |
| 15 mL tubes | Corning, NY, USA | 430791 | |
| 7.5% (w/v) Sodium bicarbonate | Sigma-Aldrich, MO, USA | S8761 | |
| ascorbic acid | Sigma-Aldrich, MO, USA | A4544 | |
| Collagen I | Thermo Fisher Scientific | A10483-01 | |
| fetal bovine serum | Thermo Fisher Scientific | 10099141 | |
| homogenizer | BiHeng Biotechnology, Shanghai, China | SKSI | |
| laser confocal fluorescence microscopy | Carl Zeiss, Oberkochen, Germany | LSM 800 | |
| Live/Dead Cell Imaging kit | Thermo Fisher Scientific | R37601 | |
| Matrigel matrix | Corning, NY, USA | 356234 | |
| MEM (10X), no glutamine | Thermo Fisher Scientific | 21430079 | |
| paraformaldehyde | Sigma-Aldrich, MO, USA | 158127 | |
| phosphate buffered saline | Hyclone | SH30256.FS | |
| stereo microscope | Carl Zeiss, Oberkochen, Germany | Zeiss Axio ZOOM.V16 | |
| Trizol | Thermo Fisher Scientific | 15596026 | |
| X-ray fluorescence | EDAX, USA | EAGLE III | |
| β-glycerophosphate | Sigma-Aldrich, MO, USA | G9422 |
Referências
- Bonewald, L. F. The amazing osteocyte. Journal of Bone and Mineral Research. 26 (2), 229-238 (2011).
- Dallas, S. L., Prideaux, M., Bonewald, L. F. The osteocyte: An endocrine cell … and more. Endocrine Reviews. 34 (5), 658-690 (2013).
- Bonewald, L. F. The role of the osteocyte in bone and nonbone disease. Endocrinology and Metabolism Clinics of North America. 46 (1), 1-18 (2017).
- Woo, S. M., Rosser, J., Dusevich, V., Kalajzic, I., Bonewald, L. F. Cell line IDG-SW3 replicates osteoblast-to-late-osteocyte differentiation in vitro and accelerates bone formation in vivo. Journal of Bone and Mineral Research. 26 (11), 2634-2646 (2011).
- Wang, J. S., et al. Control of osteocyte dendrite formation by Sp7 and its target gene osteocrin. Nature Communications. 12 (1), 6271 (2021).
- Chicatun, F., et al. Osteoid-mimicking dense collagen/chitosan hybrid gels. Biomacromolecules. 12 (8), 2946-2956 (2011).
- Kawasaki, K., Buchanan, A. V., Weiss, K. M. Biomineralization in humans: making the hard choices in life. Annual Review of Genetics. 43, 119-142 (2009).
- Bosman, F. T., Stamenkovic, I. Functional structure and composition of the extracellular matrix. Journal of Pathology. 200 (4), 423-428 (2003).
- Papadopoulos, N. G., et al. An improved fluorescence assay for the determination of lymphocyte-mediated cytotoxicity using flow cytometry. Journal of Immunological Methods. 177 (1-2), 101-111 (1994).
- Staines, K. A., et al. Hypomorphic conditional deletion of E11/Podoplanin reveals a role in osteocyte dendrite elongation. Journal of Cellular Physiology. 232 (11), 3006-3019 (2017).
- Feng, J. Q., et al. Loss of DMP1 causes rickets and osteomalacia and identifies a role for osteocytes in mineral metabolism. Nature Genetics. 38 (11), 1310-1315 (2006).
- Winkler, D. G., et al. Osteocyte control of bone formation via sclerostin, a novel BMP antagonist. EMBO Journal. 22 (23), 6267-6276 (2003).
- Riminucci, M., et al. FGF-23 in fibrous dysplasia of bone and its relationship to renal phosphate wasting. Journal of Clinical Investigation. 112 (5), 683-692 (2003).
- Dallas, S. L., Bonewald, L. F. Dynamics of the transition from osteoblast to osteocyte. Annals of the New York Academy of Sciences. 1192, 437-443 (2010).
- Robin, M., et al. Involvement of 3D osteoblast migration and bone apatite during in vitro early osteocytogenesis. Bone. 88, 146-156 (2016).
- Chen, K., et al. High mineralization capacity of IDG-SW3 cells in 3D collagen hydrogel for bone healing in estrogen-deficient mice. Frontiers in Bioengineering and Biotechnology. 8, 864 (2020).

