Assessing the Putative Anticryptococcal Properties of Crude and Clarified Extracts from Mollusks
Summary
The human fungal pathogen Cryptococcus neoformans produces a variety of virulence factors (e.g., peptidases) to promote its survival within the host. Environmental niches represent a promising source of novel natural peptidase inhibitors. This protocol outlines the preparation of extracts from mollusks and the assessment of their effect on fungal virulence factor production.
Abstract
Cryptococcus neoformans is an encapsulated human fungal pathogen with a global distribution that primarily infects immunocompromised individuals. The widespread use of antifungals in clinical settings, their use in agriculture, and strain hybridization have led to increased evolution of resistance. This rising rate of resistance against antifungals is a growing concern among clinicians and scientists worldwide, and there is heightened urgency to develop novel antifungal therapies. For instance, C. neoformans produces several virulence factors, including intra- and extra-cellular enzymes (e.g., peptidases) with roles in tissue degradation, cellular regulation, and nutrient acquisition. The disruption of such peptidase activity by inhibitors perturbs fungal growth and proliferation, suggesting this may be an important strategy for combating the pathogen. Importantly, invertebrates such as mollusks produce peptidase inhibitors with biomedical applications and anti-microbial activity, but they are underexplored in terms of their usage against fungal pathogens. In this protocol, a global extraction from mollusks was performed to isolate potential peptidase inhibitors in crude and clarified extracts, and their effects against classical cryptococcal virulence factors were assessed. This method supports the prioritization of mollusks with antifungal properties and provides opportunities for the discovery of anti-virulence agents by harnessing the natural inhibitors found in mollusks.
Introduction
Cryptococcus neoformans is a human fungal pathogen that produces severe disease in immunocompromised hosts, such as individuals living with HIV/AIDS1, and leads to approximately 19% of AIDS-related deaths2. The fungus is susceptible to several classes of antifungals, including azoles, polyenes, and flucytosine, which exert fungicidal and fungistatic activity using distinct mechanisms3,4. However, the extensive use of antifungals in clinical and agricultural settings combined with strain hybridization have amplified the evolution of resistance in multiple fungal species, including C. neoformans5.
To overcome the challenges of antifungal resistance and reduce the prevalence of fungal infections on a global scale, a promising approach is to use the virulence factors of Cryptococcus spp. (e.g., temperature adaptability, polysaccharide capsule, melanin, and extracellular enzymes) as potential therapeutic targets4,6. This approach has several advantages, as these virulence factors are well-characterized in the literature, and targeting these factors could potentially reduce the rates of antifungal resistance by imposing a weaker selective pressure through impairing virulence rather than targeting cell growth6. In this context, numerous studies have assessed the possibility of targeting extracellular enzymes (e.g., proteases, peptidases) to reduce or inhibit the virulence of Cryptococcus spp.7,8,9.
Organisms like invertebrates and plants do not possess an adaptive immune system to protect themselves from pathogens. However, they rely on a strong innate immune system with an immense array of chemical compounds to deal with microorganisms and predators10. These molecules include peptidase inhibitors, which play important roles in many biological systems, including the cellular processes of invertebrate immunity, such as the coagulation of hemolymph, the synthesis of cytokines and antimicrobial peptides, and the protection of hosts by directly inactivating the proteases of pathogens11. Thus, peptidase inhibitors from invertebrates such as mollusks possess potential biomedical applications, but many remain uncharacterized10,12,13. In this context, there are approximately 34 species of terrestrial mollusks in Ontario and 180 freshwater mollusks in Canada14. However, their in-depth profiling and characterization are still limited15. These organisms present an opportunity for the identification of new compounds with potential anti-fungal activity10.
In this protocol, methods to isolate and clarify extracts from invertebrates (e.g., mollusks) (Figure 1) followed by measuring the putative peptidase inhibitory activity are described. The antifungal properties of these extracts are then assessed by measuring their impact on C. neoformans virulence factor production using phenotypic assays (Figure 2). It is important to note that differences in antifungal properties between crude and clarified extracts may be indicative of microbial factors (e.g., secondary metabolites or toxins produced by the host microbiome) of the mollusk, which may influence experimental observations. Such findings support the need for this protocol to assess both crude and clarified extracts independently to unravel the modes of action. Additionally, the extraction process is unbiased and may enable the detection of antimicrobial properties against a plethora of fungal and bacterial pathogens. Therefore, this protocol provides an initiation point for the prioritization of mollusk species with antifungal properties against C. neoformans and an opportunity to evaluate the connections between enzymatic activity and virulence factor production through putative inhibitory mechanisms.
Protocol
1. Protein extraction from mollusks
- Collect mollusks from a designated and approved natural area (e.g., Speed River, Guelph, Ontario). For this study, both native and invasive species were selected to assess a broad range of potential antifungal effects.
- Gently break the shell of the mollusks (e.g., Cepaea nemoralis, Planorbella pilsbryi, and Cipangopaludina chinensis) using a pestle and mortar, and remove the solid pieces with a pair of tweezers. Generally, 10 mollusks are pooled for use throughout the protocol.
- Collect and weigh the organs. Approximately 15-20 g of sample is optimal. Use scissors to cut the organs into small pieces approximately 0.5-1 cm in size.
- Flash-freeze the dissected and cut organ samples with liquid nitrogen. Then, grind the organ samples into a fine powder using a pestle and mortar.
- Add the organ sample powder (approximately 15 g) into 30 mL of cold distilled water (4 °C) to achieve a 1:2 ratio (w/v).
- Pour 2 mL of the sample into high-impact 2 mL tubes. Add a scoopful of 3 mm stainless-steel beads (approximately 500 µg) to each tube, and homogenize using a blender (see Table of Materials) for 3 min at 1,200 rpm and 4 °C.
- Centrifuge at 12,000 × g for 20 min at 4 °C. Collect all the supernatants with a pipette in a fresh 50 mL tube, discarding the pellet.
- Filter-sterilize the supernatant using a 0.22 µm membrane. Store the samples on ice, and proceed to the clarification protocol (section 2), as applicable.
NOTE: The samples can be flash-frozen in liquid nitrogen and stored at −20 °C to be used as crude extracts.
2. Clarification of the mollusk extract
- Place the supernatant sample from step 1.8 in a thermal bath at 60 °C for 30 min. Then, cool the samples quickly by transferring them to a bucket with ice for 20 min.
- Centrifuge the samples at 15,000 × g for 45 min at 4 °C. Collect the supernatant in a fresh 50 mL tube, and discard the pellet. Filter-sterilize the samples using a 0.22 µm membrane filter, and then store them on ice.
- Determine the protein concentration in the samples from step 1.8 (i.e., the crude extract) and step 2.2 (i.e., the clarified extract) using a quantification assay (e.g., bicinchoninic acid assay17). The optimal protein concentration range is 4-8 mg/mL.
- Store the samples on ice for up to 1 h or flash-freeze in liquid nitrogen, and store them at −20 °C until needed.
3. Inhibitory activity assay
- Measure the inhibitory activity of the crude and clarified extracts using enzymatic assays in technical triplicate.
NOTE: Each enzymatic assay consists of an enzyme (e.g., subtilisin A, which is involved in fungal virulence16,17,18; usually around 1 x 10−9 mol/L for chromogenic-based assays), a buffer, and a substrate. The reaction starts when the substrate encounters the enzyme. The enzymatic activity is proportional to the rate of substrate conversion into the product. - Assess the effect of enzyme concentration (EC) over enzymatic activity (EA) until linear dependence is obtained between both parameters. For the next steps, use an enzyme concentration within the measured linear range.
NOTE: The enzymatic assay for the peptidase subtilisin A is detailed in the following steps. - Incubate the crude (step 1.11) or clarified (step 2.5) mollusk protein extracts (e.g., 10 µL containing 4 mg/mL protein) with 10 µL of subtilisin A (concentration determined from step 3.2), and 220 µL of 100 mM Tris-HCl at pH 8.6 (for the control, add 230 µL of Tris buffer without adding the extract) for 10 min at 25 °C in a 96-well flat-bottom plate.
- Add 10 µL of N-Succinyl-Ala-Ala-Pro-Phe-p-nitroanilide, the substrate for subtilisin A, at a final concentration of 1 Km (Michaelis-Menten constant; 0.2 mM for this substrate with subtilisin A) for a final reaction volume of 250 µL.
- Measure the enzymatic activity by monitoring the appearance of the product over time by reading the optical density (OD) at 405 nm every 15 s for 3 min.
- Determine the residual enzymatic activity by calculating the ratio of the enzymatic activity in the presence of the mollusk extract to that in the absence of the extract.
- If inhibitory activity is observed (i.e., residual activity below 1), measure the inhibitory concentration 50 (IC50) value using a range of concentrations of the extracts (e.g., a twofold dilution series from 200 µg/mL to 1 µg/mL) following standard methods19.
4. Effect of mollusk extracts on C. neoformans growth
- Use a 10 µL pipette tip to introduce a single colony of C. neoformans var. grubii H99 wild-type (WT) into 5 mL of yeast extract-peptone-dextrose (YPD) medium (10 g/L yeast extract, 20 g/L peptone, and 20 g/L dextrose, pH 6.5), and incubate for 16-18 h at 30 °C and 200 rpm. Perform the experiment in biological triplicate and technical duplicate.
- Collect 500 µL of the culture in a cuvette, and measure the growth by reading the OD at 600 nm. Dilute the culture with yeast nitrogen base (YNB) medium to a final OD600 of 0.02.
- Co-culture C. neoformans cells with different concentrations (e.g., a threefold dilution series from 440 µg/mL to 15 µg/mL protein) of the extracts (i.e., crude and clarified). To do this, mix 10 µL of the extracts with 190 µL of the diluted C. neoformans culture (step 4.2) in a 96-well plate.
- Measure the growth by reading the OD600 using a plate reader every 15 min to 1 h for 72 h at 200 rpm and 37 °C.
5. Effect of mollusk extracts on C. neoformans melanin production
- Prepare L-3,4-dihydroxyphenylalanine (L-DOPA) plates.
- Autoclave 230 mL of 14 g/L agar, and let it cool until it reaches 50-60 °C.
- Add 3.25 mL of 1 M glycine, 7.35 mL of 1 M KH2PO4, 2.5 mL of 1 M MgSO4.7H2O, 0.6 mL of 40% glucose, 70 µL of 10 mM thiamine, and 2.5 mL of 100 mM L-DOPA (filter-sterilized) to the liquid agar.
- Pour 15 mL of the mixture into Petri plates, let them dry for approximately 1 h in the dark, and store at 2-4 °C for up to 1 week. Ensure the lids are partially lifted for the agar to properly dry.
- Inoculate 5 mL of YNB medium with 50 µL of C. neoformans culture from step 4.1. Incubate for 16-18 h at 30 °C and 200 rpm.
- Spin down the cells at 1,000 × g for 5 min at 4 ˚C. Wash the cells 2x with sterile phosphate-buffered saline (PBS) at pH 7.4.
- Count the cells using a hemocytometer. Resuspend the cells in 1 mL of PBS to reach a final concentration of 106 cells/mL.
- Prepare a dilution series (e.g., twofold) of the crude and clarified mollusk extracts. Similarly, prepare a 10-fold dilution series of C. neoformans cells (step 5.4) from 1 x 106 cells/mL to 1 x 101 cells/mL using sterile PBS in 96-well plates.
- Spread 200 µL of the extracts onto Petri plates containing L-DOPA agar using a swab, and let them dry for 15 min.
- Spot 5 µL of culture from each well onto the L-DOPA plate, leaving 1 cm between drops. Let it dry in the dark for 15 min.
- Incubate the plates at 30 °C and 37 °C in a static incubator for 3-5 days, capturing images every 24 h with a camera or plate imager (e.g., scanner).
NOTE: Two temperature conditions are used here to explore the influence of temperature on the growth inhibitory effects of the mollusk extracts, with 30 °C representing environmental and 37 ˚C representing human physiological temperatures.
6. Effect of mollusk extracts on C. neoformans polysaccharide capsule production
- Prepare low iron medium (LIM) as described below.
- Dissolve 5 g of chelating resin (see Table of Materials) in 60 mL of ultrapure water, and pack into a glass column (2 cm diameter x 25 cm length). Wash the column with 100 mL of water, and discard the collected water.
- Prepare low iron water (LI-water) by passing 2 L of sterile ultrapure water through the chelating column. Store the LI-water at 4 ˚C for up to 3 months.
- Dissolve 5 g of glucose in 100 mL of LI-water. Dissolve 5 g of L-asparagine in 200 mL of LI-water in a separate beaker.
- Add the following to 500 mL of LI-water in order: 4.78 g of HEPES, 0.4 g of K2HPO4, 0.08 g of MgSO4.7H2O, 1.85 g of NaHCO3, and 0.25 g of CaCl2.2H2O.
- Combine the solutions from step 6.1.3 and step 6.1.4. Add 1 mL of 100 mM bathophenanthrolinedisulfonic acid sodium salt (BPS) to a final concentration of 100 µM.
- Adjust the pH to 7.2 with 1 M LI-MOPS. Add LI-water to a final volume of 1 L, and filter- sterilize the medium by passing it through a 0.22 µm membrane. Add 100 µL of sterile thiamine (4 mg/mL) to the medium.
- Inoculate 5 mL of YNB medium with 50 µL of the C. neoformans culture from step 4.1. Incubate for 16-18 h at 30 °C and 200 rpm. Use this culture to inoculate 5 mL of LIM to a final concentration of 105 cells/mL.
- Add appropriate volumes of the mollusk extracts (e.g., a twofold dilution or the volume determined from previous virulence assays) to reach the desired concentrations. Incubate for 72 h at 37 °C and 200 rpm with loosened caps. After incubation, collect 1 mL of cells, and wash 2x with sterile PBS, pH 7.4, at 1,500 × g for 2 min at 4 ˚C.
- Gently resuspend the cells in 1 mL of PBS. Mix the cells with India Ink at 1:1 ratio (e.g., 4 µL of India Ink with 4 µL of cell suspension) over a glass microscope slide. Place a coverslip on top of the slide.
- Visualize the samples using a differential interference contrast (DIC) microscope with a 63x oil objective (see Table of Materials). Set the contrast to automatic, and focus manually using the microscope dial. Select all the pictures, and export them in TIFF format. No filters need to be applied.
- Using the ruler tool in ImageJ20, quantify the ratio of the total cell size (including the capsule) to the cell body size.
7. Effect of mollusk extracts on C. neoformans biofilm production
- Inoculate 5 mL of YNB medium with 50 µL of C. neoformans culture from step 4.1. Incubate for 16-18 h at 30 °C and 200 rpm.
- Harvest cells from the 5 mL culture. Wash the cells with sterile PBS, pH 7.4, at 1,500 × g for 2 min at 4 ˚C.
- Count the cells using a hemocytometer. Resuspend the cells in 3 mL of Dulbecco's Modified Eagle Medium (DMEM) to a final concentration of 107 cells/mL.
- Transfer 300 µL of the cell suspension into the individual wells of a sterile, polystyrene, flat-bottom 24-well plate. Use wells with media only as a control.
- Add appropriate volumes of the mollusk extracts (e.g., a twofold dilution or the volume determined from previous virulence assays) to reach a final concentration of 10-20 µg/mL protein. Ensure that the extract volume is not greater than 10% of the total volume.
- Wrap the plates with aluminum foil (to avoid media evaporation), and incubate for 48 h using a static incubator at 30 °C and 37 °C.
- Using a bottle or dispenser wash the wells 2x with sterile water, and let them air-dry for 10-15 min at room temperature. Then, add 100 µL of 0.3% crystal violet solution to each well (including the medium-only control wells), and incubate at room temperature for 10 min.
- Wash the wells thoroughly 3x with sterile water (the biofilms will not be disrupted during washing). Add 200 µL of 100% ethanol, and incubate for 10 min at RT.
- Transfer 75 µL to a new 96-well flat-bottom plate, and read the OD550. Quantify the biofilm formation as (OD550nm of sample – OD550nm of blank).
Representative Results
The workflow described herein enables the isolation of proteins and peptides from mollusks with potential anti-virulence properties against C. neoformans. Similarly, assessing different forms of extracts (i.e., crude and clarified) allows for the semi-purification of the potential active compounds and supports downstream assessment (e.g., mass spectrometry-based proteomics). Typically, the protein extraction workflow produces homogenized solutions with protein concentrations of 4-8 mg/mL. Here, the representative results demonstrate the assessment of the enzymatic activity and antifungal properties of C. chinensis extracts.The crude and clarified extracts were able to inhibit the proteolytic activity of subtilisin A (related to virulence in C. neoformans) (Figure 3), with IC50 values of 5.3 µg/mL and 4.53 µg/mL, respectively. The activity of the C. chinensis extracts was further tested against processes associated with C. neoformans virulence factor production, including fungal growth, capsule and melanin production, and the formation of biofilms. There was a significant reduction in fungal growth at 37 °C in the presence of the crude (Figure 4A) and clarified (Figure 4B) C. chinensis extracts. Notably, there were no changes in capsule or melanin production in the presence of crude or clarified C. chinensis extracts (Figure 4C,D). However, a significant reduction of 70%-80% in biofilm formation was observed at high concentrations of the crude (Figure 4E) and clarified (Figure 4F) extracts relative to the untreated control.
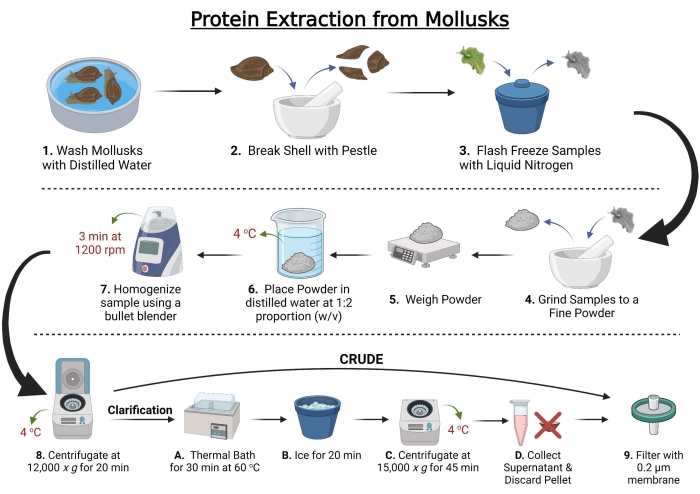
Figure 1: Workflow for total protein extraction from mollusks. Created with BioRender.com. Please click here to view a larger version of this figure.
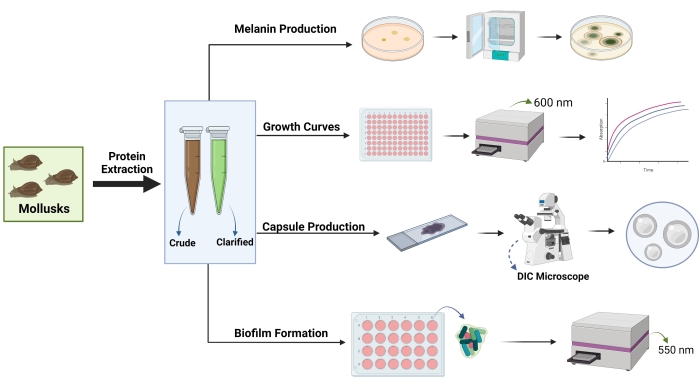
Figure 2: General strategy used to assess the effects of mollusk extracts on proteolytic activity, growth, and virulence factor production in Cryptococcus neoformans. DIC = differential interference contrast microscopy. Optical density measurement indicated with the wavelength in nanometers (nm). Created with BioRender.com. Please click here to view a larger version of this figure.
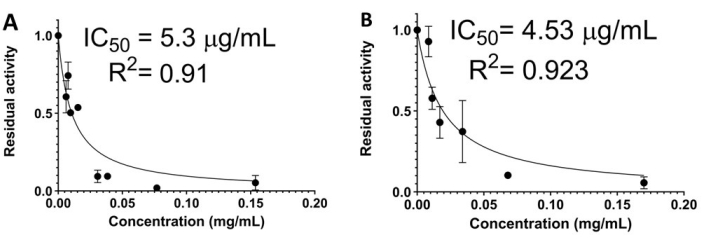
Figure 3: Representative results of the effects of mollusk protein extracts on the proteolytic activity of subtilisin A. (A) Cipangopaludina chinensis crude extracts. (B) C. chinensis clarified extracts. Each point represents the average of three replicates. Bars indicate standard deviation. IC50 = half-maximal inhibitory concentration. Please click here to view a larger version of this figure.
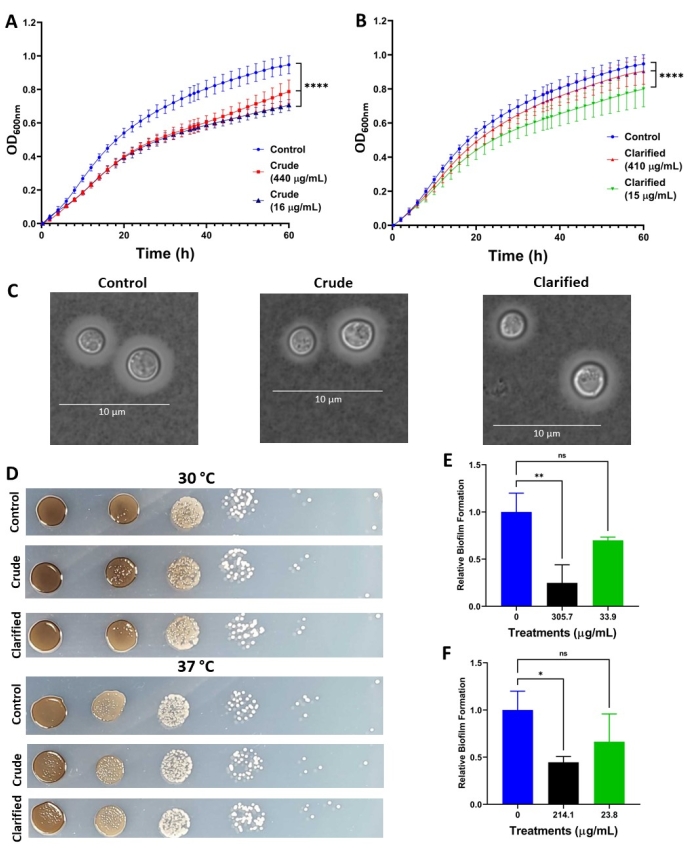
Figure 4: Representative results of the effects of C. chinensis extracts on virulence factor production in C. neoformans. (A,B) Growth of C. neoformans at 37 °C in the presence of crude and clarified C. chinensis extracts, respectively. (C) DIC microscopy images of C. neoformans showing capsule production in the presence of crude (50 µg/mL) and clarified (40 µg/mL) extracts. Scale bar = 10 µm. (D) Melanin production of C. neoformans in the presence of crude (440 µg/mL) and clarified (410 µg/mL) extracts at 30 °C and 37 °C. (E,F) Relative biofilm formation of C. neoformans in the presence of crude and clarified extracts, respectively. For statistical analysis, a one-way ANOVA test and a Dunnett's multiple comparison test were performed. *p < 0.05; **p < 0.01; and ****p < 0.0001. Each value corresponds to an average of at least five biological replicates and two technical replicates. Error bars indicate standard deviation. Melanin images shown are representative of three biological replicates and two technical replicates. Capsule images shown are representative of 40-50 cells per condition. Please click here to view a larger version of this figure.
Discussion
The extraction protocol described here outlines the isolation of compounds from mollusks collected from Ontario, Canada, and demonstrates a novel investigation of using mollusk extracts against the human fungal pathogen, C. neoformans. This protocol adds to a growing body of research investigating peptidase inhibitor activity from invertebrates13. During the extraction, some extract samples were difficult to filter-sterilize, possibly due to the presence of soluble polysaccharides and/or pigments that obstructed the filter membrane. To overcome this limitation, it is recommended to filter first through a 5 µm membrane to exclude large compounds (e.g., disrupted cell membranes, genome DNA), thus allowing the proteins to pass through the filter, and then to filter again through a 0.22 µm membrane. These steps are critical to the protocol as they restrict the presence of microbes that may contaminate the samples and interfere with downstream experiments.
During this investigation, peptidase inhibitors were detected against subtilisin A, a model enzyme for the S8 family of subtilisin. Members of this enzyme family are widely distributed among organisms and have varying roles, such as in protein processing, nutrition, and virulence mechanisms21,22, which support the phenotypic effects observed in this study. For example, a significant reduction in fungal growth was observed in the presence of both crude and clarified extracts and at relatively high and low concentrations, suggesting that the inhibitory activity was robust in the tested model. It is notable that when measuring the OD with a plate reader, the presence of the extracts caused clumping of the fungal cells at the bottom of the well, interfering with the growth measurement. This limitation can be overcome by using a high-speed shaking incubator (e.g., 900 rpm), which would avoid cell clumping.
Other technical limitations may exist that may influence the anticipated phenotypic observations. For instance, subtilisin-like peptidases are associated with melanin synthesis and quorum sensing in C. neoformans, but the crude or clarified extracts from any mollusks do not show significant effects on melanin production16,17,18. This is possibly due to the natural protective effect of melanin and capsule against external agents, which could prevent the mollusk extracts from impacting the intracellular components of the fungus. Important factors to consider when working with organic substrates include the need to dissolve them in dimethyl sulfoxide (DMSO), which may present solubility issues within the agar plate (as used in the melanin assays). To overcome this problem, the extracts may be spread along the surface of the agar plate and allowed to dry prior to spotting the fungal cells onto the plate.
Previous work demonstrated the susceptibility of C. neoformans biofilms to two antifungal agents in vitro, including amphotericin B and caspofungin; however, the fungus was resistant to fluconazole and voriconazole23. Given the importance of fungal biofilms in virulence and antimicrobial resistance, uncovering new strategies to interfere with or disrupt biofilm formation would be valuable. In the current study, crude and clarified extracts from C. chinensis impaired the formation of biofilms of cryptococcal cells with apparent dose-response behavior. These results support the impact and novelty of the approach. Notably, biofilm formation was inhibited to a greater extent on treatment with the crude compared to the clarified extract, which may be due to a loss of inhibitory compounds during the clarification process or a reduction in inhibitory function under the tested conditions. It is possible that the proteins responsible for these inhibitory effects are of high molecular weight and were lost during the clarification process or were susceptible to degradation during the thermal treatment. These results highlight the importance of using both crude and clarified extracts to detect changes in inhibitory functions, as differences related to the inhibitor source may influence the outcome.
Overall, this protocol enables the extraction of compounds from mollusks and the measurement of putative inhibitory activity against a selected peptidase with demonstrated roles in fungal virulence. In this protocol, the high yield and strong peptidase inhibitory activity of the extracts and their effect against C. neoformans growth and biofilm formation were evaluated. While further experiments are needed, these results stress the importance of mollusks as putative new sources of compounds against this dangerous fungal pathogen. Furthermore, while this protocol focuses on the extraction of inhibitors from mollusks, the methodology is adaptable to other invertebrates and can be used to assess the effect of inhibitors derived from different invertebrates on virulence factor production in a variety of microorganisms, including fungi and bacteria24,25,26. Ultimately, the extraction of compounds from natural sources can increase the repertoire of putative novel antimicrobial agents and, thus, improve our abilities to combat infectious diseases.
Declarações
The authors have nothing to disclose.
Acknowledgements
The authors thank members of the Geddes-McAlister Lab for their valuable support throughout this investigation and their manuscript feedback. The authors acknowledge the funding support from the Ontario Graduate Scholarship and International Graduate Research Award – University of Guelph to D. G.-G and from the Canadian Foundation of Innovation (JELF 38798) and Ontario Ministry of Colleges and Universities – Early Researcher Award for J. G.-M.
Materials
| 0.2 μm Filters | VWR | 28145-477 (North America) | |
| 1.5 mL Tubes (Safe-Lock) | Eppendorf | 0030120086 | |
| 2 mL Tubes (Safe-Lock) | Eppendorf | 0030120094 | |
| 3,4-Dihydroxy-L-phenylalanine (L-DOPA) | Sigma-Aldrich | D9628-5G | CAS #: 59-92-7 |
| 96-well plates | Costar (Corning) | 3370 | |
| Bullet Blender Storm 24 | NEXT ADVANCE | BBY24M | |
| Centrifuge 5430R | Eppendorf | 5428000010 | |
| Chelex 100 Resin | BioRad | 142-1253 | |
| CO2 Incubator (Static) | SANYO | Not available | |
| Cryptococcus neoformans H99 | ATCC | 208821 | |
| DIC Microscope | Olympus | ||
| DIC Microscope software | Zeiss | ||
| DMEM | Corning | 10-013-CV | |
| Glucose (D-Glucose, Anhydrous, Reagent Grade) | BioShop | GLU501 | CAS #: 50-99-7 |
| Glycine | Fisher Chemical | G46-1 | CAS #: 56-40-6 |
| GraphPad Prism 9 | Dotmatics | ||
| Hemocytometer | VWR | 15170-208 | |
| HEPES | Sigma Aldrich | H3375 | |
| Magnesium sulfate heptahydrate (MgSO4.7 H2O) | Honeywell | M1880-500G | CAS #: 10034-99-8 |
| Peptone | BioShop | PEP403 | |
| Phosohate buffer salt pH 7.4 | BioShop | PBS408 | SKU: PBS408.500 |
| Plate reader (Synergy-H1) | BioTek (Agilent) | Not available | |
| Potassium phosphate monobasic (KH2PO4) | Fisher Chemical | P285-500 | CAS #: 7778-77-0 |
| Subtilisin A | Sigma-Aldrich | P4860 | CAS #: 9014-01-01 |
| Succinyl-Ala-Ala-Pro-Phe-p-nitroanilide | Sigma-Aldrich | 573462 | CAS #: 70967-97-4 |
| Thermal bath | VWR | 76308-834 | |
| Thiamine Hydrochloride | Fisher-Bioreagents | BP892-100 | CAS #: 67-03-8 |
| Yeast extract | BioShop | YEX401 | CAS #: 8013-01-2 |
| Yeast nitrogen base (with Amino Acids) | Sigma-Aldrich | Y1250-250G | YNB |
Referências
- Derek, J., Sloan, V. P. Cryptococcal meningitis: Epidemiology and therapeutic options. Clinical Epidemiology. 6, 169-182 (2014).
- Rajasingham, R., et al. The global burden of HIV-associated cryptococcal infection in adults in 2020: a modelling analysis. The Lancet Infectious Diseases. , (2022).
- Mourad, A., Perfect, J. R. Present and future therapy of Cryptococcus infections. Journal of Fungi. 4 (3), 79 (2018).
- Bermas, A., Geddes-McAlister, J. Combatting the evolution of antifungal resistance in Cryptococcus neoformans. Molecular Microbiology. 114 (5), 721-734 (2020).
- Geddes-McAlister, J., Shapiro, R. S. New pathogens, new tricks: Emerging, drug-resistant fungal pathogens and future prospects for antifungal therapeutics. Annals of the New York Academy of Sciences. 1435 (1), 57-78 (2019).
- Kronstad, J. W., Hu, G., Choi, J. The cAMP/protein kinase A pathway and virulence in Cryptococcus neoformans. Mycobiology. 39 (3), 143-150 (2018).
- Olszewski, M. A., et al. Urease expression by Cryptococcus neoformans promotes microvascular sequestration, thereby enhancing central nervous system invasion. The American Journal of Pathology. 164 (5), 1761-1771 (2004).
- Shi, M., et al. Real-time imaging of trapping and urease-dependent transmigration of Cryptococcus neoformans in mouse brain. The Journal of Clinical Investigation. 120 (5), 1683-1693 (2010).
- Vu, K., et al. Invasion of the central nervous system by Cryptococcus neoformans requires a secreted fungal metalloprotease. mBio. 5 (3), 01101-01114 (2014).
- Gutierrez-Gongora, D., Geddes-McAlister, J. From naturally-sourced protease inhibitors to new treatments for fungal infections. Journal of Fungi. 7 (12), 1016 (2021).
- Nakao, Y., Fusetani, N. Enzyme inhibitors from marine invertebrates. Journal of Natural Products. 70 (4), 689-710 (2007).
- Reytor, M. L., et al. Screening of protease inhibitory activity in extracts of five Ascidian species from Cuban coasts. Biotecnologia Aplicada. 28 (2), 77-82 (2011).
- González, L., et al. Screening of protease inhibitory activity in aqueous extracts of marine invertebrates from Cuban coast. American Journal of Analytical Chemistry. 7 (4), 319-331 (2016).
- Brown, D. S., Werger, M. J. A. Freshwater molluscs. Biogeography and Ecology of Southern Africa. , 1153-1180 (1978).
- Forsyth, R. G., Oldham, M. J. Terrestrial molluscs from the Ontario Far North. Check List. 12 (3), 1-51 (2016).
- Eigenheer, R. A., Lee, Y. J., Blumwald, E., Phinney, B. S., Gelli, A. Extracellular glycosylphosphatidylinositol-anchored mannoproteins and proteases of Cryptococcus neoformans. FEMS Yeast Research. 7 (4), 499-510 (2007).
- Homer, C. M., et al. Intracellular action of a secreted peptide required for fungal virulence. Cell Host & Microbe. 19 (6), 849-864 (2016).
- Clarke, S. C., et al. Integrated activity and genetic profiling of secreted peptidases in Cryptococcus neoformans reveals an aspartyl peptidase required for low pH survival and virulence. PLoS Pathogens. 12 (12), 1006051 (2016).
- Copeland, R. A. . Evaluation of Enzyme Inhibitors in Drug Discovery: A Guide for Medicinal Chemists and Pharmacologists. , (2013).
- Collins, T. J. ImageJ for microscopy. Biotechniques. 43, 25-30 (2007).
- Rawlings, N. D., et al. The MEROPS database of proteolytic enzymes, their substrates and inhibitors in 2017 and a comparison with peptidases in the PANTHER database. Nucleic Acids Research. 46, 624-632 (2018).
- Gutierrez-Gongora, D., Geddes-McAlister, J. Peptidases: Promising antifungal targets of the human fungal pathogen, Cryptococcus neoformans. Facets. 7 (1), 319-342 (2022).
- Martinez, L. R., Casadevall, A. Susceptibility of Cryptococcus neoformans biofilms to antifungal agents in vitro. Antimicrobial Agents and Chemotherapy. 50 (3), 1021-1033 (2006).
- Culp, E., Wright, G. D. Bacterial proteases, untapped antimicrobial drug targets. Journal of Antibiotics. 70 (4), 366-377 (2017).
- Ruocco, N., Costantini, S., Palumbo, F., Costantini, M. Marine sponges and bacteria as challenging sources of enzyme inhibitors for pharmacological applications. Mar Drugs. 15 (6), 173 (2017).
- Costa, H. P. S., et al. JcTI-I: A novel trypsin inhibitor from Jatropha curcas seed cake with potential for bacterial infection treatment. Frontiers in Microbiology. 5, 5 (2014).

