Investigating Angiogenesis on a Functional and Molecular Level by Leveraging the Scratch Wound Migration Assay and the Spheroid Sprouting Assay
Summary
This study presents the two-dimensional (2D) scratch wound migration assay and the three-dimensional (3D) spheroid sprouting assay, along with their respective downstream analysis methods, including RNA extraction and immunocytochemistry, as suitable assays to study angiogenesis in vitro.
Abstract
Angiogenesis plays a crucial role in both physiological and pathological processes within the body including tumor growth or neovascular eye disease. A detailed understanding of the underlying molecular mechanisms and reliable screening models are essential for targeting diseases effectively and developing new therapeutic options. Several in vitro assays have been developed to model angiogenesis, capitalizing on the opportunities a controlled environment provides to elucidate angiogenic drivers at a molecular level and screen for therapeutic targets.
This study presents workflows for investigating angiogenesis in vitro using human umbilical vein endothelial cells (HUVECs). We detail a scratch wound migration assay utilizing a live cell imaging system measuring endothelial cell migration in a 2D setting and the spheroid sprouting assay assessing endothelial cell sprouting in a 3D setting provided by a collagen matrix. Additionally, we outline strategies for sample preparation to enable further molecular analyses such as transcriptomics, particularly in the 3D setting, including RNA extraction as well as immunocytochemistry. Altogether, this framework offers scientists a reliable and versatile toolset to pursue their scientific inquiries in in vitro angiogenesis assays.
Introduction
Angiogenesis, which refers to the formation of new blood vessels from pre-existing ones1, is a crucial process during physiological development and pathologic conditions. It is indispensable for providing energy to highly metabolically active tissues such as the retina2 or the developing central nervous system3 and during the healing of damaged tissue4. Abnormal angiogenesis, on the other hand, is the basis for numerous diseases. Solid tumors, such as colorectal cancer or non-small cell lung cancer, facilitate their growth and the necessary energy supply by promoting angiogenesis5. Apart from cancer, neovascular diseases of the eye like diabetic retinopathy or age-related macular degeneration, which represent leading causes of blindness in the developed world, result from aberrant vessel growth6,7. A detailed understanding of the underlying pathomechanism is crucial to comprehend how physiological angiogenesis can be enhanced, for instance, in wound healing while better controlling pathological conditions such as vasoproliferative eye diseases.
On a cellular level, vascular endothelial cells are activated by various signaling molecules in angiogenesis, initiating cell proliferation and migration8. The cells subsequently organize into a hierarchy, with non-proliferating tip cells forming filopodia at the leading edge of the developing vessel branch8. Alongside, fast-proliferating stalk cells trail the tip cells, contributing to the formation of the emerging vessel. Subsequently, other cell types, such as pericytes or smooth muscle cells, are recruited to further stabilize the nascent branch9.
To explore molecular processes at the vascular endothelial cell level, numerous in vitro protocols have been developed and recently reviewed10. These assays typically fall into two categories: more simplistic but scalable 2D approaches and more elaborate 3D protocols. In a recent project, we conducted a comprehensive comparative analysis between the 2D scratch wound migration assay and the 3D spheroid sprouting assay11 to assess the extent of their differences and their ability to model various aspects of angiogenesis12.
While both offer the advantages of being reliable and easily implementable, on a molecular level, the 3D spheroid sprouting assay was favorable in addressing key aspects of angiogenesis compared to in vivo data, such as metabolic switches or cell-matrix interactions. Since in vitro angiogenesis assays are used to evaluate the angiomodulatory potential of signaling pathways13 and to screen for therapeutic agents, transferability of in vitro results to in vivo settings is essential. Furthermore, the opportunity for omics-based analyses on the RNA and protein levels to characterize the molecular changes in response to targeted modulation of angiogenic processes under controlled conditions remains an important benefit compared to in vivo settings14,15.
In this publication, we present key assays for exploring angiogenesis-related questions through the utilization of a live-cell imaging migration assay and a spheroid sprouting assay, including subsequent molecular analyses like RNA sequencing for transcriptomic analysis and immunohistochemistry on the protein level.
Protocol
1. HUVEC cell culture
NOTE: Perform all following steps under sterile working conditions (sterile working bench).
- Expanding HUVECs
- For both angiogenesis assays, use pooled human umbilical vein endothelial cells (HUVECs) or human microvascular endothelial cells (HMVECs).
- Cultivate the cells until they reach 90% confluence in an uncoated T75 flask with endothelial cell growth medium (EGM) before performing a split. Perform medium change 3 times a week.
NOTE: To expedite growth, the medium can be changed daily. Do not use HUVECs beyond the sixth passage. The study received the most consistent results running all experiments with HUVECs of the same passage.
- Splitting HUVECs
- When HUVECs reach the desired confluence in the T75 flask, rinse them once with 5 mL of phosphate-buffered saline (PBS), followed by incubating the cells with 0.5% trypsin for 2 min in the incubator.
NOTE: Prolonged exposure to trypsin can damage HUVEC function. - Gently detach the cells by tapping on the flask. Confirm successful detachment under an inverted phase contrast microscope.
- Add 8 mL of EGM, transfer the solution to a tube, and pellet the cells by centrifugation at 250 x g for 3 min.
- If cell counting is necessary, resuspend the cells in 5 mL of EGM and use a Neubauer chamber. Otherwise, add 40 mL of EGM and distribute it onto 4 fresh T75 cell culture flasks.
- When HUVECs reach the desired confluence in the T75 flask, rinse them once with 5 mL of phosphate-buffered saline (PBS), followed by incubating the cells with 0.5% trypsin for 2 min in the incubator.
2. Scratch wound migration assay
NOTE: The scratch wound migration assay requires a duration of 3 days for completion (Figure 1). Perform all the following steps under sterile working conditions (sterile working bench).
- Plating and starving of cells (day 1)
- Using a specialized 96-well plate for live cell imaging, seed 20,000 HUVECs in 100 µL of EGM per well. Allow the cells to settle for 6 h in the incubator.
NOTE: It is advisable to optimize cell numbers considering the varying speed of vascular endothelial cell growth between suppliers and laboratories. The objective is to achieve a perfect monolayer the following day before initiating the scratch. Overpopulated wells can alter endothelial cell behavior, e.g., by contact inhibition16, while an incomplete monolayer hampers the analysis drastically. - Starve all wells overnight with 100 µL of EBM per well supplemented with 2% FBS.
NOTE: Be careful not to damage the cell monolayer, especially when using a multichannel pipette.
- Using a specialized 96-well plate for live cell imaging, seed 20,000 HUVECs in 100 µL of EGM per well. Allow the cells to settle for 6 h in the incubator.
- Preparing stimulation solutions (day 2)
- Dilute desired substances and controls in endothelial cell growth basal medium (EBM) supplemented with 2% FBS. For each well, use 100 µL of solution.
NOTE: EBM supplemented with 2% FBS serves as a negative control, while recombinant human vascular endothelial growth factor (VEGF) at 25 ng/mL can be used as a positive control. Using 96-well plates, it is recommended that the experiments be designed with a minimum of technical replicates (4 wells with identical conditions), although it is suggested that 8 be used for optimal results. Strive to minimize potential edge or corner effects when planning the layout.
- Dilute desired substances and controls in endothelial cell growth basal medium (EBM) supplemented with 2% FBS. For each well, use 100 µL of solution.
- Creating the scratch (day 2)
- Check cells under a microscope to verify a confluent cell monolayer and cell viability. Position the 96-well plate in a 96-well wound maker tool and generate the scratch by pressing down the lever of the device. Carefully lift off the lid of the wound maker tool before releasing the lever to prevent double scratching.
- Wash all wells twice using 200 µL of EBM per well supplemented with 2% FBS. Confirm under a microscope that debris was successfully removed.
- Cell stimulation (day 2)
- Add 100 µL of the prepared stimulation or control solutions per well.
- Imaging (day 2-3)
- Acquire images once per hour for a period of up to 24 h using an automated schedule provided by the live-cell imaging microscope.
NOTE: To ensure good image quality, scan every well immediately after transferring it into the incubator housing the live-cell imaging system. A 30 min acclimation time in the incubator helps to get a better picture quality. The microscope software is programmed to acquire one image per hour for each well at the midline of the defined scratch.
- Acquire images once per hour for a period of up to 24 h using an automated schedule provided by the live-cell imaging microscope.
- In the absence of a wound maker tool and a live cell imager, follow the steps described below.
- Use a pipette tip in a 24-well plate to induce a scratch. Take images with a classic cell culture microscope at specific intervals. For this purpose, utilize a motorized microscope equipped with precise x and y tracking capabilities. This enables automatic positioning, ensuring consistent image capture at predefined locations.
NOTE: When manual imaging is necessary it is a viable option to only image T0 as well as one time point 'x' hours following treatment. In the described protocol using HUVECs the time point T12 (12 h post-stimulation) was an attractive time point with a clear separation between negative and VEGF-stimulated positive control. However, running initial time course experiments with imaging every 2 h or 1 h is recommended to determine the optimal time point for the specific experimental settings in each lab.
- Use a pipette tip in a 24-well plate to induce a scratch. Take images with a classic cell culture microscope at specific intervals. For this purpose, utilize a motorized microscope equipped with precise x and y tracking capabilities. This enables automatic positioning, ensuring consistent image capture at predefined locations.
- Analysis (day 3)
- The live-cell imaging microscope software may enable direct analysis of the acquired images. Define masks delineating the cell lawn, initial scratch, and relative wound density regions. Determine cell boundaries based on contrast differences.
NOTE: A segmentation adjustment of 1.6, a hole file of 500 µm2, an adjusted size of 1 pixel, an area >500 µm, and an eccentricity >0.6 were used for experiments run with HUVECs. Thorough visual vetting of the cell detection parameters relative to actual images is essential. With optimized configurations, the software calculates relative wound density (RWD) over time automatically and generates corresponding temporal graphs with the standard deviation (SD) of the technical replicates. This data can then be used for statistical analysis.
- The live-cell imaging microscope software may enable direct analysis of the acquired images. Define masks delineating the cell lawn, initial scratch, and relative wound density regions. Determine cell boundaries based on contrast differences.
3. RNA extraction with 2D cultivated cells
NOTE: Perform all the following steps until step 3.3 under sterile working conditions (sterile working bench).
- Plating cells (day 1)
- Detach HUVECs per step 1.2 of the HUVEC cell culture protocol.
- Seed 50,000 cells per well in a 12-well plate with 2 mL of EGM per well.
- Change the medium after 6 h to EBM containing 2% FBS (starving media) for starving overnight.
- Treatment of the cells (day 2)
- Dilute agents of interest in EBM supplemented with 2% FBS. Replace starving media with the prepared dilution and incubate cells for the desired time in an incubator.
- RNA extraction (day 2-3)
- Lyse the cells with 750 µL of Trizol per well and incubate for 10 min on an orbital shaker.
- Resuspend the samples with a 1000 µL pipette a few times and then store them in specific low-binding RNA tubes (volume 1.5 mL). Store at -80 °C.
4. Immunocytochemistry with 2D cultivated cells
NOTE: Perform all following steps until step 4.4 under sterile working conditions (sterile working bench).
- Preparation of coverslips
- Incubate 100 coverslips with a diameter of 12 mm in 5% potassium hydroxide in methanol in a 50 mL tube for 30 min while shaking the applied solution.
NOTE: This is a very important step in deprotonating the surface of the coverslips and improving the conditions for coating and culture. At this point, ensuring that the coverslips are well distributed within the medium and don’t get dry and glued together is highly recommended. - Wash the coverslips for 30 min 4 times with demineralized water.
- After washing, transfer them into 70% isopropyl solution for storage.
- Incubate 100 coverslips with a diameter of 12 mm in 5% potassium hydroxide in methanol in a 50 mL tube for 30 min while shaking the applied solution.
- Coating of coverslips
- Lean the coverslips individually against the wall of a square Petri dish to allow the evaporation of isopropyl alcohol.
- After 5 min transfer the coverslips into a 24 well plate (one coverslips per well). Apply 1 mL of a solution of 10 mg/mL collagen I in PBS to each well and incubate for 60 min at 37 °C.
- Then, wash the coverslips 4-5 times for 5 min with PBS.
- Cultivating cells
- Detach HUVECs following the procedures outlined in step 1.2 of the HUVEC cell culture protocol.
- Seed 50,000 cells per well in a 24-well plate with 2 mL of EGM per well.
- After 6 h, change the medium to EBM containing 2% FBS to allow the cells to attach to the well and starve the cells overnight.
- On the next day, dilute agents of interest in EBM supplemented with 2% FBS. Replace the starving solution in wells with the prepared dilution and incubate cells for the desired time in an incubator.
- Fixation and blocking
- Cultivate the cells for the desired time, then wash the cells with PBS for 5 min.
- Fix the cells with 2% paraformaldehyde (PFA) in PBS for 20 min at room temperature (RT).
- Wash with PBS 3-4 times for 5 min.
- Incubate samples with blocking buffer containing 5% normal goat serum (NGS), 0.1% Triton-X-100 in PBS for 1 h at RT.
NOTE: At this step, it is important to wash the samples to guarantee that PFA is removed completely. Choose the species of the serum based on the origin of the secondary antibody.
- Staining
- Incubate the samples overnight at 4 °C in the blocking buffer with the primary antibody.
- On the next day, to remove any unbound primary antibody, wash the samples 3-4 times with PBS for 5 min each.
- Subsequently, incubate the samples for 1 h at RT with the corresponding secondary antibody and Phalloidin-FITC, diluted together in the blocking buffer.
- Wash an additional 3-4 times.
- Invert the coverslips onto slides with a small droplet of DAPI-containing mounting medium, allow to dry in the dark, and seal with nail polish. Store samples in the dark at 4 °C.
5. Spheroid sprouting assay
NOTE: The spheroid sprouting assay requires 3 days for completion (Figure 2). Perform all following steps until step 5.7 under sterile working conditions (sterile working bench).
- Generating spheroids in hanging drops (day 1)
- Detach HUVECs following the procedures outlined in step 1.2 of the HUVEC cell culture protocol.
- Combine 200,000 cells with 8 mL of EGM and 2 mL of methocel stock solution, ensuring thorough mixing. Please refer to Supplementary File 1 for instructions on preparing the methocel stock solution.
- Dispense 25 µL droplets of the solution onto the inverted inner surface of a large, squared Petri dish. Gently rotate the lid by 180° and position it on the bottom of the cell culture vessel so that the droplets are hanging. Place the vessel in a cell culture incubator overnight.
- Prepare the collagen gel (day 2)
- Mix thoroughly 2.3 mL of rat tail collagen type I with 0.280 mL of 10x Medium 199 until the mixture achieves a consistently even yellow hue. Keep this mixture on ice throughout the entire process.
- Titrate the collagen gel (day 2)
- Titrate the mixture to achieve a physiological pH, indicated by an orange color, using NaOH (2 N).
- Add 50 µL of HEPES buffer to the final product.
NOTE: To ensure an optimal dynamic range for the assay, it is imperative to approach as closely as possible to a physiological pH. Various batches of collagen may require different amounts of NaOH. Keep the mixture strictly on ice throughout the process and homogeneously mixed all the time.
- Harvesting spheroids (day 2)
- Rinse hanging drops from cell culture lids with 20 mL of PBS.
- Centrifuge the solution for 7 min at 250 x g. Discard the supernatant and resuspend the spheroids in 0.1 mL of FBS and 0.4 mL of EGM by gently tapping the tube.
- Add 2 mL of methocel stock solution and mix thoroughly. Use a pipette with a minimum capacity of 5 mL to avoid damaging the spheroids.
- Adding spheroids to the 3D collagen matrix (day 2)
- Add 2 mL of the prepared collagen mixture to the resuspended spheroids and mix thoroughly.
- Dispense 0.5 mL of the resulting mixture into the wells of a 24-well plate and incubate in a cell incubator for 30 min.
- Stimulating spheroids in a 3D collagen matrix (day 2)
- Prepare a dilution of desired substances in EBM. For each well, use 100 µL of solution.
NOTE: EBM serves as a negative control, while vascular endothelial growth factor (VEGF) at 25 ng/mL (final concentration in 500 µL of collagen matrix and 100 µL EBM layer) can be used as a positive control. - Incubate for the desired time.
NOTE: In a standard assay, 18-24 h is recommended, but timing needs to be carefully optimized in each lab.
- Prepare a dilution of desired substances in EBM. For each well, use 100 µL of solution.
- Imaging spheroids (day 3)
- Utilize an inverted microscope and imaging platform to capture images of all spheroids that are not in contact with the well rim, with each other, or show signs of damage. Maintain consistent settings and zoom levels across all conditions.
- Analysis (day 3)
- Utilize the measuring tool in ImageJ Fiji to ascertain the length of all sprouts from each spheroid in every image.
NOTE: Use the plugin provided in Supplementary File 2, which makes the analysis more efficient and generates an output .txt file. - Import the data into a spreadsheet, R, or any other preferred software, and use the average cumulative length of all sprouts per spheroid as the readout for each condition.
NOTE: Results from different gels cannot be combined in the form of absolute sprouting lengths. Data from each treatment group need to be normalized to the EBM or VEGF control group (relative sprouting length) before data can be merged.
- Utilize the measuring tool in ImageJ Fiji to ascertain the length of all sprouts from each spheroid in every image.
6. RNA extraction with 3D cultivated cells
- Spheroid sprouting assay
- Perform the spheroid sprouting assay as described in section 5.
- Lysis of collagen gel
- Wash spheroid gels with warm PBS for 5 min.
- Cut the spheroid gels into quarters and transfer all 4 quarters into a 50 mL tube. Use a scalpel for the mechanical dissection of the gels.
- Treat gel samples with lysis solution (containing 2 mg/mL Collagenase D in PBS) and incubate for 45 min at 37 °C on a shaker.
- Wash gels gently with PBS supplemented by 2 mM EDTA to stop the reaction of the lysis buffer.
- RNA extraction
- Resuspend the lysed gels in 750 µL of Trizol and incubate for 10 min to guarantee sufficient RNA extraction as described in step 3.3.
- Resuspend the samples with a 1000 µL pipette a few times and transfer samples to specific low-binding RNA tubes (volume 1.5 mL). Store samples at -80 °C.
7. Immunocytochemistry of spheroids in a 3D collagen matrix
- Spheroid sprouting assay
- Perform the spheroid sprouting assay as described in section 5.
- Fixation and blocking
- Fix the gels with 4% PFA in PBS for 1 h.
NOTE: To ensure good fixation of the cells in the gel, gels need to be detached carefully on the sides of the wells with a scalpel and tweezers following rinsing with PBS. - Wash gels gently 3-4 times with PBS and then fully detach them from the surface of the wells and transfer them into new 24-well plates.
- Incubate gels for 1 h at RT with the blocking solution (containing 5% NGS and 0.1% Triton-X-100 in PBS) on an orbital shaker.
- Fix the gels with 4% PFA in PBS for 1 h.
- Staining
- Incubate the samples overnight at 4 °C on an orbital shaker with the primary antibody diluted in blocking buffer.
NOTE: Turning the gels during this incubation did not improve staining quality. - The next day, wash the gels gently 4-5 times with PBS for 5 min each.
- Incubate the corresponding secondary antibody, diluted with Phalloidin-FITC in blocking buffer overnight at 4 °C on an orbital shaker.
- Conduct 4-5 additional washing steps with PBS.
- Transfer the gels onto microscope slides and mount them with two drops of DAPI-containing mounting medium on a coverslip placed onto each gel. Allow the samples to dry before sealing with nail polish and store them in the dark at 4 °C.
- Incubate the samples overnight at 4 °C on an orbital shaker with the primary antibody diluted in blocking buffer.
- Microscopy
- Image all samples on a confocal microscope. Use the 20x objective and focus on the region of interest. Acquire images as Z-stacks, as well as individual focal plane images.
NOTE: Acquiring Z-stack and focal plane images at multiple sections throughout the 3D sample is essential to capture the complete representation, especially for thick 3D samples.
- Image all samples on a confocal microscope. Use the 20x objective and focus on the region of interest. Acquire images as Z-stacks, as well as individual focal plane images.
8. Human retinal microvascular endothelial cells (HRMVECs)
NOTE: All the described steps can also be performed with microvascular endothelial cells, e.g., human retinal microvascular endothelial cells (HRMVECs). In that case, the medium needs to be switched to a specific microvascular endothelial cell medium containing 10% fetal bovine serum (FBS) for cultivation as well as specific steps in each assay. HRMVEC-specific differences to the assay protocols are outlined below:
- Scratch wound assay
- Cultivate cells in 10% FBS microvascular endothelial cell medium instead of EGM.
- Seed 20,000 cells/well.
- Do not starve the cells overnight.
- After performing the scratch, change the medium to basal endothelial cell medium containing 5% FBS instead of EBM with 2% FBS.
- Immunocytochemistry of 2D cultivated HRMVECs
- Cultivate the cells in 10% microvascular endothelial cell medium instead of EGM.
- Do not starve the cells overnight.
- Change the medium right before treatment to EBM + 5% FBS instead of EBM+2% FBS.
- Spheroid sprouting assay
- Seed the cells as hanging drops with microvascular endothelial cell medium containing 10% FBS instead of EGM.
- Immunocytochemistry of spheroids in a 3D collagen matrix
- Seed the cells as hanging drops with microvascular endothelial cell medium containing 10% FBS instead of EGM.
Representative Results
For the migration assay, it is crucial to thoroughly examine the images captured at the t = 0 h time point to ensure the presence of a fully formed cell monolayer is accurately detected by the system (Figure 1B). Additionally, the clarity and straightness of the scratch border should be confirmed (Figure 1B). The cell-free area ought to be largely free of debris. At the end of the assay, a group stimulated with, for example, 25 ng/mL VEGF as a positive control should show successful migration in the original scratched area (Figure 1B). An instance of technical issues might manifest as patchy cell growth, double scratching, or waviness in the scratch borders (Figure 1C). Furthermore, it is important to establish a substantial dynamic range between the negative control (EBM-stimulated samples) and the positive control (VEGF samples) (Figure 1D). In our experimental setup, we consider a 25% increase in RWD after 12 h as an optimal range (Figure 1D). In a technical problematic assay, positive and negative controls do not separate correctly, indicating a low dynamic range (Figure 1E).
Immunocytochemical staining of HUVECs on coverslips, visualized under a fluorescence microscope to validate successful staining (Figure 1F). It is important to survey a majority of the cells and identify a characteristic region of the slide. Staining specificity is assessed using a control slide treated only with a secondary antibody to exclude nonspecific binding (Figure 1F). Specific staining signals should be critically examined according to the known subcellular localization of the target protein. We use subcellular localization patterns along with the absence of signal in the control sample as indices of a successful immunostaining protocol.
Similarly, certain aspects must be considered for high-quality spheroid sprouting assays. The quality should, in this case, be assessed at the end of the assay at, for example, t = 24 h. The collagen matrix observed under the microscope should exhibit homogeneity and should not appear friable or rough (Figure 2B). Spheroids stimulated with just basal medium as a negative control and 25 ng/mL VEGF as a positive control should be clearly distinguishable (Figure 2B). Furthermore, spheroids should float in the gel and not drop to the ground, where they disperse, indicating low quality (Figure 2C). This issue often arises from inadequate cooling of the collagen during titration of the mixture or from over- or under-titration of the pH (Figure 2C). A minimum of 10 spheroids per well should be imageable for each well. The negative control (EBM-stimulated samples) should demonstrate a moderate baseline sprouting rate, while VEGF, serving as the positive control, ideally doubles or, preferably, triples the RSL compared to the controls (Figure 2D, left graph). The number of sprouts should also show a similar dynamic range (Figure 2D, right graph). A technical suboptimal assay will not show this dynamic range between controls and could, therefore, lead to a false interpretation of data (Figure 2E).
Analog to 2D stainings, examination of the slides under a fluorescence microscope and critical assessment of the stained structures are of great importance for successful staining (Figure 2F). If the cellular localization of the structures and the signal intensity in comparison to the control is comprehensible, it can be assumed that the staining was successful.
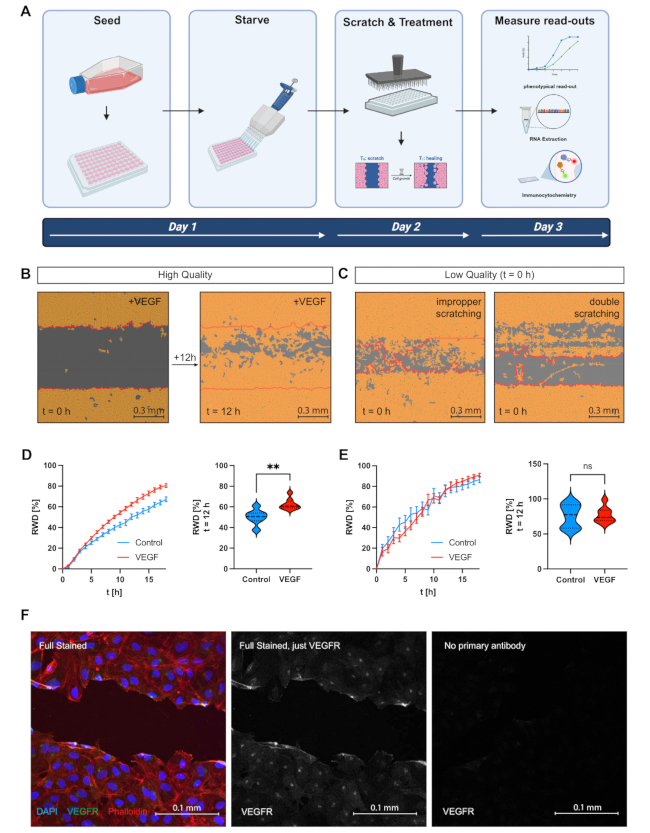
Figure 1: The live-cell imaging scratch wound migration assay. (A) Scheme of the scratch wound migration assay. (B) Images showcasing a high-quality technical replicate at t = 0 h and t = 12 h. Cells have been stimulated with 25 ng/mL VEGF to serve as a positive control. (C) Examples of two low-quality technical replicates at t = 0 h due to improper scratching or double scratching. (D) Results of a successful scratch wound migration assay. The correct separation of positive control stimulated with 25 ng/mL VEGF and negative control just treated with basal medium was further visualized by a violin plot at t = 12 h. Data include 7-8 technical replicates per group. (E) Exemplary results highlighting a failed Scratch Wound Migration Assay. Suboptimal separation of positive control stimulated with 25 ng/mL VEGF and negative control just treated with basal medium was visualized by a violin plot at t = 12 h. Data include 7-8 technical replicates per group. (F) Exemplary images of HUVECs seeded on coverslips and stained for DAPI, phalloidin, and VEGF-R2. Please click here to view a larger version of this figure.
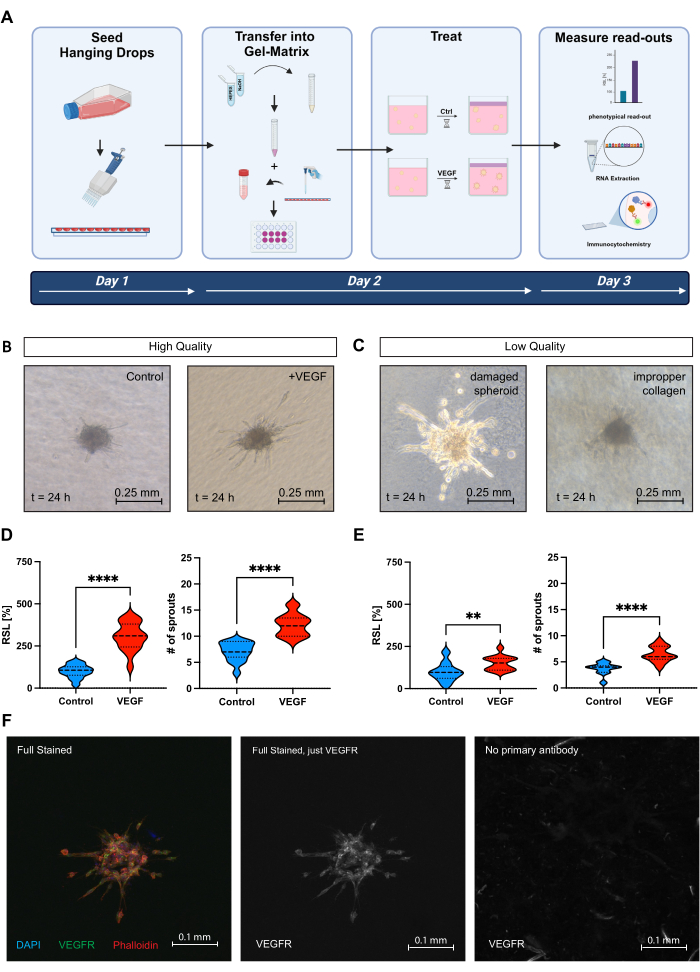
Figure 2: The spheroid sprouting assay. (A) Scheme of the spheroid sprouting assay. (B) Images showcasing the negative control (basal medium stimulated spheroids) and positive control (VEGF 25 ng/mL stimulated spheroids) of an ideal technical assay at the end at t = 24 h. (C) Examples of two low-quality spheroids at t = 24 h. Left images visualize a damaged spheroid that dropped to the bottom of the plate, which results in a loosened spheroid body and an abnormally enhanced sprouting rate. The right image visualizes collagen with noticeable color patches, creating a highly heterogeneous environment that significantly compromises the assay's quality. (D) Analysis of the RSL (left graph) and number of sprouts (right graph) of a successful assay with a good dynamic range between negative and positive control. Data is visualized using a violin plot and includes 10-25 spheroids per group. (E) Analysis of the RSL (left graph) and number of sprouts (right graph) of a technical problematic assay with a suboptimal dynamic range between negative and positive control. Data is visualized using a violin plot and includes 10-25 spheroids per group. (F) Exemplary images of HUVECs from the spheroid sprouting assay stained for DAPI, phalloidin, and VEGF-R2.c
Supplementary File 1: Instructions for preparing the methocel stock solution. Please click here to download this File.
Supplementary File 2: SpheroidCount.ijm plugin. Please click here to download this File.
Discussion
In this report, we presented a spectrum of techniques with functional and molecular readouts to study angiogenesis in vitro.
The migration assay represents a well-established technique used across all fields of wet laboratory work. We chose the commercially available live-cell imaging approach to take advantage of the 96-well format suitable for screening and dose-response experiments, the standardized and reproducible wound size created by the WoundMaker tool, the opportunity to observe the migration kinetic through time-lapse imaging over up to 24 h as well as the automated image quantification software. However, the presence of a life-cell imaging system equipped with angiogenesis analysis software in the lab or a core facility is needed. Numerous alternative experimental setups have been established that do not necessitate specific live-cell imaging microscopes or other special equipment10.
Based on our experience, it is crucial to optimize the analyzer settings, as HUVECs and other vascular endothelial cell types can tend to provide poor contrast, thereby hindering the automated readout by the microscope software. To address this issue, an additional fluorescence staining step (e.g., live-cell staining) can be introduced. Live-cell imaging systems support such staining and even multi-color staining for tracking a protein of interest. One major limitation of the assay is that differentiation between cell migration and proliferation is not always straightforward and that assay results represent a combination of both processes. The live cell imaging solution used in this study tries to tackle this issue by introducing the relative wound density (RWD). In contrast to traditionally used confluence in the scratched area, the RWD assesses the confluence within the scratched area relative to the outside scratched area. Migration in the scratched area elevates the RWD by increasing confluence within it while concurrently reducing confluence outside the scratched area due to cell migration. Conversely, cell proliferation also increases confluence outside the scratched area, thereby decreasing RWD. It is important to recognize this as a strategic analysis approach to only lessen the impact of proliferation on the final readout. Complementary proliferation assays may be needed to address that issue. Alternatively, cell cycle inhibitors such as mitomycin C can be added to the assay to block proliferation. Implementing protocols with these inhibitors needs to be carefully optimized in each lab to achieve the desired effect without reducing the dynamic range of the assay. Furthermore, by increasing the scanning frequency of each well, this setup offers an easily implementable opportunity to precisely track moving cells on an individual level if such detailed tracking is of particular interest.
While the live-cell imaging migration assay demonstrates the advantage of scalability and automated, unbiased analyses, our previous study highlighted the spheroid sprouting assay’s ability to capture more intricate details of angiogenesis. This includes fundamental aspects like tip and stalk cell formation, cell-matrix interaction, and a glycolytic switch12. To guarantee assay reproducibility, an investigator must be experienced in preparing the collagen matrix, as variations in pH and temperature during this process can impact results. Furthermore, the analysis is conducted manually. Image analysis is hence time consuming and contains the risk for bias. Proper masking of conditions during analysis is essential to prevent the introduction of bias. To address this concern, we recently proposed a neural network-based approach to identify and mitigate potential bias in the analysis of sprouting assays, which can be easily implemented17. In conclusion, the best approach to validate results is to combine both assays. However, it is crucial to acknowledge that differences may arise due to the distinct nature of 2D and 3D settings12.
As there is a wide selection of 2D and 3D in vitro angiogenesis assays, it is important to consider the advantages and disadvantages of the presented assays compared to other methods. The scratch wound assay, for example, focuses on the 2D horizontal migration on a plastic dish, while the Boyden chamber assay characterizes vertical migration through a mesh insert with a chemoattractant in the “outer chamber”. Based on its simple setup and readout, the scratch wound assay allows for high-throughput experiments, particularly in its 96-well format, providing high reproducibility as well as the possibility of visualizing the experiment and conducting a time-lapse. Unfortunately, the disadvantages of the scratch wound assay are the difficulty in differentiating between proliferation and migration, the need for adherent cells, and, in the absence of a wound-maker tool, higher variability due to unequal scratches. The readout of the Boyden Chamber Assay has the advantage of measuring the chemotactic effect of soluble substances on motile cells18 and highlights both invasion and migration. Unfortunately, disadvantages are the poor reproducibility and high variance of the assay, the inability to visualize the movement of the cells, the high number of migrated cells required to obtain a signal, the quite long and elaborate setup and readout, as well as the lack of the opportunity for a time-lapse setting19.
The spheroid sprouting assay quantifies the sprouting of endothelial cells into a 3D gel matrix, while the tube formation assay characterizes the formation of cellular tubes on the surface of a gel matrix20. The sprouting assay has the advantage of characterizing the invasion of cells into a matrix as well as proliferation and migration. Disadvantages are the higher variance of the titrated collagen gel and, based on this, the quality of spheroids. The advantage of the tube formation assay is the formation of vascular tubes after a distinct amount of time (dependent on the cytokine) and an easy approach. Disadvantages are the lack of invasion of cells and the effect of the Matrigel itself on tube formation21.
All of the presented experiments were performed with HUVECs. It is well-known that significant differences in the behavior of vascular endothelial cells from different origins (e.g., macro-vascular vs. micro-vascular) exist22,23. It may thus be necessary to validate experimental results with other vascular endothelial cell lines. The experimental settings were transferrable for human retinal microvascular endothelial (HRMVECS), except for the fact that HRMVECs require higher FBS concentrations. On the other hand, the commonly used bovine aortic endothelial cell line (BAECs) could not be used in the spheroid sprouting assay due to an excessive basal sprouting rate.
Both presented assays share the limitation that they are confined to a single cell type. Given that angiogenesis in vivo is an intricate multicellular process, this crucial aspect is not accurately captured by either assay. Co-culture variations for both assays have been developed, with a particular focus on the spheroid sprouting assay, and recently discussed10,24. Alternatively, validating the data directly in relevant in vivo angiogenesis models represents a viable option. Established assays for this purpose include the mouse model for oxygen-induced retinopathy (OIR)25, the laser-induced choroidal neovascularization (Laser-CNV) model26, and the plug assay27.
Overall, both the 2D live-cell imaging migration assay and the 3D spheroid sprouting assay, in conjunction with the presented molecular analysis tools, offer a robust platform for angiogenesis research and are widely recognized within the angiogenesis community. Combining these assays with subsequent in vivo analyses to validate findings provides a solid foundation for investigating specific angiogenesis-related questions.
Declarações
The authors have nothing to disclose.
Acknowledgements
The authors thank Sophie Krüger and Gabriele Prinz for their excellent technical support. We thank Sebastian Maier for developing the ImageJ plugin to quantify spheroid sprouts and the Lighthouse Core Facility, Zentrum für Translationale Zellforschung (ZTZ), Department of Medicine I, University Hospital Freiburg for the use of the IncuCyte system. The graphics were created with biorender.com. This work was supported by the Deutsche Forschungsgemeinschaft [Bu3135/3-1 + Bu3135/3-2 to F.B], the Medizinische Fakultät der Albert-Ludwigs- Universität Freiburg [Berta-Ottenstein-Program for Clinician Scientists and Advanced Clinician Scientists to F.B.], the Else Kröner-Fresenius-Stiftung [2021_EKEA.80 to F.B.] the German Cancer Consortium [CORTEX fellowship for Clinician Scientists to J.R.] and the Volker Homann Stiftung [to J.N.+F.B.] and the "Freunde der Universitäts-Augenklinik Freiburg e.V." [to P.L.]
Materials
| 10x Medium 199 | Sigma-Aldrich | M0650 | |
| 2-(4-(2-Hydroxyethyl)1-piperazinyl)-ethan-sulfonsäure | PAN-Biotec | P05-01100 | HEPES |
| Alexa Fluor 647-conjugated AffiniPure F(ab)‘2-Fragment | Jackson IR | 115-606-072 | |
| Axio Vert. A1 | Zeiss | ||
| CapturePro 2.10.0.1 | JENOTIK Optical Systems | ||
| Collagen Type 1 rat tail | Corning | 354236 | |
| Collagenase D | Roche | 11088858001 | |
| Endothelial Cell Basal Medium | Lonza | CC-3156 | EBM |
| Endothelial Cell Growth Medium | Lonza | CC-3162 | EGM |
| Ethylenediaminetetraacetic acid | Serva | 11290.02 | EDTA |
| Fetal bovine serum | Bio&SELL | S 0615 | FBS |
| Human Umbilical Vein Endothelial Cells, pooled | Lonza | C2519A | HUVEC |
| IncuCyte ImageLock 96-well plates | Sartorius | 4379 | |
| Incucyte S3 Live-Cell Analysis System | Sartorius | ||
| Methocel | Sigma | m-0512 | |
| Microvascular Endothelial Cell Medium with 10% FBS | PB-MH-100-4090-GFP | PELOBiotech | |
| NaOH | Carl Roth | P031.2 | |
| Phalloidin-Fluorescein Isothiocyanate Labeled (0.5 mg/mL Methanol) | Sigma-Aldrich | P5282-.1MG | Phalloidin-FITC |
| Phosphate-buffered saline | Thermo Fisher Scientfic | 14190-094 | PBS |
| Primary Human Retinal Microvascular Endothelial Cells | Cell Systems | ACBRI 181 | |
| ProLong Glass Antifade Mountant with NucBlue | Invitrogen by ThermoFisher Scientific | 2260939 | |
| QIAzol Lysis Reagent | QIAGEN | 79306 | |
| recombinant human Vascular Endothelial Growth Factor | PeproTech | 100-20 | VEGF |
| Squared petri dish | Greiner | 688102 | |
| Trizol | Qiagen | 79306 | |
| Trypsin | PAN-Biotec | P10-024100 | |
| VEGF-R2 (monoclonal) | ThermoFisher Scientific Inc. | B.309.4 | |
| WoundMaker | Sartorius | 4493 |
Referências
- Risau, W. Mechanisms of angiogenesis. Nature. 386 (6626), 671-674 (1997).
- Gariano, R. F., Gardner, T. W. Retinal angiogenesis in development and disease. Nature. 438 (7070), 960-966 (2005).
- Mancuso, M. R., Kuhnert, F., Kuo, C. J. Developmental angiogenesis of the central nervous system. Lymphat Res Biol. 6 (3-4), 173-180 (2008).
- Tonnesen, M. G., Feng, X., Clark, R. A. Angiogenesis in wound healing. J Investig Dermatol Symp Proc. 5 (1), 40-46 (2000).
- Folkman, J. Role of angiogenesis in tumor growth and metastasis. Semin Oncol. 29 (6 Suppl 16), 15-18 (2002).
- Crawford, T. N., Alfaro, D. V., Kerrison, J. B., Jablon, E. P. Diabetic retinopathy and angiogenesis. Curr Diabetes Rev. 5 (1), 8-13 (2009).
- Ng, E. W., Adamis, A. P. Targeting angiogenesis, the underlying disorder in neovascular age-related macular degeneration. Can J Ophthalmol. 40 (3), 352-368 (2005).
- Blanco, R., Gerhardt, H. Vegf and notch in tip and stalk cell selection. Cold Spring Harb Perspect Med. 3 (1), a006569 (2013).
- Hellström, M., Kalén, M., Lindahl, P., Abramsson, A., Betsholtz, C. Role of PDGF-B and PDGFR-β in recruitment of vascular smooth muscle cells and pericytes during embryonic blood vessel formation in the mouse. Development. 126 (14), 3047-3055 (1999).
- Nowak-Sliwinska, P., et al. Consensus guidelines for the use and interpretation of angiogenesis assays. Angiogenesis. 21 (3), 425-532 (2018).
- Korff, T., Augustin, H. G. Tensional forces in fibrillar extracellular matrices control directional capillary sprouting. J Cell Sci. 112 (Pt 19), 3249-3258 (1999).
- Rapp, J., et al. 2D and 3D in vitro angiogenesis assays highlight different aspects of angiogenesis. Biochim Biophys Acta Mol Basis Dis. 1870 (3), 167028 (2024).
- Rapp, J., et al. STAT3 signaling induced by the IL-6 family of cytokines modulates angiogenesis. J Cell Sci. 136 (1), jcs260182 (2023).
- Heiss, M., et al. Endothelial cell spheroids as a versatile tool to study angiogenesis in vitro. FASEB J. 29 (7), 3076-3084 (2015).
- Rapp, J., et al. Oncostatin m reduces pathological neovascularization in the retina through müller cell activation. Invest Ophthalmol Vis Sci. 65 (1), 22 (2024).
- Fagotto, F., Gumbiner, B. M. Cell contact-dependent signaling. Dev Biol. 180 (2), 445-454 (1996).
- Rapp, J., et al. Addressing bias in manual segmentation of spheroid sprouting assays with U-Net. Mol Vis. 29, 197-205 (2023).
- Boyden, S. The chemotactic effect of mixtures of antibody and antigen on polymorphonuclear leucocytes. J Exp Med. 115 (3), 453-466 (1962).
- Guy, J. B., et al. Evaluation of the cell invasion and migration process: A comparison of the video microscope-based scratch wound assay and the Boyden chamber assay. J Vis Exp. 129, 56337 (2017).
- Decicco-Skinner, K. L., et al. Endothelial cell tube formation assay for the in vitro study of angiogenesis. J Vis Exp. 91, e51312 (2014).
- Vernon, R. B., Angello, J. C., Iruela-Arispe, M. L., Lane, T. F., Sage, E. H. Reorganization of basement membrane matrices by cellular traction promotes the formation of cellular networks in vitro. Lab Invest. 66 (5), 536-547 (1992).
- Craig, L. E., Spelman, J. P., Strandberg, J. D., Zink, M. C. Endothelial cells from diverse tissues exhibit differences in growth and morphology. Microvasc Res. 55 (1), 65-76 (1998).
- Müller, A. M., Hermanns, M. I., Cronen, C., Kirkpatrick, C. J. Comparative study of adhesion molecule expression in cultured human macro-and microvascular endothelial cells. Exp Mol Pathol. 73 (3), 171-180 (2002).
- Chiew, G. G. Y., Wei, N., Sultania, S., Lim, S., Luo, K. Q. Bioengineered three-dimensional co-culture of cancer cells and endothelial cells: A model system for dual analysis of tumor growth and angiogenesis. Biotechnol Bioeng. 114 (8), 1865-1877 (2017).
- Smith, L. E., et al. Oxygen-induced retinopathy in the mouse. Invest Ophthalmol Vis Sci. 35 (1), 101-111 (1994).
- Lambert, V., et al. Laser-induced choroidal neovascularization model to study age-related macular degeneration in mice. Nat Protoc. 8 (11), 2197-2211 (2013).
- Kastana, P., et al. Matrigel plug assay for in vivo evaluation of angiogenesis. Methods Mol Biol. 1952, 219-232 (2019).

.
