The ChroP Approach Combines ChIP and Mass Spectrometry to Dissect Locus-specific Proteomic Landscapes of Chromatin
Summary
By combining native and crosslinking chromatin immunoprecipitation with high-resolution Mass Spectrometry, ChroP approach enables to dissect the composite proteomic architecture of histone modifications, variants and non-histonic proteins synergizing at functionally distinct chromatin domains.
Abstract
Chromatin is a highly dynamic nucleoprotein complex made of DNA and proteins that controls various DNA-dependent processes. Chromatin structure and function at specific regions is regulated by the local enrichment of histone post-translational modifications (hPTMs) and variants, chromatin-binding proteins, including transcription factors, and DNA methylation. The proteomic characterization of chromatin composition at distinct functional regions has been so far hampered by the lack of efficient protocols to enrich such domains at the appropriate purity and amount for the subsequent in-depth analysis by Mass Spectrometry (MS). We describe here a newly designed chromatin proteomics strategy, named ChroP (Chromatin Proteomics), whereby a preparative chromatin immunoprecipitation is used to isolate distinct chromatin regions whose features, in terms of hPTMs, variants and co-associated non-histonic proteins, are analyzed by MS. We illustrate here the setting up of ChroP for the enrichment and analysis of transcriptionally silent heterochromatic regions, marked by the presence of tri-methylation of lysine 9 on histone H3. The results achieved demonstrate the potential of ChroP in thoroughly characterizing the heterochromatin proteome and prove it as a powerful analytical strategy for understanding how the distinct protein determinants of chromatin interact and synergize to establish locus-specific structural and functional configurations.
Introduction
Chromatin is a highly dynamic nucleoprotein complex that is involved as primary template for all DNA-mediated processes. The nucleosome is the basic repeated unit of chromatin and consists of a proteinaceous octameric core containing two molecules of each canonical histone H2A, H2B, H3 and H4, around which 147 bp of DNA are wrapped1,2. All core histones are structured as a globular domain and a flexible N-terminal “tail” that protrudes outside the nucleosome. One of the major mechanisms for regulating chromatin structure and dynamics is based on covalent post-translational modifications (PTMs), which mainly occur on the N-termini of histones3,4. Histone modifications can function either by altering the higher order chromatin structure, by changing contacts between histones-DNA or between nucleosomes, and thus controlling the accessibility of DNA-binding proteins (cis mechanisms), or by acting as docking sites for regulatory proteins, either as single units, or embedded in multimeric complexes. Such regulating proteins can exert their function in different ways: by modulating directly gene expression (i.e. TAF proteins), or by altering the nucleosome positioning (i.e. chromatin remodeling complexes) or by modifying other histone residues (i.e. proteins with methyl-transferase or acetyl-transferase activity) (trans mechanisms)5. The observation that distinct PTM patterns cluster at specific chromatin loci led to the elaboration of the hypothesis that different modifications at distinct sites may synergize to generate a molecular code mediating the functional state of the underlying DNA. The "histone code hypothesis" has gained large consensus in the years but its experimental verification has been held back by technical limitations6,7.
Mass spectrometry (MS)-based proteomics has emerged as a powerful tool to map histone modification patterns and to characterize chromatin-binding proteins8. MS detects a modification as a specific Δmass between the experimental and theoretical mass of a peptide. At the level of individual histones, MS provides an unbiased and comprehensive method to map PTMs, allowing the detection of new modifications and revealing interplays among them9-14.
In recent years, a number of strategies have been developed to dissect the proteomic composition of chromatin, including the characterization of intact mitotic chromosomes15, the identification of soluble hPTM-binding proteins16-18 and the isolation and analysis of specific chromatin regions (i.e. telomeres)19,20. However, the investigation of the locus-specific synergies between histone PTMs, variants, and chromatin-associated proteins is still incomplete. Here, we describe a new approach, named ChroP (Chromatin Proteomics)21, that we have developed to efficiently characterize functionally distinct chromatin domains. This approach adapts chromatin immunoprecipitation (ChIP), a well-established protocol used in epigenetic research, for the efficient MS-based proteomic analysis of the enriched sample. We have developed two different protocols, depending on the type of chromatin used as input and the question addressed by MS; in particular: 1) ChIP of unfixed native chromatin digested with MNase is employed to purify mono-nucleosomes and to dissect the co-associating hPTMs (N-ChroP); 2) ChIP of crosslinked chromatin fragmented by sonication is used in combination with a SILAC-based interactomics strategy to characterize all co-enriching chromatin binding proteins (X-ChroP). We illustrate here the combination of N- and X-ChroP for enriching and studying heterochromatin, using H3K9me3 as bait for the immunoprecipitation steps. The use of ChroP can be extended to study either distinct regions on chromatin, or changes in the chromatin composition within the same region during the transition to a different functional state, thus paving the way to various applications in epigenetics.
Protocol
1. Cell Culture
- Standard medium for native ChIP
- Grow HeLa cells in Dulbecco's modified Eagle's medium (DMEM) supplemented with 10% Fetal Bovine Serum (FBS), 1% Glutamine, 1% Pen/Strep and 10 mM HEPES pH 7.5.
- SILAC labelling for crosslinking ChIP
- Grow HeLa cells in SILAC DMEM medium, depleted of lysine and arginine, supplemented with 10% dialyzed FBS, 1% Glutamine, 1% Pen/Strep, 10 mM HEPES pH 7.5 and either the light L-lysine (Lys 0) and L-arginine (Arg 0) or their heavy counterparts, L-lysine (Lys 8) and L-arginine (Arg 10) (Materials Table), at the final concentrations of 73 mg/L and 42 mg/L, respectively.
- Grow cells for up to eight generations in SILAC medium to ensure complete isotope-coded amino acids incorporation. Pass cells every two days, when they reach a density of 1.0-1.5 x 106 cells/ml, seeding them at a concentration of 3 x 105 cells/ml.
- Evaluate cells growth and viability in SILAC medium to individuate any possible alteration from physiology that may result from the poorer composition of the SILAC medium; to do this:
- Visually inspect cells morphology at the microscope every day during the labeling.
- Count cells and plot growth curves of cells growing in SILAC versus standard medium.
2. Native Chromatin Immunoprecipitation (N-ChIP)
- Nuclei preparation from cultured cells
- Use 1-2 x 108 unlabeled HeLaS3 cells per experiment. Harvest cells, make aliquot of 50 x 106 cells in 50 ml tubes and centrifuge at 340 x g for 10 min at 4 °C, wash them with ice-cold PBS.
- Resuspend each cell pellet in 8 ml Lysis Buffer (Table 1) and incubate for 10 min at 4 °C on a rotating wheel. Pour carefully each cellular lysate on a sucrose cushion (Table 1) and centrifuge in a swing-out rotor, at 3,270 x g for 20 min at 4 °C, to separate nuclei from cytoplasm.
- Discard supernatants, keep nuclear pellets and wash them twice in ice-cold PBS by centrifugation, discard supernatant at each wash.
- Micrococcal Nuclease (MNase) digestion (small scale) and quality control of chromatin after digestion
- Resuspend the washed nuclear pellet in 1 ml Digestion Buffer (Table 1); divide in two aliquots of 500 μl and keep on ice.
- Measure the Optical Density (OD) at 260 nm of an aliquot, diluted 1:200 in 0.2% sodium dodecyl sulphate (SDS). Note: OD = 1 corresponds to about 50 μg/ml of chromatin DNA. Typically, starting from 2 x 108 cells, the OD is in the range of 40-60.
- Take 1% of nuclei, add MNase enzyme to a final concentration of 0.005 U of enzyme/μl of nuclei and incubate at 37 °C for different time lapses (0, 10, 20, 40, 60 min).
- At each time point, collect 4 μl of digested nuclei and add 1 mM EDTA to stop the MNase reaction. Keep on ice.
- Extract DNA samples by PCR purification kit and elute them in 50 μl TE Buffer (Table 1).
- Take 20 μl of each DNA sample, add 10 μl Loading Buffer (Table 1) and load the samples on 1% (w/v) agarose gel containing Ethidium Bromide, with PCR marker as a size control.
- Check the digestion evaluating the chromatin nucleosome ladder produced by MNase incubation.
- Choose the optimal time for digestion, based on the prevalence of mono-nucleosomes in the preparation. Typically for 0.005 U of enzyme/μl of nuclei the optimal time of MNase digestion is 60 min (Figure 1B, left panel). Note: The optimal MNase concentration/time of digestion may be adjusted depending on the desired cell type and on the size of nucleosomes stretch.
- Large-scale/preparative MNase digestion and recovery of soluble chromatin fractions
- Add to each aliquot (see 2.2.1) 5 μl MNase, corresponding to a final concentration of 0.005 U of enzyme/μl of nuclei; mix gently and incubate at 37 °C for 60 min (or in general for the optimized time interval, based on the small scale test).
- Add 1 mM EDTA to stop the reaction and keep on ice. Pellet the digested nuclei by centrifugation at 7,800 x g at 4 °C for 10 min. Transfer the supernatants (fraction S1) in a new tube and store at 4 °C. Note: S1 contains the first soluble fraction of chromatin, comprising mono-nucleosomes. Add the protease inhibitors (Table 1).
- Carefully resuspend pellets in 1 ml Dialysis Buffer (Table 1) and dialyze overnight at 4 °C (cut off of the dialysis tube is 3.5 kDa), in 3 L Dialysis Buffer, with constant mild stirring. Collect dialyzed material and centrifuge at 7,800 x g for 10 min at 4 °C. Transfer the supernatant (S2 fraction) in a new Eppendorf tube and store at 4 °C. Note: S2 comprises di- to epta-nucleosomes, depending on the MNase digestion extent.
- Quality Control of chromatin before immunoprecipitation
- Take an aliquot corresponding to 5 μg of S1 and S2 chromatin fractions (quantified by measuring the OD at 260 nm in a NanoDrop system). Extract DNA by PCR purification kit and elute in 50 μl TE Buffer (Table 1).
- Take 20 μl of S1 and S2 DNA, mix with 10 μl Loading Buffer (Table 1) and load them on 1% (w/v) agarose gel. Check the quality and efficiency of MNase digestion by visual inspection of the nucleosomes ladder. Note: Fraction S1 is highly enriched in mono-nucleosomes, while fraction S2 contains mainly poly-nucleosomes (Figure 1B, right panel).
- Incubation of chromatin with antibody
- Store at -20 °C 50 μl of S1 fraction for subsequent Mass Spectrometry analysis.
- Add 1 volume of ChIP Dilution Buffer (Table 1) to fraction S1.
- Add the antibody against the hPTM (or protein) of interest; incubate overnight on a rotating wheel at 4 °C. Note: Typically, use 10 μg to 20 μg of antibody for 2 x 108 cells. The optimal ratio between amount of antibody and starting cells number must be carefully set case by case, depending on the abundance of the hPTM/protein used as bait within the sample and on the antibody efficiency. Optimization is experimental, based on the following tests:
- Compare the amount of hPTM/protein of interest between input and flow-through (FT, see 2.7.2) by Western Blot or MS to verify that the immunoprecipitation enriches a significant proportion of specific chromatin region. Note: This is generally achieved when at least 50% of the hPTM/protein bait is depleted in the FT (Figure 1C).
- Check that the immunoprecipitated protein of interest is detectable on SDS-PAGE gel, which ensures that a sufficient amount of material is available for the MS analysis. Note: The presence on the gel of the bands corresponding to the four core histones at the correct stoichiometry indicates that the intact nucleosome has been immunoprecipitated by N-ChroP (Figure 1D). Lack of the proper stoichiometry is in fact indicative of a partial disruption/unfolding of the nucleosome.
- In parallel, prepare the protein G-coupled magnetic beads (see 2.6).
- Equilibration and blocking of protein G-coupled magnetic beads
- Equilibrate 100 μl of protein G-coupled magnetic beads slurry in Blocking Solution (Table 1), washing three times and incubating them overnight at 4 °C on a rotating wheel. Note: Following the binding capacity of beads, use 100 μl of slurry for 2-20 μg of antibody.
- Wash the blocked beads once with Blocking Solution and then twice with ChIP Dilution Buffer (Table 1).
- Isolation of chromatin using magnetic beads
- Add 100 μl of blocked beads to the S1 chromatin sample and incubate them on a rotating wheel at 4 °C for 3 hr. Centrifuge at 340 x g for 1 min to spin down the sample from the lid of the tips. Put in a magnetic rack to pellet the beads.
- Transfer the supernatant to a new Eppendorf tube. This is the flow through (FT), namely the fraction of chromatin that did not bind to antibody.
- Wash the beads four times in Washing Buffer (Table 1) increasing the salt concentration at each wash (75, 125 and 175 mM NaCl).
- To elute the immunoprecipitated chromatin, incubate the beads in 30 μl of LDS Sample Buffer, supplemented with 50 mM dithiothreitol (DTT) for 5 min at 70 °C. Separate the eluted proteins on a 4-12% Bis-Tris acrylamide SDS-PAGE pre-cast gradient gel and stain the gel with Colloidal Coomassie staining kit (Figure 1D).
3. Crosslinking Chromatin Immunoprecipitation (X-ChIP)
- Crosslinking of cells with formaldehyde
- Add 0.75% formaldehyde to SILAC-labeled cells, mix briefly and incubate for 10 min at room temperature. Quench the formaldehyde by adding 125 mM glycine and incubate for 5 min at room temperature.
- Divide cells in 5 x 107 aliquots; rinse them three times with ice-cold PBS by centrifuging at 430 x g for 5 min at 4 °C and discarding the supernatants after each wash. At the last wash, discard supernatant and keep the pellet. Note: Once cells are crosslinked, they can be stored at -80 °C, if not immediately used.
- Nuclei preparation
- Resuspend each cell pellet in 10 ml Lysis Buffer (Table 1). Incubate for 10 min at 4 °C with rotation. Centrifuge at 430 x g for 5 min at 4 °C. Discard the supernatants; keep nuclear pellets.
- Resuspend each nuclear pellet in 10 ml Washing Buffer (Table 1). Incubate at room temperature for 10 min on a rotating wheel.
- Centrifuge at 430 x g for 5 min at 4 °C. Discard the supernatants and collect the pellets.
- Resuspend each nuclear pellets in 3 ml ChIP Incubation Buffer (Table 1). Keep on ice.
- Chromatin sonication and quality control
- Sonicate chromatin at 200 W (cycles of 30 sec “on” and 1 min “off”), in a cooled Bioruptor, to fragment chromatin. Note: the number of cycles and sonication intervals depends on the cell type and on the average length of nucleosomal stretch desired. Typically, 30 min of sonication are necessary to generate DNA fragments of 300-500 bp in lengths, corresponding to di- and tri-nucleosomes. The choice of the fragments size depends on the type of chromatin domain under investigation.
- Take 2% of total input, reverse the crosslinking by incubation at 65 °C for a minimum of 1 hr in ChIP Incubation Buffer. Extract the DNA by PCR purification kit, elute in 50 μl TE Buffer and load the DNA on agarose gel to check chromatin fragmentation.
- Incubation of chromatin with antibody
- Add 1/10 volume of 10% Triton X-100 to sonicated chromatin. Centrifuge at 12,000 x g for 10 min at 4 °C to pellet debris.
- Save 50 μl of chromatin input from heavy cells for testing the incorporation level of SILAC amino acids in labeled cells and for protein profiling.
- Add the antibody of choice to the remaining heavy and light labeled sonicated chromatin and incubate overnight at 4 °C on the rotating wheel. In the light channel, add also an excess fold of the soluble peptide. Incubate overnight at 4 °C on a rotating wheel. Notes: The optimal condition between the μg of antibody and the starting material of cells is chosen case by case, as discussed in 2.5.3. The optimal excess fold molarity of the soluble peptide in respect to the antibody must be carefully titrated case by case.
- Isolation of chromatin using magnetic beads
- Equilibrate and block protein G-coupled magnetic beads by following the steps described in 2.6 and using ChIP Incubation Buffer.
- Add 100 μl of blocked beads to the chromatin samples and incubate on a rotating wheel for 3 hr at 4 °C. Centrifuge at 340 x g for 1 min to spin down the sample from the lid of the tips. Put in a magnetic rack to pellet the beads. The supernatant is the flow through (FT), containing unbound nucleosomes. Wash beads four times in Washing Buffer (Table 1) with increasing salt concentration (two washes at 150 mM and two at 300 mM NaCl).
- Incubate beads in 30 μl SDS-PAGE Loading Sample Buffer (Table 1) and for 25 min at 95 °C to both elute and de-crosslink the immunoprecipitated proteins. Separate proteins on 4-12% Bis-Tris acrylamide SDS-PAGE pre-cast gels (Figure 3D).
4. Sample Preparation Prior to MS
- In-Gel digestion of histones enriched from N-ChIP
Note: During protein digestion and peptide extraction steps take care to minimize keratin contaminations that interfere with LC-MS/MS, as previously described22,23.- Cut gel slices corresponding to the core histones bands (Figure 1D). De-stain the gel pieces with 50% acetonitrile (ACN) in ddH2O, alternating with 100% ACN to shrink the gel. Repeat until gels pieces are completely de-stained and dry them in a vacuum centrifuge.
- Add D6-acetic anhydride 1:9 (v/v) in 1 M ammonium bicarbonate (NH4HCO3) (typically the final volume is 60-100 μl) and 3 μl sodium acetate (CH3COONa), as catalyzer. Incubate for 3 hr at 37 °C with strong shaking. Note: The sample may generate bubbles in the very first minutes after the assembly of the reaction: it important to handle with caution and release the gas generated, opening from time to time the tubes during incubation.
- Rinse the gel pieces several times with NH4HCO3, alternate with ACN at increasing percentage (from 50% to 100%), in order to completely eliminate residues of D6-acetic anhydride.
- Shrink the gel pieces in 100% ACN; dry them in a vacuum centrifuge to ensure complete de-hydration. Re-hydrate gel pieces with ice-cold 100 ng/μl trypsin solution in 50mM NH4HCO3 and incubate overnight at 37 °C. Note: The combination of chemical modification of lysines using deuterated acetic anhydride and trypsin digestion generates an “Arg-C like” in gel digestion pattern of histones21,24.
- Discard the excess solution and add a volume of 50 mM NH4HCO3 to completely cover the gel pieces; incubate overnight at 37 °C.
- Collect soluble digested peptides in a new Eppendorf tube. Lyophilize peptides. Resuspend them in 0.5% acetic acid/0.1% trifluoracetic acid (TFA). Desalt and concentrate peptides on a reversed phase C18/Carbon “sandwich” and ion-exchange chromatography (SCX) on hand-made microcolumns (StageTip) 25.
- Prepare the StageTip microcolumns by putting Teflon meshwork disks containing immobilized C18/Carbon (“sandwich”) and SCX beads in a 200 μl tips. Obtain the “sandwich” by loading the C18 microcolumn on top of a second tip loaded with Carbon filter. Note: The very short peptides, that are not retained on the C18 filter, pass in the flow-through that is loaded directly on the Carbon Tip, which typically captures them. SCX StageTips can enrich specific peptides, such as H3(3-8) peptide, not efficiently retained by reversed phase chromatography.
- Load 50% of peptides onto the C18/Carbon “sandwich StageTip” and 50% onto SCX StageTips. Elute them from the C18/Carbon Tips using 80% ACN/0.5% acetic acid and from the SCX Tips with 5% ammonium hydroxide (NH4OH)/30% methanol. After lyophilization, resuspend peptides in 0.1% FA and analyze by LC-MS/MS.
- In-gel Digestion of immunopurified proteins
In-gel digestion of proteins is carried out as previously described 22, with minor modifications.- Cut each lane in ten slices (Figure 3D) and each slice in small cubes of 1 mm3. De-stain the gel pieces with 50 mM NH4HCO3/50% ethanol and add absolute ethanol to shrink the gels. Repeat until the gels are completely de-stained.
- Add reduction buffer (Table 1) to the gel pieces for 1 hr at 56 °C, followed by the addition of alkylation buffer (Table 1) for 45 min at room temperature, in the dark. Wash and dry the gel pieces in vacuum centrifuge.
- Rehydrate the gel pieces with ice-cold 12.5 ng/μl trypsin solution in 50 mM NH4HCO3 and incubate on ice till complete rehydration of the gel pieces. Remove trypsin in excess.
- Add 50 mM NH4HCO3 to completely cover the gel pieces. Incubate overnight at 37 °C.
- Collect the liquid part. Add the extraction buffer (Table 1) to the gel pieces; incubate with strong agitation for 10 min at room temperature. Repeat twice.
- Incubate the gel pieces in ACN for 10 min with strong agitation. Repeat twice and pool all supernatants.
- Lyophilize the peptides. Resuspend dried peptides in 0.5% acetic acid/0.1% TFA.
- Desalt and concentrate peptides on a reversed phase C18 StageTip, as described26,27.
- Elute peptides from the C18 StageTip using 80% ACN/0.5% acetic acid. Remove the organic solvent by evaporating in a vacuum centrifuge and resuspend the peptides in 0.1% FA (typically 5-10 µl), when ready to the MS analysis.
5. LC-MS Analysis
- Liquid chromatography analysis
- Pack the analytical column in a 15 cm fused silica emitter (75 μm inner diameter, 350 μm outer diameter), using reverse-phase (RP) C18, 3 μm resin in methanol, at a constant helium pressure (50 bar) using a bomb-loader device, as described previously28.
- Couple the packed emitter (C18 RP column) directly to the outlet of the 6-port valve of the HPLC through a 20-cm long (25 μm inner diameter) fused silica without using pre-column or split device.
- Load the digested peptides in C18 RP column at flow of 500 nl/min mobile phase A (0.1% FA/5% ACN in ddH2O).
- After sample loading, separate the peptides using the following gradients:
- Apply 0–40% mobile phase B (0.1% FA/99% ACN in ddH2O) at 250 nl/min over 90 min followed by a gradient of 40-60% in 10 min and 60-80% over 5 min, for elution of peptides deriving from histones (see 2).
- Apply 0–36% mobile phase B at 250 nl/min over 120 min followed by a gradient of 36-60% in 10 min and 60-80% over 5 min, for elution of peptides deriving from immunopurified proteins (see 3).
- Mass Spectrometry analysis using LTQ-FT-ICR-Ultra mass spectrometer
- Operate in a data-dependent acquisition (DDA) mode to automatically switch between MS and MSMS acquisition. MS full scan spectra are acquired, typically in the m/z range from 200-1,650, is acquired with resolution R = 100,000 at 400 m/z. The five (Top5) most intense ions are isolated for fragmentation in the linear ion trap using a collision-induced dissociation (CID) at a target value of 5,000.
- Use the parameters listed in Table 2 for the “Tuning” acquisition file.
- Set the standard acquisition settings as listed in Table 2.
6. Data Analysis
- Label free quantification of histone modifications co-enriched in chromatin domains
- Convert the acquired raw files to mgf files using Raw2msm software (version 1.10)29.
- Search the histone modifications using Mascot Deamon (version 2.2.2), setting the parameters described in Table 3.
- On the Mascot output peptide list, remove peptides with score lower than 15 or with more than 5 putative PTMs29. Note: For each single peptide ID, select the peptide with the highest Mascot score and filter out all the other redundant peptides with same ID.
- Construct the extracted ion chromatograms (XIC) for each precursor corresponding to every modified peptides, based on the m/z value, using the QualBrowser. Calculate the area under the curve (AUC) for each peak (Figure 2A).
- Validate each identified peptides containing modifications by visual inspection of the MS/MS spectra using the QualBrowser (Figure 2B).
- Calculate the relative abundance for each modified peptide. Calculate the relative enrichment of each modification in the ChIP-ed material. Note: Relative abundance is calculated as a ratio between the AUC of each specific modified peptide over the sum of AUC of all the modified and unmodified forms of the same peptide, while relative enrichment as a ratio of the relative abundance of the specific modification in the ChIP over the Input (Figure 2C).
- Quantitative proteomic analysis of proteins co-associated within chromatin domains
- For proteins identification and quantification use the MaxQuant package (http: //www. maxquant.org/)30. Configure the built-in search engine “Andromeda” using the AndromedaConfig.exe31 and set the search parameters listed in Table 3.
- Incorporation test
- Estimate the degree of incorporation of heavy amino acids into proteins in heavy-labeled chromatin input, using MaxQuant software, as follows:
- Set the parameters as described in 6.2 but disabling the re-quantify option.
- Calculate the incorporation percentage applying the following formula to non-redundant peptide ratios: Incorporation (%) = ratio (H/L) / ratio (H/L) + 1 × 100 (Figure 3B). Note: Accept only if incorporation >95%.
- Estimate the degree of incorporation of heavy amino acids into proteins in heavy-labeled chromatin input, using MaxQuant software, as follows:
Representative Results
Chromatin immunoprecipitation is a powerful technique used to profile the localization of a protein or a histone modification along the genome. In a proteomics equivalent, ChIP is followed by MS-based proteomics to identify qualitatively and quantitatively the hPTMs, histone variants and chromatin-binding proteins that are immunoprecipitated together with the modification or protein of interest, used as “bait”. In the N-ChroP approach, outlined in Figure 1A, native ChIP, in which chromatin is digested with MNase (Figure 1B), is used as input to purify from bulk chromatin a distinct functional domain. Digested chromatin enriched in mono-nucleosomes is incubated with a specific antibody and the immunopurified proteins are separated by SDS-PAGE. In the example illustrated, H3K9me3, marker of silent chromatin32,33, is used to set up the approach. The choice is based on the fact that both its functional role as well as some of its protein interactors are well described. Moreover, a highly specific and efficient antibody optimized for ChIP is available34, as also confirmed by visual inspection of the Coomassie gel, where the intact nucleosome, with the core histones at the correct stoichiometry, is enriched in appropriate amounts for MS (Figure 1D).
The comparison between the amount of H3K9me3 present in the flow-through (FT) and input (IN) indicates that about 50% of the region of interest is immunopurified, excluding the risk of a bias due the enrichment of a minor sub-population of chromatin (Figure 1C). MS is employed to characterize the PTMs co-associated within the enriched nucleosomes: each core histone is digested using a protocol designed ad hoc in order to achieve an “Arg-C like” digestion, in polyacrylamide gel. In fact, on the one hand Arg-C is the best protease for MS analysis of hPTMs because it produces peptides of optimal length; on the other hand, it does not cleave efficiently in gel. To overcome this limitation, our tailored protocol exploits the chemical alkylation of lysine achieved by incubating histone gel bands with deuterated (D6)-acetic anhydride, followed by trypsin digestion. Since trypsin does not cleave D3-acetylated lysines, the resulting peptides mixture mimics an “Arg-C like” pattern (Figure 1E).
The addition of a D3-acetyl moiety to a lysine produces an unambiguous delta mass of 45.0294 Daltons, discriminating between native and chemically added acetylations in MS. Moreover, D3-acetylation offers two additional advantages that facilitate the discernment of isobaric modified peptides: first, the alkylation occurs only on unmodified and mono-methylated lysines, but not on di- and tri-methylated residues; as such modified peptides bearing the same total number of modifications but in different arrangements are differentially decorated by distinct set of D3-acetyl groups that produce unambiguous m/z shifts. Second, distinct patterns of D3-alkylation cause slightly different retention times in liquid chromatography on reverse phase column, which generate an additional level of separation for isobaric modified peptides21.
After validation of all hPTMs by manual inspection of the corresponding MS/MS spectra (Figure 2B), the label free quantification is achieved in two steps: first, we calculate the relative abundance of each modification using the signal intensity of the unmodified and modified species for the corresponding peptide, measured through the calculation of the extracted ion chromatograms (XIC) (Figure 2A and 2C, upper panel); second, the relative enrichment is estimated as a ratio between the relative abundance of each modification in the ChIP-ed octamer and the corresponding relative abundance from input (Figure 2C, lower panel). The analysis of the H3(9-17) peptide shows the enrichment of di- and tri-methylated K9, with the corresponding depletion of the unmodified and mono-methylated forms (Figure 2C). With this results as positive control for antibody specificity, the co-association or depletion of all other modifications can be assessed, both at the intra-molecular level on the same H3 and at the inter-molecular level, on other co-enriched histones within the same nucleosome. This allows the screening of hPTMs cross-talks within the H3K9me3 domains, the so-called “heterochromatin modificome” (Figure 2D): the significant enrichment of known markers associated with gene silencing, with the corresponding depletion of markers linked with gene activation is observed. In addition, novel associations are detected such as the enrichment of H3K18me1.
To screen for all proteins co-associating within heterochromatin, the classical crosslinking ChIP is combined with SILAC (Stable Isotope Labelling by Amino acids in Cell culture) (Figure 3A). In the SILAC experiment, the lysine and arginine (light form) are replaced with their isotopically labeled analogs (heavy form) in the culturing medium. Upon cells grown and replication in both light and heavy media, the two differentially isotope-encoded amino acids are metabolically incorporated into proteins, generating light and heavy forms of proteins, respectively, that are distinguishable by MS, due to a specific Δmass. Before starting a large-scale SILAC experiment, the labeling efficiency is evaluated, calculating the incorporation level, measured as the percentage fraction of heavy peptides versus the sum of heavy and light ones, found in the only heavy labeled sample. Incorporation superior to 95% for both heavy arginine and heavy lysine is required for accurate protein quantification (Figure 3B). After crosslinking of heavy and light cells, chromatin is fragmented by sonication. SILAC is used in conjunction with a competition assay using an excess fold of soluble H3 peptide (QTARKSTGG) that bears tri-methylation on K9 in order to discriminate specific H3K9me3 interactors from background. The soluble peptide is added in excess to one of the two SILAC ChIP experiments, where it saturates the binding capacity of the antibody, thus “competing out” the majority of the H3K9me3- nucleosomes and, accordingly, all specific interactors. In the other SILAC channel, the competing peptide is not added to the ChIP and the immunoprecipitation takes place normally. SILAC-competition experiments are typically performed in duplicate in so-called “Forward” and “Reverse” formats, where the competition with the excess soluble peptide is switched from the heavy (H) to the light (L) chromatin samples. This results in inverted complementary SILAC ratio readouts, used for discerning genuine from unspecific binders: proteins specifically enriched are present with a higher intensity in the heavy form in comparison with the light form (protein ratio H/L>1) in the experiment where excess peptide is added to the light channel (Forward) (Figure 4A, upper panel); an opposite trend (protein ratio H/L<1) is observed in the Reverse replica (Figure 4A, lower panel). Proteins whose intensity in the heavy and light form are similar in both Forward and Reverse experiments produce a constant ratio close to 1 and are classified as background (Figure 4B).
The choice of the optimal excess fold for the soluble peptide is important and must be tuned accurately. Typically, we set the correct antibody-to-peptide proportion performing a competition assay using serial dilutions of excess peptide and comparing the level of the “bait” (hPTM/protein) between the positive control ChIP, where the peptide is not added, and the different serial competition ChIPs, by Western Blot or MS. Generally, the optimal excess fold peptide reduces of about 90% the amount of hPTM/protein in the immunoprecipitated material. In fact, on the one hand, the larger the protein ratio obtained, the higher the discriminating power of SILAC; on the other hand, an excessive saturation by the peptide may evict completely the specific binders from the antibody, thus leading to missing H/L ratios for quantification and statistical analysis. For H3K9me3 antibody we defined the correct proportion by measuring the H/L ratio for the QTARK(me3)STGG peptide, in a competition test carried out in a Forward set up and we took this value as a measure of the competition efficiency (Figure 3C).
The intersection of the Forward and Reverse X-ChIP experiments leads to the identification of 635 proteins, present in both experiments and quantified with at least 2 ratio counts. The log2 plot of their H/L ratios represent the so-called “heterochromatome” (Figure 4C), where the genuine H3K9me3 interactors are unambiguously identified as the proteins present within the top 40% of the proteins ratio distributions (upper right quadrant of the scatter plot and Figure 4D).
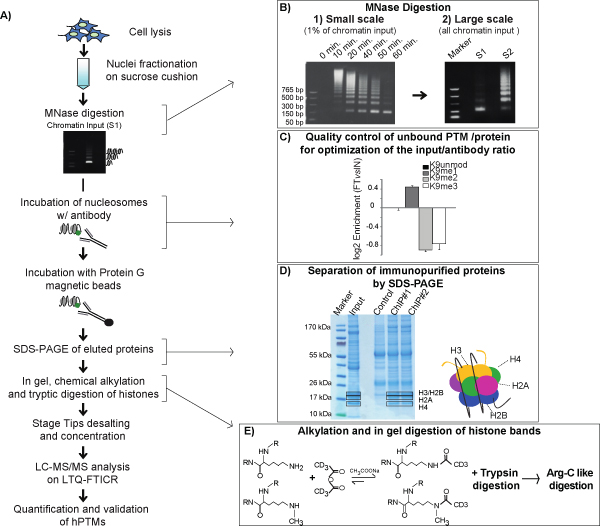
Figure 1: Schematic view of the N-ChroP workflow. A) Scheme of native ChIP combined with MS analysis. Chromatin from cells is digested with MNase and the fraction enriched in mono-nucleosomes (S1) is immunoprecipitated using an anti-H3K9me3 antibody. Immunopurified proteins are separated by SDS-PAGE and core histones are in gel-digested with an ad hoc protocol to mimic an Arg-C digestion. Peptides are analyzed by nano-LC-MS/MS. Histone PTMs are identified, validated by manual inspection of MS/MS spectra and quantified. B) Small scale MNase test: DNA resolved on a 1% agarose gel after chromatin digestion with MNase at different time lapse (left panel); large scale MNase digestion: DNA resolved on a 1% agarose gel after 60 minutes of MNase chromatin digestion and after separation of S1 fraction, containing mono-nucleosomes from S2 faction, containing poly-nucleosomes (right panel). C) Estimation of enrichment/depletion of unmodified, mono-, di-, and tri-methylated K9 in flow-through (FT) compared to input (IN). Histogram represents the average ±SEM from three independent experiments for each modification. D) SDS-PAGE of chromatin input and co-immunopurified proteins: core histones H3, H4, H2A and H2B are visible around and below the 17kDa band, with H3 and H2B co-migrating (black squares). E) Each core histone from both immunoprecipitated nucleosomes and input are chemically alkylated using deuterated (D6)-acetic-anhydride before trypsin treatment, in order to obtain an “Arg-C like” in gel digestion. The D6-acetic anhydride reacts with the Epsilon amino group of unmodified and mono-methylated lysines but not with di-methylated, tri-methylated and acetylated lysines. As result, the enzymatic activity of trypsin is blocked on all native and chemical acetylated lysine, thus producing a “Arg-C like” digestion pattern. This figure has been modified from 21, using Figure 1, Figure S1 and Figure S5 as reference. Click here to view larger image.
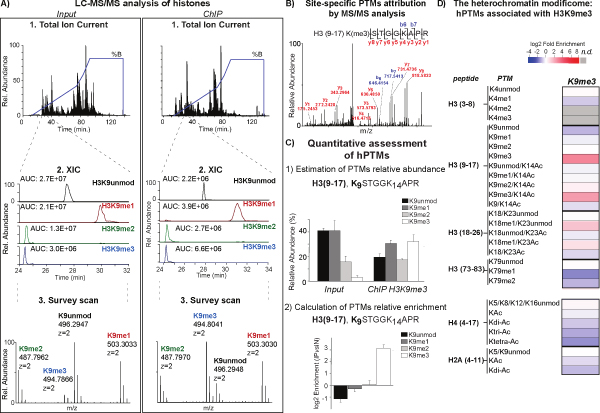
Figure 2: Mass Spectrometry analysis of the H3K9me3 “modificome”. A) Zoomed mass spectra and extracted ion chromatograms (XIC) constructed at the corresponding m/z value of the 2+ charge unmodified, mono-, di-, and tri-methylated K9 in the H3(9-17) peptide, both for input and ChIP samples. B) Representative MS/MS spectra using CID fragmentation. The b-ion and y-ion series allow to define the sequence of H3(9-17) peptide and to localize specifically the tri-methylation on K9 residue. C) Relative abundance of the different degrees of methylation on K9 in the H3(9-17) peptide, estimated by dividing the area under the curve (AUC) of each modified peptide by the sum of the areas corresponding to all observed unmodified and modified forms of that peptide, in the input and ChIP-ed octamer. Histogram represents the average ±SEM from three independent experiments for each modification (upper panel). Relative enrichment of K9 methylations in H3(9-17) peptide. The enrichment is expressed as a log2 ratio between the relative abundance of each methylation in the ChIP-ed octamer as compare to input. Histogram represents the averages ±SEM from three independent experiments (lower panel). D) Heatmap summarizing the enrichment of all co-associating hPTMs identified on histone H3, H4 and H2A. Each row corresponds to a different modification (n.d. are not detected modifications). This figure has been modified from 21, using Figure 1, Figure 2, Figure 3 and Figure S10 as reference. Click here to view larger image.
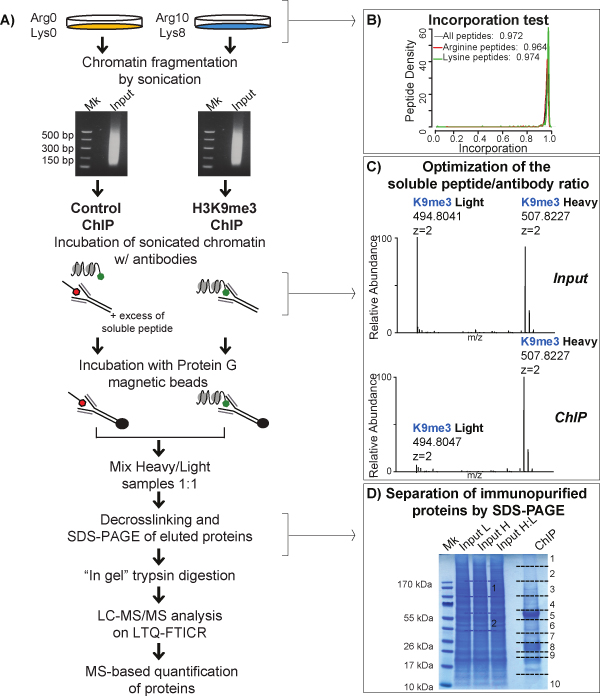
Figure 3: Schematic view of X-ChroP workflow. A) Scheme of crosslinking ChIP combined with MS analysis. Cells grown in light and heavy media are fixed with formaldehyde and chromatin inputs are fragmented by sonication to generate DNA fragments. In a Forward set up, the sonicated heavy-labeled chromatin is immunoprecipitated using anti-H3K9me3 antibody while the light-labeled chromatin is incubated with the same antibody saturated with an excess fold of soluble H3 peptide bearing K9me3. Immunoprecipitated proteins from heavy and light chromatins are pooled, extracted and separated by SDS-PAGE. Proteins are digested with trypsin and peptides are analyzed by nano-LC-MS/MS. B) Efficiency of SILAC labeling is monitored calculating the incorporation of heavy lysine (Lys8) and arginine (Arg10) into proteins. In the plot, the median of lysine and arginine peptides density distribution is equal to 0.974 (green line) and 0.964 (red line), respectively. C) Zoomed mass spectra at the corresponding m/z value of the 2+ charge tri-methylated K9 in the H3(9-17) peptide, both for light and heavy forms, in input and ChIP-ed octamer. While in input the intensities of heavy and light peptide are close to 1, in the Forward SILAC ChIP, the intensity of light peptide is much lower than the one of the heavy counterpart, thus indicating an efficient competition. D) SDS-PAGE of light (L) and heavy (H) labeled chromatin input and of co-immunoprecipitated material: the lane corresponding to the ChIP-ed material is cut in ten slices (black line), while only two slices of input are analyzed for incorporation test analysis (blue line). This figure has been modified from 21, using Figure 4 and Figure S6 as reference. Click here to view larger image.
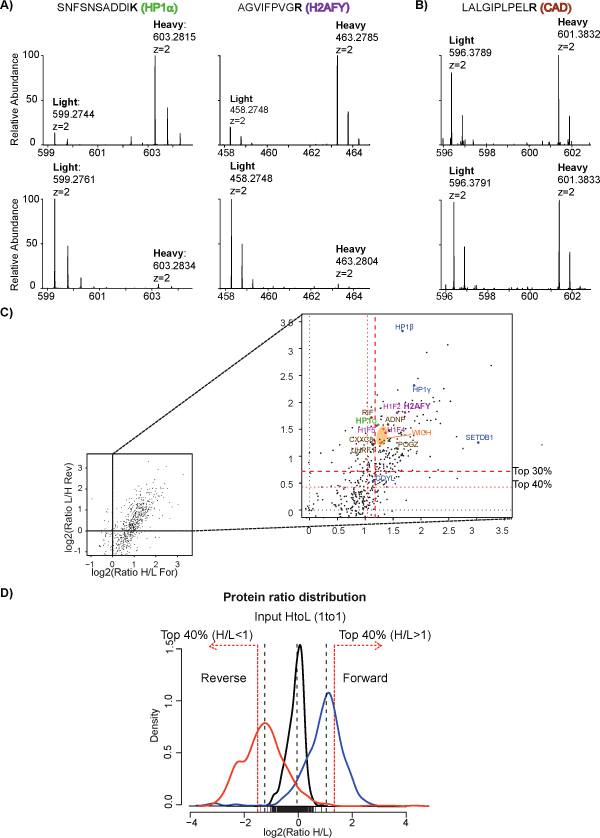
Figure 4: Mass Spectrometry analysis of the H3K9me3 “interactome”. A) Representative full spectra showing the SILAC-pair corresponding to peptides from HP1 and macro-2A proteins: the ratios H/L>1 in the Forward experiment (upper panels), mirrored by a ratios H/L<1 in the Reverse replica (lower panels), demonstrate the specific enrichment of these proteins in heterochromatin. B) Representative full spectra with SILAC-pair corresponding to peptide from one background protein: H/L ratios in Forward and Reverse replicates are equal to 1. C) Quantified proteins are distributed in a scatter plot based on their SILAC-ratios in Forward and Reverse experiments (x and y axes, respectively); red dotted lines represent the top 40% and 30% of protein ratios, as indicated. D) Protein ratios distributions from input (H/L mixed 1:1) (black), Forward (blue) and Reverse (orange) X-ChroP experiments; red dotted lines represent the top 40% of protein ratios, set as cutoffs to select the genuine interactors. This figure has been modified from21, using Figure 5 as reference. Click here to view larger image.
| Buffer for section 2 | Composition |
| Lysis Buffer | 10% sucrose, 0.5 mM EGTA pH 8.0, 15 mM NaCl, 60 mM KCl, 15 mM HEPES, 0.5% Triton, 0.5 mM PMSF, 1mM DTT, 5 mM NAF, 5 mM Na3VO4, 5mM NaButyrate, 5 mg/ml Aprotinin, 5 mg/ml Pepstatin A, 5 mg/ml Leupeptin |
| Sucrose cushion | 2 g sucrose in 20 ml of Lysis Buffer |
| Digestion Buffer | 0.32 M sucrose, 50 mM Tris-HCl pH 7.6, 4 mM MgCl2, 1 mM CaCl2, 0.1 mM PMSF |
| TE Buffer | 10 mM Tris-HCl pH 7.5, 1 mM EDTA |
| Dialysis Buffer | 10 mM Tris-HCl pH 7.6, 1 mM EDTA, 0.5 mM PMSF, 5 mM NAF, 5 mM Na3VO4, 5mM NaButyrate, protease inhibitors cocktail |
| ChIP Dilution Buffer | 100 mM Tris HCl pH 7.6, 100 mM NaCl and 10 mM EDTA |
| Blocking Solution | BSA 0.5% in PBS |
| Washing Buffer | 50 mM Tris-HCl pH 7.6,10 mM EDTA |
| Protease Inhibitors | EDTA-free protease inhibitors cocktail, 0.5 mM PMSF, 5 mM NAF, 5 mM Na3VO4, 5mM NaButyrate |
| Loading Buffer | orange loading dye, 50% (v/v) glycerol in H20 |
| Buffer for section 3 | Composition |
| Lysis Buffer | 50 mM HEPES-KOH pH 7.5, 140 mM NaCl, 1 mM EDTA, 10% glycerol, 0.5% NP-40, 0.25% Triton-100, 0.5 mM PMSF, 5 mM NAF, 5 mM Na3VO4, 5mM NaButyrate, 5 mg/ml Aprotinin, 5 mg/ml Pepstatin A, 5 mg/ml Leupeptin |
| Washing Buffer | 10 mM Tris-HCl pH 8, 200 mM NaCl, 1 mM EDTA, 0.5 mM EGTA, 0.5 mM PMSF, 5 mM NAF, 5 mM Na3VO4, 5mM NaButyrate, 5 mg/ml Aprotinin, 5 mg/ml Pepstatin A, 5 mg/ml Leupeptin |
| ChIP Incubation Buffer | 10 mM Tris-HCl pH 8, 100 mM NaCl, 1 mM EDTA, 0.5 mM EGTA, 0.1% sodium deoxycholate, 0.5% sodium lauroylsarcoside, 0.5 mM PMSF, 5 mM NAF, 5 mM Na3VO4, 5 mM NaButyrate, 5 mg/ml Aprotinin, 5 mg/ml Pepstatin A, 5 mg/ml Leupeptin |
| Washing Buffer 2 | 20 mM Tris-HCl pH 7.6, 2 mM EDTA, 0.1 % SDS,1 % Triton-100 |
| Loading Sample Buffer | 250 mM Tri-HCl pH 8.8, 0.5 M b-mercaptoethanol, 2% SDS |
| Buffer for section 4 | Composition |
| Alkylation Buffer | 55 mM iodoacetamide in 50 mM NH4HCO3 |
| Reduction Buffer | 10 mM dithiothreitol in 50 mM NH4HCO3 |
| Extraction Buffer | 30% ACN and 3% TFA in ddH20 |
Table 1. Buffers Composition.
| Tuning acquisition file | Parameters |
| FT full scan | accumulation target value 1 x 106; maximum filling time 1,000 msec |
| IT MSn | accumulation target value 10 x 104; maximum filling time 150 msec |
| Acquisition setting | Parameters |
| Electrospray voltage | 2.4 kV |
| Sheath and auxiliary gas flow | No |
| Ion transfer capillary temperature | 200 °C |
| Dynamic exclusion | up to 500 precursor ions for 60 sec upon MSMS |
| Exclusion mass width | 10 ppm |
| Normalized collision energy using wide-band activation mode | 35% |
| Ion selection thresholds | 100 counts |
| Activation q | 0.25 |
| Activation time | 30 msec |
Table 2. MS settings.
| Mascor Deamon search | Parameters | Notes |
| Database | depends on the organism (i.e. for HeLa cells: human database, version 3.68; 87,061 entries) | |
| Enzyme | Arg-C | Arg-C cleave at the C-terminal of all arginine residues |
| Variable modifications | acetyl (K), oxidation (M), D3-acetylation (K), methyl-D3-acetyl (K), dimethyl (k), trimethyl (K) | acetyl [42.010 Da], oxidation [15.995 Da], D3-acetylation [+45.0294 Da], methyl-D3-acetyl [sum of +14.016 Da and +45.0294 Da], dimethyl [28.031], trimethyl [42.046 Da] |
| Missed cleavages | up to 2 | |
| Mass accuracy of the parent ions in the search | 10 ppm | |
| Mass-accuracy for CID MSMS | 0.5 Da | |
| MaxQuant search | Parameters | Notes |
| Database | depends on the organism | |
| Enzyme | trypsin/P | Trypsin cleaves at the C-terminal of all lysine and arginine residues. The search is performed taking in account the fact that the efficiency of trypsin to cleave lysine and arginine is reduced when the next amino acid is the proline (/P). |
| Fixed modification | carbamidomethylation | |
| Variable modifications | N-acetyl (Protein), oxidation (M) | |
| Missed cleavages | up to 3 | |
| Label parameters | lys8 and arg10 | |
| Maximum label amminoacid | 3 for Trypsin | |
| Mass-accuracy of the parent ions in the initial “Andromeda” search | 20 ppm | |
| Mass accuracy of the parent ions in the main “Andromeda” search | 6 ppm | |
| Mass-accuracy for CID MSMS | 0.5 Da (six top peaks per100 Da) | |
| Peptide false discovery rates (FDR) | 0.01 | |
| Protein false discovery rates (FDR) | 0.01 | Setting the FDR for peptide and protein to 0.01 means that both peptides and proteins identified are expected to contain 1% of false positives. This value is estimated using a target-decoy database |
| Maximum posterior error probability (PEP) | 1 | PEP is the probability that an individual peptide is a false positive match. PEP equal to 1 in your setting means that all peptides will be taken irrespective of the PEP thus filtering is based exclusively on FDR. |
| Minimum peptide length | 6 | |
| Minimum number of peptides | 2 | |
| Minimum number of unique peptides | 1 | |
| Using only unmodified peptides and oxidation (M)/acetyl (Protein N-Term) | activate the option | Peptides with modifications should generally not be counted for protein quantification since their abundance may not reflect the ratio of the corresponding protein. |
| Minimum ratio count | 1 | |
| “Match between runs” | activate the option | This option matches precursor masses in a 2-min retention time window (after realignment of the runs) based on the accurate mass measurement and allows transferring the MS/MS identification between the different LC MS/MS runs. |
Table 3. Data Analysis.
Discussion
We have recently described ChroP, a quantitative strategy for the large-scale characterization of the protein components of chromatin. ChroP combines two complementary approaches used in the epigenetic field, ChIP and MS, profiting from their strengths and overcoming their respective limitations. ChIP coupled to deep sequencing (ChIP-Seq) allows the genome-wide mapping of histone modifications at the resolution of few nucleosomes35. Although advantageous for their sensitivity, antibody-based assays are limited in their capability to distinguish similar modifications and in dissecting the combinatorial aspect of the histone code36. On the other hand, while MS provides a comprehensive and unbiased analysis of hPTMs, singly and in combinations9, it has so far been applied to the analysis of bulk chromatin, with consequent lack of information on locus-specific PTMs patters. We employ a modified version of ChIP to isolate functionally distinct chromatin domains and mass spectrometry to characterize the histone PTM patterns and the non-histonic proteins specifically co-enriched.
By using N-ChroP, the analysis of hPTMs co-associated with the H3K9me3 revealed a significant enrichment of markers associated with silent chromatin, and a depletion of modifications associated with active chromatin. The agreement of these results with previous studies37 proved the robustness of the strategy. In X-ChroP of H3K9me3 domains, the SILAC-based investigation of the chromatin-interacting proteins confirmed some previously described interactors, thereby validating the method.
ChroP presents two main original aspects in respect to the strategies already available to investigate the proteomic component of chromatin: for instance, the possibility to reveal inter-molecular synergisms between modifications decorating distinct core histones within the same intact mono-nucleosome purified by N-ChIP; second the chance to assess the specific compartmentalization of histone variants and linker histone subtypes. Since the investigation of histone variants is held back by the lack of good quality reagents (i.e. antibodies), ChroP emerges as the unique tool available to assess their location and functional role.
One limitation of ChroP in its current set up lies in peptide-centric (“Bottom-up”) approach used in MS, with histones digested in short peptides and the consequent impairment in detecting the long-distance connectivity between modifications. As such, the conjugation of ChroP with “Bottom-up” MS analysis permits a partial assessment of the combinatorial aspect of the histone code. We envisage that the implementation of alternative MS strategies, such as “Middle- and Top-Down” for hPTMs mapping on longer peptides (>20 aa) up to on intact proteins38-40, will overcome this restraint.
Overall, N- and X-ChroP are highly complementary, with the possibility of dissecting the complexity of chromatin structure at functionally distinct domains, with a resolution of mono- to oligo-nucleosomes. We predict that ChroP will be useful also to characterize the composition of chromatin regions marked by the presence of specific non-histonic nuclear proteins, for example transcription factors (TFs). In addition, ChroP can be employed in functional studies to map the dynamic composition of chromatin at specific loci upon various perturbations, for instance during global transcriptional activation. For these reasons, ChroP emerges as an additional useful tool in the arsenal of analytical strategies available to dissect the proteomic landscape of chromatin.
Declarações
The authors have nothing to disclose.
Acknowledgements
This research was originally published in Mol Cell Proteomics. Soldi M. and Bonaldi T. The Proteomic Investigation of Chromatin Functional Domains Reveals Novel Synergisms among Distinct Heterochromatin Components MCP. 2013; 12: 64-80. © the American Society for Biochemistry and Molecular Biology. We thank Roberta Noberini (Italian Institute of Technology and IEO, Italy) for critical reading of the manuscript. TB work is supported by grants from the Giovanni Armenise-Harvard Foundation Career Development Program, the Italian Association for Cancer Research and the Italian Ministry of Health. MS work was supported by a FIRC fellowship.
Materials
| DMEM | Lonza | BE12-614F | |
| FBS | Invitrogen | 10270-106 | |
| SILAC DMEM | M-Medical | FA30E15086 | |
| Dialyzed FBS | Invitrogen | 26400-044 | |
| Lysine 0 (12C6 14N2 L-lysine) | Sigma Aldrich | L8662 | |
| Arginine 0 (12C6 14N4 L-arginine) | Sigma Aldrich | A6969 | |
| Lysine 8 (13C6 15N2 L-lysine) | Sigma Aldrich | 68041 | |
| Arginine 10 (13C6 15N4 L-arginine) | Sigma Aldrich | 608033 | |
| Micrococcal Nuclease | Roche | 10 107 921 001 | |
| Complete, EDTA-free Protease Inhibitor Cocktail Tablets | Roche | 04 693 132 001 | |
| Spectra/Por 3 dialysis tubing, 3.5K MWCO, 18mm flat width, 50 foot length | Spectrumlabs | 132720 | |
| QIAquick PCR purification kit | QIAGEN | 28104 | |
| Anti-Histone H3 tri-methylated K9-ChIP grade | Abcam | ab8898 | |
| Histone H3 peptide tri-methyl K9 | Abcam | ab1773 | |
| Dynabeads Protein G | Invitrogen | 100.04D | |
| NuPAGE Novex 4-12% Bis-Tris Gel | Invitrogen | NP0335BOX | |
| Colloidal Blue Staining Kit | Invitrogen | LC6025 | |
| LDS Sample Buffer | Invitrogen | NP0007 | |
| Formaldheyde | Sigma Aldrich | F8775 | |
| Aceti anhydride-d6 | Sigma Aldrich | 175641-1G | |
| Formic Acid | Sigma Aldrich | 94318-50ML-F | |
| Iodoacetamide ≥99% (HPLC), crystalline | Sigma Aldrich | I6125 | |
| DL-Dithiothreitol | Sigma Aldrich | 43815 | |
| Sequencing Grade Modified Trypsin, Frozen 100ug (5 × 20μg) | Promega | V5113 | |
| Nanospray OD 360μm x ID 75μm, tips ID 8μm uncoated Pk 5 | Microcolumn Srl | FS360-75-8-N-5-C15 | |
| ReproSil-Pur 120 C18-AQ, 3 µm 15 % C | Dr. Maisch GmbH | r13.aq. | |
| C18 extraction disk, 47 mm | Agilent Technologies | 12145004 | |
| Carbon extraction disk, 47 mm | Agilent Technologies | 12145040 | |
| Cation extraction disk | Agilent Technologies | 66889-U |
Referências
- Kornberg, R. D. Chromatin structure: a repeating unit of histones and DNA. Science. 184, 868-871 (1974).
- Luger, K., Mader, A. W., Richmond, R. K., Sargent, D. F., Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature. 389, 251-260 (1997).
- Kouzarides, T. Chromatin modifications and their function. Cell. 128, 693-705 (2007).
- Bannister, A. J., Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 21, 381-395 (2011).
- Taverna, S. D., Li, H., Ruthenburg, A. J., Allis, C. D., Patel, D. J. How chromatin-binding modules interpret histone modifications: lessons from professional pocket pickers. Nat Struct Mol Biol. 14, 1025-1040 (2007).
- Jenuwein, T., Allis, C. D. Translating the histone code. Science. 293, 1074-1080 (2001).
- Spotswood, H. T., Turner, B. M. An increasingly complex code. J Clin Invest. 110, 577-582 (2002).
- Soldi, M., Cuomo, A., Bremang, M., Bonaldi, T. Mass spectrometry-based proteomics for the analysis of chromatin structure and dynamics. Int J Mol Sci. 14, 5402-5431 (2013).
- Sidoli, S., Cheng, L., Jensen, O. N. Proteomics in chromatin biology and epigenetics: Elucidation of post-translational modifications of histone proteins by mass spectrometry. J Proteomics. 75, 3419-3433 (2012).
- Britton, L. M., Gonzales-Cope, M., Zee, B. M., Garcia, B. A. Breaking the histone code with quantitative mass spectrometry. Expert Rev Proteomics. 8, 631-643 (2011).
- Taverna, S. D., et al. Long-distance combinatorial linkage between methylation and acetylation on histone H3 N termini. Proc Natl Acad Sci U S A. 104, 2086-2091 (2007).
- Garcia, B. A., Shabanowitz, J., Hunt, D. F. Characterization of histones and their post-translational modifications by mass spectrometry. Curr Opin Chem Biol. 11, 66-73 (2007).
- Villar-Garea, A., Imhof, A. The analysis of histone modifications. Biochim Biophys Acta. 1764, 1932-1939 (2006).
- Plazas-Mayorca, M. D., et al. One-pot shotgun quantitative mass spectrometry characterization of histones. J Proteome Res. 8, 5367-5374 (2009).
- Ohta, S., et al. The protein composition of mitotic chromosomes determined using multiclassifier combinatorial proteomics. Cell. 142, 810-821 (2010).
- Vermeulen, M., et al. Selective anchoring of TFIID to nucleosomes by trimethylation of histone H3 lysine 4. Cell. 131, 58-69 (2007).
- Vermeulen, M., et al. Quantitative interaction proteomics and genome-wide profiling of epigenetic histone marks and their readers. Cell. 142, 967-980 (2010).
- Bartke, T., et al. Nucleosome-interacting proteins regulated by DNA and histone methylation. Cell. 143, 470-484 (2010).
- Dejardin, J., Kingston, R. E. Purification of proteins associated with specific genomic Loci. Cell. 136, 175-186 (2009).
- Lambert, J. P., Mitchell, L., Rudner, A., Baetz, K., Figeys, D. A novel proteomics approach for the discovery of chromatin-associated protein networks. Mol Cell Proteomics. 8, 870-882 (2009).
- Soldi, M., Bonaldi, T. The proteomic investigation of chromatin functional domains reveals novel synergisms among distinct heterochromatin components. Mol Cell Proteomics. 12, 764-780 (2013).
- Shevchenko, A., Tomas, H., Havlis, J., Olsen, J. V., Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat Protoc. 1, 2856-2860 (2006).
- Shevchenko, A., Wilm, M., Vorm, O., Mann, M. Mass spectrometric sequencing of proteins silver-stained polyacrylamide gels. Anal Chem. 68, 850-858 (1996).
- Bonaldi, T., Regula, J. T., Imhof, A. The use of mass spectrometry for the analysis of histone modifications. Methods Enzymol. 377, 111-130 (2004).
- Cuomo, A., Moretti, S., Minucci, S., Bonaldi, T. SILAC-based proteomic analysis to dissect the "histone modification signature" of human breast cancer cells. Amino Acids. 41, 387-399 (2011).
- Rappsilber, J., Mann, M., Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat Protoc. 2, 1896-1906 (2007).
- Rappsilber, J., Ishihama, Y., Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal Chem. 75, 663-670 (2003).
- Olsen, J. V., Ong, S. E., Mann, M. Trypsin cleaves exclusively C-terminal to arginine and lysine residues. Mol Cell Proteomics. 3, 608-614 (2004).
- Jung, H. R., Pasini, D., Helin, K., Jensen, O. N. Quantitative mass spectrometry of histones H3.2 and H3.3 in Suz12-deficient mouse embryonic stem cells reveals distinct, dynamic post-translational modifications at Lys-27 and Lys-36. Mol Cell Proteomics. 9, 838-850 (2010).
- Cox, J., Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol. 26, 1367-1372 (2008).
- Cox, J., et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J Proteome Res. 10, 1794-1805 (2011).
- Lachner, M., O’Carroll, D., Rea, S., Mechtler, K., Jenuwein, T. Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature. 410, 116-120 (2001).
- Bannister, A. J., et al. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature. 410, 120-124 (2001).
- Ram, O., et al. Combinatorial patterning of chromatin regulators uncovered by genome-wide location analysis in human cells. Cell. 147, 1628-1639 (2011).
- Barski, A., et al. High-resolution profiling of histone methylations in the human genome. Cell. 129, 823-837 (2007).
- Beck, H. C. Mass spectrometry in epigenetic research. Methods Mol Biol. 593, 263-282 (2010).
- Ernst, J., Kellis, M. Discovery and characterization of chromatin states for systematic annotation of the human genome. Nat Biotechnol. 28, 817-825 (2010).
- Tipton, J. D., et al. Analysis of intact protein isoforms by mass spectrometry. J Biol Chem. 286, 25451-25458 (2011).
- Young, N. L., et al. High throughput characterization of combinatorial histone codes. Mol Cell Proteomics. 8, 2266-2284 (2009).
- Boyne, M. T., Pesavento, J. J., Mizzen, C. A., Kelleher, N. L. Precise characterization of human histones in the H2A gene family by top down mass spectrometry. J Proteome Res. 5, 248-253 (2006).

