Derivation and Characterization of a Transgene-free Human Induced Pluripotent Stem Cell Line and Conversion into Defined Clinical-grade Conditions
Summary
We describe a protocol for deriving lentiviral-based reprogrammed and characterized factor-free human induced pluripotent stem cells and conversion into putative clinical-grade conditions.
Abstract
Human induced pluripotent stem cells (hiPSCs) can be generated with lentiviral-based reprogramming methodologies. However, traces of potentially oncogenic genes remaining in actively transcribed regions of the genome, limit their potential for use in human therapeutic applications1. Additionally, non-human antigens derived from stem cell reprogramming or differentiation into therapeutically relevant derivatives preclude these hiPSCs from being used in a human clinical context2. In this video, we present a procedure for reprogramming and analyzing factor-free hiPSCs free of exogenous transgenes. These hiPSCs then can be analyzed for gene expression abnormalities in the specific intron containing the lentivirus. This analysis may be conducted using sensitive quantitative polymerase chain reaction (PCR), which has an advantage over less sensitive techniques previously used to detect gene expression differences3. Full conversion into clinical-grade good manufacturing practice (GMP) conditions, allows human clinical relevance. Our protocol offers another methodology—provided that current safe-harbor criteria will expand and include factor-free characterized hiPSC-based derivatives for human therapeutic applications—for deriving GMP-grade hiPSCs, which should eliminate any immunogenicity risk due to non-human antigens. This protocol is broadly applicable to lentiviral reprogrammed cells of any type and provides a reproducible method for converting reprogrammed cells into GMP-grade conditions.
Introduction
Adult human cells have been shown to be capable of undergoing epigenetic remodeling and reprogramming, as a result of lentiviral-based expression of four key transcription factors4,5. An important advancement in the reprogramming field was the use of a single excisable lentiviral stem cell cassette (STEMCCA), which housed all four reprogramming transcription factors that allowed a precise stoichiometric ratio of protein expression6. Additionally, when transduced in specific multiplicity of infection ranges, STEMCCA can lead to predominantly single genomic integration events during the reprogramming process7. The introduction of an excisable version of STEMCCA, which utilizes Cre/loxP technology followed by excision of the reprogramming vector after derivation of the stem cell line, enabled factor-free human induced pluripotent stem cell (hiPSC) lines to be derived8. Additionally, in order to enhance therapeutic applications of hiPSCs, a novel, quick, and readily applicable methodology for good manufacturing practice (GMP)-grade cell line conversion, from xeno-containing to xeno-free conditions, needed to be implemented. Here, we discuss a relevant methodology that more precisely assesses integrated gene expression differences, specifically when integrated into an intron, and clinical-grade cell conversion into putative GMP conditions.
Previous research has used only relatively insensitive microarray transcriptional analysis to analyze gene expression differences in integrated genes after STEMCCA transduction3,9. Here, we introduce the methodology of sensitive quantitative polymerase chain reaction (PCR) analysis, to further examine integrated gene expression differences. Importantly, current safe-harbor criteria discard hiPSCs that have genes with viral integrations, thus limiting the applicability of these cells for downstream human cellular therapeutics9. We propose that the status quo may change with the use of fully characterized and transgene-free intronically reprogrammed hiPSCs. Additionally, we introduce a robust GMP-grade cell conversion protocol that can be readily applied to a variety of different cell types, which were originally derived under xeno-containing conditions10. This provides significant opportunities for the development of future cell reprogramming experiments, which require clinical-grade conditions to maintain human therapeutic relevance.
These methodologies provide a foundation upon which current safe-harbor criteria may be expanded to include characterized STEMCCA reprogrammed hiPSC lines that maintain a normal gene expression profile after STEMCCA excision from the integrated intron. Also, full conversion into clinical-grade conditions, free from non-human animal antigens, will help to incorporate many more cell types, which have previously been reprogrammed and characterized only in xeno-containing conditions. These methodologies combined, are persuasive grounds for the US Food and Drug Administration (FDA) to consider expanding their limited approval from human embryonic stem cell (ESC)-based therapeutics to hiPSC-based therapeutics11.
We recently detailed the derivation of a factor-free hiPSC line that was fully characterized and converted into putative clinical-grade conditions10. Here, we detail the protocol for hiPSC derivation by utilizing the STEMCCA lentivirus. These stem cells then undergo an excision process followed by gene expression characterization. Finally, the hiPSCs are converted over into GMP-grade conditions by a slow conversion methodology.
Protocol
NOTE: This method was used in the research reported in Awe et al.10.
1. Reprogramming Adult Human Dermal Fibroblasts with STEMCCA
- Thaw adult dermal human fibroblasts (HUFs), passage 4 or lower, in a 37 °C water bath for 2 min, taking care not to submerge the top of the vial with water.
- Place the 1-ml slurry of human fibroblasts into a 15-ml conical vial and slowly place 4 ml of standard HUF media, pre-warmed to 37 °C, in a drop-wise fashion, to dilute out dimethyl sulfoxide (DMSO) and re-suspend cells.
- Centrifuge the vial at 200 x g for 10 min at room temperature. Aspirate the supernatant and re-suspend cells in an adequate volume of media to create a suspension of 100,000 cells/ml and then add 2 ml into one well of a six-well plate, pre-coated for 20 min with 0.2% gelatin.
NOTE: A sister well with an equal number of cells per line and condition needs to be seeded for the cell counting/multiplicity of infection (MOI) calculation. - Rock the plate side to side and back and forth to evenly distribute the cells. Incubate overnight at 37 °C, 5% CO2 in a humidified incubator.
- On the day of transduction, obtain cell count(s) from the sister well(s) approximately 24 hr after initial HUF plating.
- Thaw out 2 x 108 TU, or equivalent, lentiviral concentrate (STEMCCA) and dilute to 1 x 108 TU/ml in HUF media.
- Transduce cells at a 10-MOI ratio in HUF media supplemented with 8 µg/ml transfection agent. Mix transduced wells evenly and incubate overnight at 37 °C, 5% CO2 in a humidified incubator.
- Approximately 24 hr after lentiviral transduction, switch the HUF media to standard human pluripotent stem cell (PSC) medium.
- On days 2 through 6, change media daily with human PSC medium.
- On day 6, plate two 0.2% gelatin-coated 10-cm plates of 1.5 to 1.75 x 106 MEFs derived from CF1 mice in HUF media 12.
- On day 7, being careful not to centrifuge more than 100 x g, use room temperature trypsin to lift off the day 7 reprogrammed fibroblasts from one well of the six-well plate and passage at a 1:16 ratio, by plating all cells evenly between two 10 cm plates, with fresh human PSC media, that were plated with MEFs the day before. Distribute cell aggregates in a clockwise spiral movement and place in a 37 °C, 5% CO2 humidified incubator overnight.
- Approximately 48 hr after seeding the reprogrammed cells onto MEFs and every day thereafter, change out spent human PSC media with fresh media.
- After 3 to 4 weeks, pick individual colonies with typical ESC like morphology with a 21-gauge needle and subclone out into a 24-well plate by placing approximately 8 to 10 individual pieces of specific colonies into individual wells in a 0.2% gelatin-coated 24-well plate pre-seeded with MEFs derived from CF1 mice in human PSC medium.
NOTE: After expansion of picked colonies, the colonies have to be characterized as pluripotent by embryoid body formation and expression analysis of pluripotency markers at the protein level.
2. Vector Integration Site Analysis and STEMCCA Excision
- Analyze the STEMCCA integration site by non-restrictive linear amplification PCR as described10.
- After assaying to ensure one integration into a desired area of the genome, excise STEMCCA by aspirating off the old human ESC medium. Then, combine 45 µl of concentrated Adeno-Cre-PuroR virus with 8 µg/ml transfection agent in 3 ml of standard PSC media for 24 hr.
- After the 24-hr incubation period, aspirate off the mixed viral supernatant and wash the cells twice with human PSC media. Place fresh human PSC media along with 2 µg/ml puromycin for a period of 5 days, changing the media daily with fresh media and antibiotic.
- After 5 days, subclone remaining colonies with a 21-gauge needle onto 0.2% gelatin-coated wells, in a 12-well plate pre-coated with MEFs derived from CF1mice and expand as detailed above.
NOTE: After sufficient expansion to obtain a frozen stock, conversion into feeder-free conditions is required. Expect at least 95% stem cell colony death as most cells will not be successfully transduced and will succumb to antibiotic over the 5 day period of incubation. Although Adeno-Cre-PuroR integrates at a 0.001-1% efficiency into host chromosomes of infected cells, reexposing a sample of factor-free colonies to puromycin to ensure 100% cell death will confirm no integration has occurred13. - Thaw out one vial of pre-aliquoted basement membrane matrix on ice, over a period of approximately 2 hr (manufacturer’s recommendations) until the matrix is a liquid and dilute out to a final concentration of 1:30 in cold basal media. Coat desired number of wells in a six-well plate with 1 ml per well. Let sit at room temperature for 1 hr.
- Cross-hatch (using an 18-gauge needle, or a glass or plastic “tip”) at least 20 to 30 colonies from a well previously coated with MEFs. Be careful to reduce MEF uplifting and transfer from plate.
- Generate the cross-hatching device through a number of different approaches, from the more complicated heating, pulling, and finishing of a glass Pasteur pipette on an open flame, to the simple usage of a sterile plastic pipette tip or, as in our case, 21- and/or 18-gauge needles.
- Aspirate the matrix from the coated well after incubation.
- Remove pieces of the colonies by gentle scraping from the old plate with a 200-µl pipettor and carefully place into 3 ml of fresh combined media 1 in the matrix-coated plate. Incubate overnight in a 37 °C, 5% CO2 humidified incubator. Change media 48 hr after passaging.
- Extract genomic DNA from post-excised and feeder-free converted hiPSCs, at more than 80% confluency, using commercial DNA extraction kits.
- Analyze for proper excision with primers specific for exogenous integrations of the STEMCCA lentivirus: gDNA-hendo-MycS-forward, 5′-acgagcacaagctcacctct-3′; gDNA-hWPRE-reverse, 5′-tcagcaaacacagtgcacacc-3′. Reconstitute lyophilized primers with PCR-grade water to a 10-µM stock solution.
- Run a PCR with the following five-step protocol: initial denaturation at 95 °C for 3 min; 35 cycles of each of the following: denaturation at 98 °C for 20 sec, primer annealing at 62 °C for 15 sec, and extension at 72 °C for 15 sec; followed by a single cycle final extension at 72 °C for 3 min.
- Make a 3% agarose gel by weighing out 1.5 grams of agarose and placing it into 50 ml 1x Tris-acetate-EDTA buffer. Microwave until agarose is dissolved and place in DNA gel stain at proper dilution, mix, and dispense proper volume into gel cassette to solidify for 1 hr at room temperature.
- Load 12 µl of PCR product mixed with 3 µl of loading dye into a 3% agarose gel. Run the gel for 30 min at 80 V.
- Visualize with ultraviolet light for the absence of a band indicating proper excision.
3. Quantitation of Gene Expression Differences by Using Quantitative PCR from Pre- and Post-excised hiPSCs
- Extract total RNA from post-excised and feeder-free converted hiPSCs by using commercial RNA extraction kits.
- Reverse-transcribe up to 1 µg of total RNA by using commercial kits in accordance with the instructions of the manufacturer. Use anchored-oligo(dT)18 and random hexamer primers.
NOTE: Ensure that the PCR protocol is tailored for gene transcripts of more than or less than 4 kb. Specific primers will need to be made for whichever gene STEMCCA originally integrated into. - Reconstitute the lyophilized primer with PCR-grade water to a 20 µM stock solution.
- Use 5 ng of sample per 20 µl of reaction that consists of 10 µM UPL probe, 2x LightCycler 480 Probes Master, and 20 µM forward and reverse primers.
4. Conversion into GMP-grade Conditions
- On the first day of transitioning the post-excised hiPSCs over to GMP-grade conditions, make a blend of media consisting of human PSC combined media 1 (combined media 1) and clinical grade good manufacturing practice (GMP) compliant media (combined media 2) at an 80:20 ratio.
- Aspirate off the old combined media 1 and place 3 ml of the new combined media 1 and 2, pre-warmed, at the 80:20 ratio. Change media daily for 3 days with this specific ratio. Place cells back into a 37 °C, 5% CO2 incubator.
- On day 4, make the combined media 1 and 2 at a 50:50 ratio. Aspirate off the old medium and replace with pre-warmed media, with the combined media 1 and 2 at the 50:50 ratio. Change media for 3 days with this specific ratio. Place cells back into a 37 °C, 5% CO2 incubator.
- On day 7, make the combined media 1 and 2 at a 20:80 ratio. Aspirate off the old medium and replace with pre-warmed media, with the combined media 1 and 2 at the 20:80 ratio. Change media for 3 days with this specific ratio. Place cells back into a 37 °C, 5% CO2 incubator.
- On day 10, make the combined media 1 and 2 at a 0:100 ratio. Aspirate off the old medium and replace with pre-warmed media, with the combined media 1 and 2 at the 0:100 ratio. Change media every day at this specific ratio. Cells are now in completely defined xeno-free media. Place cells back into a 37 °C, 5% CO2 incubator.
NOTE: During this entire conversion process, routine passaging with an 18-gauge needle is needed to maintain proper colony size and density. Plan on passaging cells every 4 to 5 days on the basis of colony size, confluency, and differentiation. - Coat one well in a six-well plate with a defined xeno-free substrate as recommended by the manufacturer.
- Mechanically passage a confluent well, already converted to xeno-free media conditions, from a six-well plate with an 18-gauge needle. Importantly, place new media (combined media 2) on the cells with 1x ROCK inhibitor for 1 hr, before passaging to pre-condition the media.
- Aspirate the synthetic substrate solution off the new plate that the cells are being transferred to. Remove all the media containing the stem cell clumps from the old plate and transfer to new plate.
- Place cells back into a 37 °C, 5% CO2 incubator for 2 days with minimal physical disturbance (ideally plates, once inside incubator, are not directly touched in any way) to allow maximum attachment.
NOTE: Cells will require daily media changes. Continual passaging every 4 days will be important. Using a 21-gauge needle, passage only the parts of colonies with optimal ESC-like morphology (high nucleocytoplasmic ratio). This will ensure selection for proper hiPSC colonies that will grow well and eventually yield homogenous colonies. Overall time to derivation of ESC like colonies is between 15-20 days after the initial combined media change. - Freeze down the GMP-grade post-excised hiPSCs by first cross-hatch all of the colonies by using an 18-gauge needle. Lift all the pieces off by using a 200 µl pipettor and transfer all media containing the cells to a 15 ml conical vial and centrifuge at 200 x g for 5 min at 4 °C.
- While the cells are centrifuging, make a freezing media consisting of equal volumes of cold combined media 2 and freezing medium with a final DMSO concentration of 7.5%.
- After cells are pelleted, aspirate off old medium and drop-wise re-suspend the pellet with the freezing media. Immediately transfer to freezing vials and put in a −80 °C freezer.
5. Characterizing GMP-grade Post-excised hiPSCs
- To ensure proper expression of pluripotency markers post-conversion, use QPCR to quantify expression of key pluripotency markers (SOX2, OCT4, and NANOG) by using the primers listed in Table 1 with the same steps as mentioned in step 3. QPCR methodologies follow the same steps as listed in step 3.
- Use flow cytometry to detect the non-human sialic acid Neu5Gc (N-glycolylneuraminic acid) in accordance with standard conditions listed in the kit as previously published10.
- Gently wash the cell culture plates with cells by using 1x phosphate-buffered saline (PBS).
- Dissociate colonies by using 1x dissociation reagent for 5 min at room temperature.
- During the 5-min incubation time of step 5.4, prepare the blocking solution (provided in kit) by making a 0.5% Blocking Agent in ice-cold PBS.
- Label the following set of tubes for flow cytometry analysis: Unstained; 4′,6-Diamidino-2-phenylindole (DAPI) DAPI; Control antibody 1:200; Primary antibody 1:200; Secondary antibody 1:200 Donkey Anti-Chicken IgG (H+L); Sample MEFs; Sample post-excised cells on matrix; and Sample post-excised cells on synthetic substrate.
- Gently dislodge cell aggregates from step 5.4 by adding 2 ml of blocking solution and transfer contents to a 15-ml conical tube. Centrifuge at 80 x g for 5 min at 4 °C.
- Wash the cells twice with blocking solution and centrifuge as stated in step 5.7.
- Gently re-suspend approximately 1 x 106 cells in 100 µl of blocking solution with the corresponding dilution volume of each antibody. Incubate with gentle rocking at 4 °C for 1 hr.
- Repeat washes three times as in step 5.8.
- Re-suspend the cell pellet in 400 µl of Diluent Buffer (from kit) with 1:100 of DAPI for tubes 2, 6, 7, and 8 as listed under step 5.6.
- Strain cells through a 40 µM strainer and run through flow cytometer.
Representative Results
We present a protocol for deriving clinical-grade factor-free hiPSCs by using the STEMCCA lentiviral-based reprogramming approach. Figure 1A shows a representative picture of three different pre-excised hiPSC lines, after reprogramming with the STEMCCA approach on a layer of MEFs. The primary advantage of the STEMCCA reprogramming approach lies in the consistent reprogramming success achieved by multiple scientists, in different research groups and locations. Figure 1B presents the post-excision reverse transcription-PCR gel, showing one particular subclone (2.3) which is completely free from the STEMCCA lentivirus, as evidenced by the lack of an amplicon band specific for a particular sequence endogenous to STEMCCA. Figure 2A shows the pre-converted hiPSCs on a xeno containing matrix and post-excised xeno-free GMP-grade hiPSCs on synthetic matrix. Quantitation of standard pluripotency-associated factors (SOX2, OCT4, and NANOG) by using QPCR is illustrated in Figure 2B. Pre-converted and post-converted hiPSCs to GMP-grade conditions were tested through flow cytometry for the sialic acid N-glycolylneuraminic acid, indicative of non-human antigens, as shown in Figure 2C.
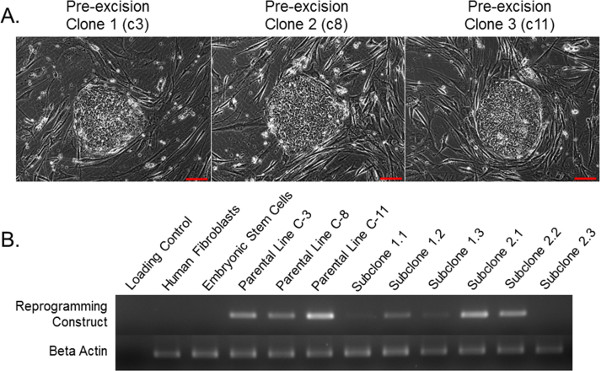
Figure 1: Representative human pluripotent stem cell cassette (hSTEMCCA)-derived human induced pluripotent stem cell (hiPSC) lines. (A) All three derived lines were analyzed with nrLAM-PCR technology and only the presented C8 line was found to have one integration into the PRPF39 gene. Only this line, due to the safe intronic STEMCCA integration, was selected to undergo Adeno-Cre-PuroR selection for Cre-mediated excision. (B) Adeno-Cre mediated STEMCCA excision out of the C8 line. Primers against unique nucleotide sequences found in STEMCCA-endo-Myc-s and A-WPRE- found that only one subclone (2.3 post-excised iPSCs) were properly excised and free of transgenic transcription factors from the integrated provirus. Bars = 100 μm. This figure has been modified from Awe et al.10.
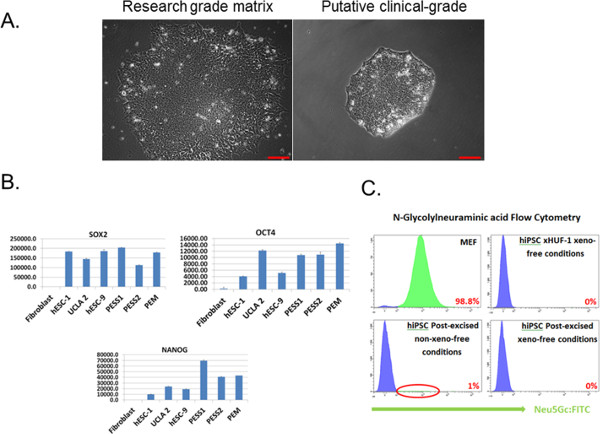
Figure 2: Conversion of hiPSCs from xeno-containing to clinical grade GMP xeno-free conditions and characterization. (A) Representative hiPSC line that has been converted from xeno-containing (research grade matrix) to xeno-free (synthetic substrate) conditions under current good manufacturing practice (GMP) conditions. (B) Quantitative polymerase chain reaction to assay for proper pluripotency associated gene expression post-conversion into GMP grade conditions. (C) In addition to the standard sterility tests to ensure GMP compatibility, a flow cytometry-based assay testing for the non-human antigen, N-glycolylneuraminic acid, is used to show that post-converted hiPSCs eliminate all sialic acid detection (1% with post-excised cells on matrix compared to 0% with post-excised cells on a synthetic substrate). Mouse embryonic fibroblasts and a hiPSC line derived under GMP conditions served as positive and negative controls, respectively, in this experiment. Bars = 100 µm. hESC, human embryonic stem cell; PESS, post-excised synthetic substrate; PEM, post-excised matrix. This figure has been modified from Awe et al.10.
| Gene Name | Forward Primer 5'-3' | Reverse Primer 5'-3' | Probe # |
| QRT-hPOU5F1 | gaagttaggtgggcagcttg | tgtggccccaaggaatagt | 13 |
| QRT-hSOX2 | gggggaatggaccttgtatag | gcaaagctcctaccgtacca | 65 |
| QRT-hNANOG | cagtctggacactggctgaa | cacgtggtttccaaacaaga | 55 |
Table 1: Primers used for quantitative PCR analysis.
Discussion
We describe a methodology of deriving factor-free hiPSCs and making them clinically relevant by converting these cells into GMP-grade conditions for downstream cell differentiation in future human therapeutics. Although this protocol is broadly applicable to a variety of cell types, we chose to reprogram human dermal fibroblasts, due to the ease of extraction from the patient and their applicability to personalized human therapeutics. Once limitations are remedied as far as full differentiation into clinically relevant cell derivatives is concerned14, the presented conversion into GMP-grade conditions will become even more relevant to a variety of different cell types.
The first part of this study involves reprogramming human fibroblasts with the STEMCCA lentivirus. This technique was chosen in large part due to its reproducibility, relatively high reprogramming efficiency (0.02%), and robust ability to reprogram across many different cell types10. This technique is superior to other reprogramming methodologies such as sendai virus, episomal plasmids, and synthetic mRNA based reprogramming that suffer from lower reprogramming efficiencies and reproducibility10. Although there is a correlation between HIV-based vectors integrating into increased gene activity hotspots15, there is no guarantee that they will integrate into a gene, much less into a safer area of a gene, the intron. Thus this obstacle represents a notable limitation to this approach. Additionally, this integration site preference into a safer genomic location decreases with more integrations throughout the genome. Therefore, it is imperative that proper lentiviral provirus integration site and integration number be properly interrogated by sensitive techniques such as nrLAM-PCR technology. Future reprogramming utilizing zinc finger nuclease technology, TALEN, or CRISPR/Cas9-based gene targeting (via homologous recombination) may further guide this field into specific loci for reprogramming16.
Once the human fibroblasts are properly reprogrammed and colonies have been derived, another critical aspect of this protocol is the Adeno-Cre-PuroR expression for selection of post-excised hiPSCs. It is important to consider re-exposing the colonies to puromycin after a few weeks of cell culture, in order to ensure that all the adenovirus has been diluted out through cell replication and has not sporadically integrated into the genome. Improper integration into the genome of the adenovirus would represent a safety concern for personalized cellular therapeutics.
Conversion into GMP-grade conditions is an important step in introducing clinical applicability to these factor-free hiPSCs. It is important to change only one condition at a time when converting these cells over to xeno-free conditions; start by converting the cells into xeno-free media through the slow conversion methodology. The hiPSCs cannot tolerate a media change and a substrate change at the same time. Once the cells are passaging regularly with normal morphology in the new media conditions, begin the transition onto the xeno-free substrate. If specific cell types are having trouble transferring over onto the recommended substrate, it is feasible to consider trying other xeno-free substrates on the market.
In conclusion, the robust and broad applicability of this reprogramming technique along with the GMP-grade conversion should allow easy reproducibility and serves as a foundation for future hiPSC-based derivative cell culture for human therapeutics.
Declarações
The authors have nothing to disclose.
Acknowledgements
We would like to thank Patrick C. Lee, Cyril Ramathal, and Saravanan Karumbayaram (SK) for their assistance in performing the iPSC derivation and characterization experiments; Aaron Cooper for performing the iPSC analysis experiments; Vittorio Sebastiano and Renee A. Reijo Pera for directing the initial reprogramming efforts; SK, William E. Lowry, Jerome A. Zack, and Donald B. Kohn for directing the establishment of the UCLA GMP facilities permitting the conversion and characterization of clinical-grade iPSCs; Gustavo Mostoslavsky for providing us with the STEMCCA polycistronic reprogramming vector. This work is based on a research collaboration with Fibrocell Science and the Clinical Investigations for Dermal Mesenchymally Obtained Derivatives (CIDMOD) Initiative to generate safe personalized cellular therapeutics. This work was supported by funding from the Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research at UCLA, The Phelps Family Foundation, Fibrocell Science, Inc., and the UCLA CTSI Scholar’s Award to JAB.
Materials
| Name | Company | Catalog Number | Comments |
| Media and reagents | |||
| DMEM/F12 (basal media) | Invitrogen (Carlsbad, CA, USA) | 11330057 | |
| Fetal bovine serum | Invitrogen | 16000044 | |
| Minimum essential medium (MEM) non-essential amino acids (NEAA), 100x | Invitrogen | 11140050 | |
| Glutamax, 100x | Invitrogen | 35050-061 | |
| PenStrep, penicillin-streptomycin, 100x | Invitrogen | 15140-122 | Thaw at 4 °C, aliquot and store at −20 °C |
| Knockout serum replacement | Invitrogen | 10828028 | Thaw at 4 °C, aliquot and store at −20 °C |
| Trypsin/EDTA, 0.5% | Invitrogen | 15400-054 | Dilute stock out to 0.05% in 1x PBS |
| Basic fibroblast growth factor | GlobalStem (Rockville, MD, USA) | GSR-2001 | Reconstitute to 10 µg/ml stock in 0.1% bovine serum albumin dissolved in 1x PBS and store at −80 °C |
| β-mercaptoethanol | Millipore (Billerica, MA, USA) | ES-007-E | |
| Matrigel (basement membrane matrix) | BD Biosciences (San Jose, CA, USA) | 356231 | Dilute stock Matrigel vial with 10 ml of DMEM/F12 while on ice for a 1:2 dilution. Aliquot and store at −20 °C |
| CELLstart (Synthetic Substrate) | Invitrogen | A1014201 | |
| Stemmolecule Y27632 | Stemgent (Cambridge, MA, USA) | 04-0012-02 | |
| Puromycin | Invitrogen | A1113802 | |
| LightCycler 480 Probes Master | Roche (Basel, Switzerland) | 4707494001 | |
| ProFreeze-CDM Medium/freezing medium | Lonza (Basel, Switzerland) | 12-769E | |
| Dimethyl sulfoxide | Sigma-Aldrich (St. Louis, MO, USA) | D8418 | |
| PBS | Invitrogen | 14190-250 | |
| 100 BP DNA Ladder | Invitrogen | 15628019 | |
| SYBR Safe DNA Gel Stain 10000x | Invitrogen | S33102 | |
| Agarose | Bio-Rad Laboratories, Inc. (Hercules, CA, USA) | 161-3101 | |
| Gelatin, from porcine skin | Sigma-Aldrich | G1890-100G | Make stock at 0.2% in PBS, autoclave and store at room temperature |
| mTeSR1 | StemCell Technologies (Vancouver, BC, Canada) | 5850 | Combine Supplement 5X with the basic medium, aliquot and store at 4 °C for up to 2 weeks. |
| Stemedia NutriStem XF/FF Culture Medium | Stemgent | 05-100-1A | Thaw at 4 °C O/N, aliquot and store at 4 °C for up to 2 weeks. |
| Primocin | InvivoGen (San Diego, CA, USA) | ant-pm-1 | |
| Accutase (Dissociation Reagent) | Invitrogen | A1110501 | |
| Donkey anti-Chicken IgG AlexaFluor 488 | Jackson ImmunoResearch (West Grove, PA, USA) | 703-546-155 | |
| Polybrene/transfection agent | Millipore | TR-1003-G | |
| Plasticware | |||
| 12-well plates | VWR (West Chester, PA, USA) | 29442-038 | |
| 6-well plates | VWR | 29442-042 | |
| 10-cm plates | Sigma-Aldrich | Z688819 | |
| 18-gauge needle | Fisher Scientific (Pittsburgh, PA, USA) | 148265D | |
| 21-gauge needle | Fisher Scientific | 14-829-10D | |
| Equipment | |||
| BD LSRII Flow Cytometer | KSystem by Nikon (Tokyo, Japan) | ||
| BD FACSDiva Version 6.1.3 Software | BD Biosciences | ||
| Kits | |||
| PureLink Genomic DNA Mini Kit | Invitrogen | K182000 | |
| KAPA HiFi Hotstart ReadyMix PCR Kit | KAPA Biosystems (Wilmington, MA, USA) | KK2601 | |
| High Pure RNA Isolation Kit | Roche | 11828665001 | |
| Transcriptor First Strand cDNA Synthesis Kit | Roche | 4379012001 | |
| Sialix anti-Neu5Gc Basic Pack Kit | Sialix (Newton, MA, USA) | Basic Pack | |
| Media | |||
| Combined media 1 | StemCell Technologies and Stemgent | Consists of equal parts mTeSR1 and Nutristem | |
| Combined media 2 | StemCell Technologies and Stemgent | Consists of equal parts TeSR2 and Nutristem | |
| HUF Media | Dulbecco’s modified Eagle’s medium/F12 [DMEM/F12] supplemented with 10% fetal bovine serum, 1x non-essential amino acids, 1x Glutamax, and 1x Primocin | ||
| Human Pluripotent Stem Cell Media | DMEM/F12 supplemented with 20% knockout serum replacement, 1x Glutamax, 1x non-essential amino acids, 1x Primocin, 1x β-mercaptoethanol, and 10 ng/ml basic fibroblast growth factor | ||
| DMEM/F12, Dulbecco’s modified Eagle’s medium/F12; PBS, phosphate-buffered saline. | |||
Referências
- Sommer, C. A., Mostoslavsky, G. The evolving field of induced pluripotency: Recent progress and future challenges. J Cell Physiol. 228, 267-275 (2013).
- Lu, H. F., et al. A defined xeno-free and feeder-free culture system for the derivation, expansion and direct differentiation of transgene-free patient-specific induced pluripotent stem cells. Biomaterials. 35, 2816-2826 (2014).
- Karumbayaram, S., et al. From skin biopsy to neurons through a pluripotent intermediate under Good Manufacturing Practice protocols. Stem Cells Transl Med. 1, 36-43 (2012).
- Mostoslavsky, G. Concise review: the magic act of generating induced pluripotent stem cells: many rabbits in the hat. Stem Cells. 30, 28-32 (2012).
- Takahashi, K., et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 131, 861-872 (2007).
- Sommer, C. A., et al. Induced pluripotent stem cell generation using a single lentiviral stem cell cassette. Stem Cells. 27, 543-549 (2009).
- Somers, A., et al. Generation of transgene-free lung disease-specific human induced pluripotent stem cells using a single excisable lentiviral stem cell cassette. Stem Cells. 28, 1728-1740 (2010).
- Sommer, C. A., et al. Excision of reprogramming transgenes improves the differentiation potential of iPS cells generated with a single excisable vector. Stem Cells. 28, 64-74 (2010).
- Papapetrou, E. P., et al. Genomic safe harbors permit high beta-globin transgene expression in thalassemia induced pluripotent stem cells. Nat Biotechnol. 29, 73-78 (2011).
- Awe, J. P., et al. Generation and characterization of transgene-free human induced pluripotent stem cells and conversion to putative clinical-grade status. Stem Cell Res Ther. 4, (2013).
- . Receives FDA Clearance to Begin World’s First Human Clinical Trial of Embryonic Stem Cell-Based Therapy. Geron Corporation Press Release. , (2009).
- Jozefczuk, J., Drews, K., Adjaye, J. Preparation of mouse embryonic fibroblast cells suitable for culturing human embryonic and induced pluripotent stem cells. J Vis Exp. 64, 3810-383791 (2012).
- Mitani, K., Kubo, S. Adenovirus as an integrating vector. Curr Gene Ther. 2, 135-144 (2002).
- Patterson, M., et al. Defining the nature of human pluripotent stem cell progeny. Cell Res. 22, 178-193 (2012).
- Schroder, A. R., et al. HIV-1 integration in the human genome favors active genes and local hotspots. Cell. 110, 521-529 (2002).
- Hockemeyer, D., et al. Genetic engineering of human pluripotent cells using TALE nucleases. Nat Biotechnol. 29, 731-734 (2011).

