High-Dimensionality Flow Cytometry for Immune Function Analysis of Dissected Implant Tissues
Summary
Isolation of cells from dissected implants and their characterization by flow cytometry can significantly contribute to understanding the pattern of immune response against implants. This paper describes a precise method for the isolation of cells from dissected implants and their staining for flow cytometric analysis.
Abstract
The success of implanting laboratory-grown tissue or a medical device in an individual is subject to the immune response of the recipient host. Considering an implant as a foreign body, a hostile and dysregulated immune response may result in the rejection of the implant, while a regulated response and regaining of homeostasis can lead to its acceptance. Analyzing the microenvironments of implants dissected out under in vivo or ex vivo settings can help in understanding the pattern of immune response, which can ultimately help in developing new generations of biomaterials. Flow cytometry is a well-known technique for characterizing immune cells and their subsets based on their cell surface markers. This review describes a protocol based on manual dicing, enzymatic digestion, and filtration through a cell strainer for the isolation of uniform cell suspensions from dissected implant tissue. Further, a multicolor flow cytometry staining protocol has been explained, along with steps for initial cytometer settings to characterize and quantify these isolated cells by flow cytometry.
Introduction
Advances in the field of medicine have led to the frequent use of implanted materials for supporting the function or re-growth of damaged tissue1,2. These include devices such as pacemakers, reconstructive cosmetic implants, and orthopedic plates used for bone fracture fixation3,4. However, the materials used to make these implants and the locations in which they are implanted play important roles in determining the success of these implants5,6,7. As foreign bodies, these implants can generate an immune response from the host that can either lead to rejection or tolerance8. This factor has driven biomaterial research to generate materials that can attract the desired immune response after implantation9,10,11,12.
The immune response is an essential requirement in the field of regenerative medicine, where a tissue or an organ is grown around a biomaterial skeleton (scaffold) in a laboratory for the replacement of a damaged tissue or organ13,14,15,16. In regenerative medicine, the goal is to replace missing or damaged tissue through the use of cells, signals, and scaffolds, each of which can be greatly modulated by immune responses17. Furthermore, even when a lack of immune response is desired, it is very rarely an absence of immune activity rather than the presence of a regulatory profile that is desired18. Techniques such as flow cytometry can play a significant role in characterizing the pattern of immune response to various biomaterials used for coating implant devices or for developing scaffolds for tissue engineering19.
This information, in turn, will ultimately help in developing biomaterials for implants that can be well-tolerated by the immune system or in developing scaffolds that can play a constructive role in tissue engineering. Proper preparation of samples for analysis by flow cytometry is an important step for avoiding inaccurate results in immune characterization via fluorescence activated cell sorting20,21. Therefore, this review presents a detailed methodology that can be utilized for the isolation of cells from scaffold tissue, staining the cell suspension, and analysis by flow cytometry.
Protocol
NOTE: Figure 1 gives an overview of the flow cytometry protocol.
1) Reagent preparation
- Prepare media for diluting enzymes and for tissue culture.
- Add 5 mL of 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES) buffer solution into 500 mL of RPMI medium and shake well. Store the medium at 4 °C until further use.
- Calculate the volume of the enzyme solution.
NOTE: The volume of the enzyme solution is the volume of the medium containing the enzymes (collagenase and DNase I) that will be needed to digest diced tissue in 6-well plates; it depends on the number of samples.- Use the following equation to calculate the required volume:
(Number of samples to be digested) × 5 mL of serum-free RPMI with 10 mM HEPES buffer
NOTE: For example, for 0.1 to 0.3 g of dissected spleen, use 5 mL of medium to prepare the enzyme solution. The enzymatic digestion protocol described in this manuscript is primarily for soft tissues (ranging from 0.1 to 1 g) dissected from mice which include the brain, kidney, skin, liver, lungs, spleen, muscles, as well as capsules around subcutaneous implants made of synthetic materials or extracellular matrix proteins. Digesting hard cartilaginous tissues might require different conditions and volumes of digestive enzymes, which can only be ascertained by optimization. - Calculate the amount of collagenase and DNase I needed for the enzyme solution.
NOTE: In this protocol, 0.25 mg/mL of collagenase and 0.2 mg/mL of DNase I were used. The working concentrations required for collagenase and DNase can vary for different types of tissues19.
- Use the following equation to calculate the required volume:
- Prepare a 1:1000 viability dye solution by adding 1 µL of the viability dye (see Table of Materials) to 999 µL of phosphate-buffered saline (PBS) and vortex the solution. Store the solution in the dark at 4 °C until further use.
- Prepare the staining buffer by adding 1 g of bovine serum albumin (BSA) to 100 mL of PBS, and vortex the solution until BSA is completely dissolved. Store the solution at 4 °C until further use.
2) Setting up enzymatic digestion plates
- Set up a 6-well plate with 70 µm cell strainers in each well. Add 3 mL of the prepared medium from step 1.1.1 into each well, and incubate on ice.
- Store the remaining media (2 mL/well) in an incubator or water bath at 37 °C for suspending the calculated amount of collagenase and DNase I (step 1.2.2).
3) Isolation of cells
- Place the dissected implants/tissue into plates, and dice finely using scissors. Alternatively, utilize a mechanical disruption method using a tissue dissociator or hand-held homogenizer. Owing to the high number of leukocytes, dissect out the spleen to utilize as a staining control in each run.
NOTE: As some materials induce higher levels of fibrosis, this protocol is applicable for both highly fibrotic and minimally fibrotic materials. However, each processing method should be evaluated for both cell yield and viability after digestion prior to staining. Avoid peripheral blood contamination during dissection. - Add collagenase and DNase I (volume calculated in step 1.2.2) to the remaining RPMI (2 mL) that has been warmed to 37 °C. Add 2 mL of the complete enzyme solution to each well.
- Place the plates in an incubated shaker for 45 min at 37 °C and 100 rpm.
NOTE: Be cautious when stacking plates as this can inhibit proper heat transfer. - Meanwhile, prepare a suspension-dispensing tip by chopping 3-4 mm from the end of a 1000 µL tip with scissors. After incubation, pipette the suspension in the wells up and down to mix it thoroughly using the chopped tip. Place the cell strainer on the top of a 50 mL conical tube and pass the digested solution through the cell strainer.
NOTE: The 70 µm cell strainer is sufficient to separate cells from implanted biomaterial, such as alginate. If greater purity is required, use processes such as density gradient separation to obtain an enriched population of leukocytes. - Rinse the wells with 1x PBS solution, and transfer the rinse volume through the strainer, followed by adding the washes of the strainer with 1x PBS solution until the volume in the tube reaches 50 mL.
NOTE: The 1x PBS must be at room temperature to prevent any increase in the viscosity of the digest and to allow cell pelleting during centrifugation. - Centrifuge the tube at 300 × g for 5 min at room temperature. Following centrifugation, aspirate the supernatant carefully using a serological pipette without disturbing the pellet. Re-suspend the pellet in 1000 µL of 1x PBS, and transfer the suspension to a microcentrifuge tube.
- Mix 10 µL of the cell suspension with 10 µL of trypan blue, and load the suspension on counting chamber slides for counting cells using an automatic cell counter or on a hemocytometer to count via the conventional method under a microscope. Alternatively, use flow cytometry cell counting beads.
4) Staining for flow cytometry
- See Figure 2 for an example plate layout for a 14-color experiment with 45 samples, fluorescence minus one (FMO) controls, compensation controls, and a fully stained spleen sample control. After estimating the total number of cells, calculate the volume of the cell suspension required for staining 1 × 106 cells for each sample, and 0.5 × 106 cells for each compensation and FMO control. Dispense the required volume of the cell suspension into the wells of a V-bottom 96-well plate, and make up the volume to 100 µL with 1x PBS.
NOTE: As an example, if 1000 µL of cell suspension obtained at step 3.6 contain 40 × 106 cells, then for 1 × 106 cells, 1000/40 × 106 = 25 µL of cell suspension will be required. - Centrifuge the plate at 300 × g for 5 min at 4 °C, aspirate the supernatant, and then resuspend the cells in the sample wells and in a compensation control well to determine viability, with 100 µL of a 1:1000 solution of the viability dye (see Table of Materials).
- For cells in the other compensation wells as well as FMO and unstained wells, resuspend them with 100 µL of 1x PBS.
- Incubate the cells in the dark for 30 min at 4 °C.
- In the meantime, prepare a surface antibody cocktail for staining the sample and FMO controls: 50 µL per sample or FMO. See Table 1 for an example of the antibody cocktail.
- After incubation, add 100 µl of 1x PBS to each well, spin down at 300 × g for 5 min at 4 °C.
- Aspirate the supernatant and resuspend in 200 µL of staining buffer (1x PBS + 1% BSA); centrifuge at 300 × g for 5 min at 4 °C.
- Aspirate the supernatant, and resuspend the cells in the compensation, FMO, and unstained wells with 50 µL of 1x PBS. Add 50 µL of the antibody cocktail to the respective sample wells and FMO wells. Add the respective antibodies to the different compensation wells.
- Incubate the plate for 45 min in the dark at 4 °C. After incubation, add 150 µL of staining buffer in each well, and centrifuge the cell suspension at 300 × g for 5 min at 4 °C.
- Aspirate the supernatant, resuspend the cells with 200 µL of staining buffer, and centrifuge the cell suspension at 300 × g for 5 min at 4 °C.
- If analyzing samples without fixation, resuspend them in 200 µL of staining buffer, and proceed to section 6 on cytometer setup and sample analysis. If fixing the cells, aspirate the supernatant after washing, and add 100 µL of a fixative such as 4% paraformaldehyde. If staining for intracellular markers, proceed to section 5 on intracellular staining, and fix and permeabilize the cells (see Table of Materials). Incubate cells for 20 min in the dark at 4 °C.
NOTE: As fixation can affect the fluorescence intensity of some fluorophores, evaluate each panel both with and without fixation. - After incubation, add 100 µL of 1x PBS, followed by centrifugation at 300 × g for 5 min at 4 °C. Aspirate the supernatant, resuspend in 1x PBS, and centrifuge at 300 × g for 5 min at 4 °C.
- Re-suspend the cells in 200 µL of staining buffer, and store at 4 °C prior to flow cytometric analysis.
5) Intracellular staining
- Continuing from step 4.11 for fixation and permeabilization for intracellular markers, add 100 µL of the appropriate buffer. Centrifuge the cells at 350 × g for 5 min at 4 °C. Aspirate the supernatant, and re-suspend the pellets in 200 µL of the fixative-permeabilizing agent solution; centrifuge the cells at 350 × g for 5 min at 4 °C. Aspirate the supernatant, and resuspend the pellets with intracellular antibodies diluted in the appropriate buffer.
NOTE: Antibody types and dilutions will depend on the target cells. For example, one common intracellular marker is forkhead box P3 for regulatory T cells, which is frequently used at a 1:100 dilution. - Incubate the cell suspension in the dark for 45 min at 4 °C. After incubation, add 150 µL of the appropriate buffer, and centrifuge the suspension at 350 × g for 5 min at 4 °C.
- Aspirate the supernatant, and re-suspend the cells in 200 µL of staining buffer followed by centrifugation at 300 × g for 5 min at 4 °C. Aspirate and re-suspend the cells in 200 µL of staining buffer for flow cytometric analysis.
6) Cytometer and compensation setup
- Immediately before running the samples, prepare compensation beads (see Table of Materials) if utilizing bead-based compensation as opposed to cell-based compensation.
- Label separate microcentrifuge tubes for each fluorochrome-conjugated antibody, and add 100 µL of 1x PBS followed by one full drop of anti-mouse negative control compensation beads and one drop of positive control compensation beads to each tube. Add 1 µL of the appropriate antibody to each separate tube. Vortex and incubate for 5 min before acquisition.
NOTE: As some dyes, such as BUV737, cannot be bound to beads, use a cell-based control.
- Label separate microcentrifuge tubes for each fluorochrome-conjugated antibody, and add 100 µL of 1x PBS followed by one full drop of anti-mouse negative control compensation beads and one drop of positive control compensation beads to each tube. Add 1 µL of the appropriate antibody to each separate tube. Vortex and incubate for 5 min before acquisition.
- Calibrate the flow cytometer before each experiment by running the cytometer setup with the tracking beads, and maintain the flow cytometer settings for each experiment.
- Before analyzing the sample, adjust the flow cytometer by running an unstained sample to adjust the cell population on a side scatter (SSC) versus forward scatter (FSC) plot so that the cell population falls in the center of the plot and is not off-scale.
- Run the stained sample briefly to adjust the voltages for FSC and SSC and for each channel to make sure that no events fall outside the logarithmic scale of each channel. Record 5,000 events for each compensation control, followed by gating of positive and negative populations. Calculate the compensation matrix and then run the samples, gathering at least 1,000 events for the populations of interest.
NOTE: Compensation on larger-color panels should be verified, as automated compensation can be prone to over- or under-compensation. Each parameter should be graphed against every other parameter to monitor for signs of over- or under-compensation, and each single-color control should be evaluated. Complex compensation of larger-color experiments should be carried out with the assistance of a collaborator or a core facility with extensive flow cytometry experience. Newer types of flow cytometers such as spectral cytometers utilize the full spectrum of a fluorophore through a process known as spectral unmixing, which can yield a cleaner distinction of fluorescent overlap as compared to standard compensation alone22.
Representative Results
The process of development of flow cytometry panels for immune analysis often relies on the comparison of results to existing data and the literature in the field. Knowledge of how populations may present in flow cytometry is critical for proper interpretation of data. Regardless, populations and cell types can appear differently in different tissues, so some variability is to be expected. In the context of well-defined control tissues, such staining optimization can be evaluated against known tissue that has well-researched cell types. Figure 3 shows the results of a 14-color FACS on control mouse tissue. In this case, the spleen was used to identify all markers that were stained for, and to show that the staining was technically functional. From this point onwards, it became easier to test in unknown conditions, be confident of the fluorophore selection, and optimize the protocol for different tissue sources. Figure 3 also shows populations of different cell types isolated from a murine spleen19.
Here, several obvious populations, such as Ly6G+ neutrophils, and different expression levels of Ly6C on different monocyte classes could be observed. CD11chiMHCII+ dendritic cells were readily apparent when gated against CD11b to rule out macrophages and other myeloid lineage cells such as neutrophils and monocytes. A subset of CD206+CD86+ dendritic cells could be found by focusing on this CD11c+ population. CD86 and CD206 were included to phenotype myeloid cells as being in a more M1-like (CD86hi) versus in a more M2-like (CD206hi) polarization state. F4/80 showed a gradient of expression, that can be commonly observed in various macrophage populations. Siglec-F was present in two populations, F4/80+ and F4/80-, most likely corresponding to a macrophage subset and eosinophils, respectively. Although these designations of cell types can be made, it is important to note that cells expressing different markers should be viewed in a functional manner as opposed to a more binary classification. Acknowledgement that what is considered a macrophage may differ in the spleen and the foreign body capsule around an implant, is important and avoids the potential misinterpretation of results.
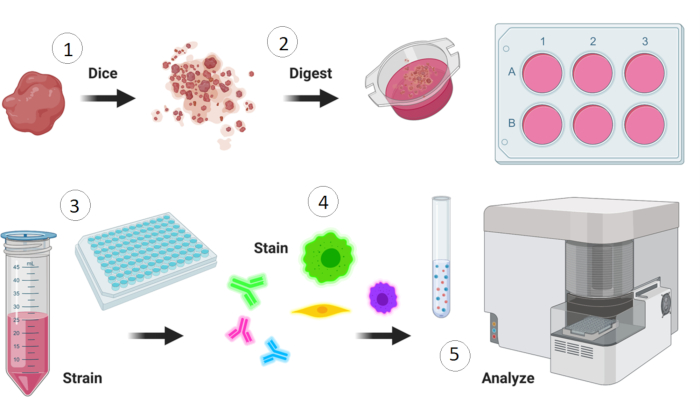
Figure 1: Overview of flow cytometry staining protocol. Preparation includes dissection of the tissue of interest followed by 1) manual dicing, 2) enzymatic digestion, 3) cell straining and washing, and 4) staining with fluorescently tagged antibodies followed by flow cytometric analysis. The illustration was made with Biorender. Please click here to view a larger version of this figure.
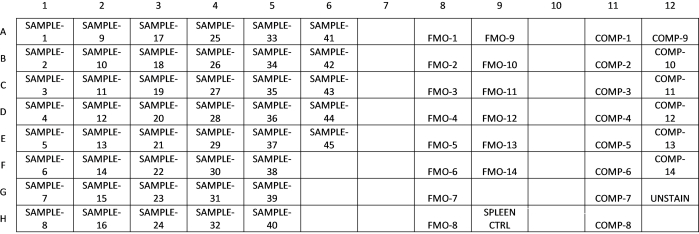
Figure 2: Example of a plate layout for flow cytometry staining. This layout includes the samples, fluorescence minus one (FMO) controls, compensation controls, and a control tissue sample (spleen). Please click here to view a larger version of this figure.
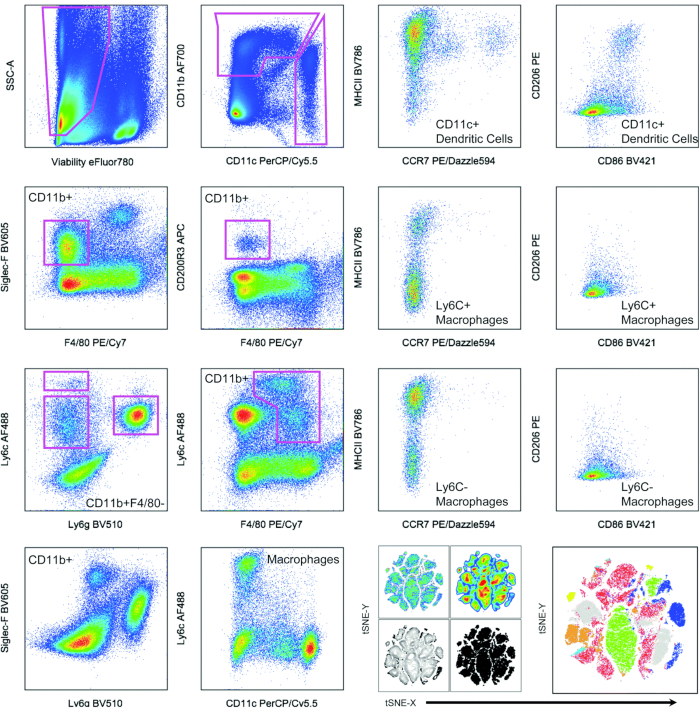
Figure 3: Representative results from 14-color FACS staining on control mouse tissue. Myeloid cell phenotyping of murine spleen cells with examples of hand-gating and automated t-stochastic neighbor-embedding (t-SNE) clustering algorithm for data display. This figure has been reproduced from Sadtler and Elisseeff19. Abbreviations: FACS = fluorescence-activated cell sorting; SSC = side scatter; CD = cluster of differentiation; PE = phycoerythrin; CCR = C-C chemokine receptor type; MHC = major histocompatibility complex; PerCP = peridinin chlorophyll protein complex. Please click here to view a larger version of this figure.
| Reagent/Antibody | µL per sample |
| CD86 BUV395 | 0.25 |
| CD45 BUV737 | 0.5 |
| CD8a BV421 | 0.25 |
| Ly6g BV510 | 0.125 |
| Siglec F BV605 | 0.25 |
| MHCII BV786 | 0.25 |
| Ly6c AF488 | 0.125 |
| CD11c PerCP/Cy5.5 | 0.2 |
| CD206 PE | 0.2 |
| CD197 PE/Dazzle594 | 0.125 |
| F4/80 PE/Cy7 | 0.25 |
| CD200R3 APC | 0.25 |
| CD11b AF700 | 0.25 |
| Fc Block | 1 |
| BD Brilliant Stain Buffer Plus | 10 |
| 1x PBS | 35.975 |
| Total Volume: | 50 µL |
Table 1: Example of surface antibody cocktail. An example antibody cocktail for myeloid phenotyping of mouse tissue.
Discussion
This review describes a detailed methodology for isolating cells from biomaterial implants to obtain a uniform cell suspension. In addition, a detailed protocol has been provided for staining the cell suspension for multicolor flow cytometry, along with the steps for configuring a flow cytometer for optimal results. Cell isolation methods can involve multiple steps, often utilizing manual tissue dissection followed by enzymatic digestion with proteolytic enzymes to dissociate the extracellular matrix in the tissue and disrupt the cell-cell junctions to liberate individual cells from the tissue. After digestion, further processing, such as cell straining, is needed to remove remaining debris and ensure a single-cell suspension. Some samples that have high levels of debris or other cell types (such as tumors) may require more thorough clean-up through the use of density separation media23. Without proper sample clean-up, data may be skewed due to debris or other cell populations obscuring the populations of interest. Excess debris can also lead to full or partial clogging of the cytometer fluidics.
While the characterization of immune cells can also be performed by other methods, such as light microscopy, by performing a differential cell count on a cell cytospin preparation, flow cytometry provides a more accurate characterization of the cells based on cell surface markers. Additionally, flow cytometry data are more precise as they can characterize and quantify millions of cells in suspension and can provide accurate estimates of distinct cell subsets much faster than manual differential counting based on only 400 cells24. The results shown in this paper rely on an iterative panel design with antibody selection done by the utilization of multiple online tools to compare the excitation and emission spectra and theoretical overlap depending on the cytometer configuration. When designing larger color experiments, it is best to start from a clean slate, as opposed to the addition to a smaller color panel, so as to fully consider antigen abundance, fluorophore brightness, and spectral overlap.
Studies analyzing distinct subsets of specific immune cells require sufficient number of overall cells to perform flow cytometry, as having a smaller number of cells can pose a significant challenge in their isolation by cell sorting or obtaining accurate estimates. This challenge can be overcome by using magnetic beads for isolating and enriching specific immune cells, such as neutrophils, in shorter periods and with limited washing25. Isolated cells from tissues of organs such as lungs can contain significant amounts of mucus, which can make the cell suspension "sticky" and result in an increased number of doublet events during flow cytometry26,27. Adding chelating agents such as 2 mM ethylenediamine tetraacetic acid to the staining buffer can prevent the aggregation of cells in such samples.
Additionally, suspending cells in an increased volume of buffer can also limit doublet creation by reducing cell-cell interactions. Staining procedures should be thoroughly optimized for each different tissue, staining protocol, flow cytometer, and antibody panel. Some fluorophores can be more sensitive to fixatives, some cell types are more sensitive to different cell isolation and processing methods, and many cell types behave differently in different tissues and different locations. With proper preparation and diligent analysis, flow cytometry can yield a detailed cellular and protein-level analysis to characterize the scaffold and biomaterial immune microenvironment.
Declarações
The authors have nothing to disclose.
Acknowledgements
This research was supported in part by the Intramural Research Program of the NIH, including the National Institute of Biomedical Imaging and Bioengineering. Disclaimer: The NIH, its officers, and employees do no recommend or endorse any company, product, or service.
Materials
| 50 mL conical tubes | Fisher Scientific | 14-432-22 | |
| 6 Well Plate | Fisher Scientific | 07-000-646 | |
| BD Brilliant Stain Buffer Plus | BD Biosciences | 566385 | |
| BD Cytofix | BD Biosciences | 554655 | For only fixing cells |
| Bovine serum albumin | Millipore Sigma | A7906 | For preparing FACS staining buffer |
| CD11b AF700 | Biolegend | 101222 | Clone: M1/70 |
| CD11c PerCP/Cy5.5 | Biolegend | 117325 | Clone: N418 |
| CD197 PE/Dazzle594 | Biolegend | 120121 | Clone: 4B12 |
| CD200R3 APC | Biolegend | 142207 | Clone: Ba13 |
| CD206 PE | Biolegend | 141705 | Clone: C068C2 |
| CD45 BUV737 | BD Biosciences | 612778 | Clone: 104/A20 |
| CD86 BUV395 | BD Biosciences | 564199 | Clone: GL1 |
| CD8a BV421 | Biolegend | 100737 | Clone: 53-6.7 |
| Comp Bead anti-mouse | BD Biosciences | 552843 | For compensation control |
| DNase I | Millipore Sigma | 11284932001 | Bovine pancreatic deoxyribonuclease I (DNase I) |
| F4/80 PE/Cy7 | Biolegend | 123113 | Clone: BM8 |
| Fc Block | Biolegend | 101301 | Clone: 93 |
| Fixation/Permeabilization Solution Kit | BD Biosciences | 554714 | For fixing and permeabilization of cells. |
| HEPES buffer | Thermo Fisher | 15630080 | Buffer to supplement cell media |
| Liberase | Millipore Sigma | 5401127001 | Blend of purified Collagenase I and Collagenase II |
| LIVE/DEAD Fixable Blue Dead Cell Stain Kit | Thermo Fisher | L23105 | Viability dye |
| Ly6c AF488 | Biolegend | 128015 | Clone: HK1.4 |
| Ly6g BV510 | Biolegend | 127633 | Clone: 1A8 |
| MHCII BV786 | BD Biosciences | 742894 | Clone: M5/114.15.2 |
| Phosphate buffer saline | Thermo Fisher | D8537 | |
| RPMI | Thermo Fisher | 11875176 | Cell culture media |
| Siglec F BV605 | BD Biosciences | 740388 | Clone: E50-2440 |
| V-bottom 96-well plate |
Referências
- Joung, Y. H. Development of implantable medical devices: from an engineering perspective. International Neurourology Journal. 17 (3), 98-106 (2013).
- Langer, R., Folkman, J. Polymers for the sustained release of proteins and other macromolecules. Nature. 263 (5580), 797-800 (1976).
- Rolfe, B., et al., Eberli, D., et al. The fibrotic response to implanted biomaterials: implications for tissue engineering. Regenerative Medicine and Tissue Engineering-Cells and Biomaterials. , (2011).
- Erdem, S., Gür, M., Kaman, M. O. Static and dynamic analyses of fracture fixation bone-plate systems for different plate materials and dimensions. Bio-Medical Materials and Engineering. 29 (5), 611-628 (2018).
- Kang, C. -. W., Fang, F. -. Z. State of the art of bioimplants manufacturing: part I. Advances in Manufacturing. 6 (1), 20-40 (2018).
- Sadtler, K., et al. Divergent immune responses to synthetic and biological scaffolds. Biomaterials. 192, 405-415 (2019).
- Sadtler, K., et al. Design, clinical translation and immunological response of biomaterials in regenerative medicine. Nature Reviews Materials. 1 (7), 16040 (2016).
- Hubbell, J. A., Thomas, S. N., Swartz, M. A. Materials engineering for immunomodulation. Nature. 462 (7272), 449-460 (2009).
- Badylak, S. F., Valentin, J. E., Ravindra, A. K., McCabe, G. P., Stewart-Akers, A. M. Macrophage phenotype as a determinant of biologic scaffold remodeling. Tissue Engineering Part A. 14 (11), 1835-1842 (2008).
- Wolf, M. T., et al. Polypropylene surgical mesh coated with extracellular matrix mitigates the host foreign body response. Journal of Biomedical Material Research Part A. 102 (1), 234-246 (2014).
- Zhang, L., et al. Zwitterionic hydrogels implanted in mice resist the foreign-body reaction. Nature Biotechnology. 31 (6), 553-556 (2013).
- Sussman, E. M., Halpin, M. C., Muster, J., Moon, R. T., Ratner, B. D. Porous implants modulate healing and induce shifts in local macrophage polarization in the foreign body reaction. Annals of Biomedical Engineering. 42 (7), 1508-1516 (2014).
- Tan, H., Marra, K. G. Injectable, Biodegradable hydrogels for tissue engineering applications. Materials. 3 (3), 1746-1767 (2010).
- Lee, D. C., Lamm, R. J., Prossnitz, A. N., Boydston, A. J., Pun, S. H. Dual polymerizations: untapped potential for biomaterials. Advance Healthcare Materials. 8 (6), 1800861 (2019).
- Sadtler, K., et al. Developing a pro-regenerative biomaterial scaffold microenvironment requires T helper 2 cells. Science. 352 (6283), 366-370 (2016).
- Gower, R. M., et al. Modulation of leukocyte infiltration and phenotype in microporous tissue engineering scaffolds via vector induced IL-10 expression. Biomaterials. 35 (6), 2024-2031 (2014).
- Graney, P. L., Lurier, E. B., Spiller, K. L. Biomaterials and bioactive factor delivery systems for the control of macrophage activation in regenerative medicine. ACS Biomaterials Science & Engineering. 4 (4), 1137-1148 (2018).
- Kontos, S., Grimm, A. J., Hubbell, J. A. Engineering antigen-specific immunological tolerance. Current Opinion Immunology. 35, 80-88 (2015).
- Sadtler, K., Elisseeff, J. H. Analyzing the scaffold immune microenvironment using flow cytometry: practices, methods and considerations for immune analysis of biomaterials. Biomaterials Science. 7 (11), 4472-4481 (2019).
- Baumgarth, N., Roederer, M. A practical approach to multicolor flow cytometry for immunophenotyping. Journal of Immunological Methods. 243 (1-2), 77-97 (2000).
- Shapiro, H. M. . Practical Flow Cytometry. , (2003).
- Nolan, J. P., Condello, D. Spectral flow cytometry. Current Protocols in Cytometry. , (2013).
- Wolf, M. T., et al. A biologic scaffold-associated type 2 immune microenvironment inhibits tumor formation and synergizes with checkpoint immunotherapy. Science Translational Medicine. 11 (477), (2019).
- Kahng, J., et al. Flow cytometric white blood cell differential using CytoDiff is excellent for counting blasts. Annals of laboratory medicine. 35 (1), 28-34 (2015).
- Sionov, R. V., et al. Isolation and characterization of neutrophils with anti-tumor properties. Journal of Visualized Experiments. (100), e52933 (2015).
- Lay, J. C., Peden, D. B., Alexis, N. E. Flow cytometry of sputum: assessing inflammation and immune response elements in the bronchial airways. Inhalation Toxicology. 23 (7), 392-406 (2011).
- Brooks, C. R., van Dalen, C. J., Hermans, I. F., Douwes, J. Identifying leukocyte populations in fresh and cryopreserved sputum using flow cytometry. Cytometry Part B: Clinical Cytometry. 84 (2), 104-113 (2013).

