Mosquito-Associated Virus Isolation from Field-Collected Mosquitoes
Summary
Numerous novel virus-like sequences have been found in mosquitoes due to the extensive use of sequencing technologies. We provide an effective procedure for isolating and amplifying viruses using vertebrate and mosquito cell lines, which might serve as the basis for future studies on mosquito-associated viruses, including mosquito-borne and mosquito-specific viruses.
Abstract
With the broad application of sequencing technologies, many novel virus-like sequences have been discovered in arthropods, including mosquitoes. The two main categories of these new mosquito-associated viruses are "mosquito-borne viruses (MBVs)" and "mosquito-specific viruses (MSVs)". These novel viruses might be pathogenic to both vertebrates and mosquitoes, or they could just be symbiotic with mosquitoes. Entity viruses are essential to confirm the biological characters of these viruses. Thus, a detailed protocol has been described here for virus isolation and amplification from field-collected mosquitoes. First, the mosquito samples were prepared as supernatants of mosquito homogenates. After centrifugation twice, the supernatants were then inoculated into either mosquito cell line C6/36 or vertebrate cell line BHK-21 for virus amplification. After 7 days, the supernatants were collected as the P1 supernatants and stored at -80 °C. Next, P1 supernatants were passaged twice more in C6/36 or BHK-21 cells while the cell status was being checked daily. When cytopathogenic effect (CPE) on the cells was discovered, these supernatants were collected and used to identify viruses. This protocol serves as the foundation for future research on mosquito-associated viruses, including MBVs and MSVs.
Introduction
Mosquitoes are a group of important pathogenic arthropod vectors. There are approximately 3,500 species of mosquitoes in the family Culicidae1,2. The development of high-throughput sequencing technologies has led to the discovery of many novel, virus-like sequences in mosquitoes from different parts of the world3. Generally, these mosquito-associated viruses can be classified into two main groups: MBVs and MSVs.
MBVs are a group of diverse viruses that are causative agents of many human or animal illnesses, such as yellow fever virus (YFV), dengue virus (DENV), Japanese encephalitis virus (JEV), West Nile virus (WNV), and Rift Valley fever virus (RVFV)4. They have seriously threatened public health by causing severe morbidity and mortality in both humans and animals across the world. MBVs naturally sustain a life cycle between diverse hosts through transmission from an infected mosquito to a naïve host, as well as from a virus-infected host and to a feeding mosquito5. Therefore, these viruses can infect both mosquito cell lines and vertebrate cell lines in the lab1.
MSVs, which include Yichang virus (YCN), Culex flavivirus (CxFV), and Chaoyang virus (CHAOV), are a subgroup of insect-specific viruses1,6,7. In recent years, there has been a rise in the discovery of novel MSVs, and some of these MSVs have been found to have an impact on the transmission of MBVs. For example, CxFV, which can be a persistent infection in Culex pipiens, could suppress WNV replication at an early stage8. Another insect-specific flavivirus, cell-fusing agent virus (CFAV), has been found to inhibit the propagation of DENV and Zika virus (ZIKV) in Aedes aegypti mosquitoes9. Thus, this protocol is a useful approach for isolating mosquito-associated viruses and can help in further research into the distribution of pathogens related to mosquitoes and the control of mosquito-borne diseases.
Protocol
1. Mosquito sampling and sorting
- Trap the adult mosquitoes through the light traps MXA-02 or carbon dioxide mosquito traps in the field.
- Kill the collected mosquitoes by dipping in liquid nitrogen10,11. Transport them to the laboratory by the cold-chain logistics system12.
NOTE: Dry ice was primarily used in the cold-chain logistics system. - (Optional) If the sample sites were near the lab, directly send the alive mosquitoes in the net traps to the laboratory and euthanize them by freezing at ≤-20 °C for 30 min4.
- Make a morphological identification of the collected mosquitoes through the tool book13 for classification and identification of mosquitoes.
NOTE: If necessary, DNA barcoding could be used to classify and identify mosquitoes14,15. - Based on sampling dates and sites, assign them into different pools, and then store them in 2-5 mL tubes.
- Draw a table to record each tube with mosquito species, number, sampling code, date, and sites.
- Print the labels with sampling codes and attach them to the tubes.
- Keep tubes with the mosquito samples at -80 °C until further analysis.
2. Mosquito grinding
- Place 20-50 mosquitoes into each 2 mL sterile tube with 3 mm ceramic beads with 1.5 mL of Roswell Park Memorial Institute (RPMI) medium containing 2% Penicillin-Streptomycin-Amphotericin B Solution.
NOTE: Keep the tubes on ice oron anice tray. - Homogenize the mosquitoes with a low-temperature tissue homogenizer by grinding for 30 s at 70 Hz at 4 °C for three cycles16.
- Centrifuge the mixtures at 15,000 × g for 30 min at 4 °C and transfer the supernatants to new tubes.
- Centrifuge the supernatants again at 15,000 × g for 10 min at 4 °C to remove the mosquito debris12,17.
NOTE: If necessary, the supernatants can be filtered using 0.22 μm filters. - Aliquot the supernatants (P0) into 2 mL screw cap storage tubes (200 µL per tube) and store them at -80 °C.
3. Preparation of cells and cell culture maintenance medium
NOTE: Mosquito cell line C6/36 (Aedes albopictus RNAi-deficient) and vertebrate cell line BHK-21 (baby hamster kidney) were used for the virus amplification and isolation.
- Inoculate 1 × 106 C6/36 cells into a 75 cm2 flask with 10 mL of RPMI medium supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin (P/S) at 28 °C in a 5% CO2 humidified incubator.
- Inoculate 1 × 106 BHK-21 cells into a 75 cm2 flask with 10 mL of Dulbecco's modified Eagle medium (DMEM) supplemented with 10% FBS and 1% P/S at 37 °C in a 5% CO2 humidified incubator.
NOTE: The cells were passaged two or three times per week17,18. - Seed the cells cultured in a 75 cm2 flask (C6/36 or BHK-21 cells) into the 24-well plates (C6/36 per well: 1.4 × 105; BHK-21 per well: 0.8 × 105).
- Place the 24-well plates at 28 °C or 37 °C overnight.
- Observe the cells under the microscope and confirm whether cell confluency in each well is 80%-90%.
- Prepare the cell culture maintenance medium (500 mL).
NOTE: For C6/36, the medium was RPMI medium supplemented with 2% FBS and 2% Penicillin-Streptomycin-Amphotericin B Solution. For BHK-21, the medium was DMEM medium supplemented with 2% FBS and 2% Penicillin-Streptomycin-Amphotericin B Solution.
4. Virus isolation
NOTE: All steps were carried out in a biosafety level 2 (BSL-2) laboratory. The safety level requirement of the biosafety laboratory was determined by the biosafety risk assessment based on the regulations of different countries and regions. The process must be performed in a biosafety cabinet.
- Remove the cell culture medium from the 24-well plates and add 100 µL of the cell culture maintenance medium and 100 µL of the supernatant of the mosquito homogenate (P0) to each well.
- Incubate the plates at 28 °C or 37 °C for 60 min and gently shake them every 15 min to prevent cell drying.
- Remove the supernatant and rinse each well gently with 600 µL of cell culture maintenance medium to remove the debris completely.
- Add 800 µL of cell culture maintenance medium to each well and keep the plates in a 5% CO2 humidified incubator at 28 °C or 37 °C for 7 days.
- Monitor the cell state of each well under the microscope daily15.
- Collect the supernatants of cells (P1 supernatants) on the 7th day and store the supernatants at -80 °C.
- Repeat steps 4.1-4.6 2x to get P2 and P3 supernatants.
NOTE: In step 4.3, rinsing each well gently with 600 µL of cell culture maintenance medium was optional for the P2 and P3. - Depending on which well shows the cytopathogenic effect (CPE) — cells die, pyknosis, and detach from the surface; fusion with adjacent cells to form syncytia; or the appearance of nuclear or cytoplasmic inclusion bodies — add 300-400 µL of the supernatant of this CPE well into each well of the new 6-well plates containing cells (the cell confluency was 80%-90%) to greatly amplify the viruses.
- Collect the supernatants and aliquot them into 2 mL screw cap storage tubes (500 µL per tube). Store them at -80 °C.
NOTE: After three generations of successive passaging in cells, the samples that did not induce CPE were discarded. If necessary, the generations of successive passaging in cells can be increased.
5. Detecting viral sequences by RT-PCR
- Extract total RNA from 200 µL of the viral supernatant using a viral RNA mini kit16.
NOTE: Here, anautomated nucleic acid extraction system was used to extract total RNA following the instrument manufacturer's instructions. - Convert the RNA to cDNA following the RT-PCR kit instructions using a PCR instrument.
- Add 15 µL of RNA mixed with 1 µL of random primers into the PCR tube. Place the tube at 70 °C for 5 min and then place it on ice for 5 min.
- Add 5 µL of 5x buffer and 25 µL of mixture (10 mM dNTP, 40 U/µL RNase inhibitor, and 200 U/µL M-MLV) into the tube and place the tube at 42 °C for 60 min and 95 °C for 10 min.
- Store the tube at -20 °C.
- Use a PCR kit and universal primers (Table 1) to detect viruses by PCR.
- For arboviruses, set up the PCR reaction system (total 30 µL) with the following components: 3 µL of 10x Taq buffer, 3 µL of dNTP (1mM), 1 µL of forward primer (10 µm), 1 µL of reverse primer (10 µm), 0.3 µL of Taq (10 U/µL), 19.7 µL of ddH2O, and 2 µL of cDNA.
- For the detection of flaviviruses, set up the reaction process as: stage 1: 35 cycles of 94 °C for 30 s, 52 °C for 30 s, and 72 °C for 30 s; stage 2: 72 °C for 10 min.
- For the detection of alphaviruses, set up the reaction process as: stage 1: 35 cycles of 94 °C for 30 s, 50 °C for 30 s, and 72 °C for 30 s; stage 2: 72 °C for 10 min.
- For the detection of bunyaviruses, set up the reaction process as: stage 1: 35 cycles of 94 °C for 30 s, 50 °C for 30 s, and 72 °C for 30 s; stage 2: 72 °C for 10 min.
- Detect the PCR amplification products by 1% agarose gel electrophoresis and send them for sequencing analysis.
NOTE: If conditions permit, the supernatants can be undergo next generation sequencing analysis to identfy new viruses and obtain the whole viral genome seqeunces.
Representative Results
After inoculation with the supernatants of the mosquito homogenates (P0), the C6/36 cells exhibited a wide intercellular space, and exfoliated cells were observed at 120 h (Figure 1A) compared to the uninoculated cells (control) at the same time (Figure 1B). After incubating the BHK-21 cells with the P3 supernatants, visible CPE was observed in the BHK-21 cells at 48 h (Figure 1C) in contrast to the control cells (Figure 1D). PCR was performed to determine viral species. The universal primers for the detection of flaviviruses, alphaviruses, and bunyaviruses were commercially synthesized (Table 1). A PCR product for the bunyavirus Ebinur Lake Virus was set as a positive control and added into Lane 1. The universal primers for bunyaviruses, flaviviruses, and alphaviruses were used to generate the PCR products for the viral supernatant, which were then added into Lane 2, 3, and 4, respectively. The estimated sizes of the PCR products for flaviviruses, alphaviruses, and bunyaviruses were 266 bp, 434 bp, and 251 bp, respectively. The bands were only visible in Lane 2 and the positive control Lane 1. As a result, the virus in the supernatants is likely to be a bunyavirus (Figure 2).
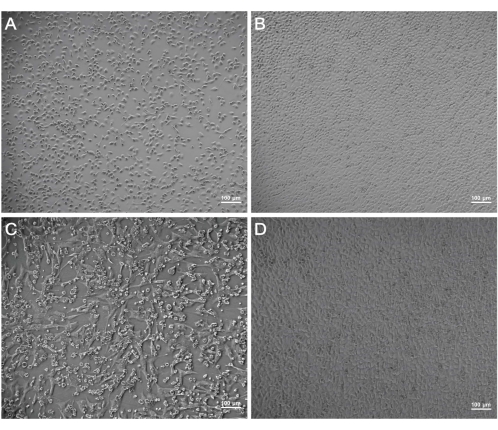
Figure 1: CPE observation of cells after the incubation with the viral supernatants. The status of C6/36 cells at 120 h post infection (CPE) (A) and that of the control cells at the same time (B). The status of BHK-21 cells at 48 hpi (C) and that of the control cells (D). Scale bars = 100 µm. Abbreviations: CPE = cytopathogenic effect; hpi = hours post infection. Please click here to view a larger version of this figure.
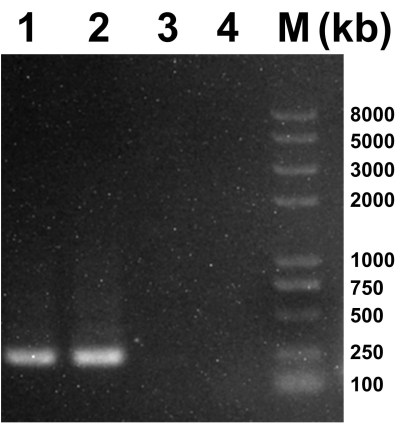
Figure 2: PCR results for viral supernatants. Lane 1 represent the positive control of bunyaviruses (Ebinur Lake Virus). Lanes 2-4 represented the PCR results of supernatants by using universal primers for bunyaviruses (251 bp), flaviviruses (266 bp), and alphaviruses (434 bp). Please click here to view a larger version of this figure.
| Virus | Primer | Oligonucleotide (5'→3') | The size of PCR product (bp) | ||
| Flaviviruses | F1 | TACAACATGATGGGAAAGAGAGAGAA | 266 | ||
| F2 | GTGTCCCAGCCGGCGGTGTCATCAGC | ||||
| Alphaviruses | M2w | YAGAGCDTTTTCGCAYSTRGCHW | 434 | ||
| cMw3 | ACATRAANKGNGTNGTRTCRAANCCDAYCC | ||||
| Bunyaviruses | BCS82C | ATGACTGAGTTGGAGTTTCATGATGTCGC | 251 | ||
| BCS332V | TGTTCCTGTTGCCAGGAAAAT | ||||
Table 1: Universal primers for the arbovirus detection.
Discussion
The objective of this method was to offer a practical way for isolating mosquito-associated viruses using various cell lines. It is critical to add the Antibiotic-Antimycotic (Penicillin-Streptomycin-Amphotericin) to the supernatants of the mosquito homogenates to avoid contamination by bacteria or fungi. Mosquitoes and viral supernatants obtained in the field must be refrigerated at -80 °C to avoid repeated freeze-thaw cycles.
Another critical step in the protocol was grinding. Mosquito samples should be thoroughly powdered and stored on ice. Insufficiently ground mosquito tissues can induce cell death and may skew the findings of CPE analysis. Prior to virus isolation, it is necessary to employ normal mosquitoes to confirm the grinding parameters. Grinding must be done at low temperatures to prevent the virus from becoming inactive during the process.
This approach can effectively isolate and amplify mosquito-associated viruses, but is only acceptable for viruses that may display CPE in mammalian or insect cell lines. This approach is inappropriate for viruses that do not induce CPE. Using the protocol, we may obtain viral candidates that contain only one virus type or a mixture with different viruses. Further investigation — viral purification, morphological identification, and plaque analysis — is still needed to obtain the purified virus.
Declarações
The authors have nothing to disclose.
Acknowledgements
This work was supported by the Wuhan Science and Technology Plan Project (2018201261638501).
Materials
| 0.22 µm membrane filter | Millipore | SLGP033RB | Polymer films with specific pore ratings.To remove cell debris and bacteria. |
| 24-well plates | CORNING | 3524 | Containers for cell |
| 75 cm2 flasks | CORNING | 430641 | Containers for cell |
| a sterile 2 mL tube with 3 mm ceramic beads | |||
| Antibiotic-Antimycotic | Gibco | 15240-062 | Antibiotic in the medium to prevent contamination from bacteria and fungi |
| Automated nucleic acid extraction system | NanoMagBio | S-48 | |
| BHK-21 cells | National Virus Resource Center, Wuhan Institute of Virology | ||
| C6/36 cells | National Virus Resource Center, Wuhan Institute of Virology | ||
| Centrifugal machine | Himac | CF16RN | Instrument for centrifugation of mosquito samples |
| CO2 | |||
| Dulbecco’s minimal essential medium (DMEM) | Gibco | C11995500BT | medium for vertebrate cell lines |
| Ebinur Lake virus | Cu20-XJ isolation | ||
| Feta Bovine Serum (FBS) | Gibco | 10099141C | Provide nutrition for cells |
| high-speed low-temperature tissue homogenizer | servicebio | KZ-III-F | Instrument for grinding |
| incubator (28 °C) | Panasonic | MCO-18AC | Instrument for cell culture |
| incubator (37 °C) | Panasonic | MCO-18AC | Instrument for cell culture |
| PCR tube | |||
| penicillin-streptomycin | Gibco | 15410-122 | Antibiotic in the medium to prevent contamination from bacteria |
| Penicillin-Streptomycin-Amphotericin B Solution | Gibco | 15240096 | |
| Refrigerator (-80 °C) | sanyo | MDF-U54V | |
| Roswell Park Memorial Institute medium (RPMI) | Gibco | C11875500BT | medium for mosiquto cell lines |
| Screw cap storage tubes (2 mL) | biofil | FCT010005 | |
| sterile pestles | Tiangen | OSE-Y004 | Consumables for grinding |
| TGrinder OSE-Y30 electric tissue grinder | Tiangen | OSE-Y30 | Instrument for grinding |
| The dissecting microscope | ZEISS | stemi508 | |
| the light traps MXA-02 | Maxttrac | ||
| The mosquito absorbing machine | Ningbo Bangning | ||
| The pipette tips | Axygen | TF | |
| The QIAamp viral RNA mini kit | QIAGEN | 52906 | |
| Tweezers | Dumont | 0203-5-PO |
Referências
- Xia, H., Wang, Y., Atoni, E., Zhang, B., Yuan, Z. Mosquito-associated viruses in China. Virologica Sinica. 33 (1), 5-20 (2018).
- Atoni, E., et al. A dataset of distribution and diversity of mosquito-associated viruses and their mosquito vectors in China. Scientific Data. 7 (1), 342 (2020).
- Atoni, E., et al. The discovery and global distribution of novel mosquito-associated viruses in the last decade (2007-2017). Reviews in Medical Virology. 29 (6), 2079 (2019).
- Xia, H., et al. Comparative metagenomic profiling of viromes associated with four common mosquito species in China. Virologica Sinica. 33 (1), 59-66 (2018).
- Ong, O. T. W., Skinner, E. B., Johnson, B. J., Old, J. M. Mosquito-borne viruses and non-human vertebrates in Australia: A review. Viruses. 13 (2), 265 (2021).
- Agboli, E., Leggewie, M., Altinli, M., Schnettler, E. Mosquito-specific viruses-transmission and interaction. Viruses. 11 (9), 873 (2019).
- Halbach, R., Junglen, S., van Rij, R. P. Mosquito-specific and mosquito-borne viruses: evolution, infection, and host defense. Current Opinion in Insect Science. 22, 16-27 (2017).
- Bolling, B. G., Olea-Popelka, F. J., Eisen, L., Moore, C. G., Blair, C. D. Transmission dynamics of an insect-specific flavivirus in a naturally infected Culex pipiens laboratory colony and effects of co-infection on vector competence for West Nile virus. Virology. 427 (2), 90-97 (2012).
- Baidaliuk, A., et al. Cell-fusing agent virus reduces arbovirus dissemination in Aedes aegypti mosquitoes in vivo. Journal of Virology. 93 (18), 00715-00719 (2019).
- Atoni, E., et al. Metagenomic virome analysis of Culex mosquitoes from Kenya and China. Viruses. 10 (1), 30 (2018).
- Xia, H., et al. First isolation and characterization of a group C Banna virus (BAV) from Anopheles sinensis mosquitoes in Hubei, China. Viruses. 10 (10), 555 (2018).
- Shi, C., et al. Stability of the virome in lab- and field-collected Aedes albopictus mosquitoes across different developmental stages and possible core viruses in the publicly available virome data of Aedes mosquitoes. mSystems. 5 (5), 00640 (2020).
- Zhou, M., Chu, H. . Handbook for Classification and Identification of Main Vectors. , (2019).
- Wang, G., et al. Identifying the main mosquito species in China based on DNA barcoding. Plos One. 7 (10), (2012).
- Ratnasingham, S., Hebert, P. D. N. Bold: The Barcode of Life Data System (www.barcodinglife.org). Molecular Ecology Notes. 7 (3), 355-364 (2007).
- Huang, Y., et al. In vitro and in vivo characterization of a new strain of mosquito Flavivirus derived from Culicoides. Viruses. 14 (6), 1298 (2022).
- Zhao, L., et al. Characterization of a novel Tanay virus isolated from Anopheles sinensis mosquitoes in Yunnan, China. Frontiers in Microbiology. 10, 1963 (1963).
- Ren, N., et al. Characterization of a novel reassortment Tibet orbivirus isolated from Culicoides spp. in Yunnan, PR China. Journal of General Virology. 102 (9), 001645 (2021).

